Abstract
Biomolecular condensates, such as stress and germ granules, often contain subcompartments. For instance, the Caenorhabditis elegans germ granule, which localizes near the outer nuclear membrane of germ cell nuclei, is composed of at least four ordered compartments, each housing distinct sets of proteins and RNAs. How these compartments form and why they are spatially ordered remains poorly understood. Here, we show that the conserved DEAD-box RNA helicase DDX-19 defines another compartment of the larger C. elegans germ granule, which we term the D compartment. The D compartment exhibits properties of a liquid condensate and forms between the outer nuclear pore filament and other compartments of the germ granule. Two nuclear pore proteins, NPP-14 and GLEL-1, are required for its formation, suggesting that the D compartment localizes adjacent to the outer nuclear membrane through interactions with the nuclear pore. The loss of DDX-19, NPP-14 or GLEL-1 leads to functional defects, including aberrant formation of the other four germ granule compartments, a loss of germline immortality and dysregulation of small RNA-based transgenerational epigenetic inheritance programs. Hence, we propose that a function of the D compartment is to anchor larger germ granules to nuclear pores, enabling germ granule compartmentalization and promoting transgenerational RNA surveillance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The MS proteomics data were deposited to the ProteomeXchange Consortium through the PRIDE partner repository with dataset identifier PXD056752. pUG RNA-seq datasets and small RNA-seq datasets were deposited to the NCBI Sequence Read Archive under BioProject accession numbers PRJNA1171951 and PRJNA1171955. Source data are provided with this paper.
Code availability
Custom scripts are available from GitHub (https://github.com/WANLAB20192/Code_DDX-19_paper).
References
Xie, Y. & Ren, Y. Mechanisms of nuclear mRNA export: a structural perspective. Traffic 20, 829–840 (2019).
Singh, G., Pratt, G., Yeo, G. W. & Moore, M. J. The clothes make the mRNA: past and present trends in mRNP fashion. Annu. Rev. Biochem. 84, 325–354 (2015).
Katahira, J. et al. The Mex67p-mediated nuclear mRNA export pathway is conserved from yeast to human. EMBO J. 18, 2593–2609 (1999).
Tseng, S. S. et al. Dbp5p, a cytosolic RNA helicase, is required for poly(A)+ RNA export. EMBO J. 17, 2651–2662 (1998).
Snay-Hodge, C. A., Colot, H. V., Goldstein, A. L. & Cole, C. N. Dbp5p/Rat8p is a yeast nuclear pore-associated DEAD-box protein essential for RNA export. EMBO J. 17, 2663–2676 (1998).
Adams, R. L., Mason, A. C., Glass, L., Aditi & Wente, S. R. Nup42 and IP6 coordinate Gle1 stimulation of Dbp5/DDX19B for mRNA export in yeast and human cells. Traffic 18, 776–790 (2017).
Lund, M. K. & Guthrie, C. The DEAD-box protein Dbp5p is required to dissociate Mex67p from exported mRNPs at the nuclear rim. Mol. Cell 20, 645–651 (2005).
Weirich, C. S. et al. Activation of the DExD/H-box protein Dbp5 by the nuclear-pore protein Gle1 and its coactivator InsP is required for mRNA export. Nat. Cell Biol. 8, 668–676 (2006).
Weirich, C. S., Erzberger, J. P., Berger, J. M. & Weis, K. The N-terminal domain of Nup159 forms a β-propeller that functions in mRNA export by tethering the helicase Dbp5 to the nuclear pore. Mol. Cell 16, 749–760 (2004).
Montpetit, B. et al. A conserved mechanism of DEAD-box ATPase activation by nucleoporins and InsP6 in mRNA export. Nature 472, 238–242 (2011).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Voronina, E., Seydoux, G., Sassone-Corsi, P. & Nagamori, I. RNA granules in germ cells. Cold Spring Harb. Perspect. Biol. 3, a002774 (2011).
Dodson, A. E. & Kennedy, S. Phase separation in germ cells and development. Dev. Cell 55, 4–17 (2020).
Gao, M. & Arkov, A. L. Next generation organelles: structure and role of germ granules in the germline. Mol. Reprod. Dev. 80, 610–623 (2013).
Wan, G. et al. Spatiotemporal regulation of liquid-like condensates in epigenetic inheritance. Nature 557, 679–683 (2018).
Phillips, C. M., Montgomery, T. A., Breen, P. C. & Ruvkun, G. MUT-16 promotes formation of perinuclear mutator foci required for RNA silencing in the C. elegans germline. Genes Dev 26, 1433–1444 (2012).
Manage, K. I. et al. A Tudor domain protein, SIMR-1, promotes siRNA production at piRNA-targeted mRNAs in C. elegans. eLife 9, e56731 (2020).
Sheth, U., Pitt, J., Dennis, S. & Priess, J. R. Perinuclear P granules are the principal sites of mRNA export in adult C. elegans germ cells. Development 137, 1305–1314 (2010).
Pitt, J. N., Schisa, J. A. & Priess, J. R. P granules in the germ cells of Caenorhabditis elegans adults are associated with clusters of nuclear pores and contain RNA. Dev. Biol. 219, 315–333 (2000).
Zhang, D. et al. The piRNA targeting rules and the resistance to piRNA silencing in endogenous genes. Science 359, 587–592 (2018).
Shen, E. Z. et al. Identification of piRNA binding sites reveals the Argonaute regulatory landscape of the germline. Cell 172, 937–951 (2018).
Ashe, A. et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Grishok, A., Tabara, H. & Mello, C. C. Genetic requirements for inheritance of RNAi in C. elegans. Science 287, 2494–2497 (2000).
Vastenhouw, N. L. et al. Gene expression: long-term gene silencing by RNAi. Nature 442, 882 (2006).
Alcazar, R. M., Lin, R. & Fire, A. Z. Transmission dynamics of heritable silencing induced by double-stranded RNA in Caenorhabditis elegans. Genetics 180, 1275–1288 (2008).
Tsai, H. Y. et al. A ribonuclease coordinates siRNA amplification and mRNA cleavage during RNAi. Cell 160, 407–419 (2015).
Preston, M. A. et al. Unbiased screen of RNA tailing activities reveals a poly(UG) polymerase. Nat. Methods 16, 437–445 (2019).
Shukla, A. et al. poly(UG)-tailed RNAs in genome protection and epigenetic inheritance. Nature 582, 283–288 (2020).
Roschdi, S. et al. An atypical RNA quadruplex marks RNAs as vectors for gene silencing. Nat. Struct. Mol. Biol. 29, 1113–1121 (2022).
Ouyang, J. P. T., Zhang, W. L. & Seydoux, G. The conserved helicase ZNFX-1 memorializes silenced RNAs in perinuclear condensates. Nat. Cell Biol. 24, 1129–1140 (2022).
Phillips, C. M. & Updike, D. L. Germ granules and gene regulation in the Caenorhabditis elegans germline. Genetics 220, iyab195 (2022).
Wan, G. et al. ZSP-1 is a Z granule surface protein required for Z granule fluidity and germline immortality in Caenorhabditis elegans. EMBO J. 40, e105612 (2021).
Burton, N. O., Burkhart, K. B. & Kennedy, S. Nuclear RNAi maintains heritable gene silencing in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 108, 19683–19688 (2011).
Wylie, C. Germ cells. Curr. Opin. Genet. Dev. 10, 410–413 (2000).
Lev, I. et al. MET-2-dependent H3K9 methylation suppresses transgenerational small RNA inheritance. Curr. Biol. 27, 1138–1147 (2017).
Shukla, A., Perales, R. & Kennedy, S. piRNAs coordinate poly(UG) tailing to prevent aberrant and perpetual gene silencing. Curr. Biol. 31, 4473–4485 e3 (2021).
Simon, M. et al. Reduced insulin/IGF-1 signaling restores germ cell immortality to Caenorhabditis elegans Piwi mutants. Cell Rep. 7, 762–773 (2014).
Barucci, G. et al. Small-RNA-mediated transgenerational silencing of histone genes impairs fertility in piRNA mutants. Nat. Cell Biol. 22, 235–245 (2020).
Strome, S.Specification of the germ line. WormBook 28, 1–10 (2005).
Spike, C. et al. Genetic analysis of the Caenorhabditis elegans GLH family of P-granule proteins. Genetics 178, 1973–1987 (2008).
Chen, W. et al. The dynamics of P granule liquid droplets are regulated by the Caenorhabditis elegans germline RNA helicase GLH-1 via its ATP hydrolysis cycle. Genetics 215, 421–434 (2020).
Spike, C. A., Bader, J., Reinke, V. & Strome, S. DEPS-1 promotes P-granule assembly and RNA interference in C. elegans germ cells. Development 135, 983–993 (2008).
Marnik, E. A. et al. The Caenorhabditis elegans TDRD5/7-like protein, LOTR-1, interacts with the helicase ZNFX-1 to balance epigenetic signals in the germline. PLoS Genet. 18, e1010245 (2022).
Patel, S. S., Belmont, B. J., Sante, J. M. & Rexach, M. F. Natively unfolded nucleoporins gate protein diffusion across the nuclear pore complex. Cell 129, 83–96 (2007).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid–liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Ribbeck, K. & Gorlich, D. The permeability barrier of nuclear pore complexes appears to operate via hydrophobic exclusion. EMBO J. 21, 2664–2671 (2002).
Schmidt, H. B. & Gorlich, D. Nup98 FG domains from diverse species spontaneously phase-separate into particles with nuclear pore-like permselectivity. eLife 4, e04251 (2015).
Duster, R., Kaltheuner, I. H., Schmitz, M. & Geyer, M. 1,6-Hexanediol, commonly used to dissolve liquid–liquid phase separated condensates, directly impairs kinase and phosphatase activities. J. Biol. Chem. 296, 100260 (2021).
Marnik, E. A. et al. Germline maintenance through the multifaceted activities of GLH/Vasa in Caenorhabditis elegans P granules. Genetics 213, 923–939 (2019).
von Moeller, H., Basquin, C. & Conti, E. The mRNA export protein DBP5 binds RNA and the cytoplasmic nucleoporin NUP214 in a mutually exclusive manner. Nat. Struct. Mol. Biol. 16, 247–254 (2009).
Schmitt, C. et al. Dbp5, a DEAD-box protein required for mRNA export, is recruited to the cytoplasmic fibrils of nuclear pore complex via a conserved interaction with CAN/Nup159p. EMBO J. 18, 4332–4347 (1999).
Noble, K. N. et al. The Dbp5 cycle at the nuclear pore complex during mRNA export II: nucleotide cycling and mRNP remodeling by Dbp5 are controlled by Nup159 and Gle1. Genes Dev. 25, 1065–1077 (2011).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Chen, X. et al. Regulation of CED-3 caspase localization and activation by C. elegans nuclear-membrane protein NPP-14. Nat. Struct. Mol. Biol. 23, 958–964 (2016).
Thomas, L., Taleb Ismail, B., Askjaer, P. & Seydoux, G. Nucleoporin foci are stress-sensitive condensates dispensable for C. elegans nuclear pore assembly. EMBO J. 42, e112987 (2023).
Tan, W., Zolotukhin, A. S., Bear, J., Patenaude, D. J. & Felber, B. K. The mRNA export in is mediated by Ce-NXF-1, an ortholog of human TAP/NXF and Mex67p. RNA 6, 1762–1772 (2000).
Batista, P. J. et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Mol. Cell 31, 67–78 (2008).
Billi, A. C., Fischer, S. E. & Kim, J. K.Endogenous RNAi pathways in C. elegans. WormBook 7, 1–49 (2014).
Claycomb, J. M. et al. The Argonaute CSR-1 and its 22G-RNA cofactors are required for holocentric chromosome segregation. Cell 139, 123–134 (2009).
Gu, W. et al. Distinct argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans germline. Mol. Cell 36, 231–244 (2009).
Lee, H. C. et al. C. elegans piRNAs mediate the genome-wide surveillance of germline transcripts. Cell 150, 78–87 (2012).
Wu, W. S. et al. piRTarBase: a database of piRNA targeting sites and their roles in gene regulation. Nucleic Acids Res. 47, D181–D187 (2019).
Bagijn, M. P. et al. Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 337, 574–578 (2012).
Tran, E. J., Zhou, Y., Corbett, A. H. & Wente, S. R. The DEAD-box protein Dbp5 controls mRNA export by triggering specific RNA:protein remodeling events. Mol. Cell 28, 850–859 (2007).
Yang, P. G. et al. G3BP1 Is a tunable switch that triggers phase separation to assemble stress granules. Cell 181, 325–345 (2020).
Sanders, D. W. et al. Competing protein–RNA interaction networks control multiphase intracellular organization. Cell 181, 306–324 (2020).
Wang, L. et al. Multiphase coalescence mediates Hippo pathway activation. Cell 185, 4376–4393 (2022).
Price, I. F., Hertz, H. L., Pastore, B., Wagner, J. & Tang, W. Proximity labeling identifies LOTUS domain proteins that promote the formation of perinuclear germ granules in C. elegans. eLife 10, e72276 (2021).
Ratke, L. & Voorhees, P. W. Growth and Coarsening: Ostwald Ripening in Material Processing 1st edn (Springer, 2002).
Singh, M. et al. Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis. Nat. Commun. 12, 3492 (2021).
Price, I. F., Wagner, J. A., Pastore, B., Hertz, H. L. & Tang, W. C. elegans germ granules sculpt both germline and somatic RNAome. Nat. Commun. 14, 5965 (2023).
Uebel, C. J., Rajeev, S. & Phillips, C. M. Caenorhabditis elegans germ granules are present in distinct configurations and assemble in a hierarchical manner. Development 150, dev202284 (2023).
Ouyang, J. P. T. et al. P granules protect RNA interference genes from silencing by piRNAs. Dev. Cell 50, 716–728(2019).
Wu, W. S. et al. pirScan: a webserver to predict piRNA targeting sites and to avoid transgene silencing in C. elegans. Nucleic Acids Res. 46, W43–W48 (2018).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Arribere, J. A. et al. Efficient marker-free recovery of custom genetic modifications with CRISPR/Cas9 in Caenorhabditis elegans. Genetics 198, 837–846 (2014).
Zhao, C. et al. HERD-1 mediates multiphase condensate immiscibility to regulate small RNA-driven transgenerational epigenetic inheritance. Nat. Cell Biol. 26, 1958–1970 (2024).
Ghanta, K. S. & Mello, C. C. Melting dsDNA donor molecules greatly improves precision genome editing in Caenorhabditis elegans. Genetics 216, 643–650 (2020).
Ishikawa-Ankerhold, H. C., Ankerhold, R. & Drummen, G. P. Advanced fluorescence microscopy techniques—FRAP, FLIP, FLAP, FRET and FLIM. Molecules 17, 4047–4132 (2012).
Lee, C. et al. Single-molecule RNA fluorescence in situ hybridization (smFISH) in Caenorhabditis elegans. Bio Protoc. 7, e2357 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Acknowledgements
We thank previous and present G.W. lab members for development and critical discussion of this work. We thank the company NanoInsights for providing the Multi-SIM microscope. We appreciate the support of J. L Xu (Core Facilities of Life Sciences, School of Life Sciences, Sun Yat‑sen University) for equipment support and technical assistance. This work was supported by the National Natural Science Foundation of China (32370729 and 32070798, to G.W.), the Shenzhen Medical Research Fund (B2302029, to G.W.) and the Guangdong Science and Technology Department (2023B1212060028). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
P.L. and B.Y.D. conducted the majority of the experiments and analyses with assistance from X.F.N., Y.H.Q., Y.T.L., Y.L.L. and Z.P.Y. X.R.L. performed the bioinformatic analysis of small RNA-seq data. G.R.T. and G.Z.L. contributed to the development of the bioinformatic analysis strategy. S.K. contributed to the editing the manuscript. G.W. conceptualized and supervised the project, interpreted the results and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Carolyn Phillips, Ekaterina Voronina and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editors: Dimitris Typas and Sara Osman, in collaboration with the Nature Structural and Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DDX-19, GLEL-1, and NPP-14 are required for RNAi inheritance targeting two somatically expressed genes.
a-b, Animals with indicated genotype were subjected to lin-15b RNAi for two generations, representative images showing F1 adult animals (a), quantification of multi-vulva animals in F1 generation (b). The white arrow indicates extruded multi-vulva. Scale bar, 100 μm. n = 3 biological replicates (For each replicate, 50 animals were scored). These experiments were performed in an eri-1(mg366) background. Student’s t-test, two-tailed. Error bar, ±s.d. c-f, animals with indicated genotype were treated with control RNAi (L4440) or dpy-11 RNAi, the progeny was grown on OP50 plates. Representative images of F1 L4 larval stage animals were shown (c, d). Body length of F1 L4 larval stage animals was measured using ImageJ (e, f). Scale bar, 100 μm. n = 3 biological replicates (For each replicate, 10 animals were scored). One-way ANOVA. Error bar, ±s.d. g, Animals with indicated genotype were subjected to lin-15b RNAi for two generations, quantification of multi-vulva animals in F1 generation. These experiments were performed in an eri-1(mg366) background. n = 4 biological replicates. Student’s t-test, two-tailed. Error bar, ±s.d. h, Animals with indicated genotype were subjected to dpy-11 RNA inheritance assay. Percentage of animals with Dumpy phenotype in F1 L4 stage was scored. n = 4 biological replicates (for each replicate, 50 animlas were scored). Student’s t-test, two-tailed. Error bar, ±s.d. Source numerical data are available.
Extended Data Fig. 2 The TEI of RNAi targeting three endogenously expressed genes was not enhanced following the loss of DDX-19.
WT and ddx-19(-) animals were subjected to oma-1 RNAi (a), mex-6 RNAi(b), and puf-5 RNAi (c) treatment. Animals from P0, F1, F2, F4, F6, and F8 generations were collected and total RNAs were extracted. Quantitative reverse transcription and PCR (qRT-PCR) were employted to detect the mRNA levels of oma-1, mex-6, or puf-5 mRNAs in the indicated generations. n = 3 biological replicates. Student’s t-test, two-tailed. Error bar, ±s.d. Source numerical data are available.
Extended Data Fig. 3 Introduce fluorescence protein genes to endogenous locus generated functional fusion proteins.
a, WT, ddx-19(-) and ddx-19::gfp animals were grown on dpy-11 RNAi, F1 progeny were grown on OP50. The percentage of Dumpy animals in inheriting F1 was scored at L4 larval stage. n = 4 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. b, WT, npp-14(-), npp-14::mCherry::ha and npp-14::gfp::3xflag animals were grown at 25 °C, brood size was counted at indicated generations. n = 3 biological replicates. Error bar, ±s.d. c, WT, glel-1(-) and ha::mCherry::glel-1 animals were subjected to dpy-11 RNAi treatment, percentage of Dpy animals were scored at F1 generation. n = 3 biological replicates. Error bar, ±s.d. d-e, WT, mut-2(-) and mut-2::mCherry animals (d) or WT, rsd-2(-) and rsd-2::gfp animals (e) were treated with pos-1 RNAi. The percentage of hatched animals was scored. n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. Source numerical data are available.
Extended Data Fig. 4 Genetic requirements of DDX-19::GFP marked D compartment.
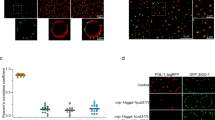
a-b, Fluorescence imaging of animals expressing DDX-19::GFP. White arrows indicate DDX-19::GFP in somatic cells. Scale bar, 5 μm for images in (a), 10 μm for images in (b). n > 3 animals. c, Fluorescence imaging of animals expressing PGL-1::TagRFP in WT, glh-1(-), and deps-1(-) animals. Scale bar, 5 μm. n > 3 animals. d-e, Fluorescence imaging of animals expressing TagRFP::ZNFX-1 (d) or DDX-19::GFP (e) in WT and zsp-1(-) animals. Scale bar, 5 μm. n > 3 animals. f, Fluorescence imaging of animals expressing GFP::ZNFX-1 in WT and lotr-1(-) animals. Scale bar, 5 μm. n > 3 animals. g, Fluorescence imaging of animals expressing MUT-2::mCherry in WT and mut-16(-) animals. Scale bar, 5 μm. n > 3 animals.
Extended Data Fig. 5 The effect of genetic mutations on protein levels.
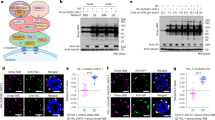
a, Schematic diagram of ddx-19 isoforms. IDR(+) indicates DDX-19 isoform 1 with complete IDR, IDR(-) indicates isoform 2-5 with truncated IDR or no IDR. b-c, Western blot to detect DDX-19::GFP isoforms in WT and DDX-19 helicase mutant young adult animals (b), quantification of DDX-19::GFP protein level normalized to Tubulin in (c). n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. d-e, Western blot to detect DDX-19::GFP isoforms in WT and npp-14(-) young adult animals (d), quantification of DDX-19::GFP protein level normalized to Tubulin in (e). n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. f-i, Western blot to detect DDX-19::GFP isoforms in WT young adult animals or embryos and young adult animals or embryos expressing NPP-14 with indicated residues mutated (f, h), quantification of DDX-19::GFP protein level normalized to Tubulin in (g, i). n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. j-m, Western blot to detect DDX-19::GFP isoforms in WT, glel-1(-) and npp-26(-) young adult animals (j) and embryos (l), quantification of DDX-19::GFP protein level normalized to Tubulin in (k, m). n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. n-q, Western blot to detect mCherry::GLEL-1 in WT, ddx-19(-) and npp-14(-) young adult animals (n) and embryos (p), quantification of mCherry::GLEL-1 protein level normalized to Tubulin in (o, q). n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. Source numerical data and unprocessed blots are available.
Extended Data Fig. 6 The localization interdependence of DDX-19, NPP-14 and GLEL-1.
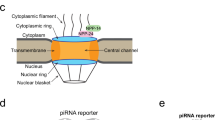
a, Fluorescence imaging of embryos expressing DDX-19::GFP in npp-14 mutants background. Scale bar, 5 μm. n > 3 animals. b-c, Fluorescence imaging of WT(b) and npp-14(-) (c) animals expressing DDX-19::GFP and PGL-1::TagRFP. n > 3 animals. d, Fluorescence imaging of animals expressing NPP-14::mCherry. Scale bar, 10 μm. White arrows indicate NPP-14::mCherry in somatic cells. n > 3 animals. e, Fluorescence imaging of animals expressing mCherry::GLEL-1. Scale bar, 10 μm. White arrows indicate mCherry::GLEL-1 in somatic cells. n > 3 animals. f, A schematic diagram of the C. elegans tissues nuclear pore. g, Spinning disc confocal imaging of animals expressing indicated proteins. Scale bar, 1 μm. n > 3 animals. h, Fluorescence imaging of WT, ddx-19(-), npp-14(-), and glel-1(-) animals expressing NPP-21::GFP, NPP-1::GFP, or TagRFP::NPP-9. Scale bar, 5 μm. n > 3 animals. i, Fluorescence imaging of animals expressing NPP-14::mCherry in WT or ddx-19(-) background. Scale bar, 5 μm. n > 3 animals. j, Fluorescence imaging of animals expressing NPP-14::mCherry in WT or glel-1(-) background. Scale bar, 5 μm. n > 3 animals.
Extended Data Fig. 7 The effect of DDX-19, NPP-14, or GLEL-1 depletion on RNA export, PZMS condensates organization, and marker proteins level.
a-b, Poly(A)+ RNAs in animals expressing PGL-1::GFP in WT or ddx-19(-) background were detected with RNA FISH. Scale bar, 5 μm for images in (a) and 10 μm for images in (b). n > 3 animals. c-d, Fluorescence imaging of animals expressing SIMR-1::TagRFP in WT, ddx-19(-), npp-14(-), and glel-1(-) background (c). Image J analysis to measure the size of SIMR-1 marked SIMR foci in (d). The y axis is expressed in log2 scale. n = 3 animals (for each animal, all granules surrounding a dozen pachytene region nuclei was quantified). Student’s t-test, two-tailed. Error bar, ±s.d. e, Fluorescence imaging of PGL-1::TagRFP, GFP::ZNFX-1, MUT-16::GFP or SIMR-1::TagRFP in rachis of germline in animals with indicated genotype. The white arrows indicate granules in the rachis. Scale bar, 5 μm. n > 3 animals. f-m, Western blotting and ImageJ quantification of GFP::ZNFX-1 and PGL-1 (f-i), MUT-16::GFP (j-k) or SIMR-1::TagRFP (l-m) after loss of DDX-19, NPP-14 or GLEL-1. n = 3 biological replicates. Error bar, ±s.d. Student’s t-test, two-tailed. n, Fluorescence imaging of another PZMS marker proteins in WT, ddx-19(-), npp-14(-) or glel-1(-) background. Scale bar, 5μm. n > 3 animals. Source numerical data and unprocessed blots are available.
Extended Data Fig. 8 Increased mixing of P and Z compartment components after loss of DDX-19, NPP-14, and GLEL-1 related to Fig. 6.
a-b, Fluorescence imaging of animals expressing GFP::ZNFX-1 and PGL-1::TagRFP in WT, ddx-19(-), npp-14(-), and glel-1(-) background (a). Image J analysis to measure the degree of colocalization between GFP::ZNFX-1 and PGL-1::TagRFP in WT and mutant animals (b). Scale bar, 1 μm for cells, 0.5 μm for inset. n = 27 (3 animals per biological replication, 3 nuclei per animal, 3 granules per nuclei). Error bar, ±s.d. Student’s t-test, two-tailed. c-d, Fluorescence imaging of animals expressing MUT-16::GFP and PGL-1::TagRFP (c) or MUT-16::GFP and TagRFP::ZNFX-1(d) in WT, ddx-19(-), npp-14(-) and glel-1(-) background. n > 3 animals. e, Fluorescence imaging of PGL-1::mCardinal, TagRFP::ZNFX-1, and MUT-16::GFP in WT, ddx-19(-), npp-14(-) and glel-1(-) animals. Scale bar, 1 μm for cells, 0.5 μm for inset. n > 3 animals. f, Quantification of the degree of colocalization in (e). n = 27 (3 animals, 3 nuclei per animal, 3 granules per nuclei). Error bar, ±s.d. Student’s t-test, two-tailed. g, ddx-19, npp-14 or glel-1 deletion were generated in animals expressing PGL-1::mCardinal, TagRFP::ZNFX-1 and MUT-16::GFP using CRISPR/Cas9. Fluorescence imaging was performed on animals expressing indicated proteins immediately at F2 generation when the heterozygous deletions became homozygous. n > 3 animals. h, Fluorescence imaging of animals expressing DDX-19::GFP and MUT-2::mCherry in WT, npp-14(-), and glel-1(-) background. n > 3 animals. i, Quantification of sphericity of DDX-19::GFP marked D compartment, PGL-1::mCardinal marked P compartment, and TagRFP::ZNFX-1 marked Z compartment in WT animals expressing DDX-19::GFP, PGL-1::mCardinal, and TagRFP::ZNFX-1. n = 27 (3 animals, 9 granule per germline). Student’s t-test, two tailed. Error bar, ±s.d. Source numerical data are available.
Extended Data Fig. 9 pUG RNAs level, length and distribution are altered after loss of ddx-19, npp-14, or glel-1 related to Fig. 7.
a, pUG PCR to detect gfp pUG RNAs in indicated generations after animals were exposed gfp RNAi. n = 3 biological replicates. b, Strategy to detect pUG RNAs using next-generation sequencing. Total RNA was extracted, and reverse transcription was performed using an (AC)9 oligo plus adaptors to generate pUG cDNA. Three rounds of PCR were subsequently conducted. In the first round, PCR was performed using a gene-specific primer and an adaptor-specific primer. For the second round of PCR, the products from the first PCR were diluted 100-fold. PCR was then performed using library primer forward 1 containing the self-index and unique molecular identifier (UMI) and library primer reverse 1. After a second 100-fold dilution, the third PCR round was performed using library primer forward 2 and library primer reverse 2 containing Illumina p5 and p7 sequences, respectively. The final PCR products were pooled, resolved on an agarose gel, and purified. The purified DNA libraries were sequenced to generate paired-end 150 bp reads. UMI, Unique Molecular Identifier. Self-index, index used for distinguishing samples. c-d, An independent replicate to detect gfp pUG RNAs in indicated generations related to Fig. 7a. e, An independent replicate to detect gfp pUG RNA tail length in indicated generations related to Fig. 7b. f, Quantification of the volume of Z compartments with or without pUG RNAs near or within. n = 27 (3 animals per biological replication, 3 nuclei per animal, 3 granules per nuclei). Error bar, ±s.d. Student’s t-test, two-tailed. g, ddx-19, npp-14, or glel-1 deletion were generated in animals expressing GFP::ZNFX-1 using CRISPR/Cas9. RNA FISH was performed to detect endogenous pUG RNAs and poly(A)+ RNAs immediately at F2 generation when the heterozygous deletions became homozygous. n > 3 animals. Source numerical data and unprocessed gels are available.
Extended Data Fig. 10 Features of DDX-19, NPP-14, and GLEL-1 Regulated Small RNAs and Working Model.
(a) Analysis of small RNA length and first nucleotide distribution in WT, ddx-19(-), npp-14(-), and glel-1(-) animals. (b) Biotype analysis of genes with significantly upregulated or downregulated targeting small RNAs in ddx-19(-), npp-14(-), and glel-1(-) animals compared with WT. (c) Representative factor analysis of genes with altered small RNAs in ddx-19(-), npp-14(-), and glel-1(-) animals against CSR-1, HRDE-1, and WAGO target lists. (d) piRNA target analysis of genes with altered small RNAs in ddx-19(-), npp-14(-), and glel-1(-) animals compared with WT. Welch’s t-test. (e) Comparison of genes with altered small RNAs in ddx-19(-), npp-14(-), and glel-1(-) animals with those in prg-1(-) or mut-16(-) mutants18. (f) Working Model. Biomolecular condensates are networks formed through protein-protein and protein-RNA interactions. Their organization depends on competing protein-protein interactions70. Homotypic interactions drive liquid-liquid phase separation, while non-competing heterotypic interactions result in co-phase separation, and competing heterotypic interactions lead to multiphase condensates71. In WT animals, the germline D compartment is anchored to the nuclear pore by NPP-14 and GLEL-1 at the cytoplasmic filament. This compartment is localized at the base of PZMS compartments, where NPP-14/GLEL-1 and P granules compete for DDX-19 interaction, facilitating D compartment formation. Loss of NPP-14 or GLEL-1 disrupts DDX-19 interactions, causing the D compartment to mix with P or Z compartments. This destabilizes the PZMS compartments, enlarging them and increasing local ZNFX-1 concentration. The elevated ZNFX-1 levels interact with more pUG RNAs, redistributing them from the M compartment to the mixed compartment and extending transgenerational epigenetic inheritance. This graph was created in BioRender. Deng, B. (2025) https://BioRender.com/w67i892.
Supplementary information
Source data
Source Data Figs. 1–4, 6 and 7 and Extended Data Figs. 1–3, 5 and 7–9 (download XLSX )
Statistical source data.
Source Data Extended Data Figs. 5, 7 and 9 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, P., Deng, B., Li, X. et al. A nuclear pore-anchored condensate enables germ granule organization and transgenerational epigenetic inheritance. Nat Struct Mol Biol 32, 1241–1254 (2025). https://doi.org/10.1038/s41594-025-01515-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41594-025-01515-7
This article is cited by
-
RNA helicase unravels the secrets of germ granules
Nature Structural & Molecular Biology (2025)
-
TurboID-based proximity labeling identifies novel germline proteins that maintain E granule integrity and small RNA homeostasis in C. elegans
Science China Life Sciences (2025)