Abstract
Neutrophil extracellular traps (NETs) constitute a vital antimicrobial defense mechanism of neutrophils, contributing to various physio-pathological processes; however, the role of plasma membrane asymmetry in this process remains unknown. Here we identify Xk-related protein 8 (XKR8), a plasma membrane phospholipid scramblase, as a pivotal regulator of NETs formation. Upon NETs induction, XKR8 is cleaved by caspase-3, thereby disrupting plasma membrane lipid asymmetry via phospholipid scrambling. Mutation of the caspase-3 cleavage site in XKR8 impairs NET formation. Inhibition of calcium signals before lipid scrambling abrogates NET formation, whereas blockade after scrambling does not. Cleaved XKR8 reorients plasma membrane lipids, altering membrane lipid tension and promoting Ca2+ signals through mechanosensitive channels. XKR8-deficient mice exhibit compromised NET formation and impaired control of Candida albicans pulmonary infection, showing that XKR8 is indispensable for neutrophil-driven immune responses in vivo. These findings define caspase-3–XKR8plasma membrane phospholipid scrambling as a central mechanism controlling NET formation and underscore its critical role in neutrophil-dependent antifungal immunity.
Similar content being viewed by others
Main
Neutrophils, the most abundant circulating leukocytes in human blood, constitute the frontline of innate immune defense against invading pathogens1,2,3. Beyond anti-infection, neutrophils have emerged as critical regulators of a broad spectrum of pathological conditions, including cancer and noninfectious diseases, such as cystic fibrosis, rheumatoid arthritis, gallstone formation, thrombosis and autoimmune disorders4,5,6,7,8. To combat pathogens, neutrophils deploy a diverse arsenal of mechanisms, including phagocytosis, the release of antimicrobial proteins through degranulation and the expulsion of NETs.
The process of NET production and release was originally characterized as a unique form of cell death, termed NETosis, which enables neutrophils to eliminate pathogens in the extracellular space9,10. NETs are web-like filamentous structures composed of decondensed chromatin, intricately decorated with antimicrobial enzymes such as neutrophil elastase and myeloperoxidase (MPO)11,12,13. These adhesive chromatin networks entrap invading microorganisms and neutralize them through locally concentrated bactericidal enzymes and elevated level of reactive oxygen species (ROS)14,15,16. Unlike other programmed cell death pathways10,17, NETosis constitutes a distinct cellular process. While NETs are professional effectors in controlling infections, accumulating evidence reveals their much broader implications in noninfectious diseases and cancer18,19,20,21,22,23.
Tight regulation of neutrophil activation is crucial to prevent excessive or uncontrolled inflammation, which can result in tissue damage and contribute to disease pathogenesis. Although a wide range of stimuli have been identified that can trigger NET formation24,25,26, the precise molecular mechanisms underlying this process remain incompletely understood. Intriguingly, despite the diversity of NET-inducing triggers, they seem to converge on a common end point (NET formation), suggesting the existence of a central signaling hub that orchestrates this response. Determining the signaling pathways and molecular players that regulate NET release is therefore critical, not only for advancing our understanding of neutrophil biology but also for identifying new therapeutic targets to treat disorders associated with aberrant NETosis; however, current knowledge regarding the molecular regulation of NET formation remains limited and fragmented, highlighting the urgent need for comprehensive mechanistic studies.
XKR8 was initially identified as a caspase-3-activated phospholipid scramblase, primarily responsible for facilitating phosphatidylserine (PS) exposure, a classic hallmark of apoptosis27,28. In contrast to the well-characterized calcium-activated scramblase TMEM16F, XKR8 mediates lipid scrambling upon caspase-3-mediated cleavage at its carboxy-terminal recognition site during apoptosis, leading to the externalization of PS on the outer leaflet of plasma membrane (PM). This disruption of PM lipid asymmetry constitutes a defining feature of apoptosis driven by XKR8 (ref. 27). At homeostasis, neutrophils maintain a tightly regulated PM lipid asymmetry that supports essential physiological functions, including controlling cell deformability during circulation and preventing premature clearance by phagocytes. While externalized PS functions as an ‘eat-me’ signal to promote the efficient clearance of the apoptotic cells by phagocytes, XKR8-mediated PS exposure is critical for the efficient removal of apoptotic neutrophils in vivo29. Given the involvement of PS exposure and PM rupture in NETosis, along with the remarkably short lifespan of circulating neutrophils30,31,32,33, we hypothesize that loss of PM lipid asymmetry may have a pivotal role in neutrophil function, particularly in the context of NET formation.
Results
XKR8 is essential for NET formation
Profound morphological alterations to the PM have been documented during the early stages of NET formation26,33. To investigate whether PM phospholipids scrambling occurs during the initiation of NET formation, we employed the classical NOX2-dependent NET activator, phorbol 12-myristate 13-acetate (PMA) in serum-free RPMI 1640 medium supplemented with 10 mM HEPES, to stimulate polymorphonuclear neutrophils (PMNs) purified from mouse bone marrow. Lipid scrambling was monitored by PS exposure detected by annexin V staining (Fig. 1a). NET formation was assessed by quantifying areas of decondensed chromatin DNA labeled with 4,6-diamidino-2-phenylindole (DAPI) (Extended Data Fig. 1a). Additionally, PM permeability was examined using propidium iodide staining in flow cytometry and SYTOX Green in live-cell imaging (Fig. 1a). Notably, we observed a progressive increase in PS exposure that preceded PM permeabilization and subsequent release of decondensed chromatin (NETs) in both mouse bone-marrow-purified neutrophils (BMNs) and human peripheral blood neutrophils (Fig. 1a and Extended Data Fig. 1b). Unless otherwise specified, all murine neutrophils used in this study were isolated from bone marrow. Consistently, lipid scrambling was also observed before NET release in all-trans retinoic acid (ATRA)-differentiated HL-60 cells (dHL-60), a neutrophil-like human myeloid leukemia cell line (Extended Data Fig. 1c,d). Notably, by sorting annexin V+PI− neutrophils (scrambled but not permeabilized), we found that only neutrophils undergoing early lipid scrambling proceeded to release NETs, suggesting that lipid scrambling marks neutrophils fated for NETosis (Fig. 1b). Collectively, these findings demonstrate that lipid scrambling precedes chromatin release and is a defining early event in NET formation.
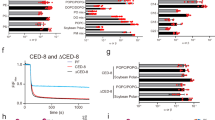
a, PS exposure and NET formation in BMNs treated with PMA (100 nM) at the indicated time points, assessed by flow cytometry (n = 3) and fluorescence microscopy (n = 6), respectively. Representative histograms of annexin V staining gated on live BMNs (propidium iodide-negative, PI−) (left) and quantification of annexin V+PI− cells and NETs (right). b, NET formation in live BMNs sorted by annexin V+ (AV+) and annexin V− (AV−) at 2 h of PMA treatment. Gating strategy (left) and quantification of NETs in defined groups at 4 h after sorting (right, n = 9 for PMA; n = 12 for AV+ and AV−). c, Representative histograms and percentages of PS exposure in WT, XKR8 KO and TMEM16F KO BMNs treated with PMA (n = 3). d, NET formation in WT and XKR8 KO BMNs treated with PMA (100 nM, 6 h), A23187 (1.25 μM, 4 h), nigericin (5 μM, 4 h) or LPS (10 μg ml−1, 2 h) (n = 10). e,f, NET formation in WT and XKR8 KO BMNs co-cultured with microorganisms (multiplicity of infection (MOI) of 10) C. albicans (1.5 h) (n = 9) (e), S. aureus, L. monocytogenes (L. M.) or E. coli for 1.5 h (n = 10) (f). g, As in d, NET formation in WT and XKR8 KO neutrophil-like HL-60 (dHL-60) cells treated with PMA, A23187 or LPS (n = 8). Representative immunofluorescence images (left) of DNA staining (DAPI) and citrullinated histone H3 (Cit-H3) in BMNs, along with quantification of NETs (right) (d–g). NETs were quantified by a nuclear area exceeding 80 µm2 in BMNs (b,d) or 100 µm2 in dHL-60 cells (g), or the ratio of Cit-H3 to DNA (e,f). Each dot represents an individual field of view (b,d–g) or a biological replicate (c). Scale bars, 10 μm (d–g). Data are represented as mean ± s.e.m. from one representative experiment out of three (b,c,f), four (e,g), six (a) and eight (d) independent experiments. Statistical analysis was performed using two-way analysis of variance (ANOVA) (c–g) and one-way ANOVA (b). P values are shown on the graphs.
Given that phospholipid scrambling is primarily mediated by either Ca2+-activated TMEM16F or caspase-3-activated XKR8, we sought to determine which scramblase is responsible for lipid scrambling during NET formation. Of note, despite being ubiquitously expressed, XKR8 expression is notably enriched in granulocytes such as neutrophils and eosinophils, both in mice and humans (Extended Data Fig. 1e,f). To investigate the functional role of XKR8, we generated Xkr8 germline knockout (KO) mice (Extended Data Fig. 2a). Depletion of Xkr8 mRNA in BMNs was confirmed by reverse transcription PCR (RT–PCR) and RNA sequencing (RNA-seq) (Extended Data Fig. 2b,c). Notably, XKR8-deficient neutrophils, but not those lacking TMEM16F, failed to externalize PS during NET induction by PMA (Fig. 1c). Of note, RNA-seq analysis of XKR8-deficient neutrophils revealed minimal changes in global gene expression, indicating that loss of XKR8 does not significantly alter the transcriptional profile of neutrophils (Extended Data Fig. 2c). Naive wild-type (WT) and XKR8 KO mice shared similar spleen and lymph node structures, comparable immune repertoire and cytokine levels (Extended Data Fig. 2d,e and Supplementary Table 1). Moreover, XKR8 KO BMNs displayed normal lobulated nuclear morphology, comparable to WT counterparts (Extended Data Fig. 2f). Expression levels of canonical neutrophil markers were also unchanged between WT and XKR8 KO mice, corroborating RNA-seq results and suggesting that XKR8 is dispensable for neutrophil development (Extended Data Fig. 2g).
Owing to the unavailability of an XKR8-specifc antibody, we further validated the loss of XKR8 function by assessing its canonical role in apoptotic lipid scrambling. As previously reported, apoptotic PS exposure is mediated by caspase-3-activated XKR8 (ref. 28). Consistent with this, XKR8 KO neutrophils exhibited marked defects in PS externalization following apoptosis induced by staurosporine (STS) or S63845 (MCL1 inhibitor), in contrast to WT controls (Extended Data Fig. 3a). These findings confirm the functional loss of XKR8 in neutrophils from XKR8 KO mice and support its essential role in both apoptotic and NETosis-associated lipid scrambling.
Of note, XKR8 KO neutrophils failed to form NETs in response to various stimuli in vitro, including PMA, calcium ionophore A23187, lipopolysaccharides (LPS), nigericin, monosodium urate (MSU) crystals and zymosan. This deficiency was evidenced by the absence of decondensed chromatin and citrullinated histone H3 (Cit-H3, a hallmark for NETs), as assessed by immunofluorescence (Fig. 1d and Extended Data Fig. 3b–d). Beyond chemical NET inducers, XKR8 KO neutrophils also exhibited severely impaired NET formation when challenged with physiological NET activators, including the fungal organism Candida albicans and various bacterial species (Staphylococcus aureus, Listeria monocytogenes and Escherichia coli) (Fig. 1e,f). Notably, XKR8 KO BMNs failed to generate both filamentous chromatin structures and the citrullinated histones, suggesting a disruption in upstream NET signaling, particularly in Ca2+ signaling pathways, given that peptidylarginine deiminase 4 (PAD4) activation is calcium-dependent34. Consistently, XKR8 KO neutrophils released significantly less extracellular double-stranded DNA (dsDNA) in the culture medium compared to WT controls during NET formation (Extended Data Fig. 3e).
Notably, other key neutrophil functions remained intact in XKR8-deficient cells. Degranulation, as measured by surface CD63 staining and phagocytosis, assessed via the uptake of fluorescent latex beads and E. coli, were comparable to WT neutrophils (Extended Data Fig. 3f–h). The extent of NET formation deficiency in XKR8-deficient neutrophils was similar to that observed in PAD4 KO cells (Extended Data Fig. 3i,j). By contrast, neutrophils lacking TMEM16F displayed normal NET formation upon PMA stimulation, indicating that TMEM16F, a Ca2+-activated scramblase, is dispensable for PMA-induced lipid scrambling and NET formation (Extended Data Fig. 3k). To further investigate the role of TMEM16F, we assessed NET formation induced by PMA and A23187 in BMNs deficient in XKR8, TMEM16F or both. While TMEM16F deficiency had no detectable effect on PMA-induced NET formation, A23187-induced NET formation was partially reduced in TMEM16F-deficient BMNs (Extended Data Fig. 3k). These findings indicate that TMEM16F contributes partially to A23187-induced NET formation but is dispensable for NET formation triggered by PMA. Moreover, in WT neutrophils, myeloperoxidase (MPO) and elastase were localized along the decondensed web-like DNA filaments, whereas this localization was absent in XKR8-deficient neutrophils, despite comparable MPO expression levels evaluated by immunoblot (Extended Data Fig. 3l,m).
To further corroborate these findings, we generated XKR8-deficient HL-60 cells by CRISPR–Cas9 gene editing. These cells exhibited impaired PS exposure upon apoptosis induction (Supplementary Fig. 1a–c). When differentiated into neutrophil-like cells (dHL-60), XKR8-deficient dHL-60 also displayed a marked inability to form NETs (Fig. 1g and Supplementary Fig. 1d–f). Overall, these findings clearly establish XKR8 as an essential regulator of NET formation in vitro and underscore its pivotal role in orchestrating the molecular events that drive NETosis.
Activation of XKR8 by caspase-3 triggers NET formation
The signaling cascades that orchestrate NET formation are extremely intricate and not yet fully understood15,26. To further determine the role of XKR8 in NET formation, we examined XKR8-mediated lipid scrambling during PMA-induced NET formation. In WT BMNs, PS exposure was significantly upregulated following PMA stimulation. By contrast, XKR8-deficient BMNs exhibited a marked impairment in PS externalization at the early stages of NET induction (Fig. 2a). A similar defect in lipid scrambling was observed in XKR8-deficient dHL-60 cells during NET formation (Extended Data Fig. 4a).
a, PS exposure during PMA-induced NET formation in WT and XKR8 KO BMNs. Representative histogram (left) and quantification (right) of PS exposure in live cells (annexin V+PI−) at the indicated time points, assessed by flow cytometry (n = 3). b, Quantification of intracellular ROS in WT and XKR8 KO BMNs treated with PMA, measured by luminol assay (n = 3). c–e, Caspase-3 activation in WT and XKR8 KO BMNs treated with PMA, assessed by CellEvent (c) (n = 3), intracellular staining of cleaved caspase-3 (d) (n = 3), and immunoblotting (e). A representative histogram (left) and quantifications (right) are shown for c and d. Raptinal is an apoptosis inducer. Cl., cleaved; s.e., short exposure; l.e., long exposure. β-actin was used as a loading control. f,g, PS exposure (f) and NET formation (g) in WT and XKR8 KO BMNs treated with PMA after preincubation with caspase inhibitor Q-VD (10 μM). Representative histogram (f, left) and quantification (f, right) of PS exposure in live cells (annexin V+PI−) (n = 3) and NET formation (g, n = 12). h, PS exposure in WT, XKR8 KO, caspase-3 KO (CASP3 KO) BMNs treated with PMA. Representative histogram (left) and quantification (right) of PS exposure in live cells (annexin V+PI−) (n = 3). i, NET formation in WT, XKR8 KO, CASP3 KO BMNs treated with PMA, A23187, nigericin or LPS (n = 8). NET formation was defined by a nuclear area greater than 80 µm2 (g,i). Each dot represents an individual field of view (g,i), or a biological replicate (a–d,f,h). Data are represented as the mean ± s.e.m. from one representative experiment out of three (g,h), four (a–c,e,i), five (f) and six (d) independent experiments. Statistical analysis was performed using a two-way ANOVA (a–d,f–i). P values are shown on the graphs.
To determine whether the observed defect in lipid scrambling resulted from disruptions in upstream signaling events, we next analyzed the early molecular pathways involved in PMA-induced NET formation. PMA is known to trigger a NOX2-dependent oxidative burst via the activation of protein kinase C (PKC) and mitogen-activated protein kinases (MAPKs), both of which are critical for NET formation35. Both WT and XKR8 KO BMNs generated comparable levels of ROS following PMA stimulation, as assessed by intracellular and extracellular ROS detection using luminol, isoluminol and DHR123 (Fig. 2b and Extended Data Fig. 4b,c). Pharmacological inhibition of ROS production using diphenyleneiodonium (DPI; an NADPH oxidase inhibitor), Tempo (an ROS scavenger) or GSK2795039 (an NOX2-specific inhibitor) effectively abrogated NET formation (Extended Data Fig. 4d–f). Additionally, inhibition of PKCs with enzastaurin and of MAPK signaling with U0126 and GW5074 also abolished NET formation (Extended Data Fig. 4g,h). Of note, phosphorylation levels of ERK1/2 (p-ERK1/2) and PKC (p-PKC) were equal between WT and XKR8 KO BMNs, indicating that these upstream signaling pathways remain intact in the absence of XKR8 (Extended Data Fig. 4i,j).
Given that XKR8 is a caspase-3-activated phospholipid scramblase, we next investigated whether caspase-3 is involved in PMA-induced NET formation. Notably, caspase-3 activation was equivalent in WT and XKR8 KO BMNs, as demonstrated by caspase-3 substrate cleavage assays (CellEvent), flow cytometric analysis of cleaved caspase-3, and immunoblotting for cleaved caspase-3 fragments (p17/p20) (Fig. 2c–e). Furthermore, the subcellular localization of active caspase-3 was comparable between genotypes during NET induction (Extended Data Fig. 4k). Of note, no activation of other apoptotic caspases, including caspase-7, caspase-8 or caspase-9, were detected during PMA-induced NET formation (Extended Data Fig. 4l,m). These findings indicate that the upstream signaling pathways necessary for NET formation are preserved in XKR8-deficient BMNs.
To further confirm the role of caspase-3 in activating XKR8-mediated lipid scrambling during NET formation, we treated neutrophils with the pan-caspase inhibitor Q-VD-OPh (Q-VD). Inhibition of caspase-3 activity effectively suppressed XKR8-mediated PS exposure upon PMA stimulation (Fig. 2f). Consequently, PMA-induced NET formation was markedly attenuated following Q-VD treatment (Fig. 2g). Similar inhibitory effects were observed upon treatment with other caspases inhibitors, including Z-VAD-FMK (Z-VAD) and emricasan in both mouse and human primary neutrophils (Extended Data Fig. 5a–e). To a broader extent, we observed comparable levels caspase-3 activation in NET formation triggered by PMA, A23187, LPS and nigericin in both WT and XKR8 KO BMNs (Extended Data Fig. 5f,g); however, PS exposure remained significantly impaired in XKR8-deficient neutrophils across all conditions except for A23187 stimulation (Extended Data Fig. 5h,i). Correspondingly, inhibition of caspase-3 activation by Q-VD substantially reduced NET formation in WT neutrophils under all four stimuli, lowering NET levels to those observed in XKR8 KO cells (Extended Data Fig. 5j). Furthermore, neutrophils derived from CASP3 KO mice exhibited profound defects in PS exposure under both apoptotic and NET-inducing conditions (Fig. 2h and Extended Data Fig. 5k–m). Notably, CASP3 KO neutrophils displayed severely impaired NET formation upon stimulation with PMA, A23187, LPS and nigericin, recapitulating the phenotype observed in XKR8-deficient cells (Fig. 2i). These results establish caspase-3 as an essential upstream activator of XKR8, which in turn mediates lipid scrambling and facilitates NET formation.
PMA-elicited NET formation rely on the ROS produced by NOX2 (ref. 36). To examine the relationship between ROS and caspase-3-mediated lipid scrambling during NET formation, we employed DPI and apocynin (NADPH oxidase inhibitors), as well as NA-11 (an inhibitor of NOX2-dependent oxidative burst in neutrophils by activating the glycolytic enzyme phosphofructokinase-1 liver type, PFKL) in PMA-induced NETs36. Treatment with these inhibitors effectively suppressed caspase-3 activation in both WT and XKR8 KO BMNs (Extended Data Fig. 5n). As a result, PS exposure and NET formation in WT neutrophils were reduced to levels comparable to those observed in XKR8-deficient cells (Extended Data Fig. 5o,p).
Together, these findings indicate that while the upstream signaling pathways responsible for initiating NETs remain intact in XKR8-deficient neutrophils, the activation of XKR8 downstream of caspase-3 is essential for phospholipid scrambling. This lipid remodeling is, in turn, a critical step in NET formation elicited by a broad range of stimuli.
XKR8-mediated extensive phospholipid re-shuffling facilitates NET formation
To uncover the mechanism of XKR8 in regulating NET formation, we generated a knock-in (KI) mouse model carrying a double-point mutation in XKR8 (the substitution of aspartic acid by alanine at positions 351 and 354 in XKR8, referred to as XKR82DA KI) (Supplementary Fig. 2a). This 2DA mutation prevents caspase-3 from cleaving XKR8, thereby abolishing the caspase-3-driven XKR8-mediated PS exposure28. Neutrophils bearing XKR82DA mutations exhibited a marked defect in PS exposure upon PMA stimulation and during apoptosis, closely resembling the phenotype observed in XKR8 KO neutrophils (Fig. 3a,b and Supplementary Fig. 2b,c). Consistent with irreversible activation of XKR8 observed in apoptosis27, caspase-mediated cleavage of XKR8 resulted in sustained PS externalization during NET formation, as demonstrated by sequential staining with distinct fluorochrome-labeled annexin V probes (Supplementary Fig. 2d). Furthermore, XKR82DA neutrophils failed to form NETs in response to diverse stimuli, mirroring the NET defects seen in XKR8-deficient cells (Fig. 3c).
a–c, PS exposure in WT, XKR8 KO, 2DA KI (XKR82DA) BMNs treated with PMA and NET formation treated with PMA (100 nM, 6 h), A23187 (1.25 μM, 4 h), nigericin (5 μM, 4 h) or LPS (10 μg ml−1, 2 h). Representative histogram (a) and quantification (b) of PS exposure in live cells (annexin V+PI−) (n = 3), and quantification of NET formation (c, n is shown on the graphs). d,e, Quantification of NETs in neutrophil-like HL-60 cells: WT, XKR8 KO and XKR8 KO cells reconstituted with full-length mouse XKR8 (KO + XKR8FL) (d), and XKR8 mutants (2DA and S/T-3A) (e) following PMA (100 nM, 6 h) treatment (n = 8). f, Immunoblot analysis of cleavage of XKR8 during PMA-induced NET formation. Mouse XKR8 (XKR8FL) and XKR82DA mutant were GFP-tagged and expressed in XKR8 KO dHL-60 cells. β-Actin was a loading control. g, Quantification of NETs in neutrophil-like HL-60 WT, XKR8 KO and XKR8 KO cells reconstituted with XKR8 mutants D30A and W45A. NETs were induced with PMA (100 nM, 6 h) (n = 8). h,i, PS and PE exposure during NET formation in WT and XKR8 KO BMNs treated with PMA (100 nM, 3 h), detected using annexin V-APC and duramycin-Cy3 staining, respectively. Representative flow cytometry plot (h) and quantification (i) of PS+/PE+ in live cells (PI−) are shown (n = 3). j, Quantification of non-extractable NBD-labeled phospholipids (PLs) translocated by activated XKR8 during PMA-induced NET formation (n = 3). Bovine serum albumin (BSA; 5 mg ml−1) was used to back-exchange the NBD-PLs from the outer layer of the PM. PC, phosphatidylcholine; SM, sphingomyelin. NETs were determined by the area of chromatin DNA (c–e,g). Each dot represents an individual field of view (c–e,g) or a biological replicate (b,i,j). Data are represented as mean ± s.e.m. from one representative experiment of three (a–c,f) and four (d,e,g,h–j) independent experiments. Statistical analysis was performed using a two-way ANOVA (b–e,g,i,j). P values are shown on the graphs.
To further investigate the essential role of XKR8-mediated lipid scrambling in NET formation, we restored the expression of full-length mouse XKR8 in XKR8 KO HL-60 cells (Extended Data Fig. 6a). Reintroduction of XKR8 reinstated PS exposure in XKR8 KO HL-60 cells during apoptosis and PMA-induced NET formation and successfully rescued NET formation (Fig. 3d and Extended Data Fig. 6b,c). Additionally, the S/T-3A mutation37, which disables serine/threonine phosphorylation sites at residues 356, 361 and 375 of XKR8, but not the 2DA mutants, enabled XKR8 KO HL-60 cells to expose PS in response to apoptosis induced by doxorubicin and NET induction by PMA, thereby restored the NET formation (Fig. 3e and Extended Data Fig. 6d–g). Mechanistically, we observed that WT XKR8, but not the 2DA mutant, underwent cleavage upon PMA stimulation in dHL-60 cells, highlighting the importance of caspase-3-dependent XKR8 activation in NET formation (Fig. 3f). Furthermore, D30A and W45A are identified to be loss-of-function and gain-of-function mutations in XKR8-mediated scrambling, respectively38. Ectopic expression of W45A mutant rescue NET formation in XKR8 KO cells, whereas D30A did not (Fig. 3g and Extended Data Fig. 6h–k).
XKR8-mediated lipid scrambling redistributes various phospholipids across the lipid bilayers in apoptosis38. In addition to PS exposure observed during NET formation, we also detected the externalization of phosphatidylethanolamine (PE) using duramycin staining, a specific probe to bind PE, in live neutrophils (Fig. 3h,i). Moreover, by employing fluorescent lipid analogs NBD-phosphatidylcholine (NBD-PC) and NBD-sphingomyelin (NBD-SM), we found that both PC and SM were translocated from outer layer to the inner leaflet of PM, indicating that extensive lipid re-distribution across the PM lipid bilayers occurs during PMA-induced NET formation (Fig. 3j). Furthermore, using Laurdan, a fluorescent dye that detects changes in membrane lipid fluidity, we observed a marked fluorescence shift in XKR8-expressing neutrophils as lipid scrambling was initiated during NET formation (Extended Data Fig. 6l).
Together, our data demonstrate that caspase-3-mediated cleavage of XKR8 triggers lipid scrambling, resulting in widespread phospholipids reorganization across the bilayers of PM during NET formation.
XKR8-mediated lipid scrambling necessitates the activation of mechanosensitive Ca2+ channels for NET formation
Given that XKR8-dependent lipid scrambling induces substantial alterations in PM phospholipids, and Ca2+ signaling is known to be critical for NET formation39, we hypothesized that Ca2+ signal triggered after lipid scrambling might be required for this process. To test this, we induced NET formation in Ca2+-free Hanks’ balanced salt solution (HBSS). Of note, PMA failed to induce NETs in both WT and XKR8-deficient BMNs in the absence of extracellular Ca2+ (Extended Data Fig. 7a). Furthermore, chelation of extracellular Ca2+ using cell-impermeable chelator EGTA or chelation of intracellular Ca2+ by cell-permeable chelator BAPTA-AM at the onset of PMA treatment, completely inhibited NET formation (Fig. 4a and Extended Data Fig. 7b); however, Indo-1 staining revealed that the initial Ca2+ influx triggered within the first 5 min of stimulation, no matter by crosslinking the Fcγ receptor CD16/32, A23187, PMA or LPS, was similar between WT and XKR8 KO BMNs. This indicates that XKR8 does not influence the early wave of Ca2+ influx during NET induction before its activation by caspase-3 (Extended Data Fig. 7c).
a,b, Quantification of PMA-induced NET formation in WT and XKR8-deficient BMNs, treated with or without EGTA from the beginning (1 mM) (n = 10) (a), or treated with EGTA (1 mM) at indicated time points after PMA stimulation (n = 9) (b). c, Evaluation of membrane tension using Flipper-TR by FLIM imaging. Representative images of Flipper-TR and annexin V in PMA-induced NETs (left) and quantification of Flipper-TR lifetime (τ) in PS-exposed (annexin V+) versus non-exposed (annexin V−) cells (right) (n = 50 in WT control, n = 57 in KO control, n = 59 in PMA-WT-annexin V+, n = 62 in PMA-WT-annexin V−, n = 53 in PMA-KO). Red indicates annexin V-AF594; green shows Flipper-TR. Scale bars, 5 μm. d, Schematic diagram showing calcium channels highly expressed on mouse neutrophils along with their respective inhibitors or agonists. e–i, Quantification of NET formation in WT and XKR8-deficient neutrophils treated with inhibitors and/or agonists listed in d. e, PMA-induced NET formation with inhibitors targeting TRPM2 (JNJ-28583113, 10 μM), TRPM4 (9-phenanthrol, 30 μM), TRPM7 (carvacrol, 100 μg ml−1), TRPV2 (SKF96365, 20 μM), TRPV4 (HC067047, 1 μM), Piezo1 (GsMTx-4, 10 μM), and TRPC3 (Pyr3, 5 μM) for 6 h (n = 10). f, PMA-induced NETs in WT and XKR8-deficient neutrophils treated with the agonists targeting TRPM2 (bilirubin, 10 μM), TRPM7 (naltriben (NTB), 50 μM), TRPV2 (cannabidiol (CBD), 10 μM), TRPV2 (probenecid, 1 mM), TRPV4 (GSK1016790A, 100 nM), Piezo1 (Yoda1, 10 μM), and TRPC3 (GSK1702934A, 100 μM) for 6 h (n = 10). g, PMA-induced NETs in WT and XKR8-deficient neutrophils with both inhibitors and agonists against TRPV2 (SKF96365 and CBD), TRPV4 (HC067047 and GSK1016790A), Piezo1 (GsMTx-4 and Yoda1), TRPC3 (Pyr3 and GSK1702934A) for 6 h (n = 9). h,i, As in f, quantification of LPS-induced NETs (h) or C. albicans-induced NETs (i) in WT and XKR8-deficient neutrophils treated with the agonists targeting TRPV2 (CBD, 10 μM), TRPV2 (probenecid, 1 mM), TRPV4 (GSK1016790A, 100 nM), Piezo1 (Yoda1, 10 μM) and TRPC3 (GSK1702934A, 100 μM) for 6 h (n = 10). NETs were measured by the area of decondensed chromatin DNA (a,b,e–i). Each dot represents one individual field of view (a,b,e–i) or a single cell (c). Data are presented as mean ± s.e.m. from one representative experiment of three (c,e–i) and four (a,b) independent experiments. Statistical analysis was performed using a two-way ANOVA (a,b,e–i) and one-way ANOVA (c). P values are shown on the graphs.
Next, we sought to determine the specific stage during NET induction at which Ca2+ chelation could still halt NET progression. To address this, we added EGTA or BAPTA-AM at various time points following PMA stimulation. Notably, chelating Ca2+ up to 2 h after PMA treatment efficiently abolished NET formation in WT neutrophils, whereas adding the chelators at 3 h after stimulation failed to do so (Fig. 4b and Extended Data Fig. 7d). This time point aligns with the peak of XKR8-mediated phospholipid scrambling, suggesting that Ca2+ signal occurring after XKR8 activation is critical for NET formation (Fig. 1a). Indo-1 staining revealed that XKR8 KO BMNs exhibited lower intracellular Ca2+ levels than WT controls at 2 h treated with PMA (Extended Data Fig. 7e). Annexin V+ BMNs displayed significantly higher intracellular Ca2+ levels than annexin V− cells among WT viable cells (gated on PI− populations) (Extended Data Fig. 7f). BAPTA-AM, a faster intracellular Ca2+ chelator compared to EGTA-AM, allowed us to further investigate the timing and localization of this Ca2+ requirement. When BMNs were treated with either BAPTA-AM or EGTA-AM simultaneously with PMA stimulation, BAPTA-AM showed superior inhibition of NET formation, highlighting that the Ca2+ signal elicited in close proximity to XKR8 activation is essential for driving NET progression (Extended Data Fig. 7g).
Our data pinpoint that Ca2+ signal following XKR8-mediated lipid scrambling is critical for NET formation. Thus we intended to figure out how lipid scrambling regulates subsequent Ca2+ signals. Using Flipper-TR, a membrane tension-sensitive fluorescent probe, we observed similar fluorescence lifetime (τ) of Flipper-TR in WT and XKR8 KO BMNs under resting conditions (Fig. 4c); however, among living neutrophils undergoing lipid scrambling (annexin V+DAPI−), WT BMNs exhibited significantly reduced Flipper-TR fluorescence lifetime compared to annexin V− neutrophils (Fig. 4c). These results support that XKR8-dependent lipid scrambling alters PM lipid tension during NET formation.
Given that Ca2+ signaling downstream of lipid scrambling is essential for NETs and that lipid scrambling alters membrane lipid tension, it is rational to postulate that mechanosensitive Ca2+ channels may be involved in XKR8-regulated NET formation. To test this, we compiled a list of candidate mechanosensitive Ca2+ channels based on their high expression in neutrophils and PM localization40. Notably, many candidates belonged to the transient receptor potential (TRP) family of channels, well-recognized for their roles in mechanosensing41 (Fig. 4d). Owing to the limited availability of specific chemical modulators for many of these channels, we conducted a small-scale pharmacological screen to identify the channels responsible for the impaired Ca2+ signaling in XKR8 KO cells during NET formation (Fig. 4d and Extended Data Fig. 8a). Several inhibitors, HC067047 (TRPV4 inhibitor), Pyr3 (TRPC3 inhibitor), SKF96365 (TRPV2 inhibitor), 9-phenanthrol (TRPM4 inhibitor), JNJ-28583113 (TRPM2 inhibitor) and GsMTx-4 (Piezo1 inhibitor) successfully suppressed PMA-induced NET formation in WT BMNs and dHL-60 cells to the levels observed in XKR8-deficient cells (Fig. 4e and Extended Data Fig. 8b). Consistently, these compounds effectively blocked the PMA-induced NET formation in primary human neutrophils, with the exception of SKF96365 (Extended Data Fig. 8c). In contrast, other inhibitors had minimal or inconsistent effects across models (Extended Data Fig. 8a,b). To complement these findings, we tested whether agonists of the candidate channels could restore NET formation in XKR8-deficient cells. Administration of the agonists CBD (TRPV2), GSK1702934A (TRPC3) and Yoda1 (Piezo1) immediately triggered strong, moderate and mild Ca2+ influx, respectively, with comparable responses observed in WT and XKR8 KO BMNs, but probenecid (TRPV2) and GSK1016790A (TRPV4) did not (Extended Data Fig. 8d). Moreover, these mechanosensitive Ca2+ channels agonists restored intracellular Ca2+ level in XKR8 KO BMNs to those observed in WT cells at 2 h following PMA treatment (Extended Data Fig. 8e). Remarkably, in the presence of PMA, these agonists successfully rescued NET formation in both XKR8 KO BMNs and dHL-60 cells (Fig. 4f and Extended Data Fig. 8f). These agonists were not able to induce NETs in the absence of PMA in either mouse or human primary neutrophils, with the sole exception of GSK1702934A, which induced NETs in primary human neutrophils (Extended Data Fig. 8g,h). The rescue effects of these agonists were specifically reversed by their corresponding inhibitors, SKF96365, HC067047, GsMTx-4 and Pyr3, further supporting the involvement of these channels in mediating the Ca2+ signal downstream of XKR8 (Fig. 4g). More broadly, agonists targeting these four channels also overcome the defective NET formation in XKR8 KO neutrophils following stimulation with LPS, nigericin and C. albicans (Fig. 4h,i and Extended Data Fig. 8i). Eventually, genetic deletion of TRPV2, TRPC3, Piezo1 and TRPV4 abrogated the NET formation upon PMA treatment in neutrophil-like HL-60 cells, without affecting PS externalization (Extended Data Fig. 8j,k). Together, these results demonstrate that XKR8-mediated lipid scrambling alters PM tension, which in turn activates the Ca2+ influx via mechanosensitive channels, thereby facilitating NET formation.
XKR8 is critical for neutrophil-mediated acute lung injury and rheumatoid arthritis
Neutrophils have a central role in the pathogenesis of human acute respiratory distress syndrome (ARDS)42. The LPS-induced lung injury in mice closely mimics the pathophysiological features of ARDS43,44 and NETs released by neutrophils are considered key contributors to the acute lung injury (ALI) triggered by LPS45,46. To assess the in vivo role of XKR8 in NET formation, we evaluated the LPS-induced ALI in both WT and XKR8 KO mice. After 24 h of intranasal administration of a proinflammatory dose of LPS (100 ng per mouse), robust neutrophil infiltration was observed in the lungs compared to the PBS group, and comparable numbers of neutrophils were detected in both bronchoalveolar lavage fluid (BALF) and lung homogenates of WT and XKR8 KO mice (Extended Data Fig. 9a,b). Despite similar levels of neutrophil recruitment, LPS triggered significantly more NET formation in WT lungs, as demonstrated by stronger Cit-H3 staining (Fig. 5a,b). In addition, BALF from WT mice contained higher levels of double-stranded DNA (dsDNA) compared to that of XKR8 KO mice (Fig. 5c). Moreover, histological analysis of lung sections demonstrated more severe pulmonary injury in WT mice compared to XKR8 KO mice (Fig. 5d).
a–k, LPS-induced ALI. a, Representative immunofluorescence staining of citrullinated histone H3 (Cit-H3, red) and myeloperoxidase (MPO, green) in lung sections from WT and XKR8 KO mice, 24 h after LPS intranasal challenge. DAPI (blue) marks nuclei. Scale bars, 10 μm. b, Quantification of NETs (Cit-H3 relative area) in lung sections (n = 5 for PBS; n = 12 for LPS). c, Detection of extracellular dsDNA in BALF supernatant from WT and XKR8 KO mice (n = 5 for PBS; n = 12 for LPS). d, Representative pictures of hematoxylin and eosin (H&E) staining (left) and histological scores (right) of pulmonary injury from lung sections in WT and XKR8 KO mice (n = 7). Scale bars, 100 μm. e, Quantification of NET formation in Xkr8fl/fl and Xkr8fl/fl Ly6gcre mice BMNs treated with PMA (100 nM, 6 h), A23187 (1.25 μM, 4 h), LPS (10 μg ml−1, 2 h) or nigericin (5 μM, 4 h), respectively (n = 10). f, Immunofluorescence staining of citrullinated histone H3 (Cit-H3; red) and myeloperoxidase (MPO; green) in lung sections from control (Xkr8fl/fl) and neutrophil-specific XKR8 KO (Xkr8fl/fl Ly6gcre) mice, 24 h after LPS intranasal challenge. DAPI (blue) marks nuclei. Scale bars, 10 μm. g, Quantification of NETs (Cit-H3 relative area) in lung sections (n = 5). h, Detection of extracellular dsDNA in BALF supernatant from Xkr8fl/fl and Xkr8fl/fl Ly6gcre mice (n = 5). i, Representative images of H&E staining (left) and histological scores (right) of pulmonary injury from lung sections in Xkr8fl/fl and Xkr8fl/fl Ly6gcre mice (n = 5). Scale bars, 100 μm. j,k, As in b and c, quantification of NETs (j) and free dsDNA in BALF supernatant (k) in WT, XKR8 KO and XKR82DA KI mice (n = 4). l–n, CAIA. l, Schematic illustration of the CAIA experimental model. ArthritoMab denotes an antibody cocktail of arthritogenic monoclonal antibodies to collagen II. m,n, Clinical scores (m) and paw thickness (n) of WT and XKR8 KO mice during CAIA (n = 4 for WT; n = 5 for KO). Each dot represents one individual mouse (b–d,g–k) or one individual field of view (e). For Cit-H3 quantification, data from at least eight images per mouse were averaged (b,g,j). All experiments were performed at least twice. Data are presented as mean ± s.e.m. Statistical analysis was performed by two-way ANOVA in (b,c,e,g,h,m,n), one-way ANOVA in (j,k) or two-tailed Mann–Whitney U-test in d,i. P values are shown on the graphs.
To further confirm the direct role of XKR8 in NET formation in vivo, we generated neutrophil-specific Xkr8 KO mice by crossing Xkr8 floxed mice (Xkr8fl/fl as the controls) with Ly6gcre KI mice, resulting in neutrophil-specific XKR8 KO mice (Xkr8fl/fl Ly6gcre) (Extended Data Fig. 9c–e). BMNs from these conditional KO mice exhibited the same defects of PS exposure and NET formation observed in germline XKR8 KO mice, as compared to control neutrophils (Fig. 5e and Extended Data Fig. 9f,g). Consistent with the results from germline XKR8 KO mice challenged with LPS, neutrophil-specific XKR8 KO mice displayed significantly reduced NET formation in the lungs, lower levels of extracellular dsDNA in BALF and mitigated lung injury, despite similar levels of neutrophil infiltration following LPS challenge (Fig. 5f–i and Extended Data Fig. 9h,i). Moreover, XKR82DA KI mice also showed impaired NET formation and reduced DNA release into the BALF, mirroring the phenotype seen in XKR8-deficient mice (Fig. 5j,k and Extended Data Fig. 9j,k). Together, these results demonstrate that neutrophil-intrinsic expression and caspase-3-dependent activation of XKR8 are critical for NET formation in vivo during LPS-induced ALI model.
Given the established role of NETs in collagen antibody-induced arthritis (CAIA), a mouse model of rheumatoid arthritis47, we evaluated the contribution of XKR8 in this autoimmune disease mouse model. Notably, XKR8 KO mice exhibited markedly attenuated disease severity compared to WT controls, as evidenced by lower clinical scores and reduced paw thickness (Fig. 5l–n). These results demonstrate that XKR8-mediated NET formation is functionally important in the pathogenesis of autoimmune disease.
XKR8 is critical for neutrophil-mediated antifungal infection
Neutrophils deploy their chromatin to trap and kill microorganisms, serving as a critical antimicrobial defense mechanism. To further investigate the role of XKR8 in neutrophils under more pathophysiological relevant conditions, we assessed its function in response to C. albicans, an opportunistic fungal pathogen known to strongly induce NET formation in vivo48. Airway infection of C. albicans causes pulmonary candidiasis, which disrupts the lung epithelial barrier and leads to extensive pulmonary hemorrhage49. NET formation is considered a key mechanism for preventing C. albicans infection in the lung48. To determine whether XKR8 in neutrophils contributes to the control of C. albicans infection in vivo, we infected WT (Xkr8fl/fl) and neutrophil-specific XKR8 KO mice (Xkr8fl/flLy6gcre) with C. albicans intranasally. Upon challenge with a lethal dose of 3 × 106 colony-forming units (c.f.u.), XKR8 neutrophil-specific KO mice displayed significantly higher mortality compared to controls (Fig. 6a). When infected with a sublethal dose (1 × 106 c.f.u.), both groups recruited similar amounts of neutrophils into the BALF and lungs; however, XKR8 neutrophil-specific KO mice manifested increased susceptibility to fungal infection, as evidenced by higher fungal burdens and more severe lung injury (Fig. 6b–d and Extended Data Fig. 10a,b). Additionally, XKR8 neutrophil-specific KO mice showed exacerbated pulmonary hemorrhage, indicating impaired fungal clearance led to lung vascular damage (Fig. 6e–g and Extended Data Fig. 10c). Meanwhile, these mice also exhibited markedly reduced NET formation upon airway infection of C. albicans compared to WT controls (Fig. 6h,i). Similar to XKR8 KO mice, XKR82DA KI mice were more susceptible to C. albicans infection, as reflected by increased fungal load in the lungs (Fig. 6j and Extended Data Fig. 10d,e). Furthermore, we performed a disseminated C. albicans infection model by tail vein injection (1 × 106 c.f.u.). Neutrophil-specific deletion of XKR8 markedly compromised antifungal immunity, resulting in significantly increased lethality in XKR8 KO mice (Fig. 6k). Following intravenous challenge, XKR8-deficient mice developed severe systemic infection, exhibiting substantially higher fungal burdens in multiple organs, including the liver, lungs, spleen and brain, along with exacerbated tissue injury (Fig. 6l and Extended Data Fig. 10f). Consistently, XKR82DA KI mice phenocopied the KO mice, demonstrating elevated C. albicans loads across these organs in the disseminated infection model (Fig. 6m). Finally, based on our in vitro data showing that activating mechanosensitive Ca2+ channels can restore NET formation in XKR8 KO BMNs (Fig. 4i), we tested whether this strategy could enhance antifungal immunity in vivo. Albeit the TRPV4 agonist GSK1016790A did not directly inhibit fungal viability in vitro (Extended Data Fig. 10g), its administration significantly reduced fungal burden and pulmonary hemorrhage in XKR8 neutrophil-specific KO mice (Fig. 6n and Extended Data Fig. 10h).
a–j, C. albicans-induced ALI. a, Survival curve of Xkr8fl/fl and Xkr8fl/flLy6gcre mice following intranasal infection with C. albicans (3 × 106 c.f.u. per mouse). Number of mice that survived out of total mice is indicated in the brackets. b–j, Sublethal dose of C. albicans infection (1 × 106 c.f.u. per mouse). b, Lung fungal burden (c.f.u.) from Xkr8fl/fl and Xkr8fl/fl Ly6gcre mice, 24 h after infection (n = 9). c,d, Assessment of lung epithelial damage by fibrin deposition. Representative pictures of H&E staining (c) and quantification of fibrin scores (d) in lung sections (n = 9). Scale bar, 100 μm. e, Erythrocytes count in BALF from Xkr8fl/fl (n = 10) and Xkr8fl/fl Ly6gcre mice (n = 11), 24 h after C. albicans challenge. f,g, As in c and d, blood vessel endothelial injury in the lung, with representative H&E staining (f) and quantification of bleeding scores (g) of lung sections in Xkr8fl/fl (n = 8) and Xkr8fl/fl Ly6gcre mice (n = 9) without bronchoalveolar lavage. Scale bar, 100 μm. h,i, Representative immunofluorescence images (h) and quantification of NETs by Cit-H3 staining (i) (n = 9). Scale bars, 10 μm. j, Lung fungal load in WT (n = 10), XKR8 KO (n = 9) and XKR82DA KI mice (n = 9). k–m, Disseminated C. albicans infection via tail vein injection. k, Survival curve of Xkr8fl/fl and Xkr8fl/fl Ly6gcre mice after intravenous administration of C. albicans (1.1 × 106 c.f.u. per mouse). The number of mice that survived out of the total is indicated in the parentheses. l, Fungal burden in the liver, lung, spleen and brain of WT and XKR8 KO mice at 24 h after infection (n = 7). m, As in l, liver, lung, spleen and brain fungal load in WT, XKR8 KO and XKR82DA KI mice (n = 8). n, Lung fungal load in Xkr8fl/fl (n = 7) and Xkr8fl/flLy6gcre (n = 8) mice with or without GSK1016790A intranasal treatment before C. albicans challenge. Pooled data from two independent experiments (a–n). Each dot represents one individual mouse (b,d,e,g,i,j,l–n). For Cit-H3 quantification, data from at least eight images per mouse were averaged (i). All experiments were performed at least twice. Data are presented as mean ± s.e.m. Statistical analysis was performed by log-rank (Mantel–Cox) test (a,k), two-tailed Mann–Whitney U-test (b,d,e,g,i, j,l,m) or two-way ANOVA (n). P values are shown on the graphs.
Overall, our data demonstrate that XKR8 is essential for NET formation in vivo. Loss of XKR8, or disruption of its activation and subsequent lipid scrambling, severely compromised the capacity of neutrophils to combat fungal infection. These findings highlight the caspase-3–XKR8-Ca2+ axis as a promising therapeutic target for conditions such as ARDS.
Discussion
Neutrophils respond to diverse stimuli by forming NETs, a process that culminates in NETosis - a distinct form of neutrophil cell death10. Despite the identification of numerous NET-inducing stimuli, the core mechanisms governing NET formation have remained poorly defined. Our findings reveal that XKR8, a PM phospholipid scramblase activated by caspase-3, is fundamentally required for NET formation. Following ROS generation from oxidative burst, caspase-3 cleaves and activates XKR8, triggering lipid scrambling that facilitates subsequent Ca2+ signaling through mechanosensitive cation channels. Our study identifies XKR8 as a central signaling hub that links ROS production and Ca2+ signaling to integrate a wide range of NET-inducing cues. Furthermore, neutrophil-specific XKR8 deficiency results in markedly impaired antifungal immunity, underscoring its essential role in host defense.
As the most abundant circulating leukocytes in human blood, neutrophils have essential roles in immune response1,2,50,51,52; however, despite the diversity in upstream signaling, neutrophils consistently form NETs in response to a wide range of stimuli, raising the question of whether a shared signaling node exists in this process. In our study, we unexpectedly discovered that various NET-inducing stimuli. All inevitably require XKR8 for NET formation. This convergence suggests that XKR8 serves as a universal signaling hub downstream of diverse stimuli. Once activated, XKR8 mediates lipid scrambling, which in turn triggers Ca2+ influx through mechanosensitive channels, providing the critical second messenger signal to propagate NET formation. It is noteworthy that this wave of Ca2+ signaling coincides with the lipid scrambling, but not the at the early stage of stimulation. Our findings reveal a unifying mechanism in which XKR8 links upstream immune signals to the intracellular machinery driving NETs. Thus, targeting the caspase–3XKR8 axis may offer a broadly applicable therapeutic strategy across a range of NET-associated pathological conditions.
Despite extensive research, the precise cellular and molecular mechanisms underlying NETosis remain incompletely understood. It has been proposed that NETosis incorporates elements of various cell death pathways, including features shared with apoptosis, necroptosis and pyroptosis53,54,55,56. Though discrepancy exists under these conditions, a common requirement of Ca2+ necessitates PAD4 activation through the pore-forming proteins, such as MLKL, GSDMD and GSDME53,54,55,56. We observed that, similar to apoptosis, neutrophils undergoing NETosis exhibit a disruption of PM lipid asymmetry. PS exposure, a marker of altered PM asymmetry, was detected as an early event during NET formation, occurring before PM permeabilization. This suggests that the critical Ca2+ required for PAD4 activation and NETosis may arise from alternative mechanisms, such as lipid scrambling mediated by XKR8.
Despite caspase-3 activation and the hallmark feature of PS exposure mediated by XKR8, commonly associated with early apoptosis, our findings highlight key distinctions between NETosis and classical apoptosis. First, during NETosis, neutrophils exhibit chromatin decondensation rather than the DNA fragmentation characteristic of apoptosis. Second, histone citrullination by PAD4, which requires intracellular Ca2+, is a defining feature of NETosis but is absent in apoptotic cells, at least under our experimental conditions. Third, the kinetics and magnitude of caspase-3 activation in NETs differ significantly from that observed in apoptosis induced by raptinal. Furthermore, other apoptotic caspases, such as caspase-7, caspase-8 and caspase-9, were not activated during NEs formation. Finally, NETosis requires both oxidative burst and PAD4 activation, events are dispensable for apoptosis. Thus, together with previous reports, our data support the conclusion that NETosis constitutes a specialized form of caspase-3-dependent cell death unique to neutrophils. Further investigations into the upstream regulation of caspase-3 activation in the context of NETs would further elucidate the molecular mechanisms governing this process.
Given that diverse extracellular cues can provoke neutrophils to form NETs, our findings suggest that ROS and Ca2+ act as two principal intracellular mediators of this process. Using PMA, a NOX2-dependent NET inducer, we identified XKR8 as a crucial molecular link between ROS production and subsequent Ca2+ signaling. Notably, recent studies have implicated Piezo1, a mechanosensitive Ca2+ channel, in shear stress-induced NET formation57, aligning with our findings that link membrane mechanics to Ca2+ influx. Collectively, our results support the concept that ROS and Ca2+ function as two fundamental pillars in facilitating NET progression, with caspase-3-dependent activation of XKR8 serving as the central integrating node. Considering ROS and Ca2+ are also core signaling modulators in various neutrophils responses, it remains an intriguing question how neutrophils prioritize downstream fates, such as phagocytosis, degranulation or releasing NETs, upon encountering the pathogens.
In the lung, neutrophils have a pivotal role in safeguarding the epithelium from pathogenic invasion. Individuals with ARDS bear cumulated neutrophils in the pulmonary vasculature, contributing to tissue damage and inflammation58. Using a murine model of LPS-induced ALI, we revealed an essential requirement for XKR8 in neutrophil-mediated defense. Similarly, in a pulmonary C. albicans infection model, XKR8 deficiency resulted in impaired antifungal immunity. Of note, treatment with a Ca2+ channel agonist would ameliorate the infection and related disease manifestation in XKR8-deficient mice, highlighting the therapeutic potential of targeting this pathway. Our study positions the caspase-3–XKR8–Ca2+ axis as a unifying mechanism underlying NET formation and provides a compelling therapeutic target for modulating NETs in both infectious and sterile inflammatory diseases.
Methods
Mice
All mice used in this study were of the C57BL/6 genetic background. Experiments were conducted using age- and sex-matched mice between 6–12 weeks of age. The XKR8 germline KO, floxed and 2DA mutant KI mice were generated for this study. To produce the Xkr8 germline KO mice, exons 2 and 3 of the WT Xkr8 allele were deleted using CRISPR–Cas9 system, as illustrated in Extended Data Fig. 1f. Littermate Xkr8+/+ mice were used as WT controls. To generate neutrophil-specific Xkr8 KO mice (Xkr8fl/flLy6gcre), Xkr8 floxed mice (Xkr8fl/fl) were crossed with Ly6gcre KI mice, as depicted in Extended Data Fig. 9c. The Ly6gcre mice were provided by Z. Zeng. The Xkr82DA (p.D351A&D354A) KI mice were generated via CRISPR–Cas9-mediated KI of the 2DA mutant cassette into exon 3 of Xkr8 gene (see Supplementary Fig. 2a for the detailed strategy). TMEM16F germline KO mice were previously described59. Caspase-3 KO mice and Padi4 KO mice were gifts from S. Shi and X. Wang, respectively. Both male and female mice were used in this study. Mice were housed under specific pathogen-free conditions, and all animal experiments were conducted in accordance with protocols approved by the Animal Care and Use Committee of Anhui Medical University (LLSC20230277).
Antibodies and reagents
Anti-citrullinated histone H3 (citrulline R2 + R8 + R17) (ab5103, 1:1,000 dilution; ab281584, 1:1,000 dilution) and anti-neutrophil elastase (ab314916, 1:100 dilution) were purchased from Abcam. Anti-MPO (AF3667) was obtained from R&D systems. Antibodies against caspase-3 (9662), caspase-7 (D6H1, 8438), caspase-8 (D5B2, 8592), caspase-9 (9504), phospho-ERKT202/Y204 (4370) and phospho-PKC (βII Ser660, 9371) were from Cell Signaling Technologies. The GFP-tag polyclonal antibody (50430-2-AP) and β-actin monoclonal antibody (66009-1-lg) were from Proteintech. Flow cytometry and surface marker antibodies, including those against mouse Ly6G (1A8), CD11a (M17/4), CD11b (M1/70), CD45 (30-F11), CD44 (IM7), CD62L (MEL-14), CD101 (Moushi101), CXCR2 (SA044G4), CXCR4 (2B11), TLR4 (SA15-21), annexin V (640941, 640945) and corresponding isotype controls, were obtained from BioLegend. Elastase G-2 (sc-55549) was obtained from Santa Cruz. Activated caspase-3 antibody (570524), CD16 (555406) and CD66b (561650) were from BD Biosciences.
Chemical compounds
To investigate the signaling pathways involved in NET formation, various chemical inducers and inhibitors were employed. Neutrophils were pretreated with inhibitors for 30 min at 37 °C in serum-free RPMI 1640 (Gibco) supplemented with 10 mM HEPES before stimulation with NET inducers. The following chemical compounds were used in the experiments: PMA (P8139), EGTA (E4378), BAPTA-AM (A1076) and TEMPO (426369) were from Sigma-Aldrich. LPS (tlrl-eblps), nigericin (tlrl-nig), monosodium urate (MSU) crystals (tlrl-msu) and zymosan (tlrl-zyn) were from Invivogen. A23187 (B6646), Q-VD-OPh (A1901), Z-VAD-FMK (A1902), apocynin (B3675), DPI (B6326), U0126 (A1337), GW5074 (B1405), EGTA-AM (C4548), doxorubicin (A1832) and staurosporine (STS, A8192) were all from APExBIO Technology. Additional inhibitors, including S62845 (HY-100741), emricasan (HY-10396), 2-APB (B6643), SKF96365 (HY-100001), HC067047 (HY-100208), Pyr3 (HY-108465), JNJ-28583113 (HY-149143), 9-phenanthrol (HY-108457), carvacrol (HY-N0711), GsMTx-4 (HY-P1410), probenecid (HY-B0545), GSK1016790A (HY-19608), GSK1702934A (HY-111098), bilirubin (HY-N0323), Yoda1 (HY-18723) and NTB (HY-101302) were obtained from MedChemExpress. CBD (M9136) was purchased from Abmole, and the compound NA-11 was previously reported36. Enzastaurin (S1055) and SAR7334 (S6634) were obtained from Selleck, and CM-4620 (CSN21369) and GSK2795039 (33777) were sourced from CSNpharm and Cayman Chemical, respectively. CellEvent Caspase-3/7 Green Detection Reagent (C10423) and Laurdan (D250) were from Invitrogen, and Flipper-TR (CY-SC020) was from Cytoskeleton.
Cell culture and differentiation
HL-60 cells (CCL-240), obtained from ATCC, were cultured at 37 °C with 5% CO2 in IMDM (Gibco) supplemented with 25 mM HEPES, 2 mM l-glutamine, 20% heat-inactivated fetal bovine serum (FBS) and 1% penicillin–streptomycin. Cells were routinely tested negative for Mycoplasma contamination. Differentiation of HL-60 cells into neutrophil-like cells was induced by treating the cells with 10 μM ATRA for 6 days. The efficiency of differentiation was regularly assessed by flow cytometry (CD16+CD66b+). Fully differentiated (dHL-60) cells were used in subsequent functional assays.
Generation of XKR8-deficient HL-60 cells
XKR8-deficient HL-60 cells were generated using CRISPR–Cas9 technology. The LentiCRISPR v2 vector (pLenti-V2, 52961) and packaging plasmids pMD2.G (12259) and psPAX2 (12260) were obtained from Addgene. Gene-specific sgRNAs were designed and cloned into LentiCRISPR v2 vector, with at least two distinct sgRNAs targeting the XKR8 gene. The lentiviral vector and packaging plasmids were co-transfected into HEK293T cells using polyethyleneimine (PEI). After 48 h, viral supernatants were collected, filtered through a 0.22-μm filter and supplemented with Polybrene (5 μg ml−1) for transduction of HL-60 cells. Transduced cells were selected using puromycin (1 μg ml−1). In addition, CRISPR–Cas9 tracrRNA (1072532), Cas9-GFP V3 (10008100) and corresponding CRISPR–Cas9 crRNA were purchased from IDT. The Cas9–GFP protein, tracrRNA and gene-specific crRNAs were co-transfected into HL-60 cells by electroporation (Neon NxT Electroporation System, Thermo Fisher Scientific). Positive cells were sorted by GFP+ (BD AriaⅢ). KO clones were further screened by PCR, and indel mutations in selected clones were verified by genomic DNA sequencing. At least three independent clones were pooled for subsequent experiments. Oligonucleotides used in this study are listed in Supplementary Table 2.
Overexpression of XKR8 and its mutants in HL-60 KO cell lines were achieved using lentivirus-based infection. The cDNA of Xkr8 and its 2DA, S/T-3A, D30A and W45A mutants were cloned into pLVX-IRES-ZsGreen1 (Clontech, Cat. 632187). Transfection and infection processes were performed as previously described. ZsGreen+ cells were sorted for further experiments.
Culture of bacteria and fungi
L. monocytogenes (10403s) was cultured overnight at 37 °C as previously described59. S. aureus (MW2) was cultured overnight at 37 °C in Brain Heart Infusion broth supplemented with erythromycin (10 μg ml−1). To prepare bacteria at the logarithmic phase, 100 μl of the overnight culture was added to 5 ml of fresh medium and incubated for 2 h at 37 °C with agitation. Bacterial cultures were collected when the optical density (OD) at 600 nm reached 0.4, washed with PBS and prepared for subsequent experiments. E. coli (CFT073) was cultured overnight at 37 °C in Luria-Bertani (LB) medium containing kanamycin (50 μg ml−1). For experiments, 100 μl bacterial culture was added to 2 ml of fresh LB medium containing with 1 μM IPTG to boost for 2 h at 37 °C. The bacterial concentration was determined using the formula: 1 OD600nm = 5 × 108 c.f.u. per ml. C. albicans (SC5314) was cultured overnight at 37 °C in yeast extract peptone dextrose (YPD) medium. To boost fungal growth, 100 μl of the overnight fungal culture was transferred to 5 ml of fresh medium and incubated for 2 h at 37 °C. Fungal cells were then collected and counted using a hemocytometer.
Isolation of neutrophils from mouse bone marrow
Neutrophils were isolated from mouse bone marrow using gradient centrifugation with Percoll (17089109-1, GE Healthcare). Bone marrow cells were collected in HBSS supplemented with 0.5% FBS and 20 mM HEPES, without Ca2+ and Mg2+. To remove red blood cells, the pellet was resuspended in 6 ml of 0.2% NaCl for 45 s, followed by the addition of 14 ml of 1.2% NaCl to restore isotonicity. After centrifugation at 1,000g for 30 min at room temperature, the neutrophil-enriched cell pellet, appearing as a cloudy layer at the bottom of the tube, was collected and washed with HBSS. The purity of the neutrophil population was typically greater than 90%, as determined by morphological examination with Giemsa staining and flow cytometry, based on CD11b+ (1:500, clone M1/70) and Ly6G+ (1:500, clone 1A8) surface markers.
Isolation of neutrophils from human peripheral blood
Peripheral blood from healthy donors were obtained from the Department of Clinical Laboratory, the First Affiliated Hospital of Anhui Medical University, with the approval of the Institutional Review Board for Clinical Research of The First Affiliated Hospital of Anhui Medical University. For the human blood samples used, the written informed consent was given to donors indicating that the blood left after the health examination was going to be used for research. Granulocytes were isolated using density gradient centrifugation with Percoll. To prepare the gradient, 2.5 ml of 75% Percoll was added to the bottom of a 15-ml centrifuge tube, followed by 2.5 ml of 62% Percoll layered on top. Fresh blood (1–3 ml) was diluted with serum-free RPMI 1640 to a total volume of 5 ml and carefully overlaid onto the 62% Percoll layer. The gradient was centrifuged at 200g for 25 min, followed by a second centrifugation at 400g for 15 min. Neutrophils were collected from the interface between the 62% and 75% Percoll layers.
To assess NET formation and PS exposure, isolated neutrophils were treated with 100 nM PMA. Neutrophils were seeded either onto glass slides for immunofluorescence staining or into 96-well plates for flow cytometry. At the designated time points, cells on glass slides were fixed with 4% paraformaldehyde (PFA) for immunofluorescence staining. Alternatively, cells were collected, washed with PBS and stained with annexin V-APC along with either propidium iodide (1 μg ml−1) or DAPI (40 nM) for flow cytometry analysis.
NET induction and quantification
Freshly isolated neutrophils were stimulated immediately after purification with chemical reagents or microorganisms in serum-free RPMI 1640 medium supplemented with 10 mM HEPES. For chemical induction of NETs, neutrophils were treated with PMA (100 nM) for 6 h, LPS (10 μg ml−1) for 2 h or A23187 (1.25 μM), nigericin (5 μM), zymosan (25 μg ml−1) and MSU (100 μg ml−1) for 4 h in a cell culture incubator. For microbial induction, neutrophils (1.5 × 105 cells per well) were co-cultured with E. coli, L. monocytogenes, S. aureus or C. albicans at a MOI of 10 for 1.5 h. After stimulation, cells were fixed in 4% PFA for 10 min and permeabilized with 0.1% Triton X-100 in PBS for 20 min at room temperature. Fixed cells were blocked with 1% BSA and 10% goat serum or donkey serum for 1 h at room temperature before immunofluorescence staining. Neutrophils undergoing NET formation were labeled using antibodies against citrullinated histone H3 (Cit-H3, 1:1,000, ab5103, Abcam), MPO (1:1,000, AF3667, R&D system) or elastase (1:100, ab314916, Abcam), diluted in 1% BSA in PBS. NETs were visualized by mounting cells with DAPI and imaging using confocal microscopy (Olympus FV3000 and Nikon Ti2-E).
NETs were defined based on the DNA area observed in the fluorescence images. Semi-automated image analysis of DAPI-stained neutrophils was performed using Fiji software (v.2.14/1.54f), and calculated using MATLAB-based customer-written scripts ‘gating’ (Supplementary Materials 1), as outlined in Extended Data Fig. 1a. Image stacks were processed and converted in eight-bit grayscale. To quantify neutrophil numbers and measure the DNA area and fluorescence intensity of each cell, DAPI images were thresholded using ‘RenyiEntropy’ algorithm, followed by conversion to a mask and watershed segmentation to separate touching particles. NETs were recognized as structures with a DNA area greater than 80 μm2 in BMNs or 100 μm2 in dHL-60 cells. The percentage of NET formation was calculated by dividing the number of NETs by the total number of neutrophils in the field of view.
ROS measurements
ROS production was assessed using a luminol- and isoluminol-dependent chemiluminescence assay to measure intracellular and extracellular ROS, respectively, as previously described60. For intracellular ROS measurement following stimulation with PMA (100 nM), luminol (50 μM, A8515, Sigma-Aldrich) was added to the cell suspension along with membrane-impermeable superoxide dismutase (SOD; 50 U, S8160, Sigma-Aldrich) and catalase (2,000 U, C9322, Sigma-Aldrich) to eliminate extracellularly released O2− and H2O2. Chemiluminescence was recorded at 1-min intervals for 40 min at 37 °C using a BioTek Synergy HT microplate reader.
Extracellular ROS production was assessed by stimulating neutrophils with PMA in the presence of isoluminol (50 μM, A8264, Sigma-Aldrich) and exogenous horseradish peroxidase (HRP; 8 U, P8375, Sigma-Aldrich). The chemiluminescent signal was measured under the same conditions as described for intracellular ROS.
In addition, intracellular ROS levels were quantified using the fluorogenic probe dihydrorhodamine 123 (DHR123; 10 μM, C5313, APExBIO). Neutrophils were incubated with DHR123 for 30 min at 37 °C, followed by stimulation with PMA. The resulting fluorescence intensity was measured using flow cytometry, providing a quantitative assessment of intracellular ROS production.
To investigate the effects of ROS inhibitors or scavengers on NET formation, neutrophils were pretreated with NA-11 (1 μM), apocynin (30 μM), DPI (10 μM) or a dimethylsulfoxide (DMSO) vehicle control for 30 min before PMA (100 nM) stimulation for 6 h.
Evaluation of caspase-3 activation
Caspase-3 activation during the early stages of PMA-induced NET formation was evaluated using the CellEvent Caspase-3/7 Green Detection Reagent (C10423, Invitrogen) and an active caspase-3-specific antibody (570524, BD Biosciences). For the CellEvent assay, neutrophils were stained with CellEvent (1 μM) and subsequently stimulated with PMA (100 nM) in RPMI 1640 medium. Fluorescent signals were measured at defined time points by flow cytometry (LSR Fortessa, BD). Active caspase-3 was further detected using an antibody that specifically recognizes the cleaved form of caspase-3. Neutrophils undergoing NET formation were stained with Ly6G-AF488 (BioLegend, 127626) and Fixable Viability Dye eFluor 780 (eBioscience, 65-0865-18) for 15 min at room temperature to distinguish viable cells. Cells were then fixed with BD Cytofix Fixation Buffer (554655) and permeabilized with BD Perm/Wash buffer (554723). Subsequently, cells were stained with a rabbit anti-active caspase-3 antibody (1:1,000 dilution), followed by secondary staining with either goat anti-rabbit PE or F(ab’)2-goat anti-rabbit AF594 (1:1,000 dilution, A13203, Invitrogen). Stained samples were analyzed using a LSR Fortessa flow cytometer (BD) or sorted on an Aria III cell sorter (BD). Only eFluor 780-negative cells were sorted onto adhesive slides. These slides were then examined using a confocal microscope (Nikon Ti2-E) to visualize localization of activated caspase-3 during NET formation.
To investigate the role of caspase-3 on NET formation, mouse neutrophils were pretreated with caspase-3 inhibitors, including Q-VD (10 μM), Z-VAD (30 μM), emricasan (10 μM) or a DMSO vehicle control for 30 min, followed by PMA stimulation (100 nM) for 6 h at 37 °C. Human primary neutrophils were treated in the same manner, except that PMA stimulation was performed at 5 nM for 4 h. The impact of these inhibitors on NET formation was then assessed using confocal microscopy and flow cytometry.
Measurements of lipid scrambling
PS exposure was measured using annexin V staining. Following NET or apoptosis induction, neutrophils were pelleted and resuspended in 1× annexin V binding buffer (eBioscience), containing annexin V (5 ng ml−1) and either propidium iodide (100 ng ml−1, P4170, Sigma) or DAPI (40 nM, 422801, BioLegend). The samples were then analyzed using either a LSR Fortessa or a Symphony A5 flow cytometer (BD). To monitor lipid scrambling from the outer to the inner leaflet of PM, cells were stained with NBD-labeled phospholipid analogs, including NBD-phosphatidylcholine (NBD-PC; 810132P, Sigma) and NBD-sphingomyelin (NBD-SM; 810218 P, Sigma), as described previously38. After staining, an equal volume of cell suspension and prechilled RPMI1640 containing 5 mg ml−1 BSA was mixed to extract any unincorporated fluorescent lipid analogs. DAPI was added to exclude dead cells from the analysis. Additionally, PE exposure during lipid scrambling was evaluated using the PE-binding probe duramycin (Molecular Targeting Technologies). The stained samples were analyzed by flow cytometry on a LSR Fortessa flow cytometer (BD). Gating strategies are shown in Supplementary Fig. 3. Data from all experiments were processed and analyzed using FlowJo software v.10.
Live-cell imaging assay
To monitor chromatin area changes during NET formation, mouse bone-marrow-derived neutrophils were plated on a 35-mm glass-bottom imaging dish. After 30 min of incubation, attached cells were treated with 200 μl of RPMI 1640 with HEPES containing 20 μg ml−1 Hoechst 33342 (2031173, Invitrogen), 100 nM SYTOX Green (S7020, Invitrogen) and 100 nM PMA. For image analysis, regions of interest (ROIs) corresponding to imaged cells and Hoechst 33342-stained nuclei in each frame were automatically segmented by Cellpose algorithm using pretrained and custom-trained model ‘nucleus’. The nuclear area of each cell was extracted from the segmented ROIs and calculated using MATLAB-based custom-written scripts ‘NETs’ (Supplementary Materials 1). Time-lapse imaging data were analyzed using ImageJ, Cellpose v.2.2.3 and MATLAB (2023a).
Fluorescence and lifetime imaging
Fluorescence and lifetime imaging were performed on a laser confocal microscope (Leica TCS SP8X). For fluorescence lifetime imaging of Flipper-TR (CY-SC020, Cytoskeleton) in neutrophils, cells were first stained with Hoechst 33342 (20 μg ml−1), seeded in a four-chamber glass-bottom dish and treated with PMA (100 nM) at 37 °C in a 5% CO2 incubator for 2 h. Neutrophils were subsequently stained with Flipper-TR (1 μM), DAPI (40 nM) and Alexa Fluor 594-conjugated annexin V (A13203, Invitrogen) at 37 °C for 15 min before imaging. Lifetime imaging was conducted using Leica’s 480 nm pulsed laser for excitation, with photon collection in the 582–670 nm range. Only DAPI-negative cells were included in the subsequent analysis. For lifetime imaging analysis, the fluorescence lifetime (τ) of the cell membrane was measured, with analysis focused on the PM. Cells were categorized as annexin V+ or annexin V−, and the mean fluorescence lifetime was calculated using Leica software (Leica Application Suite X).
Calcium influx and cytosolic Ca2+ level detection
Calcium influx was detected using Indo-1 (65-0856-39, eBioscience). Calcium influx by Indo-1 was reported previously59. In brief, neutrophils were collected and resuspended in calcium-free HBSS containing 1 μM Indo-1, then incubated at 37 °C for 30 min. The cells were subsequently washed twice with calcium-free HBSS and resuspended in HBSS supplemented with 1 mM CaCl2 and MgCl2. Neutrophils were stimulated with LPS (10 μg ml−1), PMA (100 nM), A23187 (1.25 μM) or crosslinking CD16 (2.4G2, 553142, BD) with a secondary goat anti-rat IgG (112-005-062, Jackson ImmunoResearch). For calcium channel agonists experiments, WT and XKR8 KO neutrophils were loaded with Indo-1 and treated with probenecid (1 mM), GSK1016790A (100 nM), GSK1702934A (100 μM), Yoda1 (10 μM) or CBD (5 μM). Calcium influx was measured using flow cytometry for 5 min on a BD FACSymphony A5 flow cytometer, and calcium flux was quantified as the ratio of Indo-1 emission at 400 nm (bound Ca2+) to 475 nm (free Ca2+).
To evaluate intracellular Ca2+ levels during NET formation, 2 × 105 neutrophils labeled with Indo-1 were seeded in 96-well plates and induced by PMA (100 nM) in the presence or absence of agonists for 2 h at 37 °C in RPMI 1640. Subsequently cells were resuspended in 1× binding buffer containing annexin V (5 ng ml−1) and propidium iodide (100 ng ml−1) and analyzed on a BD Symphony A5. Neutrophils were gated into annexin V+ or annexin V− populations, and intracellular Ca2+ levels was determined from the ratio of bound to free Ca2+ Indo-1.
Phagocytosis and degranulation assay
Neutrophils were seeded into 96-well plates, and GFP-expressing E. coli or FITC-labeled latex beads (L4530, Sigma-Aldrich) were opsonized through preincubation in RPMI 1640 medium containing 10% mouse serum at 37 °C for 30 min. For the phagocytosis assay, 1 × 105 neutrophils per well were co-cultured with opsonized E. coli or beads at an MOI of 10 for 30 min in a cell culture incubator. After incubation, neutrophils were washed and stained with antibodies targeting CD11b (1:500 dilution, 101224, BioLegend) and Ly6G (1:500 dilution, 127608, BioLegend). Phagocytosis was quantified by flow cytometry (BD LSR Fortessa). For the degranulation assay, neutrophils were stained with anti-CD63 (1:500 dilution, 143906, BioLegend) following co-culture with bacteria. The mean fluorescence intensity of CD63 was measured as an indicator of neutrophil degranulation.
Mouse serum cytokine profiling
Whole blood samples from naive mice were collected and centrifuged at 3,000g for 20 min to obtain serum. Cytokine levels were quantified using a Luminex multiplex immunoassay (LX-MultiDTM-23, Bio-Rad).
Acute lung injury mouse models
ALI was induced in 6–8-week-old mice via intranasal administration of LPS (100 ng in 50 μl PBS per mouse). Mice receiving an equal volume of PBS served as unchallenged control group. At 24 h after treatment, BALF and lung tissue samples were collected. The acellular fraction of BALF was obtained through double centrifugation, and the supernatant was used for further analysis. dsDNA levels in the BALF were quantified using the dsDNA HS Assay kit (Q32851, Invitrogen).
For the C. albicans-induced ALI model, anesthetized mice were challenged with 1 × 106 or 3 × 106 (for survival monitoring) C. albicans in 50 μl PBS via the intranasal route. BALF was collected 24 h after infection and lung tissues were isolated and homogenized. Serial dilutions of the homogenates were plated on YPD agar plates and incubated at 37 °C for 24 h to quantify fungal burden. To evaluate the effect of GSK1016790A, Xkr8fl/flLy6gcre mice were anesthetized and then treated intranasally with 50 μl of 10 μM GSK1016790A before C. albicans infection.
For C. albicans systemic infection, mice were intravenously injected via the tail vein with 1 × 106 C. albicans (or 1.1 × 106 for survival studies) in 200 μl PBS. At 24 h after infection, the lungs, brain, liver and spleen were collected and homogenized. Homogenates were plated on YPD agar for 24 h to determine fungal burden. Portions of each organ were fixed in PFA for subsequent histopathological analysis.
To detect NETs in lung tissues, frozen and fixed lung sections (5-μm thick) were prepared for immunofluorescence staining. Sections were permeabilized in 0.5% Triton X-100 for 20 min and blocked in PBS containing 2% BSA and 3% of donkey serum (D9663, Sigma-Aldrich) for 1 h at RT. Samples were incubated overnight at 4 °C with anti-citrullinated histone H3 (1:1,000) and anti-MPO (1:1,000), followed by incubation with secondary antibodies (1 h at room temperature in the dark): goat anti-rabbit IgG-A488 (1:1,000, A11008, Invitrogen) and donkey anti-goat IgG-A647 (1:1,000, A32849, Invitrogen). Nuclei were counterstained with DAPI. Images were acquired using a Nikon confocal microscope equipped with a ×60 oil objective lens. For quantification of NETs, at least eight random fields per sample were analyzed. Image stacks were processed and converted to eight-bit grayscale. Each channel of images was thresholded using the ‘Default’ method in ImageJ, followed by conversion to masks, and the total area of Cit-H3-positive signal was measured. NETs formation was expressed as the ratio of Cit-H3 area to DAPI area in the field of view.
Collagen antibody-induced arthritis mouse model
To assess the role of XKR8 in autoimmune disease, a CAIA model was established using 8–9-week-old WT and XKR8 KO mice maintained under specific pathogen-free conditions. On day 0, mice received an intraperitoneal (i.p.) injection of 2.5 mg per mouse of anti-type II collagen 5-clone monoclonal antibody cocktail (Chondrex). On day 3 after antibody injection, to boost the inflammatory response, mice were administered an i.p. injection of LPS (50 μg in 100 μl PBS per mouse; Chondrex). Arthritis severity was monitored daily from day 3 to day 14 using a semiquantitative clinical scoring system (0–4 points per paw; maximum total score 16) based on the following criteria: 0, no visible swelling, erythema, or joint deformity; 1, localized swelling or erythema confined to the knuckle joints; 2, mild swelling extending to the ankle; 3, severe swelling involving the entire paw (digits to ankle); and 4, paw rigidity, joint deformation or impaired weightbearing. Hind paw thickness was measured daily with a digital vernier caliper.
Histological analyses of tissue damage
For histological analyses, the left lung lobes were removed and fixed in 4% neutral formaldehyde and embedded in paraffin for tissue sectioning. The embedded waxes were cut into 10-μm slices and flattened on glass slides, followed by rehydration and H&E staining. Evaluation of lung damage for LPS ALI model was performed using a reliable scoring system as previously reported44. To assess the severity of lung damage, five randomly chosen fields were evaluated according to five independent variables: neutrophils in the alveolar space, neutrophils in the interstitial space, hyaline membranes, proteinaceous debris filling airspaces and alveolar septal thickening. Each item was scored 0, 1 or 2 according to the severity, with differential weighting according to the relevance of these variables. The sum of the weighted variables was averaged according to the number of determined fields, yielding a final score between 0 and 10. Scoring of tissue damage for C. albicans infection was performed as previously described48, ten random images of each section were analyzed by fibrin deposition and bleeding in the tissue and scored 0, 1, 2 or 3 according to the severity. The sum of the variables was averaged according to the number of determined fields.
Immunoblot
Cells were lysed in RIPA buffer containing protease and phosphatase inhibitor cocktails (R0278, Sigma-Aldrich). Protein concentrations were determined using a BCA assay (71285-M, Sigma-Aldrich). Lysates were denatured at room temperature or by boiling for 10 min in 1× sample buffer. To assess phosphorylation of ERK and PKC, caspases or XKR8 (XKR8-GFP) activation, 30 μg of total protein per sample was resolved by SDS–PAGE and transferred onto PVDF membranes. Membranes were incubated overnight at 4 °C with the following primary antibodies: phospho-ERKT202/Y204 (1:1,000), phospho-PKC (1:1,000), caspase-3 (1:1,000), caspase-7 (1:1,000), caspase-8 (1:1,000), caspase-9 (1:1,000) and GFP-tag (1:2,000). After primary antibodies incubation, membranes were probed with HRP-conjugated secondary antibodies for 1 h at room temperature. Protein bands were visualized using an ECL substrate (34580, Thermo Scientific) and detected using the ChemiDoc XRS+ Imaging System (Bio-Rad).
Statistical analysis
For comparisons involving more than two groups, one-way ANOVA or two-way ANOVA with Tukey’s multiple-comparisons test was used. All statistical analyses were performed using GraphPad Prism (v.10). Each experiment was independently repeated at least three times, with detailed statistical information provided in the corresponding figure legends. n denotes the number of animals used, technical replicates or quantified images, as specified in the legends. Data are expressed as mean ± s.e.m. P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data are deposited in BioProject (accession PRJNA1415834). All other data are available in the article and its Supplementary files. Source data are provided with this paper.
Code availability
Codes for the imaging analysis are available in the Supplementary files.
References
Amulic, B., Cazalet, C., Hayes, G. L., Metzler, K. D. & Zychlinsky, A. Neutrophil function: from mechanisms to disease. Annu. Rev. Immunol. 30, 459–489 (2012).
Burn, G. L., Foti, A., Marsman, G., Patel, D. F. & Zychlinsky, A. The neutrophil. Immunity 54, 1377–1391 (2021).
Mantovani, A., Cassatella, M. A., Costantini, C. & Jaillon, S. Neutrophils in the activation and regulation of innate and adaptive immunity. Nat. Rev. Immunol. 11, 519–531 (2011).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Jaillon, S. et al. Neutrophil diversity and plasticity in tumour progression and therapy. Nat. Rev. Cancer 20, 485–503 (2020).
Jorch, S. K. & Kubes, P. An emerging role for neutrophil extracellular traps in noninfectious disease. Nat. Med. 23, 279–287 (2017).
Munoz, L. E. et al. Neutrophil extracellular traps initiate gallstone formation. Immunity 51, 443–450 e444 (2019).
Nemeth, T., Sperandio, M. & Mocsai, A. Neutrophils as emerging therapeutic targets. Nat. Rev. Drug Discov. 19, 253–275 (2020).
Brinkmann, V. et al. Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535 (2004).
Fuchs, T. A. et al. Novel cell death program leads to neutrophil extracellular traps. J. Cell Biol. 176, 231–241 (2007).
Daniel, C. et al. Extracellular DNA traps in inflammation, injury and healing. Nat. Rev. Nephrol. 15, 559–575 (2019).
Papayannopoulos, V., Metzler, K. D., Hakkim, A. & Zychlinsky, A. Neutrophil elastase and myeloperoxidase regulate the formation of neutrophil extracellular traps. J. Cell Biol. 191, 677–691 (2010).
Sollberger, G., Tilley, D. O. & Zychlinsky, A. Neutrophil extracellular traps: the biology of chromatin externalization. Dev. Cell 44, 542–553 (2018).
Honda, M. & Kubes, P. Neutrophils and neutrophil extracellular traps in the liver and gastrointestinal system. Nat. Rev. Gastroenterol. Hepatol. 15, 206–221 (2018).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147 (2018).
Winterbourn, C. C., Kettle, A. J. & Hampton, M. B. Reactive oxygen species and neutrophil function. Annu. Rev. Biochem. 85, 765–792 (2016).
Remijsen, Q. et al. Dying for a cause: NETosis, mechanisms behind an antimicrobial cell death modality. Cell Death Differ. 18, 581–588 (2011).
Adrover, J. M., McDowell, S. A. C., He, X. Y., Quail, D. F. & Egeblad, M. NETworking with cancer: The bidirectional interplay between cancer and neutrophil extracellular traps. Cancer Cell 41, 505–526 (2023).
Binet, F. et al. Neutrophil extracellular traps target senescent vasculature for tissue remodeling in retinopathy. Science 369, eaay5356 (2020).
Gungabeesoon, J. et al. A neutrophil response linked to tumor control in immunotherapy. Cell 186, 1448–1464 e1420 (2023).
He, X. Y., Ng, D. & Egeblad, M. Caught in a web: emerging roles of neutrophil extracellular traps in cancer. Annu. Rev. Canc. Biol. 6, 223–243 (2022).
Hirschhorn, D. et al. T cell immunotherapies engage neutrophils to eliminate tumor antigen escape variants. Cell 186, 1432–1447 e1417 (2023).
Wigerblad, G. & Kaplan, M. J. NETs spread ever wider in rheumatic diseases. Nat. Rev. Rheumatol. 16, 73–74 (2020).
Kenny, E. F. et al. Diverse stimuli engage different neutrophil extracellular trap pathways. eLife 6, e24437 (2017).
Poli, V. & Zanoni, I. Neutrophil intrinsic and extrinsic regulation of NETosis in health and disease. Trends Microbiol. 31, 280–293 (2023).
Thiam, H. R., Wong, S. L., Wagner, D. D. & Waterman, C. M. Cellular mechanisms of NETosis. Annu. Rev. Cell Dev. Biol. 36, 191–218 (2020).
Nagata, S. Apoptosis and clearance of apoptotic cells. Annu. Rev. Immunol. 36, 489–517 (2018).
Suzuki, J., Denning, D. P., Imanishi, E., Horvitz, H. R. & Nagata, S. Xk-related protein 8 and CED-8 promote phosphatidylserine exposure in apoptotic cells. Science 341, 403–406 (2013).
Kawano, M. & Nagata, S. Lupus-like autoimmune disease caused by a lack of Xkr8, a caspase-dependent phospholipid scramblase. Proc. Natl Acad. Sci. USA 115, 2132–2137 (2018).
Bratton, D. L. & Henson, P. M. Neutrophil clearance: when the party is over, clean-up begins. Trends Immunol. 32, 350–357 (2011).
Casanova-Acebes, M. et al. Rhythmic modulation of the hematopoietic niche through neutrophil clearance. Cell 153, 1025–1035 (2013).
Summers, C. et al. Neutrophil kinetics in health and disease. Trends Immunol. 31, 318–324 (2010).
Thiam, H. R. et al. NETosis proceeds by cytoskeleton and endomembrane disassembly and PAD4-mediated chromatin decondensation and nuclear envelope rupture. Proc. Natl Acad. Sci. USA 117, 7326–7337 (2020).
Arita, K. et al. Structural basis for Ca(2+)-induced activation of human PAD4. Nat. Struct. Mol. Biol. 11, 777–783 (2004).
Hakkim, A. et al. Activation of the Raf-MEK-ERK pathway is required for neutrophil extracellular trap formation. Nat. Chem. Biol. 7, 75–77 (2011).
Amara, N. et al. Selective activation of PFKL suppresses the phagocytic oxidative burst. Cell 184, 4480–4494 e4415 (2021).
Sakuragi, T., Kosako, H. & Nagata, S. Phosphorylation-mediated activation of mouse Xkr8 scramblase for phosphatidylserine exposure. Proc. Natl Acad. Sci. USA 116, 2907–2912 (2019).
Sakuragi, T. et al. The tertiary structure of the human Xkr8-Basigin complex that scrambles phospholipids at plasma membranes. Nat. Struct. Mol. Biol. 28, 825–834 (2021).
Douda, D. N., Khan, M. A., Grasemann, H. & Palaniyar, N. SK3 channel and mitochondrial ROS mediate NADPH oxidase-independent NETosis induced by calcium influx. Proc. Natl Acad. Sci. USA 112, 2817–2822 (2015).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Koivisto, A. P., Belvisi, M. G., Gaudet, R. & Szallasi, A. Advances in TRP channel drug discovery: from target validation to clinical studies. Nat. Rev. Drug Discov. 21, 41–59 (2022).
Williams, A. E. & Chambers, R. C. The mercurial nature of neutrophils: still an enigma in ARDS?. Am. J. Physiol. Lung Cell. Mol. Physiol. 306, L217–L230 (2014).
Fan, E., Brodie, D. & Slutsky, A. S. Acute respiratory distress syndrome: advances in diagnosis and treatment. JAMA 319, 698–710 (2018).
Matute-Bello, G., Frevert, C. W. & Martin, T. R. Animal models of acute lung injury. Am. J. Physiol. Lung Cell. Mol. Physiol. 295, L379–L399 (2008).
Liu, S. et al. Neutrophil extracellular traps are indirectly triggered by lipopolysaccharide and contribute to acute lung injury. Sci. Rep. 6, 37252 (2016).
Radermecker, C. et al. Locally instructed CXCR4(hi) neutrophils trigger environment-driven allergic asthma through the release of neutrophil extracellular traps. Nat. Immunol. 20, 1444–1455 (2019).
Malamud, M. et al. Recognition and control of neutrophil extracellular trap formation by MICL. Nature 633, 442–450 (2024).
Branzk, N. et al. Neutrophils sense microbe size and selectively release neutrophil extracellular traps in response to large pathogens. Nat. Immunol. 15, 1017–1025 (2014).
He, Y., Liu, J., Chen, Y., Yan, L. & Wu, J. Neutrophil extracellular traps in Candida albicans infection. Front. Immunol. 13, 913028 (2022).
Aroca-Crevillen, A., Vicanolo, T., Ovadia, S. & Hidalgo, A. Neutrophils in physiology and pathology. Annu. Rev. Pathol. 19, 227–259 (2024).
Ley, K. et al. Neutrophils: new insights and open questions. Sci. Immunol. 3, eaat4579 (2018).
Soehnlein, O., Steffens, S., Hidalgo, A. & Weber, C. Neutrophils as protagonists and targets in chronic inflammation. Nat. Rev. Immunol. 17, 248–261 (2017).
Chen, K. W. et al. Noncanonical inflammasome signaling elicits gasdermin D-dependent neutrophil extracellular traps. Sci. Immunol. 3, eaar6676 (2018).
D’Cruz, A. A. et al. The pseudokinase MLKL activates PAD4-dependent NET formation in necroptotic neutrophils. Sci. Signal 11, eaao1716 (2018).
Sollberger, G. et al. Gasdermin D plays a vital role in the generation of neutrophil extracellular traps. Sci. Immunol. 3, eaar6689 (2018).
Zhu, Y. P. et al. NET formation is a default epigenetic program controlled by PAD4 in apoptotic neutrophils. Sci. Adv. 9, eadj1397 (2023).
Baratchi, S. et al. Piezo1 expression in neutrophils regulates shear-induced NETosis. Nat. Commun. 15, 7023 (2024).
Summers, C. et al. Pulmonary retention of primed neutrophils: a novel protective host response, which is impaired in the acute respiratory distress syndrome. Thorax 69, 623–629 (2014).
Wu, N. et al. Critical role of lipid scramblase TMEM16F in phosphatidylserine exposure and repair of plasma membrane after pore formation. Cell Rep. 30, 1129–1140 e1125 (2020).
Dahlgren, C. & Karlsson, A. Respiratory burst in human neutrophils. J. Immunol. Methods 232, 3–14 (1999).
Acknowledgements
We thank W. Liu for his advice on the project. D. Kuang helped with the histological analysis. This work was supported by grants from the National Key Research and Development Program of China to N. Wu (2022YFF0710600) and the National Natural Science Foundation of China to N. Wu (32070911).
Author information
Authors and Affiliations
Contributions
W.L., J.P., L.D., W.H., Z.L. and N.W. conceived the project, designed the experiments and wrote the manuscript. W.L., J.P. and L.D. conducted the experiments and data analysis. K.W., J.X. and N.P. conducted molecular cloning, viral infection and CRISPR–Cas9 KO cells. M.Y., H.T., Z.J.L., W.W., T.L. and Z.D. helped with the in vitro and in vivo experiments. Y.H. wrote the scripts for the imaging analysis. X.J. and W.H. helped with the interpretation of results. N.W. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Jatin Vyas, Borko Amulic, Gordon Brown, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Paula Jauregui, in collaboration with the Nature Immunology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PS externalization during NETs formation and the generation of XKR8-deficient mice.
a, Quantification of NETs based on DAPI fluorescence intensity and area. Cells with nuclear area greater than 80 µm2 were classified as NETs. b, PS exposure and NETs formation in human peripheral blood neutrophils (PMNs) from healthy donors treated with PMA (100 nM) at indicated time points, assessed by flow cytometry and fluorescence microscopy. Percentage of Annexin V+ DAPI− cells (representing PS externalization) and NETs per field (quantified by ImageJ software, and an average of ten different fields per donor) are shown. Each dot represents an individual donor (n = 8). c, d, As in b, PS exposure and NETs formation in ATRA-differentiated neutrophil-like HL-60 cells (dHL-60) treated with PMA (100 nM) for 1-4 h. Annexin V staining in live cells (PI−) assessed by flow cytometry. Representative histograms of Annexin V staining (c) and quantifications of Annexin V+PI− cells (n = 3) and NETs (n = 8) (d). e, f, Normalized XKR8 gene expression in human (e) or mouse (f) immune cell subsets, analyzed using data from the Human Protein Atlas (HPA) and Immunological Genome Project (ImmGen), respectively. NETs were defined by a nuclear area greater than 80 µm2 in human PMNs (b) or 100 µm2 in dHL-60 cells (d). Each dot represents an individual donor (b) or a biological replicate (d). Data are represented as mean ± SEM from one representative experiment (d). All experiments were performed for at least two independent times.
Extended Data Fig. 2 Generation and characterization of XKR8-deficient mice.
a, b, Strategy for generating Xkr8 germline knockout mice (a), and reverse transcription PCR (RT-PCR) analysis of Xkr8 expression in BMNs from WT and XKR8 KO mice (b). c, RNA-Seq analysis of BMNs from WT and XKR8 KO mice. d, Representative pictures of H&E staining of spleen and lymph nodes sections from naïve WT and XKR8 KO mice (n = 5). Scale bars, 200 µm. e, Serum cytokine levels in naïve WT and XKR8 KO mice (n = 5). f, Giemsa staining of WT and XKR8 KO BMNs to assess nuclear morphology. Scale bars, 10 µm. g, Representative flow cytometry histograms of surface markers expression in BMNs from WT (blue) and XKR8 KO (red) mice. The gray lines indicate the isotype control. Each dot represents an individual mouse (e). Data are represented as mean ± SEM from one representative experiment (e). Statistical analysis was performed by two-sided Mann-Whitney U test, and adjusted P value was calculated using the Benjamini-Hochberg (BH) procedure to control the false discovery rate (FDR) (c). All experiments were performed independently for at least twice.
Extended Data Fig. 3 Impaired NETs formation in XKR8-deficient neutrophils.
a, PS exposure in WT and XKR8 KO BMNs treated with staurosporine (STS, 10 µM) or S63845 (10 µM) to induce apoptosis. Representative histogram (left) and quantifications (right) of PS exposure in living cells (Annexin V+ PI−) are shown (n = 3). b, Representative live cell imaging in WT and XKR8 KO BMNs stained with DNA (Hoechst 33342, pseudocolor) during NETs formation. c, NETs quantification in WT and XKR8 KO BMNs treated with PMA (100 nM) based on fluorescence intensity of DAPI and nuclear area. Cells with nuclear area greater than 80 µm2 were defined as NETs. d, Quantification of NETs in WT and XKR8 KO BMNs treated with MSU (100 µg/mL, 6 h) or Zymosan (25 µg/mL, 4 h) (n = 10). e, Quantification of dsDNA in the supernatant of WT and XKR8 KO BMNs treated with PMA (100 nM, 6 h), A23187 (1.25 µM, 4 h), LPS (10 µg/mL, 2 h) or nigericin (5 µM, 4 h) (n = 3). f, Neutrophils degranulation assay in WT and XKR8 KO BMNs co-cultured with E. coli-GFP (MOI 10) for 0.5 h (n = 3). CD63 staining was used to detect degranulation. Representative histogram (left) and quantification (right) are shown. g, h, Phagocytosis assays for WT and XKR8 KO BMNs using opsonized E. coli-GFP (MOI 10) (g) or FITC-labeled latex beads (1:100) (h) for 0.5 h. Representative flow cytometry plots (left) and quantification (right) of phagocytic neutrophils (n = 3). i, j, Quantification of NETs in WT, XKR8 KO and PAD4 KO BMNs treated with PMA (100 nM, 6 h), A23187 (1.25 μM, 4 h), LPS (10 μg/mL, 2 h), and nigericin (5 μM, 4 h) or co-cultured with L. monocytogenes (L. M.) (MOI 10), E. coli (MOI 10) for 1.5 h (n = 10-13). k, Quantification of NETs in WT, XKR8 KO, TMEM16F KO and XKR8/TMEM16F double KO (DKO) BMNs treated with PMA (100 nM, 6 h) or A23187(1.25 μM, 4 h) (n = 10). l, Representative confocal microscopy images of neutrophils stained with elastase (red), MPO (green), and DNA (DAPI, blue). Scale bars, 10 µm. m, Immunoblots of MPO in WT and XKR8 KO BMNs stimulated with PMA (100 nM) at the indicated time points. β-actin as the loading control. NETs were defined by a nuclear area greater than 80 µm2 (d, i, k), or the ratio of citrullinated histone to DNA (j). Each dot represents one individual field of view (d, i, j, k) or a biological replicate (a, e–h). Data are represented as mean ± SEM from one representative experiment of three (a, d–k) independent experiments. Statistical analysis was performed using two-way ANOVA (a, d–k). P values are shown on the graphs.
Extended Data Fig. 4 Upstream signaling pathways of NETs formation remain unaltered in XKR8-deficient neutrophils.
a, Flow cytometry analysis of PS exposure in ATRA-differentiated HL-60 (dHL-60) WT and XKR8 KO cells treated with PMA (100 nM). Representative histograms of Annexin V staining gated on live cells (PI−) (left) and quantification of Annexin V+PI− cells (right) (n = 3). b, c, Quantification of extracellular and intracellular ROS levels in WT and XKR8 KO neutrophils treated with PMA (100 nM), measured using isoluminol (b) and DHR123 assay (c), respectively (n = 3). d–h, Quantification of PMA-induced NETs formation (100 nM, 6 h) in WT and XKR8-deficient neutrophils after pre-incubation with various inhibitors: Tempol (30 μM) (d, n = 12), DPI (10 μM) (e, n = 11), GSK2795039 (10 μM) (f, n = 12), Enzastaurin (10 μM) (g, n = 12), U0126 (10 μM), GW5074 (10 μM) (h, n = 12). i, j, Immunoblots of phosphorylated ERK1/2 (p-ERK1/2) (i) and phosphorylated PKC (p-PKC, Ser660) (j) in WT and XKR8 KO mice BMNs stimulated with PMA (100 nM). k, Representative confocal images of neutrophils stained with activated caspase-3 (red), Ly6G (green), and DNA (DAPI, blue). Scale bars, 10 µm. l, m, Immunoblot analysis of cleaved caspase-9 (l), caspase-7, and caspase-8 (m) in WT and XKR8 KO BMNs treated with PMA (100 nM) or Raptinal (10 μM). β-actin as the loading control. NETs were quantified by a nuclear area greater than 80 µm2 (d–h). Each dot represents an independent biological replicate (a, c), the average from three technical replicates (b), or one individual image (d–h). Data are represented as the mean ± SEM from one representative experiment of three (a–d, f–h), or four (e) independent experiments. Experiments were performed at least twice for i–m. Statistical analysis was performed using two-way ANOVA (a, c–h). P values are shown on the graphs.
Extended Data Fig. 5 XKR8-mediated lipid scrambling activated by caspase-3 triggers NETs formation.
a–d, Quantification of PS exposure (a, b, n = 3) and NETs formation (c, n = 8; d, n = 12) in WT and XKR8 KO BMNs treated with PMA (100 nM) for 6 h, in the presence or absence of caspase inhibitors Z-VAD (30 µM) or Emricasan (10 µM). e, Quantification of NETs formation in human primary neutrophils from healthy donors (HD, n = 8) treated with PMA (5 nM) for 4 h, in the presence or absence of caspase-3 inhibitors QVD (10 µM), Z-VAD (30 µM) or Emricasan (10 µM) (n = 8). f–i Quantification of caspase-3 activation by CellEvent (f, g) and PS exposure (h, i) in WT and XKR8 KO BMNs treated with PMA (100 nM), A23187 (1.25 µM), LPS (10 µg/mL) or nigericin (5 µM). Representative histograms (f, h) and quantifications (g, i) of CellEvent+ or Annexin V+ cells in live cells (PI−) (n = 3). j, Quantification of NETs in WT and XKR8-deficient neutrophils treated with PMA (100 nM, 6 h, n = 12), A23187 (1.25 µM, 4 h, n = 8), LPS (10 µg/mL, 2 h, n = 12) or nigericin (5 µM, 4 h, n = 12) after incubation with QVD (10 µM), respectively. k, Immunoblot analysis of caspase-3 in WT and caspase-3 KO BMNs. β-actin as a loading control. l, m, PS exposure in WT and caspase-3 KO BMNs were treated with Raptinal (10 µM) or S63845 (10 µM). Representative histograms (l) and quantification of Annexin V+PI− cells (m) (n = 3). n–p, Representative histograms and quantification of caspase-3 activation (n, n = 3), PS externalization (o, n = 3) and NETs formation (p, n = 8) in WT and XKR8 KO neutrophils treated with PMA (100 nM), with or without Apocynin (30 µM), DPI (10 µM) or NA-11 (1 µM). NETs were defined by a nuclear area greater than 80 µm2 (c–e, j, p). Each dot represents one individual field of view (c–e, j, p) or a biological replicate (a, b, g, i, m–o). Data are represented as mean ± SEM from one representative experiment of three (b, d–p), and four (a, c) independent experiments. Statistical analysis was performed using two-way ANOVA (a–d, g, i, j, m–p) or one-way ANOVA (e). P values are shown on the graphs.
Extended Data Fig. 6 Extensive lipid modification by XKR8-mediated scrambling during NETs formation.
a, Immunoblot of full-length XKR8 overexpression in XKR8 KO HL-60 cells (KO + XKR8FL). Flag-tag was added to the C-terminus of XKR8. β-actin as the loading control. b, c, Quantification of PS exposure in dHL-60 WT, XKR8 KO and KO + XKR8FL cells treated with doxorubicin (10 µM) (b) or PMA (100 nM) (c) (n = 3). d, e Immunoblot of mouse XKR8 mutants (2DA and S/T-3A, Flag-tagged) overexpression in HL-60 XKR8 KO cells. β-actin as the loading control. f, g, Quantification of PS exposure in dHL-60 WT, KO, and XKR8 mutant cells (KO + XKR82DA, KO + XKR8S/T-3A) treated with doxorubicin (10 µM) (f) or PMA (100 nM) (g) (n = 3). h, i, Immunoblot of mouse XKR8 mutants (D30A and W45A, Flag-tagged) in HL-60 XKR8 KO cells. β-actin as the loading control. j, k, As in f, g, PS exposure was assessed in dHL-60 WT, XKR8 KO, and mutant cells (KO + XKR8D30A, KO + XKR8W45A) treated with doxorubicin (10 µM) (j) or PMA (100 nM) (k) (n = 3). l, Gating strategy representative histograms and quantifications (l) of Laurdan dye signals in BMNs treated with PMA (100 nM) for 2 h (n = 3). Each dot represents a biological replicate (b, c, f, g, j, k, l). Data are represented as mean ± SEM from one representative experiment of three independent experiments (b, c, f, g, j, k, l). Statistical analysis was performed using two-sided unpaired Student t-test (l), or two-way ANOVA (b, c, f, g, j, k,). P values are shown on the graphs.
Extended Data Fig. 7 XKR8-mediated lipid scrambling regulates intracellular Ca2+ for NETs formation.
a, b, Quantification of NETs formation in WT and XKR8 KO BMNs treated with PMA (100 nM, 6 h) in RPMI 1640 or Ca2+-free HBSS (n = 10) (a), or in the presence of BAPTA-AM (25 μM, n = 10) (b). c, Calcium influx was evaluated in WT and XKR8 KO BMNs by Indo-1. Indo-1 signals were immediately recorded after adding PMA (100 nM), A23187 (1.25 μM), LPS (10 μg/mL), or anti-CD16 crosslinking for 5 min. d, Quantification of PMA-induced NETs in WT and XKR8 KO neutrophils treated with BAPTA-AM (25 μM) at indicated time points after PMA stimulation (n = 10). e, f, Intracellular Ca2+ levels in WT and XKR8 KO BMNs was assessed by Indo-1. Indo-1 fluorescence signals were measured 2 h after treatment with PMA (100 nM), A23187 (1.25 μM) or PMA in the presence of BAPTA-AM (25 μM) (n = 3) (e), and quantification of intracellular Ca2+ levels in PS-exposed (Annexin V+) versus non-exposed (Annexin V−) cells at 2 h following PMA stimulation (n = 3) (f). Intracellular Ca2+ levels was determined from the ratio of bound to free Ca2+ (e, f). g, Quantification of NETs in WT and XKR8-deficient neutrophils treated with PMA (100 nM, 6 h) in the presence of BAPTA-AM (25 μM) or EGTA-AM (25 μM) (n = 12). NETs were defined by a nuclear area greater than 80 µm2 (a, b, d, g). Each dot represents one individual field of view(a, b, d, g) or a biological replicate (e, f). Data are presented as mean ± SEM from one representative experiment of three (a, b, d–g) independent experiments. Statistical analysis was performed using two-way ANOVA (a, b, d, e, g) or one-way ANOVA (f). P values are shown on the graphs.
Extended Data Fig. 8 XKR8-mediated lipid scrambling necessitates the activation of mechanosensitive Ca2+ channels for NETs formation.
a, b, Quantification of NETs in WT and XKR8-deficient BMNs or HL-60 cells treated with PMA (100 nM) in the presence of inhibitors listed in Fig. 4d. a, PMA-induced NETs formation with inhibitors targeting TRPM2 (glutathione, GSH, 10 μM; JNJ-28583113, 10 μM), TRPM4 (9-phenanthrol, 30 μM), TRPM7 (Carvacrol, 100 μg/mL), SOCE (2-APB, 25 μM; CM-4620, 10 μM), TRPV2 (SKF96365, 20 μM), TRPV4 (HC067047, 1 μM), Piezo1 (GsMTx-4, 10 μM), TRPC3 (Pyr3, 5 μM), TRPC6 (SAR7334, 1 μM) (n = 8-9). b, As in a, Quantification of PMA-induced NETs in WT and XKR8-deficient dHL-60 cells with inhibitors HC067047 (1 μM), Pyr3 (5 μM), SKF96365 (20 μM), 9-phenanthrol (30 μM), JNJ-28583113 (10 μM), Carvacrol (100 μg/mL) (n = 8). c, As in a, quantification of NETs in human primary neutrophils isolated from healthy donors (HD, n = 8) treated with PMA (10 nM) in the presence of mechanosensitive Ca2+ channel inhibitors. d, Calcium influx was evaluated in WT and XKR8 KO BMNs by Indo-1. Indo-1 signals were recorded immediately after adding the agonists targeting TRPV2 (Cannabidiol, CBD, 5 μM), TRPC3 (GSK1702934A, 100 μM), TRPV4 (GSK1016790A, 100 nM), Piezo1 (Yoda1, 10 μM) for 5 min. A23187 (1.25 μM) as a positive control. e, Quantification of intracellular Ca2+ levels in WT and XKR8 KO BMNs by Indo-1. Indo-1 fluorescence signals were measured at 2 h after treatment with PMA (100 nM) and in the presence of the agonists targeting TRPV2 (CBD, 5 μM), TRPC3 (GSK1702934A, 100 μM), TRPV4 (GSK1016790A, 100 nM), Piezo1 (Yoda1, 10 μM) (n = 3). f, Quantification of PMA-induced NETs in WT and XKR8-deficient dHL-60 cells with agonists targeting TRPV2 (CBD, 5 μM; Probenecid, 1 mM), TRPC3 (GSK1702934A, 100 μM), TRPV4 (GSK1016790A, 100 nM), TRPM7 (Naltriben Mesylate, NTB, 50 μM) (n = 8). g, h, Quantification of NETs in human primary neutrophils (g, n = 8) or mouse BMNs (h, n = 10) treated with agonists in the absence of PMA. i, Quantification of nigericin-induced NETs in WT and XKR8-deficient BMNs in the presence of the agonists targeting TRPV2 (CBD, 5 μM), TRPC3 (GSK1702934A, 100 μM), TRPV4 (GSK1016790A, 100 nM), Piezo1 (Yoda1, 10 μM) (n = 10). j, k, Quantification of PMA-induced NETs (j) and PS exposure (k) in TRPV2, TRPC3, TRPV4, and Piezo1-deficient dHL-60 cells (n = 10-12). NETs were quantified by a nuclear area greater than 80 µm2 for BMNs (a, c, g–i) or 100 µm2 for HL-60 cells (b, f, j). Each dot represents one individual field of view (a–c, f–j) or a biological replicate (e, k). Data are presented as mean ± SEM from one representative experiment of three (a–k) independent experiments. Statistical analysis was performed using two-way ANOVA (a, b, f, h–k), two-sided unpaired Student’s t-test (e), or one-way ANOVA (c, g). P values are shown on the graphs.
Extended Data Fig. 9 XKR8 plays a critical role in neutrophil defense against LPS-induced acute lung injury.
a, b, Flow cytometry analysis of neutrophils (CD45+ CD11b+ Ly6G+) in lung (a) and bronchoalveolar lavage fluid (BALF) (b) from WT and XKR8 KO mice after intranasal LPS challenge (100 ng per mouse). The gating strategy (a, b, left panels) and quantifications (a, b, right panels) were shown (n = 5 for PBS; n = 12 for LPS). c, Strategy for generating neutrophil-specific XKR8 KO (Xkr8fl/fl Ly6GCre) mice. d, e, Xkr8 expression in BMNs from Xkr8fl/fl and Xkr8fl/fl Ly6GCre mice using RT-PCR and qPCR (n = 3), with 18S as the internal reference. f, g, PS exposure in Xkr8fl/fl and Xkr8fl/fl Ly6GCre BMNs treated with STS (10 μM), S63845 (10 μM) (f) or PMA (100 nM) (g). Representative histograms (left) and quantification (right) of Annexin V+ PI− cells assessed by flow cytometry (n = 3). h, i, Flow cytometry analysis of neutrophils (CD45+ CD11b+ Ly6G+) in lung (h) and BALF (i) following LPS-induced ALI in Xkr8fl/fl and Xkr8fl/fl Ly6GCre mice. Gating strategy (left) and quantifications (right) are shown (n = 5). j, k, Quantification of neutrophil percentages in the lung and BALF after LPS-induced ALI in WT, XKR8 KO and XKR82DA KI mice (n = 4). Each dot represents one individual mouse in (a, b, h–k) or a biological replicate (e–g). Data are presented as mean ± SEM. Statistical analysis was performed by two-way ANOVA in (a, f, g, h, i), two-tailed Mann-Whitney test in (b, e) or one-way ANOVA in (j, k). P values are shown on the graphs.
Extended Data Fig. 10 XKR8 is critical for neutrophils-mediated antifungal infection.
a, b, Flow cytometry analysis of neutrophils (CD45+ CD11b+ Ly6G+) in the lung and BALF after C. alb (1×106 CFU per mouse) intranasal challenge in Xkr8fl/fl and Xkr8fl/fl Ly6GCre mice. Gating strategy (a) and quantifications (b) were shown (n = 5 for BALF; n = 9 for lung in WT and n = 8 for lung in KO). c, Representative pictures of BALF from Xkr8fl/fl (top) and Xkr8fl/fl Ly6GCre (bottom) mice after C. alb infection (n = 5). d, Quantification of neutrophils in lung and BALF from WT, XKR8 KO and XKR82DA KI mice after C. alb infection (n = 9). e, Representative image of BALF from WT (top), 2DA KI (middle) and XKR8 KO mice (bottom) after C. alb infection (n = 5). f, Representative H&E staining of liver and lung sections from WT and XKR8 KO mice after infection of C. alb intravenously (1×106 CFU per mouse). Scale bars, 100 μm. g, In vitro C. alb growth in the presence of GSK1016790A (n = 6). h, Number of erythrocytes in BALF from Xkr8fl/fl and Xkr8fl/fl Ly6GCre mice with or without GSK1016790A intranasal treatment before C. albicans challenge (n = 7-8). Pooled data from two independent experiments (b, d, h). Each dot represents one individual mouse (b, d, h), or a biological replicate (g). Data are represented as mean ± SEM. Statistical analysis was performed using two-tailed Student’s t-test (b), two-tailed Mann-Whitney test (g, h), and one-way ANOVA (d). P values are shown on the graphs.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2 and Supplementary Figs. 1–3.
Supplementary Code (download ZIP )
Code 1 was used to segment the ROIs of Hoechst 33342-stained nuclear via Cellpose with the pretrained model ‘nuclear’. Code 2 and 3 were used to calculate area and mean fluorescence intensity using MATLAB-based customer-written scripts ‘NETs’ and ‘gating’. Detailed descriptions of these procedures are provided in Supplementary Materials 1.
Supplementary Table (download PDF )
Characteristics of the human research participants, including age, sex and ethnic group.
Source Data Supplementary Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Supplementary Data Fig. 2 (download XLSX )
Statistical Source Data.
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Fig. 3 (download XLSX )
Statistical Source Data.
Source Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Fig. 5 (download XLSX )
Statistical Source Data.
Source Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical Source Data.
Source Data (download PDF )
Unprocessed immunoblots.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, W., Ping, J., Deng, L. et al. Lipid asymmetry disruption by XKR8 orchestrates neutrophil extracellular trap formation and inhibits fungal infection. Nat Immunol (2026). https://doi.org/10.1038/s41590-026-02456-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41590-026-02456-z