Abstract
Integrated chicken and fish farming systems, common in Bangladesh, present significant public health risks due to the spread of antimicrobial resistance genes (ARGs) and virulence factors (VFGs) through mobile genetic elements (MGEs). This study employs metagenomic sequencing to explore the diversity and abundance of ARGs, VFGs, and MGEs in various environmental samples from these farming systems. A total of 384 ARGs were detected, with tetracycline resistance genes such as tetM and tetX being the most abundant, alongside macrolide-lincosamide-streptogramin and aminoglycoside resistance genes. Droppings harbored the highest proportion of ARGs (62.2%), whereas sediment served as a reservoir for multi-metal resistance genes. Virulence factors associated with immune modulation, such as pvdL and tssH, and biofilm formation genes like algC were particularly prevalent in sediment and droppings. Among MGEs, plasmids and transposons like Tn6072 and Tn4001 were the most abundant, playing a critical role in horizontal gene transfer. Bacterial genera including Bacteroides, Clostridium, and Escherichia were strongly associated with MGEs, indicating their role in the dissemination of resistance and virulence traits. Statistical analyses revealed significant differences in the abundance of ARGs, VFGs, and MGEs across sample types, with sediment and droppings identified as hotspots for gene exchange. These findings underscore the urgent need for improved antibiotic stewardship and waste management practices to limit the spread of antimicrobial resistance and pathogenic bacteria within integrated farming environments.
Similar content being viewed by others
Introduction
The intensification of agricultural practices, particularly within integrated farming systems, has emerged as a pivotal strategy to meet the escalating global demand for food1. Integrated chicken-fish farming systems, prevalent in Bangladesh, exemplify an approach that enhances resource optimization and agricultural productivity2. However, such systems also pose critical public health challenges, primarily linked to the propagation of antimicrobial resistance genes (ARGs)3. The indiscriminate and excessive use of antibiotics in livestock sectors, notably in poultry and aquaculture, has been identified as a principal catalyst for the emergence and proliferation of antimicrobial resistance (AMR)4. This phenomenon has facilitated the selection and dissemination of resistant bacterial strains, exacerbating the global AMR crisis—a threat that the World Health Organization (WHO) has ranked among the top ten public health concerns worldwide5.
In integrated farming systems, the close interaction between terrestrial and aquatic environments creates unique conditions that can facilitate the transfer and dissemination of ARGs6. Shared resources, such as water and feed, coupled with the recycling of animal waste, create conditions conducive to the spread of ARGs across microbial communities7. Mobile genetic elements (MGEs), such as plasmids, transposons, and insertion sequences (IS elements), play a critical role in this process by facilitating the horizontal gene transfer (HGT) of ARGs, further compounding the risks posed by antimicrobial resistance8,9. Virulence factors, which contribute to bacterial pathogenicity, are also a concern in these interconnected systems, as they can exacerbate the impact of resistant infections10. The presence of virulence factors alongside antimicrobial resistance genes can increase the pathogenic potential of bacterial strains, heightening the risk of severe infections that are more difficult to treat11.
Despite increasing awareness of the risks linked to integrated farming systems, a substantial knowledge gap persists regarding the prevalence and dissemination of antimicrobial resistance genes (ARGs), mobile genetic elements (MGEs), and virulence factors within these environments. This issue is particularly alarming in regions such as South Asia, where integrated farming practices are pervasive, and agricultural antibiotic use remains largely unregulated12. The potential for these systems to act as hotspots for the emergence and spread of AMR requires urgent attention.
Recent advancements in metagenomic technologies have provided powerful tools for studying complex microbial communities and their associated resistomes and virulomes13. Metagenomic approaches enable the detection of both culturable and non-culturable organisms, offering a comprehensive overview of the microbial diversity within integrated farming systems14. By identifying the presence of ARGs and virulence genes, as well as their potential for dissemination through MGEs, these analyses provide critical insights into the public health risks posed by these agricultural practices.
Given the significant implications of ARGs and virulence factors for global health, it is essential to investigate their dissemination within integrated farming systems. Understanding the pathways through which ARGs and virulence genes spread can help develop strategies to mitigate these risks15. This study explores the ARGs, Virulence Genes and Mobile Genetic Elements (MGEs) within integrated chicken and fish farming systems in Bangladesh, focusing on the potential for these systems to serve as reservoirs and transmission hubs. The findings of this research will contribute to broader efforts to combat AMR and inform policies aimed at promoting sustainable and safe agricultural practices16.
Result
Abundance of ARGs across samples
Metagenomic analysis identified a diverse array of Antimicrobial Resistance Genes (ARGs) across the entire integrated chicken and fish farming system. In total, 384 distinct resistance genes were detected, indicating the potential for antimicrobial resistance to emerge throughout the farming system.
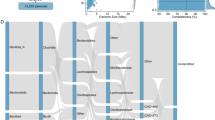
All the identified genes belonged to 40 different resistance classes where Tetracyclines were the most prevalent AMR (Antimicrobial Resistance) class, comprising 20.4% of the total resistance genes detected. This was followed by macrolide-lincosamide-streptogramin (17.6%) and aminoglycosides (15%). Droppings accounted for the highest proportion of AMR genes, contributing 62.2% of the total, while Sediment accounted for 31.5%. Chicken Gut, Feed, and Fish Intestine accounted for 5.3%, 0.9%, and 0.1%, respectively. Other AMR classes were detected at lower frequencies, including multi-drug resistance (4.3%), sulfonamides (3.9%), rifampin (3.8%), and phenicol (2.9%). Classes related to biocide and metal resistance, such as drug and biocide resistance (9.5%) and multi-metal resistance (2.5%), were also observed. Additionally, resistance to beta-lactams (1.4%), glycopeptides (0.4%), and fluoroquinolones (2.9%) was detected, albeit at lower levels (Fig. 1).
Chord diagram illustrating the distribution of antimicrobial resistance genes (ARGs) across different sample types. Droppings harbored the highest proportion of ARGs, followed by Sediment. The most prevalent resistance class was Tetracyclines, with macrolide-lincosamide-streptogramin and aminoglycosides also showing significant presence.
In terms of analyzing sample-wise relative abundance of ARGs, it was found that in the Feed sample, the most prevalent AMR classes were macrolide-lincosamide-streptogramin (28.60%), aminoglycosides (20.19%), and oxazolidinone (17.97%). Notable, though less abundant, were cationic antimicrobial peptides (6.00%) and fluoroquinolones (6.15%). The Chicken Gut sample demonstrated a dominant presence of macrolide-lincosamide-streptogramin (33.23%) and tetracyclines (22.07%), followed closely by aminoglycosides (19.12%). Sulfonamides (5.96%) and phenicol (5.29%) were also present, suggesting a complex resistance profile in the gut microbiome. Droppings exhibited the highest concentration of tetracyclines (28.43%) among all samples, with significant levels of macrolide-lincosamide-streptogramin (19.41%) and multi-drug resistance (9.63%). The presence of phenicol (5.96%) and sulfonamides (5.96%) indicates a broad spectrum of resistance within this sample. In the Fish Intestine sample, oxazolidinone (13.55%) and macrolide-lincosamide-streptogramin (14.53%) were the predominant AMR classes, with aminoglycosides (10.84%) also contributing notably. Fluoroquinolones (7.88%) were observed at lower levels, indicating a distinct resistance profile compared to other samples. The Sediment sample displayed a unique pattern, with multi-metal resistance (34.73%) being the most prevalent, followed by macrolide-lincosamide-streptogramin (10.95%) and aminoglycosides (10.99%). The significant presence of rifampin (9.11%) and phenicol (5.95%) further highlighted the diversity of resistance genes in this environmental reservoir (Fig. 2A).
(A) Relative abundance plot showing the sample-wise distribution of antimicrobial resistance genes (ARGs). In the Feed sample, the most prevalent AMR classes included macrolide-lincosamide-streptogramin, aminoglycosides, and oxazolidinone. The Chicken Gut sample demonstrated dominance of macrolide-lincosamide-streptogramin and tetracyclines, while Droppings exhibited the highest concentration of tetracyclines. Sediment displayed a unique profile with multi-metal resistance being the most prevalent class. (B) Bar plot illustrating the statistical analysis of antimicrobial resistance gene (ARG) abundance across different samples. The F-Welch test indicated a significant overall difference in ARG abundance among the samples. Subsequent Games-Howell pairwise analysis revealed statistically significant differences with a high effect size, suggesting substantial variability in resistance levels. Bayesian analysis further supported these findings, reinforcing the robustness of the observed differences.
Since resistance was detected across all samples within the farming system, statistical analyses were conducted to assess whether significant differences existed among the samples in terms of ARG presence. To address this, the distribution of antimicrobial resistance genes (ARGs) was analyzed using the F-Welch test, which is robust against unequal variances and sample sizes. The F-Welch test revealed a significant overall difference in ARG abundance among the samples [FWelch(4, 51.76) = 7.60, p = 6.77e-05]. To further explore these differences, the Games-Howell pairwise analysis was conducted, which also adjusts for unequal variances. Games-Howell test showed statistically significant differences with the effect size (ω2p = 0.42, CI95% [0.31, 1.00]), indicating that the magnitude of these differences are high. The Bayesian analysis provided a loge(BF01) of -0.41, with a posterior probability of 0.05 and a 95% credible interval [0.00, 0.15], along with an rCauchy (Random Cauchy) JZS (Jeffreys-Zellner-Siow) value of 0.71 further support the result (Fig. 2B).
To identify the most abundant ARGs, 30 most abundant genes are represented in Table 1 along with their class, function and abundance.
Virulence factors
A total of 445 types of virulence factor-associated genes were identified, belonging to 12 different mechanism classes. Among these, immune modulation was most abundant in Fish Intestine (67.01%) and Sediment (3.07%). The effector delivery system was highest in Chicken Gut (33.77%) and Sediment (34.82%). biofilm formation showed the greatest abundance in Feed (27.52%), followed by Chicken Gut (11.4%) and Droppings (9.55%). Motility was most prevalent in Droppings (20.1%) and Chicken Gut (13.4%). Other mechanisms such as Adherence were prominent in Droppings (18.05%) and Chicken Gut (14.43%). Nutritional/Metabolic factors were more abundant in Sediment (23.63%) and Droppings (14.03%). Exotoxin was mainly present in Fish Intestine (5.56%), and invasion was detected only in Droppings (1.54%). Mechanisms such as exoenzyme and Nutritional/Metabolic factors showed minimal presence across samples (Fig. 3A).
(A) Relative abundance plot depicting the distribution of virulence factor-associated genes across different samples. A total of 445 types of virulence factor-associated genes were identified, with immune modulation being most abundant in Fish Intestine and Sediment. The effector delivery system was highest in Chicken Gut and Sediment, while biofilm formation was predominant in Feed. Additionally, motility and adherence mechanisms were notable in Droppings and Chicken Gut, highlighting the diverse virulence profiles across samples. (B) Bar plot illustrating the abundance of virulence factor-associated genes across different sample types. Significant differences in virulence factor abundance were observed among the samples, with Sediment exhibiting the highest count compared to Feed, Chicken Gut, Fish Intestine, and Droppings. Droppings also showed significant differences when compared to Feed and Chicken Gut, while Chicken Gut had notable differences compared to Fish Intestine. The number of virulence factors identified in each sample varied, highlighting the diverse virulence profiles present.
There were significant differences in virulence factor abundance among the different sample types. Sediment had significantly higher virulence factor abundance compared to Feed (pHolm-adj. = 6.43e-13), Chicken Gut (pHolm-adj. = 5.88e-12), Fish Intestine (pHolm-adj. = 7.20e-12), and Droppings (pHolm-adj. = 1.71e-12). Droppings also showed significant differences compared to Feed (pHolm-adj. = 0.01) and Chicken Gut (pHolm-adj. = 0.00). Additionally, Chicken Gut exhibited significant differences when compared to Fish Intestine (pHolm-adj. = 0.03). The number of virulence factors present in each sample were: Chicken Gut (n = 39), Droppings (n = 206), Feed (n = 32), Fish Intestine (n = 26), and Sediment (n = 314) (Fig. 3B).
The top 25 most abundant virulence genes were detected exclusively in Droppings and Sediment samples, with no presence in Feed, Chicken Gut, or Fish Intestine samples. Certain genes, including pvdL, pvdI, pchF, tssH-5/clpV, pvdJ, icmF1/tssM1, cheA, tssM, and tsr, were found solely in Sediment with 100% abundance. Other genes displayed varying distributions between Droppings and Sediment. For instance, clpV1 (32.38% in Droppings, 67.62% in Sediment), ppkA (28.11% in Droppings, 71.89% in Sediment), pilT (41.06% in Droppings, 58.94% in Sediment), phzE1 (27.99% in Droppings, 72.01% in Sediment), algC (34.03% in Droppings, 65.97% in Sediment), and pilJ (37.59% in Droppings, 62.41% in Sediment) showed higher abundance in Sediment than in Droppings. Motility-related genes like flhA (38.95% in Droppings, 61.05% in Sediment) and fleQ (32.82% in Droppings, 67.18% in Sediment) followed a similar pattern, favoring Sediment (Table 2).
Abundance of mobile genetic elements
In the integrated chicken and fish farming system, a total of 2887 mobile genetic elements (MGEs) were identified, divided into six distinct types: Plasmids (n = 1,912), Insertion Sequences (IS) (n = 882), Integrative and Conjugative Elements (ICE) (n = 39), Transposons (Tn) (n = 26), Chromosomal Islands (CN) (n = 18), and Integrative Mobilizable Elements (IME) (n = 10). Plasmids, which are autonomous extrachromosomal DNA molecules, were the most prevalent, while IMEs, responsible for genome integration and mobilization, were the least common.
A Kruskal-Wallis test revealed significant overall differences between the six MGE types (χ²(5) = 115.01, p = 3.58e-23). Pairwise comparisons using Dunn’s test with Holm adjustment showed several significant differences. Notably, ICE and IME differed significantly from other types, with p-values as low as 1.21e-14. Additionally, IS and Plasmids, Tn and ICE, CN and IME, and other pairings showed significant variations with adjusted p-values ranging from 8.07e-04 to 6.30e-17. These results highlight a complex and distinct distribution of MGE types, reflecting their varied roles in horizontal gene transfer and ecological adaptation within the farming system (Fig. 4A).
(A) Bar plot displaying the distribution of mobile genetic element (MGE) types. A Kruskal-Wallis test indicated significant overall differences among the six MGE types. Pairwise comparisons revealed notable differences. Other significant variations were observed among IS and Plasmids, Tn and ICE, and CN and IME, emphasizing the complex roles of these elements in horizontal gene transfer and ecological adaptation within the farming system. (B) Bar plot illustrating the abundance of mobile genetic elements (MGEs) across different sample types. A Kruskal-Wallis test revealed significant overall differences in MGE abundance among the samples. Pairwise comparisons highlighted significant differences, particularly between Droppings and other sample types. Additional significant differences were observed between Sediment and Feed, as well as Chicken Gut and Fish Intestine, indicating distinct patterns of MGE distribution and suggesting varied levels of gene exchange and microbial activity in the environment.
Abundance of (MGEs) were identified across five sample types in the farming system counting as: Droppings (n = 1,823), Sediment (n = 1,321), Chicken Gut (n = 465), Feed (n = 227), and Fish Intestine (n = 141). Droppings and Sediment exhibited the highest abundance of MGEs, while the Fish Intestine had the lowest.
A Kruskal-Wallis test indicated significant overall differences in MGE abundance among the sample types (χ²(4) = 261.61, p = 2.06e-55). Pairwise comparisons using Dunn’s test with Holm adjustment revealed several significant differences, particularly between Droppings and other sample types, with p-values as low as 3.80e-38. Additional significant differences were observed between Sediment and Feed (p = 7.84e-12), Chicken Gut and Fish Intestine (p = 1.91e-16), and multiple other comparisons, with adjusted p-values ranging from 1.12e-03 to 4.06e-17. These findings demonstrate distinct patterns of MGE distribution across the different environmental samples, suggesting varied levels of gene exchange and microbial activity (Fig. 4B).
Correlation among organisms, ARGs and virulence genes
The correlation analysis between the top 25 most abundant MGEs and ARGs, using Spearman correlation with a threshold of r > 0.7 and p < 0.05, reveals several strong positive correlations. MGEs such as Tn6072, Tn6246, and Tn4001 exhibit significant positive correlations with ARGs like tetM, sulI, and tetX. Similarly, ICEBfrYCH46-1 shows a strong positive correlation with tetW and gyrA, while Tn6191 correlates positively with tetQ. Other notable positive correlations include CTn341 with rpoB, ISLjo5 with tetL, and CTnHyb with sulII. These results indicate a widespread co-association between MGEs and ARGs, suggesting the potential role of MGEs in the dissemination of antibiotic resistance within microbial communities (Fig. 5A).
(A) Heatmap depicting the correlation between the top 25 most abundant mobile genetic elements (MGEs) and antimicrobial resistance genes (ARGs). Correlation analysis using Spearman correlation revealed several strong positive correlations, with MGEs such as Tn6072, Tn6246, and Tn4001 significantly correlating with ARGs like tetM, sulI, and tetX. Additionally, ICEBfrYCH46-1 showed a strong positive correlation with tetW and gyrA, highlighting the widespread co-association between MGEs and ARGs, which suggests their potential role in the dissemination of antibiotic resistance within microbial communities. (B) Heatmap illustrating the correlation between the top 25 virulence genes and mobile genetic elements. Correlation analysis revealed that the transposon element Tn125 and the plasmid Nonomuraea gerenzanensis isolate nono1 plasmid I exhibited positive correlations with multiple virulence genes, including pvdL, pvdI, chpA, and pchF, among others.
The correlation analysis between the top 25 virulence genes revealed that the transposon element Tn125 and the plasmid Nonomuraea gerenzanensis isolate nono1 plasmid I showed a positive correlation with multiple virulence genes, including pvdL, pvdI, chpA, pchF, tssH-5/clpV, pchE, pvdD, icmF1/tssM1, pvdJ, tse5/rhsP1, cheA, tsr, and tssM (Fig. 5B).
A Spearman correlation analysis was conducted between microbial taxa and Mobile Genetic Elements (MGEs), focusing on the top 700 most abundant Operational Taxonomic Units (OTUs) out of 5796 identified OTUs. The analysis filtered results to retain only significant correlations where the correlation coefficient (r) was greater than 0.7 and the p-value was less than 0.05. A total of 521 OTUs, representing 260 taxonomic genera, were found to be significantly correlated with 453 MGEs.
The genus Bacteroides was strongly correlated with multiple MGEs, including ISEc36, ISKpn6, and ISVsa2. Similarly, Clostridium exhibited notable correlations with IS116, ISKpn26, and ISLsa11, indicating a close association with these genetic elements. The genus Escherichia was linked to several MGEs, such as ISEc52, ISKpn43, and ISPa42, showing its potential role in disseminating these elements across microbial communities. Spirochaeta demonstrated strong correlations with ISPme5 and ISXyl3, while Actinomyces was associated with ISXyl2 and ISEc54, highlighting their connections to distinct MGEs. Further correlations were observed between Lactobacillus and IS26, Prevotella and ISGca8, and Streptococcus with both IS26 and IS903, suggesting widespread genetic exchanges involving these genera. Additionally, Bifidobacterium was correlated with ISKpn7, and Pseudomonas showed a strong correlation with ISEcp1.
Moreover, Enterococcus was significantly correlated with IS26, IS903, and ISEc54, while Faecalibacterium displayed associations with IS26 and IS200, reinforcing the importance of these MGEs in microbial gene flow. Lastly, Roseburia was linked to ISEcp1, and Bifidobacterium was correlated with ISKpn7 and IS903, further demonstrating the widespread interaction between these bacterial taxa and MGEs. These correlations indicate substantial interactions between the microbial communities and MGEs, contributing to the potential dissemination of genetic material. Figure 6A, alongside of highlighting correlation between MGEs and organisms, also highlights the correlation among most significant (p < 0.01) ARGs and Virulence genes with the MGEs.
(A) Chord diagram illustrating the correlations between mobile genetic elements (MGEs) and various microbial genera, along with significant antimicrobial resistance genes (ARGs) and virulence genes. The analysis reveals strong correlations between the genus Bacteroides and multiple MGEs, such as ISEc36 and ISKpn6, while Clostridium showed notable associations with IS116 and ISKpn26. Escherichia, Spirochaeta, and Actinomyces also exhibited significant links to various MGEs, indicating their roles in genetic dissemination. The diagram further highlights the interactions between these bacterial taxa and significant ARGs and virulence genes, underscoring the potential for widespread genetic exchange within microbial communities. (B) Relative abundance plot illustrating the distribution of microbial phyla associated with mobile genetic elements (MGEs) across different samples. Pseudomonadota was the most abundant phylum, particularly in Chicken Gut and Fish Intestine. Actinomycetota exhibited high prevalence in Feed and Sediment, while Bacillota was notably abundant in Sediment and Droppings. Bacteroidota was also significant in Droppings and Fish Intestine. Other phyla, such as Thermodesulfobacteriota and Spirochaetota, showed lower abundance, with Mycoplasmatota and Campylobacterota being nearly absent across all samples.
The relative abundance of microbial phyla associated with MGEs shows distinct patterns in terms of presence throughout the whole farming system. Pseudomonadota was the most abundant, especially in Chicken_Gut (44.62%) and Fish_Intestine (29.62%). Actinomycetota was prevalent in Feed (28.05%) and Sediment (32.37%), while Bacillota was abundant in Sediment (25.93%) and Droppings (16.26%). Bacteroidota was notable in Droppings (18.92%) and Fish_Intestine (15.25%). Thermodesulfobacteriota, Spirochaetota, and other phyla showed lower abundance, with Mycoplasmatota and Campylobacterota nearly absent across all samples (Fig. 6B).
The correlation analysis revealed a significant association between various Mobile Genetic Elements (MGEs), bacterial genera, and their capacity to disseminate antimicrobial resistance genes (ARGs) and virulence genes. Among the most abundant MGEs, ICE SGB76 was linked to 12 genera and associated with ARGs such as lsa, eca, and tuf, alongside virulence genes like xcpR and tssC. Similarly, IME GISul2 was correlated with 13 genera, disseminating ARGs like cat, ber, and gtet, as well as virulence genes such as waaP and mucA. Transposon Tn558 was found to be connected to 11 genera, spreading ARGs like dfr and gtet and virulence genes such as xcpU and pilD. Tn4351 and Tn6031, both linked to 13 genera, were identified to disseminate various ARGs including erm and fol, with virulence genes like tssC and xcpA. Conjugative elements like Tn6191 and Tn6246 were associated with 12 genera, transmitting virulence genes such as mucA and rpoN. The study also highlighted Tn5432 and Tn4001, which are linked to the spread of ARGs such as aph and ble across 13 and 11 genera, respectively. Other significant MGEs, such as ICEBsa18170-1, tISCpe8, and CTn341, were also associated with the transfer of ARGs and virulence genes, indicating their crucial role in microbial gene dissemination (Table 3).
Discussion
The metagenomic analysis of antimicrobial resistance genes (ARGs) across the integrated chicken and fish farming system highlights the complex interplay between agricultural practices and the dissemination of resistance genes. The widespread detection of ARGs across different samples reflects the pervasive nature of resistance in such environments, with implications for both environmental and public health17.
The dominance of tetracycline resistance genes, particularly in droppings, is indicative of the selective pressures from tetracycline use in animal farming18. Tetracyclines have long been used as growth promoters and for disease prevention in poultry, which has led to their persistence in the environment and subsequent spread through manure and other waste products19. The detection of these genes in sediment further emphasizes the role of environmental reservoirs in maintaining and disseminating ARGs, as sediments often serve as sinks for pollutants, including antibiotics and resistance genes, especially in integrated farming systems where water flow connects different environments20,21.
The presence of macrolide-lincosamide-streptogramin and aminoglycoside resistance genes across various samples, particularly in the chicken gut and feed, suggests ongoing exposure to these antibiotic classes22. Macrolides and aminoglycosides are commonly used in livestock farming, and their resistance genes can be readily transferred between bacteria through horizontal gene transfer23. The detection of oxazolidinone resistance genes, though less common, raises concerns as oxazolidinones are often considered last-resort antibiotics in human medicine, and the presence of resistance genes in the farming environment could contribute to the diminishing efficacy of these drugs24,25.
Among the detected ARGs, those encoding resistance to last-resort antibiotics such as, TETW, MLS23S, RPOB, and GYRA belong to high-risk antimicrobial resistance genes (ARGs)26,27,28. These genes are linked to resistance against tetracyclines, macrolides, rifampin, and fluoroquinolones, respectively, and pose significant threats to both public and veterinary health6,18. When expressed at high levels in animal environments, such as feed, droppings, and sediment, these genes can act as reservoirs for resistance, potentially transferring to humans either directly through consumption or indirectly via environmental exposure29. Medium-risk ARGs, such as ERMB and PARE, also warrant attention despite their less widespread expression30,31. ERMB, involved in macrolide resistance, has been detected in animal environments, indicating a potential for resistance spread32. Similarly, PARE, related to aminocoumarin resistance, could contribute to DNA topoisomerase resistance, though its impact is still emerging33. These genes, though not as pervasive as their high-risk counterparts, still contribute to the broader AMR issue.
The identification of biocide and multi-metal resistance genes, particularly in sediment, highlights the role of environmental contaminants beyond antibiotics in selecting for resistance34. Heavy metals and biocides are often co-selectors for antibiotic resistance35, as bacteria that develop resistance to one type of stressor may simultaneously acquire resistance to others36. This co-selection process complicates efforts to control the spread of ARGs, as it suggests that reducing antibiotic use alone may not be sufficient to curb resistance in such complex environments37.
While this study provides insights into the presence of ARGs across various environmental samples, it is important to consider that the mere detection of ARGs does not confirm their expression or activity. Recent findings38 indicate that a substantial proportion of ARGs in animal and human gut microbiota are transcriptionally inactive, which underscores the need for integrative metagenomic and metatranscriptomic approaches to discern the functional relevance of detected resistance genes. Incorporating such analyses in future studies will provide a clearer understanding of the active resistome and its implications for antimicrobial resistance (AMR) risk assessment.
The presence of virulence factors poses a significant risk, as these genes can enhance bacterial pathogenicity, compounding the threat posed by antimicrobial resistance genes (ARGs)39. The identification of virulence factors associated with immune modulation, effector delivery systems, and biofilm formation is particularly concerning40. Immune modulation mechanisms can facilitate bacterial evasion of host immune responses, leading to persistent infections and increased transmission risks within farming environments41. The abundance of biofilm formation factors, which enhance bacterial adherence and resilience, indicates a potential for increased environmental persistence and resistance to disinfectants, further complicating disease control42. It is also evident that antimicrobial resistance is highly correlated with biofilm formation43.
Significant differences in virulence factor abundance among samples are primarily attributed to the distinct environmental conditions and microbial dynamics within each sample type. For instance, sediment provides a stable, nutrient-rich environment conducive to the proliferation of bacteria with diverse virulence traits44. This stability supports the maintenance and dissemination of both ARGs and virulence factors, explaining the higher abundance of immune modulation and effector delivery system factors in sediment45. Conversely, the high prevalence of biofilm formation factors in feed and specific gastrointestinal samples can be linked to the constant microbial exposure and physical conditions that favor biofilm development46. This enhances bacterial persistence and complicates control efforts47.
The discovery of motility and adherence mechanisms, particularly in samples from droppings and chicken gut, suggests that bacteria in these environments are equipped to colonize and spread within the host and across environmental surfaces48,49. These traits could lead to the dissemination of both pathogenic and resistant bacteria within and beyond the farming system, representing a public health risk50. Nutritional and metabolic factors, more abundant in sediment, highlight the role of environmental reservoirs in sustaining bacterial populations and serve as hotspots for the persistence and spread of both ARGs and virulence factors51.
The detection of exotoxin and invasion-related genes, although at lower levels, underscores the potential for harmful infections that could result from the spread of these virulence factors to humans or other animals52. The distinct distribution of virulence genes across different environmental compartments within the farming system reinforces the notion that environmental factors play a crucial role in the maintenance and dissemination of bacterial virulence and resistance53.
In the context of the integrated chicken and fish farming system, the analysis of mobile genetic elements (MGEs) provides critical insights into microbial dynamics and mechanisms of horizontal gene transfer54. The predominance of plasmids among the identified MGEs underscores their pivotal role in horizontal gene transfer, particularly in the dissemination of antimicrobial resistance genes55. Their substantial presence within the system underscores their central function in propagating genetic traits across bacterial communities, thereby contributing to dissemination of genes including ARGs56.
Insertion Sequences (IS) and Integrative and Conjugative Elements (ICE) are also evident within the system, albeit in lesser quantities relative to plasmids. IS elements are fundamental to the genome plasticity of microorganisms, driving genetic variability through insertional mutagenesis and subsequent genomic rearrangements57. On the other hand, ICEs, which possess the capacity to integrate into host genomes and facilitate conjugative transfer, serve a specialized role in gene transfer58. Despite their lower frequency, ICEs are integral to both vertical and horizontal gene dissemination processes within microbial communities59.
Additionally, Transposons (Tn), Chromosomal Islands (CN), and Integrative Mobilizable Elements (IME), which further augment the diversity of MGEs. The distribution patterns of these MGEs across varied sample types reveal considerable environmental influence on their prevalence. The elevated abundance of MGEs in droppings and sediment suggests these environments act as prominent hotspots for gene exchange and microbial activity, likely attributable to elevated microbial biomass and metabolic activity60. Conversely, the reduced MGE abundance in the fish intestine points to a diminished role in gene transfer within this specific environment. This variability in MGE distribution underscores the influence of environmental factors on microbial gene dynamics and elucidates the intricate interactions between microbial populations and their respective habitats61.
Transposons, such as Tn6072, Tn6246, and Tn4001, are critical vectors for the horizontal dissemination of antimicrobial resistance genes (ARGs)62. Their ability to transpose across different genomic locations facilitates the spread of resistance determinants such as tetM, sulI, and tetX, thereby contributing to the persistence and proliferation of antimicrobial resistance within microbial populations63. Integrative and conjugative elements (ICEs), including ICEBfrYCH46-1, play a crucial role in both vertical and horizontal gene transfer64. Their capacity for genome integration and conjugative transfer highlights their importance in the exchange of resistance genes across bacterial communities, further reinforcing the genetic adaptability and survival of microbial populations in the farming environment. The role of transposons and plasmids extends to the transfer of virulence genes, with elements such as Tn125 and the plasmid Nonomuraea gerenzanensis isolate nono1 plasmid I facilitating the dissemination of various virulence factors65.
The association between specific microbial taxa and MGEs further elucidates the mechanisms of genetic exchange within microbial communities. Genera such as Bacteroides, Clostridium, and Escherichia exhibit significant interactions with MGEs, indicating their integral role in the propagation and distribution of these genetic elements60. The distribution patterns of microbial phyla associated with MGEs reveal differential impacts on gene transfer processes across various environmental contexts. For instance, the predominance of Pseudomonadota in specific environments suggests a significant role in MGE-mediated genetic exchange66, while the limited presence of other phyla, such as Mycoplasmatota and Campylobacterota, indicates a more restricted involvement in these processes67.
This study provides a comprehensive reference genome for the chicken gut microbiome, offering critical insights into the diversity and functional potential of microbial communities within integrated farming systems. This resource serves as a foundation for comparative analyses across bacterial, archaeal, and viral domains, facilitating an in-depth exploration of the interplay between these microbial kingdoms within the gut environment. Such integrative analyses are essential for advancing our understanding of host-microbiota interactions and their roles in health and disease regulation in poultry systems68.
The reference genome also holds significant potential for microbiome-based interventions, including the development of probiotics, precision feed additives, and targeted prebiotics to enhance poultry health and productivity69,70. For example, probiotics specifically designed to align with the microbial composition of the chicken gut could effectively modulate the microbiota, reducing pathogenic colonization while improving nutrient absorption71,72. Moreover, this genomic dataset enables the identification of auxiliary metabolic genes involved in the degradation of complex feed substrates, offering new avenues for optimizing feed formulations72.
Integrating this reference genome with multi-omics approaches could facilitate the identification of microbial markers for antimicrobial resistance (AMR) surveillance and management in farming systems73. Such markers are instrumental in tracking the spread of resistance genes across interconnected environments, as highlighted in recent studies linking ARGs to specific microbial hosts and mobile genetic elements74. Additionally, the inclusion of viral genomic data underscores the pivotal role of bacteriophages as vectors for horizontal gene transfer, thereby amplifying the dissemination of ARGs within the gut microbiome75,76.
In contrast to earlier studies that predominantly explored the resistome diversity within chicken gut microbiomes, such as the work documenting the diversity of antibiotic resistance genes in live poultry markets and associated workers77 or the comprehensive multi-kingdom microbiome catalog of the chicken gastrointestinal tract78, this study broadens the scope by investigating the interplay of antimicrobial resistance genes (ARGs), virulence factors (VFGs), and mobile genetic elements (MGEs) across interconnected terrestrial and aquatic environments.
Additionally, this study uniquely emphasizes the contribution of environmental reservoirs, such as sediment, to the persistence and spread of ARGs, highlighting their role as long-term sinks for resistance genes. Unlike previous works that focused on specific host-associated microbiomes79,80, this study integrates cross-compartmental analysis, revealing how interconnected environments facilitate horizontal gene transfer via MGEs. Furthermore, the comprehensive assessment of VFGs alongside ARGs provides novel insights into the dual risk posed by pathogens capable of both resistance and heightened virulence, which has not been extensively explored in earlier studies.
By employing robust statistical analyses, such as the F-Welch and Games-Howell tests, this research also offers a quantitative evaluation of ARG distribution across sample types, strengthening the evidence for significant environmental differences in resistance gene abundance. This multi-dimensional approach contributes to a deeper understanding of how integrated farming practices shape microbial community dynamics and resistance profiles, setting this study apart from previous investigations.
In summary, the role of MGEs in the dissemination of ARGs and virulence genes is integral to understanding microbial adaptability and the dynamics of genetic exchange within integrated farming systems. The findings highlight the essential contribution of MGEs to microbial gene flow and the ecological complexity of microbial communities.
Conclusion
In conclusion, this reveals a concerning prevalence of antimicrobial resistance genes (ARGs) and virulence factors throughout the farming system, with MGEs playing a pivotal role in their dissemination. Poultry droppings emerge as the most significant source of both ARGs and virulence factors, underscoring the critical need for meticulous management of waste. In contrast, fish intestines and feed contribute minimally to the spread of resistance genes. Therefore, stringent antibiotic stewardship and careful handling of poultry droppings are essential measures recommended by this study.
Methodology
Ethical approval
The study was approved by the Ethical Review Committee of the Faculty of Biological Sciences at Jashore University of Science and Technology, Jashore, Bangladesh (certification number: ERC/FBST/JUST/2024–200). All experimental protocols were reviewed and approved by this committee. All methods were carried out in accordance with the ARRIVE (Animal Research: Reporting of In Vivo Experiments) and AVMA (American Veterinary Medical Association) guidelines. All possible measures were taken to minimize pain and harm to the experimental animals. All experimental methods were carried out in accordance with relevant guidelines and regulations.
Sample collection and total DNA extraction
Three integrated fish farms incorporating chicken farming, located in Solua Bazar, Jhikargacha, and Khajura within the Jashore district of Bangladesh, were selected for this study, designated as Farm-1, Farm-2, and Farm-3, respectively. Five distinct sample types were collected from each farm, including feed (n = 1), chicken gut (n = 2), chicken droppings (n = 2), fish intestine (n = 2), and pond sediment (n = 2), resulting in nine samples per farm and 27 samples in total. Feed, chicken droppings, and sediment were aseptically collected using sterile containers to prevent contamination. Chicken droppings were sampled immediately post-defecation using sterile spatulas. Chicken euthanasia was performed following the AVMA Guidelines for the Euthanasia of Animals (2020 Edition). Birds were placed in a CO₂ chamber prefilled to 30% of its volume, with CO₂ concentration increased at a rate of 30–70% of chamber volume per minute. Death was confirmed through the cessation of respiratory movements and absence of the corneal reflex. Dissection was performed using sterile instruments to obtain gut sections.
Fish euthanasia also adhered to AVMA guidelines, using a CO₂-enriched tank with gradual gas diffusion to ensure sedation and euthanasia. Fishes were selected and acclimated in laboratory tanks maintained at 26 ± 2 °C, with pH 7.0–7.5 and dissolved oxygen levels above 6 mg/L observations included loss of equilibrium, cessation of opercular movement, and secondary decapitation for confirmation of death. Intestine samples were dissected using sterile instruments. Pond sediment samples were collected from multiple pond locations using sterile scoops. All samples were stored at 4 °C immediately after collection and transported to the laboratory within 1–2 h. For downstream analysis, samples of the same type were pooled within each farm, resulting in five composite samples per farm. DNA extraction was performed using the DNeasy PowerSoil Pro Kit and QIAamp Fast DNA Stool Mini Kit (QIAGEN) per manufacturer protocols. Extracted DNA was stored at -20 °C for sequencing. Detailed sampling site and sample descriptions are available in Supplementary Table 1.
Library Preparation and shotgun metagenomic sequencing
Shotgun metagenomic libraries were prepared using the iNextEra library preparation protocol. Briefly, the DNA was tagmented with Illumina bead-linked transposomes (Illumina DNA library prep, Illumina, Inc., San Diego, CA, USA) and 2X TMP buffer81 in a total volume of 6 µL (3 µL of 2X TMP buffer, 0.5 µL of Bead-linked transposome and 2.5 µL of DNA) and incubation at 53 °C for 30 min. Following tagmentation, a limited-cycle PCR step completed the Illumina-compatible adapters with dual-indexes using custom iNextEra primers for sequencing of pooled libraries. Each 51.5 µL of PCR master mix included 40.25 µL HyClone water (Cytiva Life Sciences, Marlborough, MA, USA), 9.5 µL of Q5 Reaction Buffer (New England Biolabs, Ipswich, MA, USA), 1.25 µL of 10mM dNTPs (Roche Diagnostic Corporation, Indianapolis, IN, USA), and 0.5 µL of Q5 High-Fidelity DNA Polymerase (New England Biolabs, Ipswich, MA, USA). The master mix was then added to 6 µL of tagmentation mix. Finally, 2.5 µL of 5 µM iNextEra5 index primer and 2.5 µL of 5 µM iNextEra7 index primer were added to the PCR mix, creating unique index combinations in a total PCR volume of 62.5 µL. The limited-cycle PCR was done using a T100 thermal cycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA) with an initial strand repair at 72 °C for 3 min, denaturation at 98 °C for 3 min followed by 12 cycles consisting of 45 s at 98 °C, 30 s at 62 °C, and 2 min at 72 °C, concluding with a final extension at 72 °C for 1 min. After PCR, 5 µL of PCR product was run on an agarose gel to ensure each sample amplified, yielding a smear in the desired size-range (300–700 bp). Then, the libraries were quantified using Qubit Flex (Thermo Fisher Scientific, Waltham, MA USA), normalized, pooled and purified using AMPure XP beads (Beckman Coulter, Inc., Brea, CA, USA). The final pool was quantified and sent to Novogene (Novogene Corporation Inc., Sacramento, CA, USA) to obtain 2 × 150 bp paired-end reads using Illumina NovaSeq X plus sequencing system (Illumina, Inc., San Diego, CA, USA).
Quality control and identification of MGEs, ARGs, VFGs and bacterial taxa
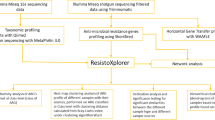
The generated FASTQ files were evaluated for quality using FastQC v0.11.682. Adapter sequences, and low-quality ends per read were trimmed by using Trimmomatic v0.3983 with a sliding window size of 4; a minimum average quality score of 20; minimum read length of 40 bp (base pairs). After quality control, there were an average of 129.63 million pairs of reads (Minimum = 89.15 million and Maximum = 204.47 million reads). The abundance of mobile genetic elements (MGEs), antimicrobial resistance genes (ARGs), and virulence factors were identified by AMR + + bioinformatic pipeline (https://github.com/Microbial-Ecology-Group/AMRplusplus)84. In this pipeline, the Burrows-Wheeler Aligner (BWA) was used to align sequence reads to host genomes, such as the chicken (Gallus gallus) GRCg6 and the common carp (Cyprinus carpio) GCF_018340385.1, in order to filter out these reads. Then the reads were aligned with the MobileElementFinder MGEdb database (v1.0.2)85, the MEGARes database (v2.0)84, the updated Virulence Factor Database (VFDB)86 as of August 2024, plasmid detection PLSDB (v. 2023_11_03_v2)87. Only genes that met a stringent identification threshold of 90%—defined as the percentage of genes matched by at least one sequencing read—were retained for analysis using the Resistome Analyzer (https://github.com/cdeanj/resistomeanalyzer). Bacterial taxonomic classification was conducted using CZID88 formerly known as IDseq, where only sequences with an NT% identity above 90% and alignment length (NRL) greater than 50 were considered. This classification process was further refined using Taxonkit89 for accurate taxonomic resolution.
Statistical analysis and data visualization
Statistical analyses and visualizations were performed using R Studio version 4.3.390. To ensure the normality of our data, we applied the Total Sum Scaling (TSS) normalization method. The TSS normalization equation is expressed as follows:
Where:
\(\:{X}_{ij}\) : The raw count or value of feature j in sample i. \(\:{\sum\:}_{j=1}^{n}\:{X}_{ij}\) : The sum of all feature values in sample i. \(\:{X}_{ij}^{TSS}\): The normalized value of feature j in sample i.
A range of plots was employed to analyze and present the data comprehensively. A chord diagram was constructed using the “circlize”91 package, with the “RColorBrewer”92 package providing enhanced color schemes. For visualizing distributions and conducting statistical comparisons, violin plots were created with the “ggstatsplot”93 package. This package allowed for the integration of statistical tests, including Welch’s test68, Kruskal-Wallis test94, Games-Howell test95, Dunn96 and Bayesian analysis97. Welch’s test was utilized to compare means between two groups assuming unequal variances, while the Games-Howell test was used for pairwise comparisons in cases of unequal variances and sample sizes. Bayesian analysis provided a probabilistic approach to inferential statistics, incorporating prior knowledge and estimating the credibility of hypotheses. Heatmaps were generated using the “pheatmap”98 package, with the “RColorBrewer” package contributing color schemes and clustering functionalities. Correlation heatmaps, depicting Spearman correlations99, were plotted with the “ggcorrplot”100 package, supported by the “ggplot2”101 package for enhanced visualization. Spearman correlation analysis was employed to assess monotonic relationships between variables, providing a non-parametric measure of association. The relative abundance plot was created using the “ggplot2” package, which facilitated the graphical representation of the proportion of different components within the dataset.
Limitations and future prospects
This study’s reliance on metagenomic analysis offers a broad genetic overview but lacks insight into the functional activity of identified genes. While the detection of antimicrobial resistance genes (ARGs) and virulence factors (VFGs) provides valuable information, the inability to validate their expression or phenotypic implications limits our understanding of their real-world impact. The research is geographically limited, focusing solely on integrated farming systems in Bangladesh, which may constrain the broader applicability of its findings to other regions with differing practices or environmental conditions. Additionally, the cross-sectional design captures only a single time point, potentially overlooking temporal variations in gene abundance and dynamics across seasons or farming cycles. Future studies should aim to integrate metagenomic data with transcriptomic, proteomic, or metabolomic analyses to link gene presence with functional activity and phenotypic outcomes. Expanding the scope to diverse geographical regions and varying integrated farming practices will provide a more comprehensive understanding of ARG and VFG dissemination. Additionally, adopting longitudinal study designs will help elucidate the temporal dynamics of these genes, offering deeper insights into their persistence and spread. Exploring the role of environmental co-factors, such as heavy metals and biocides, alongside ARGs, could also shed light on additional drivers of resistance proliferation.
Data availability
The data that support the findings of this study are openly available in NCBI BioProject at https://www.ncbi.nlm.nih.gov/bioproject/ reference number PRJNA1165771.
References
Bhagat, R. et al. The integrated farming system is an environmentally friendly and cost-effective approach to the sustainability of agri‐food systems in the modern era of the changing climate: A comprehensive review. Food Energy Secur. 13 (1), e534 (2024).
Al Mamun, S., Nasrat, F. & Debi, M. R. Integrated farming system: prospects in Bangladesh. J. Environ. Sci. Nat. Resour. 4 (2), 127–136 (2011).
Miller, S. A., Ferreira, J. P. & LeJeune, J. T. Antimicrobial use and resistance in plant agriculture: a one health perspective. Agriculture 12 (2), 289 (2022).
Pham, M. N. et al. Recent Advancement of Eliminating Antibiotic Resistance bacteria and Antibiotic Resistance Genes in Livestock Waste: A Review. 103751 (Environmental Technology & Innovation, 2024).
Zhou, N. et al. Global antimicrobial resistance: a system-wide comprehensive investigation using the global one health index. Infect. Dis. Poverty 11 (1), 92 (2022).
Manyi-Loh, C. et al. Antibiotic use in agriculture and its consequential resistance in environmental sources: potential public health implications. Molecules 23 (4), 795 (2018).
Lima, T., Domingues, S., Da, G. J. & Silva Manure as a potential hotspot for antibiotic resistance dissemination by horizontal gene transfer events. Veterinary Sci. 7 (3), 110 (2020).
Meng, M., Li, Y. & Yao, H. Plasmid-mediated transfer of antibiotic resistance genes in soil. Antibiotics 11(4): 525. (2022).
Che, Y. et al. Conjugative plasmids interact with insertion sequences to shape the horizontal transfer of antimicrobial resistance genes. Proc. Natl. Acad. Sci. 118 (6), e2008731118 (2021).
Geisinger, E. & Isberg, R. R. Interplay between antibiotic resistance and virulence during disease promoted by multidrug-resistant bacteria. J. Infect. Dis. 215 (suppl_1), S9–S17 (2017).
Schroeder, M., Brooks, B. D. & Brooks, A. E. The complex relationship between virulence and antibiotic resistance. Genes 8 (1), 39 (2017).
Thapa, S. P., Shrestha, S. & Anal, A. K. Addressing the antibiotic resistance and improving the food safety in food supply chain (farm-to-fork) in Southeast Asia. Food Control. 108, 106809 (2020).
Kanyerezi, S. et al. Metagenomics insights into the microbial resistome and virulome composition of Kampala’s wastewater. Open. Res. Afr. 7, 8 (2024).
Nwachukwu, B. C. & Babalola, O. O. Metagenomics: a tool for exploring key Microbiome with the potentials for improving sustainable agriculture. Front. Sustainable Food Syst. 6, 886987 (2022).
Nguyen, A. Q. et al. Monitoring antibiotic resistance genes in wastewater treatment: current strategies and future challenges. Sci. Total Environ. 783, 146964 (2021).
Majumder, M. A. A. et al. Antimicrobial stewardship: fighting antimicrobial resistance and protecting global public health. Infection and drug resistance. 4713–4738. (2020).
Ben, Y. et al. Human health risk assessment of antibiotic resistance associated with antibiotic residues in the environment: A review. Environ. Res. 169, 483–493 (2019).
Zalewska, M. et al. Antibiotics and antibiotic resistance genes in animal manure–consequences of its application in agriculture. Front. Microbiol. 12, 610656 (2021).
Ljubojević, D. et al. Resistance to Tetracycline in Escherichia coli isolates from poultry meat: epidemiology, policy and perspective. World’s Poult. Sci. J. 73 (2), 409–417 (2017).
Chen, H. et al. Characterization and source identification of antibiotic resistance genes in the sediments of an interconnected river-lake system. Environ. Int. 137, 105538 (2020).
Vera-Herrera, L., Romo, S. & Soria, J. How agriculture, connectivity and water management can affect water quality of a mediterranean coastal wetland. Agronomy 12 (2), 486 (2022).
Paul, S. S. et al. Effects of dietary antimicrobial growth promoters on performance parameters and abundance and diversity of broiler chicken gut Microbiome and selection of antibiotic resistance genes. Front. Microbiol. 13, 905050 (2022).
Page, S. & Gautier, P. Use of antimicrobial agents in livestock. Revue Scientifique Et Technique-OIE 31 (1), 145 (2012).
Nasibullah, M. et al. Recent developments in Oxazolidinones as potent antibacterials. Adv. Sci. Eng. Med. 7 (2), 91–111 (2015).
Terreni, M., Taccani, M. & Pregnolato, M. New antibiotics for multidrug-resistant bacterial strains: latest research developments and future perspectives. Molecules 26 (9), 2671 (2021).
Osei Sekyere, J. Current state of resistance to antibiotics of last-resort in South Africa: a review from a public health perspective. Front. Public. Health 4, 209 (2016).
Mohapatra, S. S., Dwibedy, S. K. & Padhy, I. Polymyxins, the last-resort antibiotics: mode of action, resistance emergence, and potential solutions. J. Biosci. 46 (3), 85 (2021).
Li, W. et al. Evaluation of culturable ‘last-resort’antibiotic resistant pathogens in hospital wastewater and implications on the risks of nosocomial antimicrobial resistance prevalence. J. Hazard. Mater. 438, 129477 (2022).
Schwarz, S., Loeffler, A. & Kadlec, K. Bacterial resistance to antimicrobial agents and its impact on veterinary and human medicine. Adv. Vet. Dermatol. 8, 95–110 (2017).
Zhou, L. J. et al. Trends in the occurrence and risk assessment of antibiotics in shallow lakes in the lower-middle reaches of the Yangtze river basin, China. Ecotoxicol. Environ. Saf. 183, 109511 (2019).
Li, J. et al. Global review of macrolide antibiotics in the aquatic environment: sources, occurrence, fate, ecotoxicity, and risk assessment. J. Hazard. Mater. 439, 129628 (2022).
Fletcher, S. Understanding the contribution of environmental factors in the spread of antimicrobial resistance. Environ. Health Prev. Med. 20, 243–252 (2015).
Schmutz, E. et al. Resistance genes of aminocoumarin producers: two type II topoisomerase genes confer resistance against coumermycin A1 and clorobiocin. Antimicrob. Agents Chemother. 47 (3), 869–877 (2003).
Amarasekara, N. R. et al. Exploring the co-occurrence of antibiotic, metal, and biocide resistance genes in the urban agricultural environment. J. Agric. Food Res. 11, 100474 (2023).
Pal, C. Effects of biocides and metals on antibiotic resistance: a genomic and metagenomic perspective. (2017).
Harms, A., Maisonneuve, E. & Gerdes, K. Mechanisms of bacterial persistence during stress and antibiotic exposure. Science 354 (6318), aaf4268 (2016).
Li, L. G. et al. Source tracking of antibiotic resistance genes in the environment—Challenges, progress, and prospects. Water Res. 185, 116127 (2020).
Wang, Y. et al. Integrated metagenomic and metatranscriptomic profiling reveals differentially expressed resistomes in human, chicken, and pig gut microbiomes. Environ. Int. 138, 105649 (2020).
Adeyemi, F. M. et al. Integrated poultry-fish farming system encourages multidrug-resistant gram-negative bacteria dissemination in pond environment and fishes. Aquaculture 548, 737558 (2022).
Ren, C. et al. Distribution and pathogenic relationship of virulence associated genes among Vibrio alginolyticus from the mariculture systems. Mol. Cell Probes 27 (3–4), 164–168 (2013).
Hornef, M. W. et al. Bacterial strategies for overcoming host innate and adaptive immune responses. Nat. Immunol. 3 (11), 1033–1040 (2002).
Römling, U. & Balsalobre, C. Biofilm infections, their resilience to therapy and innovative treatment strategies. J. Intern. Med. 272 (6), 541–561 (2012).
Qi, L. et al. Relationship between antibiotic resistance, biofilm formation, and biofilm-specific resistance in Acinetobacter baumannii. Front. Microbiol. 7, 483 (2016).
Zhang, H. et al. Pollution gradients shape the co-occurrence networks and interactions of sedimentary bacterial communities in Taihu lake, a shallow eutrophic lake. J. Environ. Manage. 305, 114380 (2022).
Zhang, Q. Q., Tian, G. M. & Jin, R. C. The occurrence, maintenance, and proliferation of antibiotic resistance genes (ARGs) in the environment: influencing factors, mechanisms, and elimination strategies. Appl. Microbiol. Biotechnol. 102, 8261–8274 (2018).
von Rosenvinge, E. C. et al. Microbial biofilms and Gastrointestinal diseases. Pathogens Disease 67 (1), 25–38 (2013).
Michiels, J. E. et al. Molecular mechanisms and clinical implications of bacterial persistence. Drug Resist. Updates 29, 76–89 (2016).
Mappley, L. J. et al. Lactobacilli antagonize the growth, motility, and adherence of Brachyspira pilosicoli: a potential intervention against avian intestinal spirochetosis. Appl. Environ. Microbiol. 77 (15), 5402–5411 (2011).
Costerton, J. et al. The role of bacterial surface structures in pathogenesis. CRC Crit. Reviews Microbiol. 8 (4), 303–338 (1981).
Ahmad, N., Joji, R. M. & Shahid, M. Evolution and implementation of one health to control the dissemination of antibiotic-resistant bacteria and resistance genes: A review. Front. Cell. Infect. Microbiol. 12, 1065796 (2023).
Wobus, A. et al. Microbial diversity and functional characterization of sediments from reservoirs of different trophic state. FEMS Microbiol. Ecol. 46 (3), 331–347 (2003).
Huynh, M. H., Harper, J. M. & Carruthers, V. B. Preparing for an invasion: charting the pathway of adhesion proteins to Toxoplasma micronemes. Parasitol. Res. 98, 389–395 (2006).
Bengtsson-Palme, J., Kristiansson, E. & Larsson, D. J. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol. Rev. 42 (1), fux053 (2018).
Li, S. et al. Comprehensive insights into antibiotic resistance gene migration in microalgal-bacterial consortia: Mechanisms, factors, and perspectives. Sci. Total Environ. 166029. (2023).
Michaelis, C. & Grohmann, E. Horizontal gene transfer of antibiotic resistance genes in biofilms. Antibiotics 12 (2), 328 (2023).
Jian, Z. et al. Antibiotic resistance genes in bacteria: occurrence, spread, and control. J. Basic Microbiol. 61 (12), 1049–1070 (2021).
Vandecraen, J. et al. The impact of insertion sequences on bacterial genome plasticity and adaptability. Crit. Rev. Microbiol. 43 (6), 709–730 (2017).
Cury, J. et al. Host range and genetic plasticity explain the coexistence of integrative and extrachromosomal mobile genetic elements. Mol. Biol. Evol. 35 (9), 2230–2239 (2018).
Cury, J. et al. Host range expansion and genetic plasticity drive the trade-off between integrative and extrachromosomal mobile genetic elements. bioRxiv 250266. (2018).
Aminov, R. I. Horizontal gene exchange in environmental microbiota. Front. Microbiol. 2, 158 (2011).
Javvadi, Y. & Mohan, S. V. Temporal dynamics and persistence of resistance genes to broad spectrum antibiotics in an urban community. Npj Clean. Water 7 (1), 56 (2024).
Babakhani, S. & Oloomi, M. Transposons: the agents of antibiotic resistance in bacteria. J. Basic Microbiol. 58 (11), 905–917 (2018).
Scott, K. The role of conjugative transposons in spreading antibiotic resistance between bacteria that inhabit the Gastrointestinal tract. Cell. Mol. Life Sci. CMLS 59, 2071–2082 (2002).
Bean, E. L. et al. Biology and engineering of integrative and conjugative elements: construction and analyses of hybrid ICEs reveal element functions that affect species-specific efficiencies. PLoS Genet. 18 (5), e1009998 (2022).
Drongitis, D. et al. Roles of transposable elements in the different layers of gene expression regulation. Int. J. Mol. Sci. 20 (22), 5755 (2019).
Panwar, S. et al. Toxin-linked mobile genetic elements in major enteric bacterial pathogens. Gut Microbiome 4, e5 (2023).
Oh, S., Lee, H. J. & Park, K. U. Metagenomic characterization of the microbiomes in five different body habitats of otherwise healthy individuals with periodontal disease. Front. Cell. Infect. Microbiol. 13, 1257816 (2023).
Lu, Z. & Yuan, K. H. Welch’s t test pp. 1620–1623. (2010).
Al-Nijir, M. et al. Metabolic modelling uncovers the complex interplay between fungal probiotics, poultry microbiomes, and diet. Microbiome 12 (1), 267 (2024).
Dasriya, V. L. et al. Modulation of gut-microbiota Through Probiotics and Dietary Interventions To Improve Host Health. (Journal of the Science of Food and Agriculture, 2024).
Fathima, S. et al. Gastrointestinal microbiota and their manipulation for improved growth and performance in chickens. Foods 11 (10), 1401 (2022).
Shehata, A. A. et al. Probiotics, prebiotics, and phytogenic substances for optimizing gut health in poultry. Microorganisms 10 (2), 395 (2022).
Bartelo, N. et al. Future Prospective of Omics-System Biology To Control AMR: Recommendations and Directions, in Antimicrobial Resistance: Factors To Findings: Omics and Systems Biology Approaches. 415–449 (Springer, 2024).
Mao, X. et al. Standardization in global environmental antibiotic resistance genes (ARGs) surveillance. Critical Reviews in Environmental Science and Technology. pp. 1–18. (2024).
Ji, Y. et al. Metagenomics analysis reveals potential pathways and drivers of piglet gut phage-mediated transfer of ARGs. Sci. Total Environ. 859, 160304 (2023).
Kang, Y. et al. Unveiling the genomic diversity and ecological impact of phage communities in hospital wastewater. J. Hazard. Mater. 477, 135353 (2024).
Wang, Y. et al. More diversified antibiotic resistance genes in chickens and workers of the live poultry markets. Environ. Int. 153, 106534 (2021).
Wang, Y. et al. The multi-kingdom Microbiome catalog of the chicken Gastrointestinal tract. Biosaf. Health 6 (02), 101–115 (2024).
Inda-Díaz, J. S. et al. Latent antibiotic resistance genes are abundant, diverse, and mobile in human, animal, and environmental microbiomes. Microbiome 11 (1), 44 (2023).
Gan, D. et al. Housefly gut microbiomes as a reservoir and facilitator for the spread of antibiotic resistance. ISME J. 18 (1), wrae128 (2024).
Jones, A. et al. Cost-conscious generation of multiplexed short-read DNA libraries for whole-genome sequencing. Plos One 18 (1), e0280004 (2023).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. Cambridge, UK: Babraham Bioinformatics, Babraham Institute. (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30 (15), 2114–2120 (2014).
Bonin, N. et al. MEGARes and AMR++, v3. 0: an updated comprehensive database of antimicrobial resistance determinants and an improved software pipeline for classification using high-throughput sequencing. Nucleic Acids Res. 51 (D1), D744–D752 (2023).
Johansson, M. H. et al. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: mobileelementfinder. J. Antimicrob. Chemother. 76 (1), 101–109 (2021).
Chen, L. et al. VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res. 33 (suppl_1), D325–D328 (2005).
Schmartz, G. P. et al. PLSDB: advancing a comprehensive database of bacterial plasmids. Nucleic Acids Res. 50 (D1), D273–D278 (2022).
Simmonds, S. E. et al. CZ ID: a cloud-based, no-code platform enabling advanced long read metagenomic analysis. bioRxiv, 2024-02 (2024).
Shen, W. & Ren, H. TaxonKit: A practical and efficient NCBI taxonomy toolkit. J. Genet. Genomics 48 (9), 844–850 (2021).
Team, R. S. RStudio: Integrated Development Environment for R. (2020).
Gu, Z. et al. Circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Neuwirth, E. RColorBrewer: colorbrewer palettes.
Patil, I. Visualizations with statistical details: the ‘ggstatsplot’ approach. J. Open. Source Softw. 6 (61), 3167 (2021).
McKight, P. E. & Najab, J. Kruskal-wallis test. The corsini encyclopedia of psychology. 1–1. (2010).
Games, P. A. & Howell, J. F. Pairwise multiple comparison procedures with unequal N’s and/or variances: A Monte Carlo study. J. Educ. Stat. 1 (2), 113–125 (1976).
Dinno, A. Nonparametric pairwise multiple comparisons in independent groups using Dunn’s test. Stata J. 15 (1), 292–300 (2015).
Kleibergen, F. & Zivot, E. Bayesian and Classical Approaches To Instrumental Variable Regression (Department of Economics at the University of Washington, 1998).
Kolde, R. & Kolde, M. R. Package ‘pheatmap’. R Package 1 (7), 790 (2015).
Spearman rank correlation coefficient. In The Concise Encyclopedia of Statistics. Springer New York: New York, NY. pp. 502–505. (2008).
Kassambara, A. & Kassambara, M. A. Package ‘ggcorrplot’. R package version 0.1. 3(3): 908 (2019).
Wickham, H., Chang, W. & Wickham, M. H. Package ‘ggplot2’. Create elegant data visualisations using the grammar of graphics. Version 2 (1), 1–189 (2016).
Acknowledgements
This study received valuable instrument support from the Department of Microbiology, Jashore University of Science and Technology. Reagents and consumables were supplied through partial grants from the Ministry of Science and Technology, Government of Bangladesh, and the Faculty of Biological Sciences, Jashore University of Science and Technology. We sincerely thank Dr. Travis C. Glenn (Department of Environmental Health Science, University of Georgia, USA) for reviewing the iNextEra library preparation protocol and making key suggestions.
Funding
This research was funded by Jashore University of Science and Technology, Jashore-7408, Bangladesh (Grant number: JUST/Research Cell/Research Project/2022-23/22-FoBST 05). We also extend our gratitude to the Ministry of Science and Technology (MOST, Fiscal Year 2022-23, Project ID: SRG-221252) for their support of this project.
Author information
Authors and Affiliations
Contributions
SMK carried out the bioinformatics analysis, generated figures, curated data, and drafted the original manuscript. KTI performed the analysis of ARGs and MGEs, formatted taxonomic data, and contributed to manuscript writing. RNR analyzed virulence genes, prepared data for plotting, and contributed to manuscript writing. MIUB performed shotgun sequencing and contributed to data analysis, manuscript review, and editing. TC participated in data analysis, manuscript reviewing, and editing. MSR OKI and MTI developed the research concept, finalized the bioinformatics pipelines, interpreted the results, and contributed to manuscript review and editing. MTI conceived the research, secured reagent support, thoroughly reviewed the manuscript, and led the overall project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Declaration of generative AI and AI-assisted technologies in the writing process
During the preparation of this work the author(s) used ChatGPT 4.0/Paraphrasing service in order to improve writing quality. After using this service, the author(s) reviewed and edited the content as needed and take(s) full responsibility for the content of the publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kador, S.M., Islam, K.T., Rubaiyat, R.N. et al. Abundance and transmission of antibiotic resistance and virulence genes through mobile genetic elements in integrated chicken and fish farming system. Sci Rep 15, 20953 (2025). https://doi.org/10.1038/s41598-025-92921-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92921-w