Abstract
Pregnant women undergoing assisted reproductive technology (ART) have an increased risk of gestational diabetes mellitus (GDM), potentially linked to altered immune regulation, but the underlying circulating immune features remain unclear. Here we show an integrated single-cell transcriptomic, immune receptor repertoire, and plasma proteomic analysis of peripheral blood from 32 women with GDM and 31 normal pregnancies after ART. GDM is associated with increased proportions of CD8+ T cells and elevated plasma levels of CD6, CXCL5, MMP10, and 4E-BP1. Cytotoxic CD8+ T cell subsets display enhanced effector and cytotoxic activity, while B cells, monocytes, and natural killer cells exhibit activated phenotypes. Transcription factors from the FOS and JUN families and KLF6 are implicated in immune activation, accompanied by distinct T cell and B cell receptor repertoire features. The hypothesis-generating multi-omic results highlight potential therapeutic targets and offered insights for future research and management ART-associated pregnancy complications.
Similar content being viewed by others
Introduction
Gestational diabetes mellitus (GDM) refers to abnormal glucose intolerance that that first develops or is first recognized during pregnancy but falls below the diagnostic thresholds for overt diabetes1. As one of the most common metabolic complications of pregnancy, GDM is estimated to affect approximately 14% of pregnant women worldwide2. Hyperglycemia during pregnancy has been shown to be strongly associated with adverse maternal and neonatal outcomes, including spontaneous abortion, gestational hypertension, preeclampsia/eclampsia, cesarean section, preterm birth, large-for-gestational-age (LGA) offspring, perinatal asphyxia, neonatal respiratory distress, shoulder dystocia/birth trauma, neonatal hypoglycemia, neonatal jaundice, admission to the neonatal intensive care unit, and even stillbirth/neonatal death3,4,5,6,7. Beyond these immediate pregnancy complications, GDM is also associated with long-term metabolic sequelae, including disorders of glucose metabolism, dyslipidemia, cardiometabolic diseases, and increased mortality risk in both mothers and offspring8,9,10,11.
Infertility represents a growing global health challenge, affecting approximately 17.5% of the adult population worldwide according to a recent World Health Organization (WHO) report12. Assisted reproductive technology (ART) has become an effective therapeutic approach for addressing fertility issues13, with over 10 million children estimated to have been born through ART worldwide14. Notably, women with singleton pregnancies conceived through ART exhibit a significantly higher risk of GDM compared with those who conceive spontaneously15,16,17. Multiple factors contribute to this increased risk, including underlying subfertility-related conditions such as advanced maternal age, elevated body mass index (BMI), polycystic ovary syndrome (PCOS), and other endocrine disorders that are more prevalent in women seeking ART18,19. Beyond these established risk factors, emerging evidence suggests that exposure to fertility-enhancing drugs is associated with a significantly higher risk of developing GDM compared to spontaneous conception20. Retrospective cohort study of women undergoing ART have demonstrated that hormonal disturbances during the ART cycle significantly elevated the risk of GDM21. However, the underlying mechanisms driving the increased GDM risk in ART pregnancies remain poorly understood.
Accumulating evidence indicates that immune dysregulation plays a critical role in GDM pathogenesis. Alterations in immune cell populations, elevated levels of circulating proinflammatory cytokines, and chronic low-grade inflammation have been observed in GDM22. For example, increased proportions of natural killer (NK) cells and macrophages have been documented in patients with GDM23. Single-cell RNA sequencing (scRNA-seq) analysis of regulatory T (Treg) cells isolated from peripheral blood mononuclear cells (PBMCs) revealed no significant difference in the proportions of Treg subtypes between GDM patients and controls24. However, both naïve and effector Treg cells exhibited reduced expression levels of the transcription factors JUN, FOS, and EGR1, which are crucial for Treg proliferation and differentiation24. GDM patients also showed significant increases in the proportions of IL-8+ IL-1β+ monocytes and CXCL8+ FCGR3B+ neutrophils25. Furthermore, these monocytes and neutrophils exhibited a proinflammatory transcriptional profile with upregulated genes related to the TNF-α, NF-κB, and chemokine signaling pathways25.
The intersection of infertility, ART treatment, and immune dysregulation presents a complex pathophysiological landscape. Factors related to infertility and reproductive disorders, as well as ART treatments themselves, may contribute to immune alterations through shared hormonal abnormalities and inflammatory pathways26,27,28. Studies using mouse models of assisted reproduction have reported enhanced placental immune inflammation and oxidative stress responses29. Similarly, human placental samples from ART pregnancies exhibit differential gene expression profiles enriched in inflammation, oxidative stress, and metabolic pathways compared with those from spontaneously conceived pregnancies30. Importantly, the relationship between immune dysregulation and GDM is likely bidirectional: while pre-existing immune alterations may contribute to metabolic dysfunction, hyperglycemia and insulin resistance can also directly impact maternal immune responses through multiple mechanisms, including altered glucose metabolism in immune cells, production of advanced glycation end products, and activation of inflammatory signaling pathways31,32,33. Despite these insights, the specific composition of immune cell subsets, immunological functions, and cytokine profiles associated with GDM in women undergoing ART remain largely uncharacterized.
To address this knowledge gap, we employed scRNA-seq to generate a comprehensive peripheral blood immune cell atlas of pregnant women (24–28 weeks of gestation) undergoing ART. In parallel, we prospectively profiled inflammatory markers in women with GDM and matched controls with normal pregnancies (NP) undergoing ART across three critical time points: the pre-pregnancy period, 4–6 weeks of gestation, and 24–28 weeks of gestation, using the Olink 96 inflammation panel. This integrated approach, combining single-cell and plasma proteomic analyses, provides a comprehensive characterization of the distinct circulating immunological signatures associated with GDM following ART.
Results
Study design and clinical characteristics of the participants
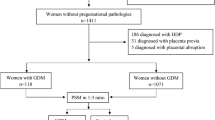
In this study, we recruited a total of 63 primiparous participants with singleton pregnancies undergoing ART from the Ningbo Birth Cohort of Population Undergoing Assisted Reproductive Technology (NBart)34, including 32 pregnant women diagnosed with GDM and 31 matched women with NP (Fig. 1A). The participants were enrolled prior to the initiation of ART, and peripheral blood samples and clinical data were collected dynamically during the baseline (pre-pregnancy), clinical pregnancy (4–6 weeks of gestation), and second trimester (24–28 weeks of gestation). We performed a single-cell RNA sequencing analysis and 5′ V(D)J sequencing on PBMCs obtained from 10 women with GDM and 10 matched women with NP at 24 to 28 weeks of gestation to delineate the distinct circulating immune features of GDM patients after ART. Additionally, we quantified the plasma levels of 92 inflammatory factors using the Olink 96 inflammation panel in all participants (32 with GDM and 31 with NP) throughout the three periods. The baseline and pregnancy characteristics of the participants in both groups were comparable, except for slightly elevated baseline fasting blood glucose (FBG) levels and blood pressure among women with GDM (Table 1).
A Schematic overview of the study design. B Uniform manifold approximation and projection (UMAP) plot depicting 26 clusters of high-quality PBMCs (n = 158,241) identified by unsupervised clustering analysis. PBMCs were obtained from ten women with GDM and ten women with NP at 24–28 weeks of gestation. C UMAP plot showing distinct immune cell types annotated based on canonical marker genes. DCs dendritic cells, mDCs myeloid dendritic cells. D UMAP plots illustrating the expression levels of canonical marker genes for distinct cell types. E Proportions of each immune cell type across study participants. Bars are color-coded by cell type. F Comparison of the percentages of distinct immune cell types between women with GDM (n = 10) and NP (n = 10). p values were determined using the Wilcoxon rank-sum test. Boxplots show the median (center line), interquartile range (box), and whiskers indicating the minimum and maximum values. Each dot represents one biologically independent sample.
Single-cell transcriptomic profiling of heterogeneous peripheral immune cells in patients with GDM
Single-cell expression profiles were generated using the 10× Genomics platform for PBMCs obtained from 10 participants with GDM and 10 participants with NP at 24–28 weeks of gestation, with well-matched demographic and clinical characteristics between the two groups (Supplementary Table 1). After the removal of low-quality cells (Supplementary Figs. 1 and 2), a total of 158,241 PBMCs were obtained for further analysis, including 91,055 cells from the GDM group and 67,186 cells from the NP group (Supplementary Table 2). The high-quality cells were integrated, normalized, and subsequently subjected to principal component analysis (PCA) and harmony batch effect correction processing (Supplementary Fig. 3).
High-quality cells were divided into 26 distinct clusters using uniform manifold approximation and projection (UMAP) (Fig. 1B). These clusters were further annotated into 12 main immune cell types according to the expression of canonical marker genes (Fig. 1C, D and Supplementary Fig. 4), including CD4+ T cells (CD3D+, CD3G+, CD3E+, CD4+, 20.01%)35, CD8+ T cells (CD3D+, CD3G+, CD3E+, CD8A+, CD8B+, 24.20%)35, γδ T cells (CD3D+, CD3G+, CD3E+, TRDV2+, TRGV9+, 4.06%)36, B cells (CD19+, MS4A1+, CD79A+, CD79B+, 6.88%)37, plasma B cells (MZB1+, JCHAIN+, 0.42%)37, CD14+ monocytes (CD68+, CD14+, LYZ+, 25.42%)36,38, CD16+ monocytes (CD68+, FCGR3A+, 3.45%)36,38, dendritic cells (DCs) (LYZ+, CST3+, 2.58%)39, myeloid dendritic cells (mDCs) (FCER1A+, CD1C+, LYZ+, 0.27%)37, megakaryocytes/platelets (PPBP+, PF4+, GP9+, 2.92%)37,39, natural killer (NK) cells (KLRF1+, KLRD1+, NCAM1+, 6.63%)37, proliferative T (pro T) cells (MKI67+, TYMS+, 0.35%)36,37, and several mixed cells (2.81%). We calculated the proportions of the main cell types in each subject and compared their compositions between the GDM and NP groups to evaluate the circulating immune states of women with GDM after undergoing ART (Fig. 1E and Supplementary Data 1). The results revealed an increased proportion of CD8+ T cells in women with GDM, whereas no significant differences were observed in other immune cell types (Fig. 1F).
T cells are the main circulating immune cell type and are indispensable in the adaptive immune response throughout pregnancy39. In this study, T cells (n = 76,942) were separately classified into 14 subpopulations based on their transcriptomic signatures. (Supplementary Figs. 5 and 6). Using specific marker genes, we annotated 11 distinct T cell subtypes, including CD4+ naïve T cells (CD4+, CCR7+, SELL+)37,40, CD4+ memory T cells (CD4+, S100A4+, TNFRSF4+, TIMP1+, GPR183+)36,37,41, CD4+ Treg (CD4+, FOXP3+) cells36,37, CD4+ exhausted-like T cell (CD4+, FOXO1+, IL7R high, CCR7−)42, CD8+ naïve T cells (CD8A+, CD8B+, CCR7+, SELL+)36, CD8+ cytotoxic T lymphocyte (CTL)-GZMK (CD8A+, CD8B+, GZMA+, GZMK+)39,43,44,45, CD8+ CTL-GNLY (CD8A+, CD8B+, GZMA+, GNLY+)36,44,45, CD8+ mucosal-associated invariant T (MAIT) cells (CD8A+, TRAV1-2+, SLC4A10+), CD8+ NKT cells (CD8B low, TYROBP+, KLRD1+, KLRF1+)36,37, pro T cells (MKI67+, TYMS+), and γδ T cells (TRDV2+, TRGV9+) (Fig. 2A, B). The distribution and proportions of T cell subsets in each subject are presented in Fig. 2C; however, no statistically significant differences were observed between the GDM group and the NP group (Fig. 2D and Supplementary Data 1).
A Uniform manifold approximation and projection (UMAP) plot illustrating 11 T cell subtypes annotated based on canonical marker genes. B Violin plots showing the expression of selected canonical marker genes across T cell subtypes. C Relative proportions of each T cell subtype among study participants. Bars are color-coded by T cell subtype. D Comparison of the percentage of distinct T cell subtypes between women with GDM (n = 10) and NP (n = 10). p values are determined using the Wilcoxon rank-sum test. Boxplots indicate the median (center line), interquartile range (box), and minimum and maximum values (whiskers). Each dot represents one biologically independent sample. E Pseudo-time trajectory of CD8+ T cell subtypes. Cells were ordered along an inferred pseudotime trajectory and visualized in reduced-dimensional space. Each dot represents a single cell. In the upper panel, cells are colored by pseudotime, indicating a continuous transition from early to late cellular states. In the middle panel, cells are colored by inferred cell states along the trajectory. In the lower panel, cells are colored by annotated CD8⁺ T cell subtypes, illustrating the distribution of different functional subsets along the differentiation trajectory. F Scatter plots with fitted curves showing the expression dynamics of canonical marker genes for CD8+ T cells along the pseudotime. Each dot represents a single cell, colored by cell state as defined in (E). Black curves indicate smoothed expression trends fitted along pseudotime. G Heatmap illustrating dynamic expression changes of differentially expressed transcription factor (TF) genes along pseudotime. Cells were ordered from early to late states along the horizontal axis according to pseudotime. Each row represents a gene, and gene expression levels were scaled by Z-score across pseudotime for visualization. Genes were grouped into clusters based on the similarity of their expression dynamics along pseudotime, representing distinct gene modules with coordinated temporal regulation. Color intensity indicates relative expression levels, with blue representing lower expression and yellow to red representing higher expression. White horizontal lines separate different gene clusters.
Additionally, we next evaluated the differentiation states of CD4+ T cell and CD8+ T cell subtypes by performing a pseudotime analysis. We observed a continuous state of differentiated CD8+ T cells and a trajectory of CD8+ T cells initiated from CD8+ naïve T cells, with the differentiation stage of CD8+ CTL-GZMK T cells in an intermediate state bridging between naïve T cells and more highly activated CD8+ CTL-GNLY T cells and CD8+ NKT cells (Fig. 2E). In accordance with the functional characteristics of cells, early markers such as CCR7 were expressed at the beginning of this trajectory, while GZMK expression was rapidly upregulated upon activation but quickly decreased. Conversely, GZMA, PRF1, FGFBP2, CCL5, GNLY, and NKG7 exhibited peak expression during the intermediate stages of the trajectory in CD8+ CTL-GNLY T and CD8+ NKT cells (Fig. 2F). Transcription factors (TFs) exhibited differential expression patterns with a continuous trajectory in CD8+ T cells. Specifically, in the intermediate cell state, JUND, FOS, CHD2, ZEB1, ETS1, and CUX1 were relatively highly expressed in CD8+ T cells, suggesting their potential roles in regulating immune molecules within these cells (Fig. 2G). The trajectory of CD4+ T cells started with CD4+ naïve T cells, followed by transitional T cells such as CD4+ memory T cells and CD4+ Tregs, and eventually led to CD4+ exhausted-like T cells (Supplementary Fig. 7A). We identified 31 TFs that affected the pseudotime trajectory of CD4+ T cells. Among these TFs, LEF1, FOXP1, FOXO1, ETS1, MAX, BPTF, BCLAF1, FOXN3, TCF12, ZEB1, JAZF1, RUNX2, STAT1, FLI1, ELF1, RORA, IRF1 and RAD21 were progressively activated during the differentiation process of CD4+ T cells (Supplementary Fig. 7B). Notably, overfitting problem was seen in the trajectory analysis as CD8+ naive cells were splitted into two clusters in which the minor cluster was probably formed by lower quality cells (Fig. 2E). Similar issue was seen in the trajectory analysis of CD4+ cells (Supplementary Fig. 7A).
Enhanced T cell immune response in GDM patients
We further analyzed the differentially expressed genes in T cells between the GDM group and the NP group and performed a functional enrichment analysis to characterize the T cell immune features associated with GDM. The results revealed significant transcriptomic changes in the T cells from the GDM group compared with those from the NP group (Fig. 3A, Supplementary Fig. 8 and Supplementary Data 1), and the gene set enrichment analysis (GSEA) revealed activated immune and inflammatory responses in the GDM group (Fig. 3B, C). We next performed gene set scoring to assess immune and inflammatory processes at the single-cell level, thereby evaluating the immune status of each T cell subtype and its role in GDM. Compared with total T cells, CD8+ CTL-GNLY, CD8+ NKT cells and CD8+ CTL-GZMK presented apparently higher gene set scores for defined effector genes, defined cytotoxicity genes, the immune response-regulating signaling pathway, the regulation of T cell activation, the positive regulation of lymphocyte activation, the T cell receptor signaling pathway, leukocyte activation involved in the inflammatory response, and the immune response regulating the cell surface receptor signaling pathway (Fig. 3D–H and Supplementary Fig. 9F–H). Additionally, CD8+ CTL-GNLY cells presented higher gene set scores for cytokine signaling pathways, including the Th1_th2 pathway, the IL17 pathway and the defined cytokine score (Supplementary Fig. 9B–D). CD8+ CTL-GZMK presented elevated scores for specific gene sets associated with inflammation, cytokine production, and TNF signaling (Supplementary Fig. 9A, B, E). Although γδ T cells and CD8+ MAIT cells constituted relatively minor proportions of the total population, they presented more pronounced immune features (Fig. 3D–H and Supplementary Fig. 9). This study revealed that relatively high gene set scores were associated with inflammation, cytokine production, and TNF signaling in CD4+ memory T cells (Supplementary Fig. 9).
A Volcano plot depicting differentially expressed genes (DEGs) in T cells between the GDM and NP groups. Genes with |log2(FC)| ≥0.25 and adjusted p < 0.05 are labeled with their respective gene symbols. B Gene Ontology (GO) enrichment analysis of DGEs in total T cells between the GDM and NP groups. C Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of DGEs in T cells between the GDM and NP groups. Gene Set Enrichment Analysis (GSEA) was performed using GO terms and KEGG pathways. D Boxplots showing defined-effector scores across different T cell subtypes. E Boxplots showing defined-cytotoxicity scores across different T cell subtypes. F Boxplots showing immune response regulating signaling pathway scores across different T cell subtypes. G Boxplots showing regulation of T cell activation scores across different T cell subtypes. H Boxplots showing positive regulation of lymphocyte activation scores across different T cell subtypes. I Comparison of defined effector scores differences between the GDM and NP groups in CD8⁺ NKT, CD8⁺ CTL-GZMK, and CD8⁺ CTL-GNLY cell subtypes. J Comparison of defined-cytotoxicity scores differences between the GDM and NP groups in CD8⁺ NKT, CD8⁺ CTL-GZMK, and CD8⁺ CTL-GNLY cell subtypes. K Comparison of immune response regulating signaling pathway scores differences between the GDM and NP groups in CD8⁺ NKT, CD8⁺ CTL-GZMK, and CD8⁺ CTL-GNLY cell subtypes. L Comparison of regulation of T cell activation scores differences between the GDM and NP groups in CD8+ NKT, CD8+ CTL-GZMK and CD8+ CTL-GNLY cell subtypes. M Comparison of positive regulation of lymphocyte activation scores between the GDM and NP groups in CD8+ NKT, CD8+ CTL-GZMK and CD8+ CTL-GNLY cell subtypes. The Wilcoxon rank-sum test was used to assess score differences between each T cell subtype and total T cells, as well as between the GDM and NP groups. Statistical significance is indicated above the boxplots. In D–H, the dotted line represents the average score of total T cells.
We further analyzed the differences in immune characteristics among T cell subtypes during the second trimester between subjects with GDM and those with NP. As expected, the CD8+ CTL-GNLY gene expression scores of the immune response regulating signaling pathway, the regulation of T cell activation, the positive regulation of lymphocyte activation (Fig. 3K–M), and the defined-cytokine, defined-inflammatory, Th1_th2 pathway, IL17 pathway, TNF signaling, T cell receptor signaling pathway, leukocyte activation involved in inflammatory response (Supplementary Figs. 12, 13, 17–20, 22) increased significantly in subjects with GDM. The CD8+ CTL-GZMK gene expression scores of defined effector functions, defined cytotoxicity, defined cytokines, defined inflammation, positive regulation of lymphocyte activation, the Th1_th2 pathway, TNF signaling, leukocyte activation involved in the inflammatory response, and the immune response regulating the cell surface receptor signaling pathway were elevated in GDM patients (Figs. 3I, J, M and Supplementary Figs. 10–13, 16, 17, 19, 21, 22). Moreover, the expression levels of genes related to immune-inflammatory processes in CD8+ NKT, γδ T, CD8+ MAIT, and CD4+ memory T cells were significantly increased in the GDM group compared with those in the NP group (Supplementary Figs. 10–22). These findings suggest that the predominant T cell subtypes are involved in immune-inflammatory responses during the second trimester and that these specific T cell subtypes significantly enhance immune responses in subjects with GDM.
Transcription factors serving as the hub genes of T cell function in GDM patients
The functional analysis of T cells mentioned above revealed significant associations between the immune-inflammatory function of circulating CD4+ and CD8+ T cells during the second trimester in women with GDM who underwent ART. Therefore, we further identified key genes expressed in T cell subsets related to GDM by analyzing the differentially expressed genes (DEGs) between the GDM and NP groups using the criteria of an absolute value of log2FC > 0.25 and a corrected p value < 0.05. Our findings revealed 216, 402, 45, 57, 235, 188, and 215 DEGs among CD8+ CTL-GNLY, CD8+ CTL-GZMK, CD8+ NKT, CD8+ MAIT, CD8+ naïve T, CD4+ memory T and CD4+ naïve T cells, respectively, between participants with GDM and those with NP (Supplementary Data 2). Based on intergroup comparisons of DGEs in T cell subsets, we constructed protein-protein interaction (PPI) networks using the STRING database. Subsequently, Cytoscape software (v3.10.3) was used to analyze the protein-protein interaction networks and identify the hub genes. In subjects with GDM, the upregulated STRING modules in CD8+ CTL-GZMK included several transcription factors (TFs), such as FOS, JUN, STAT3, JUNB, JUND, FOSB, CEBPB, and ZNF683. Additionally, key genes involved in inflammation activation, including GZMB, FCGR3A, PRF1, CCL5, CCL4, TYROBP, GZMH, CXCR4, KLRD1, and CD69, were also present within the upregulated modules of CD8+ CTL-GZMK in GDM patients (Fig. 4A). The maximum clique centrality (MCC) algorithm of the CytoHubba plugin was subsequently utilized to identify the top 10 hub genes, including CCL5, CCL4, CD69, PRF1, GZMB, FCGR3A, KLRD1, NKG7, TYROBP, and NCR3, which are associated with the immune-inflammatory functions of CD8+ CTL-GZMK (Fig. 4B). These results suggest that the hub inflammatory gene set was specifically activated in CD8+ CTL-GZMK under GDM conditions. Moreover, the TFs FOS, JUNB, JUND, KLF6, DUSP1, and ZFP36 were also identified as hub genes for GDM in the upregulated STRING modules of CD8+ CTL-GNLY, CD8+ NKT, CD8+ MAIT, CD8+ naïve T, CD4+ memory T, and CD4+ naïve T cells. Additionally, we observed that the NFKBIA gene, a crucial component of the NF-κB signaling pathway, served as a hub gene in functional pathways associated with GDM (Fig. 4A, B).
A Protein-protein interaction (PPI) networks of upregulated differentially expressed genes (DEGs) in each T cell subtype in the GDM group. B Top 10 hub genes among upregulated DEGs in each T cell subtype in the GDM group. C Differential transcription factor (TF) activity and specificity scores in CD8⁺ T cells between the GDM and NP groups. D Differential TF activity and specificity scores in CD4⁺ T cells between the GDM and NP groups. E TF-regulated intergroup differential genes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment in CD8+ T cells. F TF-regulated intergroup differential genes and KEGG pathway enrichment in CD4+ T cells.
Hub gene identification and analysis revealed that genes encoding TFs potentially affect the immune-inflammatory function of T cells in patients with GDM. We next conducted a single-cell regulatory network inference and clustering (SCENIC) analysis and calculated the TF activity scores and regulon specificity scores (RSSs) at the single-cell level. The distinct clustering features of the activity scores and RSSs of the TFs revealed the unique immune functions associated with each T cell subtype (Supplementary Figs. 23 and 24). Additionally, we compared the activity scores and RSSs of TFs in CD8+ T cells and CD4+ T cells between the GDM group and the NP group. We observed slightly higher activity scores and RSSs for the TFs FOSB, CREM, JUNB, JUND, and FOS in CD8+ T cells from the GDM group than in those from the NP group (Fig. 4C and Supplementary Data 1). We subsequently identified target genes regulated by these five TFs and intersected them with the differentially upregulated genes in CD8+ T cells from the GDM group. The findings revealed 34, 21, 125, 97, and 80 shared upregulated genes in CD8+ T cells that were targets of FOSB, CREM, JUND, JUNB, and FOS, respectively (Supplementary Fig. 25 and Supplementary Data 1). Furthermore, these shared genes were enriched in immune-inflammatory pathways, such as the TNF signaling pathway, the IL-17 signaling pathway, the T cell receptor signaling pathway, and Th1/Th2 cell differentiation (Fig. 4E). Similarly, greater activation and specificity were observed for six TFs (FOS, JUNB, JUND, FOSB, JUN and KLF6) in CD4+ T cells from the GDM group than in those from the NP group (Fig. 4D and Supplementary Data 1). Furthermore, the genes upregulated by these TFs in the CD4+ cells of the GDM group were significantly enriched in both the TNF signaling pathway and the IL-17 signaling pathway (Fig. 4F and Supplementary Fig. 26). These findings suggest that TFs specifically expressed in the GDM group play a central role in regulating the immune status during the second trimester under GDM conditions.
Features of T cell immune repertoires and specific usage of V(D)J genes in GDM patients who underwent ART
We analyzed single-cell TCR sequencing data and identified high-quality TCR sequences that matched more than 70% of the T cells in each subject to evaluate the clonal characteristics of T cells and V(D)J gene usage in GDM subjects during their second trimester (Supplementary Table 3). The TCR matching rate exceeded 80% for the major αβ T cell subtypes, with the exception of CD8+ NKT, exhausted CD4+ T and pro T cells (Fig. 5A). We observed that during the second trimester of pregnancy, unique clones of CD4+ T cells were predominant, whereas CD8+ T cells represented an apparent proportion of clonal cells, particularly the CD8+ CTL-GNLY T and CD8+ CTL-GZMK T subtypes (Fig. 5B, C and Supplementary Fig. 27). However, no statistically significant differences in the proportions of expanded CD4+ and CD8+ T cells were observed between the GDM group and the NP group; likewise, no significant disparities were found in terms of the TCR distribution, abundance, and diversity between these two groups (Supplementary Figs. 27 and 28). We specifically selected expanded T cells (with a clonotype ≥2) to further investigate the DEGs and biological functions of these T cells under GDM conditions (Fig. 5D–F and Supplementary Data 1). The findings suggest that the DEGs in expanded CD8+ T cells were significantly enriched in Th17 proliferation, TNF signaling, Th1 and Th2 cell proliferation, IL-17 signaling, and T cell receptor signaling pathways (Fig. 5F). Additionally, we observed the enrichment of DEGs related to the IL-17 signaling pathway in expanded CD4+ T cells (Fig. 5E). These results indicate that expanded CD8+ and CD4+ T cells play a key role in the immune-inflammatory response in patients with GDM.
A Uniform manifold approximation and projection (UMAP) plot of αβ T cell receptor (TCR)-matched T cells. B Bar plot illustrating the distribution of TCRs across distinct T cell subtypes. C Dot plot showing TCR clonal expansion across distinct T cell subtypes. D Differential expression gene (DEGs) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of expanded T cells in the GDM group. E DEGs and KEGG pathway enrichment analyses of expanded CD8+ T cells in the GDM group. F DEGs and KEGG pathway enrichment analyses of expanded CD4+ T cells in the GDM group. Volcano plots are shown above (D–F), and KEGG enrichment results are shown below. G Usage preferences of T cell receptor alpha variable (TRAV) genes (top), T cell receptor alpha joining (TRAJ) genes (middle), and T cell receptor beta variable (TRBV) genes (bottom) in the GDM group. p values were calculated using the Wilcoxon rank-sum test. A red asterisk indicates a statistically significant difference between the GDM and NP groups.
We then compared the distributions of the complementarity determining region 3 (CDR3) length and V-J gene usage in the CDR3 region for both the TCR α and TCR β chains between the GDM and NP groups. The results revealed no significant difference in the distribution of the CDR3 length between the two groups (Supplementary Fig. 29). The analysis of the V-J gene usage preference in the TCR α and TCR β chains indicated a decrease in the expression of the TRAV19 and TRAJ30 genes but a significant increase in the expression of the TRBV5-6 and TRBV6-1 genes in the GDM group (Fig. 5G). Moreover, the usage rates of V-J pairs were also significantly different between the GDM and NP groups (Supplementary Fig. 30). The preferential bias in V-J gene rearrangement observed in GDM patients may impact the adaptive immune function of T cells through alterations in immunodominant epitopes. The top 10 TRA and TRB CDR3 AA sequences were further extracted from the GDM and NP groups, respectively. Only two TRA CDR3 sequences, namely, CAVRDSNYQLIW and CAVMDSNYQLIW, were shared between the groups (Supplementary Fig. 31).
Moreover, the findings revealed that 1429 GDM-specific CDR3 AA sequences were identified (Supplementary Data 3). Among the 1016 shared CDR3 sequences, 13 CDR3 AA sequences showed statistical significance, with 10 being enriched in the GDM group and 3 being enriched in the NP group (Supplementary Data 4 and Supplementary Table 4). Next, antigen epitopes were predicted for enriched and specifically expressed TCR CDR3 AA sequences from the GDM group using public databases. The VDJdb database was used to obtain a comprehensive set of immune epitope information. Potential epitopes recognized by the TCR CDR3 sequences were inferred by utilizing known TCR-antigen epitope interactions (Supplementary Data 5 and 6). The predicted antigen epitopes were then compared with those available in the IEDB database. This comparison specifically focused on TCR antigen epitopes associated with diabetes-related diseases. The IEDB database contained extensive data on immune epitopes and provided valuable insights, particularly regarding immune responses in diabetes. This step enabled the confirmation of the possible immune relevance of these epitopes. Ultimately, two antigen epitopes corresponding to TCR CDR3 sequences enriched in the GDM group were identified. TCR CDR3 sequence CAVMDSNYQLIW was matched with the antigen epitope GLCTLVAML, and TCR CDR3 sequence CAVLDSNYQLIW was matched with the antigen epitopes GLCTLVAML and GILGFVFTL (Supplementary Table 5).
Differentially expressed functional genes and immune pathways of B cells in GDM patients
Given the critical role of B cells in humoral immunity, we separately reclustered the B cells. Based on the established marker genes, we subdivided B cells into six subtypes: naïve B cells (TCL1A+IGHD+IL4R+CD27−, 44.35%)36,37,44, memory B-CD1C cells (CD38−CD27+CD24+AIM2+TNFRSF13B+CD1Clow, 18.12%), memory B-CRIP1 cells (CD38-CD27+CD24+AIM2+IGHG1+IGHA1+CRIP1+, 17.50%)37,44, memory B-AIF1 cells (CD38lowCD27lowCD24lowAIF1+, 7.27%), memory B-IL32 cells (CD38lowCD27lowCD24lowIL32+, 7.79%), and plasma cells (MZB1+JCHAIN+, 4.96%)44,46 (Fig. 6A, B). Significant difference in the distributions and proportions of B-cell subgroups was not observed between the GDM group and the NP group (Supplementary Fig. 32). We performed a pseudotime analysis of B cells to investigate the progression of various B-cell subtypes. Our results revealed that peripheral B-cell subtypes originated from naïve B cells and progressively transitioned toward memory B cells and plasma cells during the second trimester (Supplementary Fig. 33). Consistent with this finding, cells at the initial stage of this trajectory presented increased expression levels of naïve markers (TCL1A and IGHD) (Fig. 6C). Additionally, several memory B-cell marker genes (TNFRSF13B, IGHG1, CRIP1, IL32, and AIF1) were highly expressed during the intermediate differentiation stage (Fig. 6C). Furthermore, we noted that the expression of FOS gradually increased along the B-cell differentiation trajectory, whereas the expression of the B-cell activation marker CD69 gradually decreased (Fig. 6C). This differentiation trajectory also reflected the distinct functional states of several B-cell subsets during mid-pregnancy.
A Uniform manifold approximation and projection (UMAP) plot showing distinct B cell subtypes. B UMAP plots illustrating the expression of canonical marker genes across distinct B cell subtypes. C Scatter plots with fitted curves showing the expression dynamics of canonical marker genes for B cells along pseudotime. Each dot represents a single cell, colored by cell state. Black curves indicate smoothed expression trends fitted along pseudotime. D Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of differential expression gene (DEGs) in B cells between the GDM and NP groups. E Protein-protein interaction (PPI) networks of upregulated DEGs in B cells between the GDM and NP groups. F Differential activity and specificity scores of transcription factors (TFs) in B cells between the GDM and NP groups. TF activity scores are shown on the left and TF specificity scores on the right. G TF-regulated intergroup differential genes in B cells. H KEGG pathway enrichment analysis of DEGs upregulated by TFs in GDM.
In addition, we compared the transcriptomic profiles of B cells from subjects with GDM to those from subjects with NP, and the DEGs were significantly enriched in the TNF signaling pathway, the IL-17 signaling pathway, the Toll-like receptor signaling pathway, and the NOD-like receptor signaling pathway (Fig. 6D, Supplementary Figs. 34–39 and Supplementary Data 1). These pathways are integral to both adaptive and innate immune responses, indicating enhanced activation of B cells and proinflammatory functions in GDM patients.
Based on the STRING PPI network and hub gene screening, we determined that the top 10 upregulated hub genes in the B cells of GDM patients were FOS, NFKBIA, TNFAIP3, JUN, JUNB, CCL5, DUSP1, CXCR4, REL, and CD69 (Fig. 6E and Supplementary Fig. 34). We further performed a SCENIC analysis to explore the transcriptomic changes and functional roles of TFs in B cells. The findings revealed distinct TF expression profiles for each B-cell subtype (Supplementary Fig. 40). Moreover, subjects with GDM exhibited higher expression levels of several TFs, such as SPIB, BCLAF1, TAF7, XBP1, IRF9, MYC, FOSB, IRF1, JUND, JUNB, KLF6, and JUN (Fig. 6F and Supplementary Data 1). Notably, three of the most distinguishing TFs (FOS, JUN and JUNB) associated with GDM activated several immune-inflammatory responses, including the TNF signaling pathway, the IL-17 signaling pathway, the Toll-like receptor signaling pathway, and the NOD-like receptor signaling pathway, through the upregulation of their target genes (Fig. 6G, H and Supplementary Fig. 41).
Characterization of BCRs and specific rearrangements of heavy chain V(D)J genes in subjects with GDM
We explored the clonal proliferation of BCRs in patients with GDM by conducting a single-cell clonotype analysis of BCR V(D)J genes. In this study, a total of 25,036 B cells were analyzed, with 14,534 cells in the GDM group (58%) and 10,502 cells in the control group (42%). A total of 16,858 unique BCR CDR3 sequences were detected. Excluding plasma cells, the coverage of BCRs in each T cell subset with varying degrees of expansion exceeded 75% (Fig. 7A, B and Supplementary Fig. 42). Naïve B cells exhibited a more pronounced state of clonal expansion, whereas other B-cell subtypes predominantly expressed distinct BCRs (Fig. 7C). We conducted a quantitative analysis of the diversity and abundance of BCR clonotypes, revealing increased BCR diversity in subjects with GDM (Fig. 7F). However, no significant difference in the proportion of expanded clonotypes was observed between the GDM and NP groups (Supplementary Fig. 42C).
A Uniform manifold approximation and projection (UMAP) plot of αβ B cell receptor (BCR)-matched B cells. B Bar plot illustrating the distribution of BCRs across distinct B cell subtypes. C Dot plot showing BCR clonal expansion across distinct B cell subtypes. D Volcano plot illustrating differentially expressed genes (DEGs) in expanded B cells between the GDM and NP groups. Genes with |log2(FC)| ≥0.25 and adjusted p < 0.05 are labeled with their respective gene symbols. E Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of intergroup DEGs in expanded B cells. F Clonal abundance and diversity of BCR between the GDM and NP groups. G U Usage preferences of immunoglobulin heavy chain variable (IGHV) genes and immunoglobulin heavy chain joining (IGHJ) genes in the GDM group. p values are calculated using the Wilcoxon rank-sum test. A red asterisk indicates a statistically significant difference between the GDM group and NP group. H Bar plot showing the top 10 immunoglobulin heavy chain complementarity-determining region 3 (IGH CDR3) amino acid sequences (AA) sequences in the GDM and NP groups.
We conducted a DEG analysis to compare these expanded B cells between the GDM group and the NP group and to elucidate the functional characteristics of expanded B cells (clonotype ≥2) in patients with GDM. The analysis of DEGs revealed that the upregulated genes in expanded B cells were enriched in several immune-inflammatory signaling pathways, including the TNF signaling pathway, the IL-17 signaling pathway, the Toll-like receptor signaling pathway, the NOD-like receptor signaling pathway, the B-cell receptor signaling pathway, and the Th1 and Th2 cell differentiation pathways (Fig. 7D, E).
Next, we compared the usage of V(D)J genes between the GDM group and the NP group (Fig. 7G and Supplementary Figs. 43 and 44). Our findings indicated significantly increased expression of the heavy chain IGHV4-34 among subjects with GDM (Fig. 7G). Previous studies reported that such heightened gene usage was associated with autoimmune dysregulation47. In contrast, no significant difference was observed in the usage of B-cell light chain V(D)J genes across the two groups (Supplementary Fig. 43).
In accordance with the usage bias observed in V-J genes, a notable disparity existed in the distribution of CDR3 heavy chain amino acid (CDR3 AA) sequences between the GDM group and the NP group. The top 10 CDR3 AA sequences selected for each group were entirely distinct (Fig. 7H). However, the distributions of the top 10 light chain CDR3 aa sequences were largely consistent between the two groups (Supplementary Fig. 45). No significant differences were observed in the length distributions of the CDR3 aa and nucleotide (nt) sequences between the two groups (Supplementary Fig. 46). The results also revealed that 161 GDM-specific BCR CDR3 AA sequences were identified (Supplementary Data 7). Among the shared CDR3 sequences, just 2 BCR CDR3 sequences showed statistical significance after frequency difference testing and FDR correction (Supplementary Data 8). To investigate whether the expanded BCR repertoire in GDM patients might confer cross-reactive immunity to known pathogens or antigens, we performed systematic sequence alignment of GDM-specific CDR3 sequences against two complementary public databases: the SARS-CoV-2 antibody repertoire (CoV-AbDab) and epitope-specific antibodies (IEDB) using Levenshtein distance metrics. Analysis of CoV-AbDab alignments revealed varying degrees of sequence similarity (Supplementary Figs. 47 and 48), with maximum homology of 84% observed in light chain CDR3 sequences (CQAWDSSTVVF, CGTWDSSLSVVVF) and 72% in heavy chain sequences (CARDPEGTYDFWSGYNYYYYGMDVW) when compared to documented SARS-CoV-2-reactive antibodies. Similarly, alignment with IEDB epitope-specific antibodies demonstrated comparable patterns (Supplementary Figs. 49 and 50), with peak light chain similarity of 91% (CMQALQTPFTF with Levenshtein distance of 1) and heavy chain similarity of 67% (CARGAHYDFWSGYPLDYW and CARDPEGTYDFWSGYNYYYYGMDVW). Notably, the majority of identified matches demonstrated low to moderate epitope similarity in both databases. Overall, the distinct clonal diversity, preferential usage of V-J genes and CDR3 sequences, and heightened immune activation of B cells collectively indicate a correlation between BCR clonal expansion and immune function in patients with GDM.
Features of innate immune cells and their interactions with the adaptive immune response
Monocytes and NK cells serve as the principal components of the innate immune system. Our findings revealed no significant difference in the proportion of circulating innate immune cells between the GDM and NP groups (Fig. 8A). However, the DEG analysis conducted on monocytes from the GDM and NP groups demonstrated a distinct immune response in monocytes (Supplementary Fig. 51 and Supplementary Data 1). The upregulated genes of monocytes in the GDM group were predominantly enriched in TNF signaling, cytokine-cytokine receptor interaction, NOD-like receptor signaling, IL-17 signaling, NF-kappa B signaling, chemokine signaling, and Toll-like receptor signaling (Fig. 8B, C). Similarly, NK cells from the GDM group presented several activated immune-inflammatory signaling pathways (Fig. 8D). Moreover, the innate immune cells in the GDM group presented enhanced biological functions related to inflammatory responses, as well as improved chemotactic and migratory abilities of immune cells (Supplementary Fig. 52). In the STRING network and hub gene analysis, we found that key genes, including TNF, CXCL8, NFKBIA, IL1B, ICAM1, and FOS, were expressed at higher levels in innate immune cells than in other peripheral immune cells (Fig. 8E and Supplementary Figs. 53 and 54). Inflammation scores based on a single-cell analysis of PBMCs also revealed that the inflammation scores for CD14+ monocytes and CD16+ monocytes were notably elevated compared with those of other immune cell subsets (Fig. 8F). Additionally, the inflammatory and cytokine scores for monocytes in the GDM group were significantly higher than those observed in the NP group (Fig. 8G).
A Uniform manifold approximation and projection (UMAP) plot of natural killer (NK) cells and monocytes. B Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of differentially expressed genes (DEGs) in CD16+ monocytes between the GDM and NP groups. C KEGG pathway enrichment analysis of DGEs in CD14+ monocytes between the GDM and NP groups. D KEGG pathway enrichment analysis of DGEs in NK cells between the GDM and NP groups. E UMAP plots illustrating the expression of hub genes in innate immune cells. F UMAP plots depicting defined inflammatory scores in PBMCs. G Boxplots showing defined inflammatory and defined cytokine scores in innate immune cells between the GDM and NP groups. The Wilcoxon rank-sum test was used to assess group differences. H Circos plot depicting the differential cell-cell communication networks among main immune cell types between the GDM and NP groups. Different colored dots represent distinct cell types. Red lines indicate significantly increased intercellular communication in the GDM group, whereas blue lines indicate increased communication in the NP group. I Dot plot showing the significantly increased ligand-receptor signaling pairs among main immune cell types in the GDM group.
We next conducted an analysis of cell-cell communication among distinct subpopulations of peripheral immune cells. The findings indicated that the GDM and NP groups presented distinct patterns of immune cell interactions (Fig. 8H and Supplementary Fig. 55). From the NP state to the GDM state, a reduction in the outgoing interaction strength was observed for B cells, plasma cells, CD8+ T cells, CD14+ monocytes, and CD16+ monocytes, whereas the outgoing interaction strength of CD4+ T cells was augmented (Supplementary Fig. 56). Notably, CD8+ T cells displayed the strongest incoming interaction strength in the second trimester (Supplementary Fig. 56), which aligns with their role in the adaptive immune response during pregnancy. Moreover, a pathway analysis of the communication probability revealed that signaling pathways involving macrophage migration inhibitory factor (MIF), the C-type lectin domain family (CLEC), adhesion G protein-coupled receptor E5 (ADGRE5), low-affinity immunoglobulin E receptor (CD23), and TNF were enhanced from the NP to the GDM stage (Supplementary Fig. 57). In the subsequent ligand-receptor analysis, a significant increase in the MIF-(CD74+CXCR4) ligand-receptor signaling network was observed in the GDM group (Fig. 8I). This increase was particularly evident in the interactions between B cells and CD8+ T cells, between CD14+ monocytes and CD8+ T cells, and between CD16+ monocytes and CD8+ T cells (Fig. 8I). The results suggested that CD8+ T cells in patients with GDM are concurrently regulated by B cells, CD14+ monocytes, and CD16+ monocytes through the MIF signaling pathway.
Integrated single-cell transcriptomic and Olink proteomic analyses reveal the consistent upregulation of the immune-inflammatory response in patients with GDM
The single-cell transcriptome abundance is not a complete reflection of actual protein expression. Consequently, we conducted dynamic assessments of the serum protein expression levels of 92 inflammatory factors using the Olink 96 inflammation panel in 32 GDM patients and 31 patients with NP, including 10 patients with GDM and 10 patients with NP who underwent single-cell sequencing, at three distinct stages: baseline, clinical pregnancy, and the second trimester (Supplementary Table 6 and Supplementary Figs. 58–60). The results indicated that in the GDM group, a significant increase in the levels of inflammatory markers was not observed prior to pregnancy or during the early stages of clinical pregnancy. However, in the second trimester, the protein levels of CD6, CXCL5, MMP10, and 4E-BP1 were significantly increased in the GDM group (Supplementary Table 7). The multivariate logistic regression analysis showed that, after adjusting for covariates, the second-trimester levels of MCP-1, MCP-4, CCL11, CXCL5, and MMP-10 remained significantly associated with GDM (Supplementary Table 8). The correlation analysis revealed that elevated baseline levels of CD8A, IL20RA, CCL11, and CST5, as well as uPA levels during 4-6 weeks of gestation were positively associated with hyperglycemia in the second trimester (Fig. 9A). Similarly, the elevated levels of CXCL5, CXCL6, CXCL10, CXCL11, IL12B, IL8, CCL20, MMP10, TNF-α, IFN-γ, VEGFA, AXIN1, and STAM-BP observed during the second trimester were positively correlated with the results of the oral glucose tolerance test (OGTT) (Fig. 9A). We performed GO and KEGG enrichment analyses of the identified inflammatory factors associated with glucose metabolism to elucidate their potential functions. The results indicated that glucose metabolism-related inflammatory factors were enriched in the adaptive immune response and innate immune response pathways (Supplementary Fig. 61). Additionally, the significantly different GO terms were predominantly involved in the chemotaxis and migration of immune cells (Supplementary Fig. 62). These results were consistent with the findings from the single-cell transcriptomic analysis.
A Correlation between inflammatory factor levels and second-trimester blood glucose levels. Fasting plasma glucose (FPG), 1-h oral glucose tolerance test (OGTT 1 h), and 2-h oral glucose tolerance test (OGTT 2 h) represent glucose measurements obtained between 24–28 weeks of gestation. P0, P1, and P2 denote detection periods for inflammatory factors during the pre-pregnancy period, 4–6 weeks of gestation, and 24–28 weeks of gestation, respectively. Pearson correlation analysis was performed. B Correlation between inflammatory factor levels and T cell subtype proportions. Mantel’s test was used to evaluate correlations between inflammatory factor protein levels and T cell subtype ratios. Inflammatory factors CD6, C-X-C motif chemokine ligand 5 (CXCL5), matrix metallopeptidase 10 (MMP10), and eukaryotic translation initiation factor 4E-binding protein 1 (4E-BP1) were assessed during the second trimester. An asterisk indicates a significant pairwise correlation. C Uniform manifold approximation and projection (UMAP) plot showing Olink inflammatory factor scores in single cells from the GDM and NP groups. Scores were calculated by mapping 92 inflammatory factor-encoding genes at the single-cell level using the AddModuleScore function. D Comparison of Olink inflammatory factor-encoding gene scores in PBMCs between the GDM and NP groups. The Wilcoxon rank-sum test was applied. E Three-dimensional scatter plots illustrating correlations among inflammatory factor levels, reproductive hormone levels, and blood glucose. Point size indicates blood glucose levels, and color indicates whether values met the diagnostic criteria for GDM. FT3 free triiodothyronine, PRL prolactin, LH luteinizing hormone.
Next, we analyzed the correlations between glucose metabolism-related inflammatory factors and the proportions of immune cell subsets using Mantel’s test. The findings revealed that the sets of inflammatory factors present during the pre-pregnancy period and the second trimester were associated with CD8+ naïve T cells and CD8+ NKT cells (Supplementary Fig. 63). Specifically, during the second trimester, the levels of CD6 and 4E-BP1 were positively correlated with the proportions of the CD8+ CTL-GNLY and CD8+ CTL-GZMK cell subsets, respectively (Fig. 9B). Finally, we mapped the genes encoding the 92 aforementioned inflammatory factors to the single-cell transcriptomic data and calculated the Olink inflammatory factor scores using the AddModuleScore function (Fig. 9C). We observed that the intracellular inflammatory factor gene expression levels in the GDM group during mid-pregnancy were significantly higher than those in the NP group (Fig. 9D), indicating a consistent correlation between the intracellular and extracellular findings.
Specific clinical features of women undergoing ART in relation to the immune response and glucose metabolism
Women undergoing ART may exhibit more complex clinical features due to both the ART intervention and underlying reproductive endocrine disorders. The potential impacts on the immune response and glucose metabolism remain to be further investigated. We analyzed the ratios of peripheral immune cell subsets and the serum levels inflammatory markers in pregnant women, comparing both the IVF and ICSI groups, as well as the frozen embryo transfer and fresh embryo transfer groups (Supplementary Tables 9–12). Our study revealed that pregnant women who underwent frozen-thawed embryo transfer presented a greater proportion of CD4+ memory T cells and elevated levels of CCL20 during the second trimester than did those who underwent fresh embryo transfer (Supplementary Tables 11 and 12). Furthermore, pregnant women in the ICSI group had significantly higher levels of several inflammatory factors than those in the IVF group (Table S10). The findings also indicated that several inflammatory factors associated with glucose metabolism were significantly correlated with reproductive endocrine hormones (Supplementary Figs. 64–67). Specifically, IL12B, CXCL11, and CD6 levels were negatively correlated with baseline free triiodothyronine (FT3), prolactin (PRL), and progesterone levels, respectively (Fig. 9E). Conversely, VEGFA levels were significantly positively correlated with luteinizing hormone (LH) levels (Fig. 9E). These results suggest that the ART intervention and reproductive endocrine function may have potential regulatory effects on the immune status of pregnant women undergoing ART.
Discussion
The maternal immune system undergoes systemic immunological adaptations throughout pregnancy39. The homeostasis of the maternal immune system confers immune tolerance to semi-allogeneic fetuses and is crucial for maintaining a normal pregnancy39,48. Conversely, the dysregulation of immune mechanisms during pregnancy serves as a pathological basis for adverse pregnancy outcomes49,50. Female infertility and ART treatment may trigger maternal inflammatory and oxidative stress pathways, which is consistent with the significantly increased risk of adverse pregnancy outcomes observed in women who conceived through ART26. Therefore, pregnant women undergoing ART are categorized as being at “high risk” for pregnancy complications18,51. Previous studies have focused primarily on immune mechanisms at the maternal-fetal interface52. With the advancement of multiomics technologies, transcriptomic analysis of peripheral blood has emerged as a robust method for elucidating the immune landscape associated with pregnancy complications53,54. In this study, we investigated the immune characteristics of women undergoing ART by integrating single-cell sequencing and targeted inflammatory proteomic analyses. Our analysis dynamically profiled the immune status of women with GDM and those with NP who underwent ART across three critical stages, namely, pre-pregnancy, early clinical pregnancy (4–6 weeks of gestation), and mid-pregnancy (24–28 weeks of gestation). We examined the multidimensional associations between immune characteristics and GDM, including the immune cell composition, the immune repertoire diversity, inflammatory factor alterations, and underlying molecular functions.
We conducted a longitudinal comparison of the plasma expression levels of 92 inflammatory factors between the GDM and the NP groups after matching for demographic and clinical characteristics. Our findings revealed no significant intergroup differences in the levels of inflammatory factors between the GDM group and the NP group during the pre-pregnancy and early clinical pregnancy stages. However, pre-pregnancy plasma concentrations of inflammatory markers such as CD8A, IL20RA, CCL11, and CST5, along with uPA levels during the clinical pregnancy period, were positively correlated with elevated blood glucose levels in the second trimester of gestation. During the second trimester (24–28 weeks) of gestation, the levels of inflammatory markers such as CD6, CXCL5, MMP10, and 4E-BP1 in the GDM group were significantly elevated compared with those in the NP group. Furthermore, multiple inflammatory markers, including MCP-1, MCP-4, CXCL6, CXCL10, CXCL11, IL12B, IL8, CCL20, TNF-α, IFN-γ, VEGFA, AXIN1, and STAM-BP, were positively correlated with increased blood glucose levels during this period. These glucose metabolism-related inflammatory factors are implicated in multiple immune response pathways. Notably, the immune imbalance during the second trimester was significantly associated with GDM. Previous studies have indicated that the pathophysiological features of GDM are characterized by a state of chronic low-grade inflammation. This condition is primarily associated with elevated levels of circulating proinflammatory cytokines, including IL-1β, IL-6, TNFα, MMP-10, MCP-1, and VEGFA, as well as reduced levels of anti-inflammatory molecules such as IL-4 and IL-1055,56,57. In vitro studies have demonstrated that low-grade inflammation, accompanied by a modest elevation in circulating levels of proinflammatory cytokines, can impair beta-cell function and survival58,59,60. Furthermore, in a hyperglycemic environment, macrophages trigger the NLRP3 inflammasome upon engulfment of apoptotic pancreatic beta cells61,62. This activation facilitates the phenotypic shift of macrophages from an anti-inflammatory state to a proinflammatory state61,62. The cascade exemplifies the detrimental cycle in which chronic low-grade inflammation, impaired pancreatic beta-cell function, and a high-glucose environment synergistically induce immune-mediated inflammation, thereby increasing the risk of adverse perinatal outcomes and long-term metabolic disorders in both mothers and offspring.
Alterations in immune phenotypes associated with GDM are also characterized by heterogeneity in the distribution of immune cells and intercellular interactions. However, previous studies that relied on blood cell counts and flow cytometry have failed to construct a comprehensive atlas of peripheral cellular and molecular immune responses. In this study, we conducted single-cell resolution mapping of 12 immune cell types associated with GDM in pregnant women receiving ART. Our findings indicate that T cells, B cells, monocytes, and NK cells are involved in the dysregulated immune responses observed during pregnancy. T cells are essential components of adaptive immune responses during pregnancy, and we identified and annotated eleven distinct circulating T cell subsets. The findings revealed a significant increase in the overall proportion of peripheral CD8+ T cells in patients with GDM. Notably, CD8+ CTL-GNLY, CD8+ NKT, and CD8+ CTL-GZMK cells exhibited enhanced cellular effector and cytotoxic functions. Furthermore, in patients with GDM, CD8+ CTL-GNLY, CD8+ CTL-GZMK, and CD8+ NKT cells exhibited enhanced activation of immune functions. This increase was characterized by elevated gene expression scores in multiple immune-related pathways, including the Th1/Th2 pathway, the IL-17 signaling pathway, the TNF signaling pathway, the T cell receptor signaling pathway, the regulation of cell surface receptor signaling pathways involved in immune responses, and leukocyte activation associated with inflammatory responses. Research has indicated that T cell responses are attenuated during pregnancy63. A single-cell analysis of dynamic T cell immunity during normal pregnancy revealed that the suppression of T cell function is characterized primarily by a marked reduction in components essential for T cell activation, including the TCR-CD3 complex (TRAC, TRBC1, TRBC2, CD3D, CD3E, and CD3G) and costimulatory molecules (CD2 and CD27)39. Concurrently, the expression levels of genes associated with T cell cytotoxicity are downregulated39. In contrast, in this study, we found that T cell cytotoxicity and inflammatory functions were enhanced in GDM patients, directly reflecting the circulating immune activation characteristics of GDM patients.
Despite their small proportions among circulating PBMCs, γδ T cells and MAIT cells are functionally potent innate-like lymphocytes that play critical roles in immune surveillance, tissue homeostasis, and inflammatory responses. In this study, we found that both cell subsets exert important immune functions in GDM. γδ T cells represent a unique lymphoid lineage positioned at the interface between innate and adaptive immunity. These cells are capable of rapid cytokine production and cytotoxic responses without requiring conventional antigen presentation64,65. During pregnancy, γδ T cells contribute to maternal-fetal immune tolerance and placental development66,67. Recent evidence has shown that γδ T cell levels were higher in women with GDM68. However, the role of γδ T cells in GDM pathogenesis remains poorly understood and requires further investigation. MAIT cells have also emerged as critical players in metabolic disorders. Multiple studies have demonstrated profound alterations in circulating MAIT cell frequency and function in obesity and type 2 diabetes, characterized by reduced circulating numbers and functional exhaustion. Paradoxically, MAIT cells show enhanced IL-17 production, which contributes to systemic inflammation and insulin resistance69,70,71. Our findings extended these observations to GDM. We demonstrated that CD8+ MAIT cells and γδ T cells displayed marked immune activation and elevated immune-inflammatory gene expression signatures during the second trimester in women with GDM following ART.
While T cells play central roles in GDM pathogenesis, recent evidence has revealed that B cells are equally important, particularly through their coordinated activation with T cells. Both T cells and B cells showed enrichment of shared inflammatory signaling pathways, including TNF signaling, IL-17 signaling, and Toll-like receptor signaling. Moreover, the ligand-receptor analysis identified increased MIF-(CD74+ CXCR4) signaling networks in GDM, with enhanced interactions between B cells and CD8+ T cells, suggesting direct cellular crosstalk between immune compartments. MIF is a pro-inflammatory cytokine that functions through its receptor CD74 and co-receptor CXCR4, which together form functional heterocomplexes on immune cells72. Recent studies have demonstrated that MIF promotes B-cell chemotaxis through cooperative engagement of CXCR4 and CD74 via a ZAP-70-dependent pathway73, while MIF also influences CD8⁺ T cell function through CD74/CXCR4 signaling complexes expressed on activated T cells74. This shared signaling pathway provided a unifying molecular mechanism for the coordinated B-cell and CD8⁺ T cell activation observed in our study. Furthermore, B cells could directly regulate CD8+ T cell responses by functioning as antigen-presenting cells, expressing MHC class I molecules that can present antigens to CD8+ T cells and influencing their activation and memory formation75,76. The coordinated activation of cellular and humoral immunity has important pathophysiological implications for GDM. The simultaneous activation of multiple immune pathways likely contributes to the chronic inflammatory state that exacerbates insulin resistance and metabolic dysfunction77,78. Both CD8⁺ T cells and B cells produce pro-inflammatory cytokines that promote tissue inflammation, potentially creating a self-reinforcing cycle of immune activation and metabolic deterioration79,80. Notably, studies have shown that altered T cell and B-cell profiles in GDM persist postpartum, suggesting long-lasting immune dysregulation that may contribute to increased risk of type 2 diabetes and cardiovascular disease later in life81. Understanding this coordinated immune activation provides critical insights into GDM pathophysiology and may inform the development of more targeted therapeutic strategies that address multiple immune pathways simultaneously82. It is important to acknowledge that our characterization of cell-cell communication networks, particularly the identified MIF-(CD74+ CXCR4) signaling axis between B cells, monocytes, and CD8+ T cells, is based on computational predictions derived from single-cell transcriptomic data using CellPhoneDB v2.0 and protein-protein interaction databases. While these bioinformatic approaches provide valuable hypotheses regarding potential intercellular crosstalk mechanisms, they have inherent limitations that must be recognized. Specifically, the detection of gene expression does not guarantee functional protein expression, secretion, or biological activity, as numerous post-transcriptional and post-translational regulatory mechanisms can modulate protein abundance, stability, localization, and activation states83,84,85. Therefore, our findings regarding cell-cell communication networks should be interpreted as predictive models that require rigorous experimental validation. Future studies should employ complementary functional approaches to validate these computational predictions. Despite these limitations, our computational predictions were supported by existing literature demonstrating functional MIF-CD74/CXCR4 signaling in both B cells and CD8+ T cells73,74, and the biological plausibility of this pathway in coordinating immune responses. These predictions thus provide a strong foundation and clear direction for future mechanistic investigations aimed at understanding the molecular basis of immune dysregulation in GDM.
To explore potential cross-reactive immunity, we aligned GDM-specific BCR CDR3 AA sequences against SARS-CoV-2 antibodies (CoV-AbDab) and epitope-specific antibodies (IEDB). Moderate sequence homology was observed, with maximum similarity of 84–91% for light chains and 67–72% for heavy chains, though the majority of matches ranged between 50–75%. These similarities fall below the threshold typically associated with functional cross-reactivity (≥ 85–90% identity) and more likely reflect convergent BCR evolution under chronic inflammatory conditions characteristic of GDM rather than pathogen-specific immunity86. Importantly, CDR3 sequence homology alone cannot predict functional binding specificity, as complete paratope architecture depends on CDR1/2, framework regions, and heavy-light chain pairing. Future studies incorporating functional analysis, including antigen-binding assays, neutralization studies, and full-length antibody characterization, are warranted to establish any potential cross-protective immunity87.
Notably, the aberrant involvement of human peripheral monocytes in insulin resistance has been implicated in the pathogenesis of GDM88. Previous studies have suggested that during pregnancy, NK cell function is similar to that of T cells, which are in a suppressed state (except for IFN/virus responses) to promote maternal-fetal balance39. Several cross-sectional studies have shown that the proportions of monocytes and NK cells in the peripheral blood of GDM patients are significantly higher than those in pregnant women with normal glucose levels88,89,90,91. Although significant differences in the distributions of monocytes and NK cells were not observed between pregnant women with GDM and pregnant women with NP, the monocytes and NK cells in GDM patients exhibited consistent immune activation, reflecting the proinflammatory characteristics of NK cells and monocytes in GDM patients.
This study further revealed that TFs function as pivotal regulatory elements in modulating the differentiation and immune responses of T cell and B-cell subsets in patients with GDM. For example, TFs such as FOS, JUN, JUNB, JUND, FOSB, and KLF6 serve as key regulators of T cell activities in patients with GDM by mediating the TNF signaling pathway, IL-17 signaling pathway, T cell receptor signaling pathway, and Th1/Th2 cell differentiation pathways. Similarly, the expression levels of the TFs FOS, JUN, and JUNB in B cells were significantly elevated in the GDM group. The target genes regulated by these transcription factors were predominantly enriched in several key signaling pathways, including the TNF signaling pathway, the IL-17 signaling pathway, the Toll-like receptor signaling pathway, and the NOD-like receptor signaling pathway. TFs can trigger cellular activation programs upon reaching a critical concentration within the nucleus92. Fos and Jun are important TFs involved in the activation of human T and B lymphocytes. They are rapidly upregulated in lymphocytes upon stimulation of the antigen receptor, forming Fos-Jun heterodimers that interact with regulatory elements such as nuclear factor of activated T cells (NFAT) and activator protein-1 (AP-1) within the nucleus93,94. This interaction modulates lymphocyte activation, proliferation, and cytokine secretion92.
In this study, we performed V(D)J sequencing on PBMCs from the GDM and NP groups at weeks 24-28 of pregnancy in women who conceived through assisted reproductive technology and analyzed the clonal expansion and diversity of TCRs and BCRs, as well as the preference for V(D)J gene segment usage. No significant differences were observed in the abundance or diversity of TCRs between the GDM and NP groups. This finding aligns with prior studies on TCR clonal expansion during natural pregnancies50. During mid-pregnancy, the distribution of unique TCR receptors in CD4+ T cells is predominant. In contrast, CD8+ T cells exhibit a higher proportion of clonal expansion, particularly among cytotoxic T cell subsets such as CD8+ CTL-GNLY and CD8+ CTL-GZMK T cells. CD8+ T cells perform proinflammatory and cytotoxic functions95. However, animal studies have demonstrated that the proliferative capacity of CD8+ T cells is preserved during the dynamic process of normal pregnancy96. The homeostasis of CD8+ T cells during pregnancy depends on receptor affinity, which is modulated by costimulatory molecules, cytokines, and the intrauterine microenvironment96. Meanwhile, the CDR3 amino acid sequence distributions of the TRA and TRB chains showed limited overlaps between the GDM and NP groups. This observation underscores the structural disparities in the complementarity-determining regions of TCRs between these groups, which may indicate differences in T cell immune function. The expansion of BCR clones in the GDM group also indicated immune-related inflammatory activation. In particular, the usage ratio of the IGHV4-34 genes in the GDM group was significantly increased. IGHV4-34 is typically upregulated in association with the activation and clonal expansion of autoreactive B cells87. In ART procedures, factors such as ovarian hyperstimulation, changes in hormone levels, and embryo implantation may alter immune tolerance, leading to B cell dysfunction and the expansion of autoreactive B cells97,98. This immune reprogramming may lower the selection threshold for B cells, biasing them toward the extrafollicular pathway, which further promotes the selection and expansion of IGHV4-34 clones. On the other hand, GDM is characterized by a low-grade systemic inflammatory response, which has been shown to disrupt maternal immune homeostasis and promote the activation of polyreactive B cells68,99. This may further create a favorable immune environment for the expansion of autoreactive IGHV4-34 clones. Therefore, we hypothesize that the combined effects of immune modulation induced by ART and inflammation associated with GDM may contribute to the elevated expression of IGHV4-34.
Previous studies have demonstrated that controlled ovarian hyperstimulation (COH) potentiates a state of systemic inflammation100, with gonadotropin stimulation altering the intrafollicular milieu such that immune cell profiles and cytokine concentrations differ significantly between conventional IVF and natural cycle IVF101. Our data provide molecular-level support for this hormonal modulation: we observed positive correlations between LH and VEGFA levels, consistent with evidence that exogenous LH/hCG stimulation of luteinized granulosa cells upregulates VEGF and contributes to ovarian hyperstimulation syndrome, a condition characterized by elevated VEGF alongside pro-inflammatory cytokines such as IL-6, IL-1β, and TNF-α102,103. Conversely, progesterone concentrations showed a significant negative correlation with CD6, a T cell surface receptor that fine-tunes T cell activation and effector responses104, indicating that rising progesterone levels promote a shift toward a less pro-inflammatory, more tolerogenic immune state. Mechanistically, progesterone induces progesterone receptors on activated lymphocytes and drives production of progesterone-induced blocking factor (PIBF), which promotes a Th2-biased cytokine profile and inhibits NK-cell degranulation105. Critically, these hormonal effects may be modulated by the temporal interval between stimulation and embryo transfer. In fresh embryo transfers, the supraphysiological estradiol and progesterone levels from ovarian stimulation create a potentially suboptimal hormonal environment for implantation, as the endometrium may be less receptive when still under hormonal stress, and the immediate post-retrieval period is characterized by elevated pro-inflammatory cytokines following hCG administration106. In contrast, frozen embryo transfer (FET) occurs after a significant recovery period, allowing hormone levels to return to physiological states and inflammatory markers to resolve, thereby creating a milieu that more closely mimics natural conception. Our data showing elevated CD4+ memory T cells and CCL20 levels in the FET group during the second trimester support this concept, as does recent evidence that patients with successful FET outcomes had significantly lower IL-13 levels on transfer day, indicating a healthier inflammatory environment107. These findings have important clinical implications, suggesting that optimizing both the hormonal stimulation protocol to minimize inflammatory triggers and the transfer timing to allow adequate immune recovery may synergistically improve ART outcomes. While our correlational data support this dual-mechanism model, we acknowledge that prospective studies with longitudinal cytokine profiling across the stimulation-transfer timeline would provide more definitive evidence for the proposed pathways.
This study has several notable strengths. Our prospective longitudinal design with repeated measurements across pre-pregnancy, early pregnancy, and mid-pregnancy identified that immune-inflammatory divergence primarily occurs at 24–28 weeks gestation, which guided strategic deployment of single-cell multi-omics sequencing at this critical window to comprehensively characterize ART-GDM-specific immune features. The integration of single-cell transcriptomics, immune repertoire profiling, and targeted proteomics provided unprecedented resolution of peripheral immune dysregulation in GDM. To minimize confounding biases, we matched demographic characteristics, infertility factors, reproductive endocrinology, ART interventions, and pregnancy clinical characteristics across groups, and conducted comparative analyses between fresh versus frozen embryo transfer and IVF versus ICSI subgroups.
However, the following limitations warrant consideration in this hypothesis-generating single cell-omic study. First, the present analysis was conducted exclusively within a population of pregnant women undergoing assisted reproductive technology (ART), and did not include spontaneously conceived GDM or non-GDM pregnancies as a comparison group. Although this design allowed us to characterize immune alterations associated with GDM within a well-defined ART cohort, it limits the ability to disentangle immune changes driven by GDM per se from those potentially attributable to ART exposure, or from interactions between the two. Consequently, the observed immune differences should be interpreted as reflecting GDM-associated immune dysregulation in the context of ART, rather than being generalized to all GDM pregnancies. Direct comparative studies across ART and non-ART reproductive contexts are required to comprehensively delineate the independent and synergistic effects of ART and GDM on maternal immune remodeling. Second, although this study represents one of the most comprehensive longitudinal immune profiling efforts in ART pregnancies to date, the relatively modest sample size may have limited statistical power to fully capture immune heterogeneity and to perform detailed subgroup analyses. Larger, multi-center cohorts will be essential to validate the robustness and generalizability of the identified immune signatures. Third, our analyses focused on peripheral blood immune cells and circulating inflammatory mediators. While peripheral blood offers clear advantages, including minimal invasiveness, clinical accessibility, and suitability for longitudinal monitoring, this approach precludes direct assessment of immune processes at the maternal-fetal interface. Emerging evidence indicates coordinated immune dysregulation between peripheral and placental compartments in GDM, whereby systemic inflammation may promote placental macrophage infiltration and metabolic dysfunction108,109. Integrating placental immune profiling with peripheral immune analyses in future studies would provide critical mechanistic insights into how systemic immune alterations translate into placental pathology. Finally, despite the application of an integrated multi-omics strategy combining single-cell transcriptomics, immune repertoire profiling, and targeted inflammatory proteomics, the current study is largely descriptive. Default options of statistical analysis were used in each bioinformatic tool which may generate false-positive results if Type I error was not well-controlled by such default options. The absence of functional immune assays limits causal inference and mechanistic interpretation of the observed transcriptomic and proteomic alterations. Future investigations incorporating functional validation experiments will be necessary to confirm the biological roles of the identified immune signatures and to explore their potential relevance for risk stratification, prevention, and management of GDM in ART pregnancies.
In summary, we conducted dynamic monitoring of peripheral circulating immune phenotypes in pregnant women undergoing ART, providing a comprehensive overview of the immune landscape associated with GDM from multiple dimensions, including peripheral immune cell types, immune cell activation status, immune repertoire diversity, and levels of inflammatory factors. Adaptive and innate immune cells in the GDM group collectively exhibited an upregulated proinflammatory state. The transcription factors FOS, JUN, JUNB, JUND, FOSB, and KLF6 were determined to be critical regulators of the immune response in patients with GDM. This study enhances our understanding of the molecular mechanisms underlying GDM in pregnant women undergoing assisted reproduction and provides a robust theoretical foundation for the management of high-risk pregnancies associated with ART.
Methods
Study participants
All participants were pregnant women who conceived via ART and were recruited from the NBart cohort between January 1, 2022, and June 30, 202434. Participants were enrolled prior to the initiation of ART and prospectively followed until undergoing a 75-g OGTT at 24–28 weeks of gestation. Based on OGTT results, participants were classified into the GDM group or the NP group.
Notably, this study did not include a spontaneous conception control group, either with or without GDM. Consequently, it is not possible to disentangle whether the observed immune alterations are attributable primarily to GDM pathophysiology, to metabolic and hormonal perturbations associated with ART procedures, or to their combined effects.
The eligible participants were defined as follows: women aged 20 to 34 years; primiparous women who underwent IVF or ICSI; and those with singleton pregnancies. The exclusion criteria included a pre-pregnancy body mass index (BMI) of less than 18.5 or 28 or higher; a history of pre-pregnancy diabetes, pre-diabetes, or impaired fasting glucose; other pre-pregnancy endocrine disorders, such as hypogonadotropic hypogonadism, hyperprolactinemia, and thyroid diseases; chronic conditions prior to pregnancy, including hypertension, cardiovascular diseases, renal diseases, autoimmune diseases, liver function abnormalities, neurological disorders, or malignancies; uncontrolled severe infections; psychological or psychiatric conditions that would impede participation in cohort studies during pregnancy; no pregnancy or miscarriage occurring after ART; and other pregnancy complications, such as gestational hypertension (GH), preeclampsia, eclampsia, hypothyroidism, and hyperthyroidism identified during follow-up. A total of 32 GDM patients were ultimately included in the study, along with 31 matched women with NPs. The two groups were matched for age, pre-pregnancy BMI, early pregnancy BMI, and type of ART treatment.
Ethical approval
Written informed consent was obtained from all participants. This study was approved by the Ningbo University Medical Science Research Ethics Committee. All ethical regulations relevant to human research participants were followed.
Sample collection and isolation
Peripheral blood samples were collected from all participants during three distinct periods: the baseline period (pre-pregnancy), early clinical pregnancy period (4–6 weeks of gestation), and second trimester (24–28 weeks of gestation). From these samples, plasma and PBMCs were isolated and preserved. The participants were instructed to provide 10 mL of peripheral blood in two EDTA tubes before 10 AM while they were in a fasting state (with no food intake after 10 p.m. the previous evening). The blood samples were gently inverted 5–10 times to ensure adequate mixing, allowed to rest for 30 min at room temperature, and subsequently centrifuged at 4 °C with a relative centrifugal force of 1600 × g for 10 min. The resulting plasma was then collected and stored at −80 °C. Ficoll-Hypaque density gradient centrifugation was used to isolate the PBMCs from the remaining blood. First, an equal volume of PBS was added to a 5 mL blood sample to achieve a 1:1 dilution. Following thorough mixing, the diluted sample was carefully transferred to a centrifuge tube containing 5 mL of Ficoll-Paque. Next, the samples were centrifuged at room temperature at 1500 × g for 30 min with no brake. The upper layer containing the serum and platelets was carefully discarded. A Pasteur pipette was used to gently aspirate the PBMC layer, which was subsequently transferred to a new centrifuge tube. The PBMCs were washed with 10 mL of PBS and centrifuged at 350 × g for 10 min at 4 °C. The supernatant was discarded, and the cell pellet was resuspended in an appropriate buffer. Finally, the recovered PBMCs were counted using a hemocytometer or automated cell counter and stored in liquid nitrogen.
Preparation of single-cell suspensions
Single-cell suspensions were prepared in accordance with the 10x Genomics Cell Preparation Guide (CG00053) and Guidelines for Optimal Sample Preparation (CG000126) to minimize cell aggregation and multiplet formation. Cells were resuspended in ice-cold phosphate-buffered saline (PBS) supplemented with 0.04% bovine serum albumin (BSA).
Cell suspensions were examined microscopically using a hemocytometer or a Countess II automated cell counter to ensure single-cell dispersion and the absence of visible aggregates. When cell clumping was observed, suspensions were filtered through 40-μm cell strainers. Cell viability was assessed using SYTOX dye staining, and samples with viability below 70% underwent dead-cell removal using magnetic bead-based kits. Only samples with post-processing viability exceeding 80% were used for downstream library construction.
Cell concentrations were measured in triplicate, maintaining a coefficient of variation below 10%. Final cell concentrations were adjusted to 300–600 viable cells/μL prior to loading onto the Chromium chip.
Single-cell capture and GEM generation
Single-cell gel bead-in-emulsions (GEMs) were generated using the Chromium Single-Cell Controller with the Single-Cell 5′ Library and Gel Bead Kit (10x Genomics, Cat. No. 1000006) and the Chromium Single-Cell A Chip Kit (Cat. No. 120236), following the manufacturer’s standard protocol.
Approximately 6000 cells were loaded per channel to achieve an expected recovery of ~3000 cells. Cell loading concentrations were determined based on the Poisson distribution governing cell capture efficiency and multiplet formation. Under these conditions, the estimated multiplet rate was approximately 0.8%.
All cell suspensions were maintained at 4 °C throughout preparation and were loaded onto the Chromium Controller within 30 min of final preparation. Immediately before loading, suspensions were gently mixed by half-volume pipetting to ensure homogeneous cell distribution without inducing aggregation.
Reverse transcription and cDNA amplification
Following GEM generation, captured cells were lysed, and released RNA molecules were reverse-transcribed and barcoded within individual GEMs. Reverse transcription was performed using a Bio-Rad S1000 Touch Thermal Cycler under the following conditions: 53 °C for 45 min, 85 °C for 5 min, followed by a hold at 4 °C.
Subsequent cDNA amplification and library quality assessment were performed according to the manufacturer’s protocol. cDNA size distribution and concentration were evaluated using the Agilent 4200 TapeStation system.
Library construction and sequencing
Single-cell RNA-sequencing libraries were constructed using the Single-Cell 5′ Library and Gel Bead Kit in combination with V(D)J enrichment kits specific for human T cells (Cat. No. 1000005) and human B cells (Cat. No. 1000016).
Sequencing was performed on the Illumina NovaSeq 6000 platform using a paired-end 150-bp (PE150) configuration. A minimum sequencing depth of 100,000 reads per cell was achieved for gene expression libraries.
Single-cell RNA-sequencing data preprocessing
Raw sequencing data were processed using the Cell Ranger pipeline (version 8.0.0). Reads were demultiplexed, aligned to the human reference genome (GRCh38), and quantified to generate gene-barcode matrices. Unique molecular identifiers (UMIs) were used to collapse PCR duplicates and obtain accurate transcript counts for each cell.
All single-cell library preparation, sequencing, and initial data preprocessing were performed by CapitalBio Technology (Beijing, China).
Quality control and data integration
Downstream data processing and analysis were conducted using R software (version 4.3.1) with the Seurat package (version 5.1.0). Genes detected in fewer than three cells were excluded. Cells meeting any of the following criteria were removed from subsequent analyses: fewer than 200 or more than 9000 detected genes, mitochondrial gene expression exceeding 25%, or red blood cell gene expression exceeding 3%.
Gene expression matrices were normalized using the NormalizeData function. The 3000 most variable genes were identified using FindVariableFeatures. Principal component analysis (PCA) was performed using the RunPCA function, and batch effects across samples were corrected using the RunHarmony algorithm110.
Unsupervised clustering analysis and dimensionality reduction
Using the scree plot derived from PCA and the visualization results obtained from the clustree function111, we determined the most appropriate dimensional range and resolution for clustering cell subtypes. Accordingly, FindNeighbors (dims = 1:30) and FindClusters (resolution = 0.6) were selected for PBMC subpopulation clustering. Additionally, two-dimensional UMAP was used for nonlinear dimensionality reduction.
T cells and B cells/plasma cells were extracted based on the annotations of the main cell types in the PBMCs. These two primary cell types were then separately integrated for further subpopulation clustering, gene expression normalization, PCA, and clustering.
Cell type annotation and identification of marker genes
The FindAllMarkers function in Seurat was used to identify marker genes for each cell subtype. The parameters for marker gene selection were configured as follows: genes must be detected in a minimum of 25% of the cells; a differential expression analysis was conducted using the Wilcoxon rank-sum test, which requires a log fold change (logFC) ≥ 0.25; and p values < 0.05 following the Bonferroni correction. Thereafter, the cell subtypes were annotated based on canonical markers of specific cell types.
Differential gene expression and functional enrichment analyses
The FindMarkers function from the Seurat R package was used to identify DEGs between the GDM and NP groups. DEGs were defined as those with an absolute log2FC ≥ 0.25 and adjusted p values <0.05 following the Bonferroni correction. Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of the DEGs were performed using the enrichGO and enrichKEGG functions. Moreover, gene set enrichment analysis (GSEA) was performed with GO terms and KEGG pathways using clusterProfiler 4.0112.
Protein-protein interaction (PPI) analysis
The PPI network of DEGs was constructed using the STRING database (http://string-db.org/), and the hub genes within this PPI network were identified using the CytoHubba plugin in Cytoscape version 3.10.3.
T cell immune-inflammatory state score
The AddModuleScore function was utilized to evaluate the state scores of predefined gene sets within T cell subsets. These gene sets were derived from the Molecular Signatures Database (MSigDB) and defined T cell functional gene sets35,39,56,60,113,114,115,116,117,118. The scores of the immune response-regulating signaling pathway (GO:0002764), regulation of T cell activation (GO:0050863), positive regulation of lymphocyte activation (GO:0051251), biocarta_Th1th2 pathway, biocarta_IL17 pathway, reactome TNF signaling, T cell receptor signaling pathway (GO:0050852), leukocyte activation involved in the inflammatory response (GO:0002269), immune response regulating the cell surface receptor signaling pathway (GO:0002768), defined T cell effector-related genes, cytotoxicity-related genes, cytokine-related genes, and inflammation-related genes were calculated to evaluate the immune-inflammatory state of the T cell subtypes.
Analysis of variance (ANOVA) was employed to examine the differences in gene set scores across various T cell subtypes. The Wilcoxon rank-sum test was performed to assess the differences in functional gene set scores between the GDM and NP groups. Visualization was performed using the ggplot2 and ggboxplot functions.
Single-cell TCR/BCR repertoire analysis
The Chromium Single-Cell V(D)J Enrichment Kit (10x Genomics, PN-1000005 for TCR and PN-1000016 for BCR) was used to enrich the TCR/BCR V(D)J gene segments by PCR amplification of the 5’ end cDNA library. Using the Cell Ranger v.8.0.0 pipeline, TCR/BCR sequences were assembled, followed by deduplication, gene quantification, and clonotype assignment. This process yielded barcoded TCR/BCR CDR3 sequences and rearranged full-length BCR/TCR V(D)J segments along with their respective clonotype frequencies, all of which were referenced to the GRCh38 genome36,119.
The T/B receptor clonotypes were subsequently mapped to T and B cells for further analysis. Initially, we quantified the clonal expansion ratios for T cell and B-cell subsets in each participant, categorized by the clone size (unique, 2–5, and ≥6). The Wilcoxon rank-sum test was employed to assess the differences in TCR/BCR clonal expansion between the GDM group and the NP group. Next, we identified the DEGs in clonally expanded T cells and B cells (clonotype frequency ≥2) between the GDM and NP groups and conducted functional enrichment analyses. Additionally, we examined the difference in the usage frequency of V genes, J genes, and V-J pairings between the two groups using the Wilcoxon rank-sum test. Finally, we defined the clonal abundance and diversity of TCRs/BCRs by calculating the number of similar TCR/BCR sequences within each cluster and applying the Hill diversity index, respectively. The clonal abundance and diversity were examined using the R packages alakazam function and clonotypeR function119.
Identification of GDM-associated TCR CDR3 sequences and antigen epitope annotation
A total of 115,702 T cells were analyzed, including 69,427 cells from the gestational diabetes mellitus (GDM) group and 46,275 cells from the normal pregnancy (NP) control group. TCR repertoire profiling yielded 80,168 unique TCR CDR3 amino acid (AA) sequences.
To identify TCR clonotypes uniquely expanded in the GDM group, CDR3 sequences that appeared exclusively in GDM samples and had a minimum clonal size ≥3 cells were defined as GDM-specific. This threshold ensured the detection of stable, biologically relevant clonal expansions.
For CDR3 sequences detected in both GDM and NP groups, differential abundance was evaluated using a 2 × 2 contingency table followed by Fisher’s exact test. Effect size was quantified by calculating the log2 fold change and odds ratio (OR). Multiple hypothesis testing was controlled by the Benjamini–Hochberg false discovery rate (FDR), and sequences with FDR < 0.05 were considered significantly enriched in either group.
Antigen epitope prediction for GDM-associated CDR3 sequences
Antigen epitope prediction for GDM-specific and GDM-enriched TCR CDR3 AA sequences was performed using the VDJdb database (https://vdjdb.cdr3.net/). VDJdb contains curated TCR–epitope interaction records. By querying CDR3 sequences against VDJdb, potential cognate antigen epitopes were inferred based on known TCR–antigen recognition patterns.
To determine the immune relevance of predicted epitopes in the context of diabetes, all VDJdb-inferred epitopes were cross-referenced with diabetes-related epitope records in the Immune Epitope Database (IEDB) (https://www.iedb.org/). This comparison enabled confirmation of potential associations between identified TCR CDR3 sequences and antigen epitopes implicated in metabolic and autoimmune pathways relevant to gestational diabetes.
Through combined VDJdb prediction and IEDB validation, antigen epitopes associated with GDM-related TCR CDR3 sequences were identified. These epitopes were subsequently used for biological interpretation and functional inference regarding immune mechanisms contributing to GDM pathogenesis.
GDM-specific BCR CDR3 sequences were defined as those that appear exclusively in the GDM group with a minimum cell count of 3, marking stable clonal expansion signals specific to the GDM group. For the shared CDR3 sequences (those occurring in GDM and NP groups), a 2 × 2 contingency table was constructed and Fisher’s exact test was used to assess frequency differences between groups, while log2 fold change and odds ratio were calculated to measure the effect size. Multiple comparisons were controlled using the Benjamini–Hochberg method to adjust for the false discovery rate (FDR), with FDR <0.05 used to determine statistical significance. The comprehensive sequence alignment analysis between GDM-specific BCR CDR3 sequences and publicly available SARS-CoV-2 antibody repertoires from the CoV-AbDab database (https://opig.stats.ox.ac.uk/webapps/covabdab/), as well as epitope information from the IEDB database (https://www.iedb.org/), using Levenshtein distance-based alignment.
Pseudotime analysis of the T and B subtypes
Monocle 2 was used to construct single-cell trajectories for CD4+ T cell subtypes, CD8+ T cell types and B-cell subtypes120. The expression matrix, gene data frame, and phenotypic information from the annotated cells in the Seurat object were extracted to construct a CellDataSet object. Gene expression across cells was normalized using the estimateSizeFactors function, and the data dispersion was calculated using the estimateDispersions function. Genes expressed in fewer than 10 cells were filtered based on the criteria defined by detectGenes (object, min_expr=0.1) and row.names (subset(fData(object), num_cells_expressed≥10)). Finally, dimensionality reduction, clustering, and trajectory inference were performed on the top 2000 DEGs using the reduceDimension function with the DDRTree method.
Single-cell regulatory network inference and clustering (SCENIC) analysis
SCENIC analysis was performed to identify gene regulatory networks specific to different cell subtypes and disease states. This analysis was implemented using the R/bioconductor package SCENIC, along with GENIE3, RcisTarget, and AUCell121. GENIE3 was employed to infer coexpression modules between TFs and their candidate target genes. RcisTarget was subsequently used for the TF motif enrichment analysis to identify direct targets known as regulons. Finally, AUCell was used to quantify the activity of each regulon in every cell, generating regulon activity scores (RASs) and regulon specificity scores (RSSs). The differentially expressed TFs and TF-regulated target genes were also selected for GO and KEGG enrichment analyses.
GENIE3 was applied to infer co-expression relationships between transcription factors and their candidate target genes. GENIE3 uses a random forest-based regression approach to predict the expression of each target gene based on the expression profiles of all candidate TFs. Specifically, for each gene, ensemble tree models were trained using TF expression as predictor variables, and TF importance scores were calculated based on their contribution to explaining target gene expression variance. This procedure generated a ranked list of TF-target gene associations, representing putative regulatory interactions based on co-expression patterns. GENIE3 analysis was performed separately within major immune cell subsets, including CD4⁺ and CD8⁺ T cells, to ensure cell-type-specific regulatory network inference.
To distinguish direct TF-target interactions from indirect co-expression relationships, RcisTarget was used to perform transcription factor binding motif enrichment analysis. For each TF-centered co-expression module inferred by GENIE3, RcisTarget assessed the enrichment of known TF binding motifs within the promoter regions and distal regulatory elements of candidate target genes, using curated motif databases. Genes showing significant enrichment for the corresponding TF motif were defined as direct regulatory targets. A regulon was defined as a transcription factor together with its set of validated direct target genes that satisfied both co-expression and motif enrichment criteria. Only regulons meeting the following quality control criteria were retained for downstream analysis: (1) significant motif enrichment (adjusted p < 0.05), and (2) a minimum regulon size of at least 10 target genes. This filtering strategy ensured high-confidence reconstruction of TF regulatory programs.
To quantify regulon activity in individual cells, AUCell (Area Under the recovery Curve) analysis was performed. For each cell, all detected genes were ranked in descending order according to their expression levels, generating a cell-specific gene expression ranking. For each regulon in each cell, AUCell computed a recovery curve representing the cumulative number of regulon target genes recovered along the ranked gene list. The area under this recovery curve (AUC) was calculated to quantify regulon activity, with higher AUC values indicating coordinated high expression of regulon target genes within a cell. This analysis produced a continuous regulon activity matrix (cells × regulons), representing regulon activity scores at single-cell resolution.
To distinguish biologically meaningful regulon activation from background noise, regulon-specific AUC thresholds were determined using the default data-driven SCENIC approach. For each regulon, the distribution of AUC values across all cells was modeled to identify an inflection point separating low-activity background from high-activity states.
These regulon-specific thresholds were optimized to minimize false-positive activation calls while preserving cell-type-specific regulatory signals. Based on these thresholds, regulon activity was binarized into active or inactive states for each cell, generating a binary regulon activity matrix.
Two complementary metrics were derived from AUCell analysis: Regulon activity scores, defined as continuous AUC values representing the overall activity level of each regulon in individual cells. These scores were used for quantitative comparisons of regulon activity between GDM and NP groups. Regulon specificity scores (RSS), calculated to identify regulons preferentially active in specific cell subtypes or disease states. RSS values quantify the degree of activity specificity using distribution-based divergence metrics, such as Jensen-Shannon divergence.
Differential regulon activity between GDM and NP groups was assessed within each immune cell subtype using Wilcoxon rank-sum tests applied to regulon AUC score distributions. p values were adjusted for multiple testing using the Benjamini–Hochberg method. Regulons with an absolute log₂ fold change greater than 0.25 and adjusted p < 0.05 were considered differentially active.
Differentially expressed transcription factors and TF-regulated target genes were further subjected to GO and KEGG pathway enrichment analyses to characterize the biological functions and pathways associated with disease-specific regulatory programs. Regulon activity patterns were visualized using heatmaps to display group-wise differences and UMAP plots to overlay regulon activity onto low-dimensional single-cell embeddings.
Cell-cell communication analysis
The potential cell-cell communication between CD4+ T cells, CD8+ T cells, B cells, plasma B cells, NK cells, CD14+ monocytes, and CD16+ monocytes in the GDM group and the NP group was examined using CellPhoneDB v2.0122,123.
CellPhoneDB (v2.0) was used to identify statistically significant ligand-receptor interactions between immune cell subtypes based on curated databases of multi-subunit ligand-receptor complexes. For each cell subtype, the average expression levels and the proportion of cells expressing each ligand or receptor gene were calculated. Only ligand-receptor pairs expressed in more than 10% of cells in at least one interacting cell type were considered for downstream analysis.
To establish a null distribution for statistical testing, CellPhoneDB performed 1000 random permutations of cell subtype labels, recalculating the average expression of ligands and receptors within each permuted interaction pair. This permutation procedure generated a background (zero) distribution for each ligand–receptor pair across all pairwise cell-type comparisons.
The significance of each ligand-receptor interaction was assessed by comparing the observed average expression values to the corresponding null distribution, yielding cell-type-specific empirical p values. Ligand-receptor pairs with statistically significant enrichment were considered biologically relevant and retained for subsequent analyses and visualization.
To complement the statistical inference provided by CellPhoneDB, CellChat was applied to quantify the strength, directionality, and pathway-level organization of cell-cell communication. CellChat infers intercellular communication probabilities based on the law of mass action, integrating gene expression data with prior knowledge of ligand-receptor interactions, including multi-subunit receptor complexes and cofactors. For each ligand-receptor pair between sender cell type i and receiver cell type j, CellChat computed communication probabilities by integrating the average expression levels of signaling ligands in sender cells, cognate receptors in receiver cells, and relevant receptor cofactors when applicable. A trimmed mean approach was applied, removing the highest 10% of expression values to reduce the influence of outliers and rare highly expressing cells.
The resulting interaction scores were normalized to generate communication probabilities ranging from 0 to 1, where higher values indicate stronger predicted signaling from sender to receiver cell types. These probabilities were aggregated to generate a weighted, directed communication network, represented as an adjacency matrix in which each element reflects the overall interaction strength from cell type i to cell type j.
CellChat enabled hierarchical analysis of cell-cell communication at two complementary levels: (1) Ligand-receptor pair level, quantifying individual molecular interactions and identifying dominant ligand-receptor pairs mediating communication between specific cell types. (2) Signaling pathway level, aggregating functionally related ligand-receptor pairs into canonical signaling pathways and quantifying overall pathway activity and information flow between cell populations. This multi-level framework facilitated the identification of key signaling pathways and interaction patterns dysregulated in GDM compared with NP.
To identify disease-specific alterations in intercellular communication, comparative analyses were performed between GDM and NP groups. Differential interaction analyses were conducted to compare ligand-receptor interaction strengths and signaling pathway activities between groups.
Quantification of plasma inflammatory proteins
Plasma levels of inflammatory proteins were quantified using the Olink 96 Inflammation panel, which simultaneously measures 92 inflammation-related proteins based on proximity extension assay (PEA) technology. Plasma samples were obtained from 63 participants at three time points: pre-pregnancy, early clinical pregnancy (4–6 weeks of gestation), and the second trimester (24–28 weeks of gestation). All proteomic assays were performed by CapitalBio Technology (Beijing, China) following the manufacturer’s standard protocols.
Briefly, pairs of oligonucleotide-labeled antibodies specifically bound to target proteins in plasma samples. Upon dual recognition, the oligonucleotides were brought into proximity and extended by DNA polymerase to generate amplifiable DNA templates. The resulting DNA amplicons were quantified using real-time quantitative PCR (qPCR). Raw cycle threshold (Ct) values were processed using Olink’s standardized pipeline and converted into normalized protein expression (NPX) values on a log2 scale, which were used for all downstream analyses.
Proteomics quality control and data normalization
Quality control of the Olink proteomics data was conducted using the platform’s built-in internal and external control system, rather than traditional technical replicates or spike-in controls. This system includes four internal controls that monitor key assay steps: incubation controls assessing antibody binding performance, extension controls evaluating technical variation during oligonucleotide extension and amplification, and detection controls providing stable signals independent of immune reactions.
In addition, external controls-including inter-plate controls, negative controls, and recommended reference samples-were incorporated to ensure sample integrity and to correct for potential batch effects and inter-plate variability. Data normalization and quality assessment were performed according to Olink’s standardized normalization workflow (Data normalization v2.2), ensuring consistency and comparability of NPX values across samples and assay plates.
Differentially expressed inflammatory proteins between groups were identified using the Wilcoxon rank-sum test, with a significance threshold of p <0.05. Pearson correlation analysis was performed to assess associations between individual inflammatory protein levels and glucose-related traits.
To examine the relationship between inflammatory protein profiles and immune cell composition, Mantel’s test was conducted using distance matrices derived from inflammatory protein levels and immune cell subset proportions. Functional enrichment analyses, including GO and KEGG pathway analyses, were performed for inflammatory proteins significantly associated with glucose-related traits.
Integration with single-cell transcriptomic data
To integrate plasma proteomic data with single-cell transcriptomic profiles, genes encoding the 92 inflammatory proteins were mapped to the single-cell RNA-sequencing dataset. Inflammatory factor activity at the single-cell level was quantified by calculating module scores using the AddModuleScore function implemented in the Seurat package.
Differences in inflammatory factor module scores between the gestational diabetes mellitus (GDM) group and the normoglycemic pregnancy (NP) group were assessed using the Wilcoxon rank-sum test.
Statistics and reproducibility
A total of 63 primiparous women with singleton pregnancies undergoing ART were included in this study, comprising 32 women with GDM and 31 matched NP controls. Longitudinal plasma inflammatory profiling using the Olink 96 Inflammation panel was performed in all participants at three time points (pre-pregnancy, early pregnancy, and second trimester). Single-cell RNA sequencing and 5′ V(D)J repertoire sequencing were conducted on PBMCs obtained at 24–28 weeks of gestation from a subset of 10 women with GDM and 10 matched NP controls.
Each participant was considered an independent biological replicate. No technical replicates were generated for single-cell RNA sequencing, V(D)J sequencing, or proteomic analyses. Repeated measurements collected from the same individual at different time points were treated as related observations. Experiments were not randomized, and investigators were not blinded to group allocation.
Continuous variables were assessed for normality prior to analysis. Differences between the GDM and NP groups were evaluated using the unpaired Student’s t test or the Wilcoxon rank-sum test, as appropriate. Categorical variables were compared using the chi-square test. Correlations between continuous variables were assessed using Pearson’s or Spearman’s correlation coefficients. Mantel’s test was applied to evaluate correlations between matrices of correlated variables. All statistical tests were two-tailed, and a p value <0.05 was considered statistically significant.
All statistical analyses and data visualization were performed using R software (version 4.3.1). Bioinformatic analyses were conducted using standard workflows and default parameters unless stated otherwise. Detailed information on statistical analyses and sample sizes is provided in the “Methods”, figure legends, and table footnotes.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2025) in the National Genomics Data Center (Nucleic Acids Res 2025), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA-Human: HRA015794), which are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human. Numerical source data underlying figures can be found in Supplementary Data 1. All other data are available from the corresponding author on reasonable request.
Code availability
The code used to perform the analyses in this study is available on GitHub at https://github.com/zhuhui-library/GDM_scRNAseq_ART and has been archived in Zenodo with the https://doi.org/10.5281/zenodo.18195363124.
References
Sweeting, A. et al. Epidemiology and management of gestational diabetes. Lancet 404, 175–192 (2024).
Wang, H. et al. IDF diabetes atlas: estimation of global and regional gestational diabetes mellitus prevalence for 2021 by International Association of Diabetes in Pregnancy Study Group’s Criteria. Diab. Res. Clin. Pr. 183, 109050 (2022).
Simmons, D. et al. Treatment of gestational diabetes mellitus diagnosed early in pregnancy. N. Engl. J. Med. 388, 2132–2144 (2023).
Sweeting, A. et al. Relationship between early-pregnancy glycemia and adverse outcomes: findings from the TOBOGM study. Diab. Care 47, 2085–2092 (2024).
Simmons, D. et al. Perinatal outcomes in early and late gestational diabetes mellitus after treatment from 24-28 weeks’ gestation: a TOBOGM secondary analysis. Diab. Care 47, 2093–2101 (2024).
Metzger, B. E. et al. Hyperglycemia and adverse pregnancy outcomes. N. Engl. J. Med. 358, 1991–2002 (2008).
Riskin-Mashiah, S., Younes, G., Damti, A. & Auslender, R. First-trimester fasting hyperglycemia and adverse pregnancy outcomes. Diab. Care 32, 1639–1643 (2009).
Lowe, W. L. Jr. et al. Association of gestational diabetes with maternal disorders of glucose metabolism and childhood adiposity. JAMA 320, 1005–1016 (2018).
Lowe, W. L. Jr. et al. Hyperglycemia and adverse pregnancy outcome follow-up study (HAPO FUS): maternal gestational diabetes mellitus and childhood glucose metabolism. Diab. Care 42, 372–380 (2019).
Ying, Q. et al. Gestational diabetes mellitus and risk of long-term all-cause and cardiac mortality: a prospective cohort study. Cardiovasc. Diabetol. 23, 47 (2024).
Lowe, L. P. et al. Associations of glycemia and lipid levels in pregnancy with dyslipidemia 10-14 years later: the HAPO follow-up study. Diab. Res. Clin. Pr. 185, 109790 (2022).
Organization, W. H. 1 in 6 people globally affected by infertility: WHO. Saudi Med. J. 44, 524–525 (2023).
Herbert, M., Choudhary, M. & Zander-Fox, D. Assisted reproductive technologies at the nexus of fertility treatment and disease prevention. Science 380, 164–167 (2023).
Pinborg, A., Wennerholm, U. B. & Bergh, C. Long-term outcomes for children conceived by assisted reproductive technology. Fertil. Steril. 120, 449–456 (2023).
Vergani, M. et al. Prevalence of gestational diabetes mellitus risk factors in singleton pregnancies obtained by assisted reproductive technology: an observational, retrospective, real-world study from a pregnancy registry. Diab. Res. Clin. Pr. 210, 111654 (2024).
Bosdou, J. K. et al. Risk of gestational diabetes mellitus in women achieving singleton pregnancy spontaneously or after ART: a systematic review and meta-analysis. Hum. Reprod. Update 26, 514–544 (2020).
Bianchi, C. et al. Assisted reproductive technology, risk of gestational diabetes, and perinatal outcomes in singleton pregnancies. Diab. Metab. Res. Rev. 39, e3625 (2023).
Holst, S., Kjær, S. K., Jørgensen, M. E., Damm, P. & Jensen, A. Fertility problems and risk of gestational diabetes mellitus: a nationwide cohort study. Fertil. Steril. 106, 427–434.e421 (2016).
Anagnostis, P., Tarlatzis, B. C. & Kauffman, R. P. Polycystic ovarian syndrome (PCOS): long-term metabolic consequences. Metabolism 86, 33–43 (2018).
Sachdev, D., Sauer, M. V. & Ananth, C. V. Gestational diabetes mellitus in pregnancies conceived after infertility treatment: a population-based study in the United States, 2015-2020. F. S. Rep. 5, 102–110 (2024).
He, Y. C. et al. Serum anti-Müllerian hormone levels are associated with perinatal outcomes in women undergoing IVF/ICSI: a multicenter retrospective cohort study. Front. Endocrinol. 14, 1081069 (2023).
Tanaka, T. et al. Oestrogen receptor α in T cells controls the T cell immune profile and glucose metabolism in mouse models of gestational diabetes mellitus. Diabetologia 64, 1660–1673 (2021).
Yang, Y. et al. Transcriptomic profiling of human placenta in gestational diabetes mellitus at the single-cell level. Front. Endocrinol. 12, 679582 (2021).
Shangaris, P. et al. Single-cell transcriptomics reveals markers of regulatory T cell dysfunction in gestational diabetes mellitus. Preprint, 1-21 https://doi.org/10.21203/rs.3.rs-3773991/v1 (2024).
Yin, M. et al. Adverse effects of gestational diabetes mellitus on fetal monocytes revealed by single-cell RNA sequencing. iScience 27, 108637 (2024).
Vannuccini, S. et al. Infertility and reproductive disorders: impact of hormonal and inflammatory mechanisms on pregnancy outcome. Hum. Reprod. Update 22, 104–115 (2016).
Brouillet, S. et al. C-reactive protein and ART outcomes: a systematic review. Hum. Reprod. Update 26, 753–773 (2020).
Parisi, F. et al. The pathophysiological role of estrogens in the initial stages of pregnancy: molecular mechanisms and clinical implications for pregnancy outcome from the periconceptional period to end of the first trimester. Hum. Reprod. Update 29, 699–720 (2023).
Raunig, J. M., Yamauchi, Y., Ward, M. A. & Collier, A. C. Placental inflammation and oxidative stress in the mouse model of assisted reproduction. Placenta 32, 852–858 (2011).
Zhang, Y. et al. Altered global gene expressions of human placentae subjected to assisted reproductive technology treatments. Placenta 31, 251–258 (2010).
Omazić, J. et al. Metabolic and immune parameters in pregnant women with impaired glucose metabolism—a pilot study. Metabolites 14, https://doi.org/10.3390/metabo14100551 (2024).
Chen, J., Kong, W. & Zhong, J. Editorial: Cross talk between the immune system and metabolism. Front. Endocrinol. 11, 588 (2020).
Kabat, A. M., Pearce, E. L. & Pearce, E. J. Metabolism in type 2 immune responses. Immunity 56, 723–741 (2023).
Zhu, H. et al. Cohort profile: the Ningbo birth cohort of population undergoing assisted reproductive technology (NBart). Int. J. Epidemiol. 53, https://doi.org/10.1093/ije/dyae086 (2024).
Zhang, C. et al. Single-cell RNA sequencing reveals intrahepatic and peripheral immune characteristics related to disease phases in HBV-infected patients. Gut 72, 153–167 (2023).
Zhang, J. Y. et al. Single-cell landscape of immunological responses in patients with COVID-19. Nat. Immunol. 21, 1107–1118 (2020).
Wang, Z. et al. Single-cell RNA sequencing of peripheral blood mononuclear cells from acute Kawasaki disease patients. Nat. Commun. 12, 5444 (2021).
Huang, L. et al. Dynamic blood single-cell immune responses in patients with COVID-19. Signal Transduct. Target Ther. 6, 110 (2021).
Chen, D. et al. Single-cell atlas of peripheral blood mononuclear cells from pregnant women. Clin. Transl. Med. 12, e821 (2022).
Terekhova, M. et al. Single-cell atlas of healthy human blood unveils age-related loss of NKG2C(+)GZMB(-)CD8(+) memory T cells and accumulation of type 2 memory T cells. Immunity 56, 2836–2854.e2839 (2023).
Cano-Gamez, E. et al. Single-cell transcriptomics identifies an effectorness gradient shaping the response of CD4(+) T cells to cytokines. Nat. Commun. 11, 1801 (2020).
Carmona, E. G. et al. Single-cell transcriptomic profiling reveals a pathogenic role of cytotoxic CD4(+) T cells in giant cell arteritis. J. Autoimmun. 142, 103124 (2024).
Wang, T. et al. Single-cell RNA sequencing reveals the sustained immune cell dysfunction in the pathogenesis of sepsis secondary to bacterial pneumonia. Genomics 113, 1219–1233 (2021).
Zhu, H. et al. Human PBMC scRNA-seq-based aging clocks reveal ribosome to inflammation balance as a single-cell aging hallmark and super longevity. Sci. Adv. 9, eabq7599 (2023).
Lee, A. H. et al. Neoadjuvant PD-1 blockade induces T cell and cDC1 activation but fails to overcome the immunosuppressive tumor associated macrophages in recurrent glioblastoma. Nat. Commun. 12, 6938 (2021).
Xia, J. et al. Single-cell landscape and clinical outcomes of infiltrating B cells in colorectal cancer. Immunology 168, 135–151 (2023).
Ota, M. et al. Multimodal repertoire analysis unveils B cell biology in immune-mediated diseases. Ann. Rheum. Dis. 82, 1455–1463 (2023).
Thiele, K., Diao, L. & Arck, P. C. Immunometabolism, pregnancy, and nutrition. Semin. Immunopathol. 40, 157–174 (2018).
Deer, E. et al. The role of immune cells and mediators in preeclampsia. Nat. Rev. Nephrol. 19, 257–270 (2023).
Zhu, H. et al. Comprehensive landscape of the T and B-cell repertoires of newly diagnosed gestational diabetes mellitus. Genomics 115, 110681 (2023).
Stern, J. E. et al. Assisted reproductive technology treatment increases obstetric and neonatal risks over that of the underlying infertility diagnosis. Fertil. Steril. 117, 1223–1234 (2022).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Förger, F. & Villiger, P. M. Immunological adaptations in pregnancy that modulate rheumatoid arthritis disease activity. Nat. Rev. Rheumatol. 16, 113–122 (2020).
Zhou, W. et al. Characterizing immune variation and diagnostic indicators of preeclampsia by single-cell RNA sequencing and machine learning. Commun. Biol. 7, 32 (2024).
Lorenzo, P. I., Martín-Montalvo, A., Cobo Vuilleumier, N. & Gauthier, B. R. Molecular modelling of islet β-cell adaptation to inflammation in pregnancy and gestational diabetes mellitus. Int. J. Mol. Sci. 20, https://doi.org/10.3390/ijms20246171 (2019).
Pan, L. et al. The causal relationship between 91 inflammatory cytokines and gestational diabetes mmellitus: a bidirectional two-sample Mendelian randomization study. Diab. Res Clin. Pr. 216, 111838 (2024).
Di Pietrantonio, N. et al. The inflammatory and oxidative phenotype of gestational diabetes is epigenetically transmitted to the offspring: role of methyltransferase MLL1-induced H3K4me3. Eur. Heart J. 45, 5171–5185 (2024).
O’Neill, C. M. et al. Circulating levels of IL-1B+IL-6 cause ER stress and dysfunction in islets from prediabetic male mice. Endocrinology 154, 3077–3088 (2013).
Eguchi, K. et al. Saturated fatty acid and TLR signaling link β cell dysfunction and islet inflammation. Cell Metab. 15, 518–533 (2012).
Pinto, Y. et al. Gestational diabetes is driven by microbiota-induced inflammation months before diagnosis. Gut 72, 918–928 (2023).
Ward, M. G., Li, G. & Hao, M. Apoptotic β-cells induce macrophage reprogramming under diabetic conditions. J. Biol. Chem. 293, 16160–16173 (2018).
Gayatri, V. et al. Crosstalk between inflammasomes, inflammation, and Nrf2: implications for gestational diabetes mellitus pathogenesis and therapeutics. Eur. J. Pharm. 963, 176241 (2024).
Erlebacher, A. Mechanisms of T cell tolerance towards the allogeneic fetus. Nat. Rev. Immunol. 13, 23–33 (2013).
Chien, Y. H., Meyer, C. & Bonneville, M. γδ T cells: first line of defense and beyond. Annu. Rev. Immunol. 32, 121–155 (2014).
Nielsen, M. M., Witherden, D. A. & Havran, W. L. γδ T cells in homeostasis and host defence of epithelial barrier tissues. Nat. Rev. Immunol. 17, 733–745 (2017).
PrabhuDas, M. et al. Immune mechanisms at the maternal-fetal interface: perspectives and challenges. Nat. Immunol. 16, 328–334 (2015).
Mor, G., Aldo, P. & Alvero, A. B. The unique immunological and microbial aspects of pregnancy. Nat. Rev. Immunol. 17, 469–482 (2017).
Yu, W., Miao, H. & Gong, Y. Lymphocyte subsets and cytokine changes in women with gestational diabetes mellitus: a systematic review. J. Diab. Res. 2025, 3494697 (2025).
Magalhaes, I. et al. Mucosal-associated invariant T cell alterations in obese and type 2 diabetic patients. J. Clin. Invest. 125, 1752–1762 (2015).
Carolan, E. et al. Altered distribution and increased IL-17 production by mucosal-associated invariant T cells in adult and childhood obesity. J. Immunol. 194, 5775–5780 (2015).
Rouxel, O. et al. Cytotoxic and regulatory roles of mucosal-associated invariant T cells in type 1 diabetes. Nat. Immunol. 18, 1321–1331 (2017).
Djudjaj, S. et al. Macrophage migration inhibitory factor mediates proliferative GN via CD74. J. Am. Soc. Nephrol. 27, 1650–1664 (2016).
Klasen, C. et al. MIF promotes B cell chemotaxis through the receptors CXCR4 and CD74 and ZAP-70 signaling. J. Immunol. 192, 5273–5284 (2014).
Zhang, L. et al. CD74 is a functional MIF receptor on activated CD4(+) T cells. Cell Mol. Life Sci. 81, 296 (2024).
Van Meerhaeghe, T., Néel, A., Brouard, S. & Degauque, N. Regulation of CD8 T cell by B-cells: a narrative review. Front. Immunol. 14, 1125605 (2023).
Bevan, M. J. Helping the CD8(+) T-cell response. Nat. Rev. Immunol. 4, 595–602 (2004).
Kampmann, U., Knorr, S., Fuglsang, J. & Ovesen, P. Determinants of maternal insulin resistance during pregnancy: an updated overview. J. Diab. Res. 2019, 5320156 (2019).
Plows, J. F., Stanley, J. L., Baker, P. N., Reynolds, C. M. & Vickers, M. H. The pathophysiology of gestational diabetes mellitus. Int. J. Mol. Sci. 19, https://doi.org/10.3390/ijms19113342 (2018).
Pantham, P., Aye, I. L. & Powell, T. L. Inflammation in maternal obesity and gestational diabetes mellitus. Placenta 36, 709–715 (2015).
McElwain, C. J., McCarthy, F. P. & McCarthy, C. M. Gestational diabetes mellitus and maternal immune dysregulation: what we know so far. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms22084261 (2021).
Gillespie, S. L., Porter, K. & Christian, L. M. Adaptation of the inflammatory immune response across pregnancy and postpartum in Black and White women. J. Reprod. Immunol. 114, 27–31 (2016).
Ray, G. W. et al. Genetic and inflammatory factors underlying gestational diabetes mellitus: a review. Front. Endocrinol. 15, 1399694 (2024).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Maier, T., Güell, M. & Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 583, 3966–3973 (2009).
Buccitelli, C. & Selbach, M. mRNAs, proteins and the emerging principles of gene expression control. Nat. Rev. Genet. 21, 630–644 (2020).
Briney, B., Inderbitzin, A., Joyce, C. & Burton, D. R. Commonality despite exceptional diversity in the baseline human antibody repertoire. Nature 566, 393–397 (2019).
Kunik, V., Peters, B. & Ofran, Y. Structural consensus among antibodies defines the antigen binding site. PLoS Comput. Biol. 8, e1002388 (2012).
Huang, X. et al. Decreased monocyte count is associated with gestational diabetes mellitus development, macrosomia, and inflammation. J. Clin. Endocrinol. Metab. 107, 192–204 (2022).
Wang, J., Zhu, Q. W., Cheng, X. Y., Sha, C. X. & Cui, Y. B. Clinical significance of neutrophil-lymphocyte ratio and monocyte-lymphocyte ratio in women with hyperglycemia. Postgrad. Med. 132, 702–708 (2020).
Angelo, A. G. S. et al. Monocyte profile in peripheral blood of gestational diabetes mellitus patients. Cytokine 107, 79–84 (2018).
Lobo, T. F. et al. Impaired Treg and NK cells profile in overweight women with gestational diabetes mellitus. Am. J. Reprod. Immunol. 79, https://doi.org/10.1111/aji.12810 (2018).
Qiao, X., Pham, D. N., Luo, H. & Wu, J. Ran overexpression leads to diminished T cell responses and selectively modulates nuclear levels of c-Jun and c-Fos. J. Biol. Chem. 285, 5488–5496 (2010).
Yaseen, N. R., Park, J., Kerppola, T., Curran, T. & Sharma, S. A central role for Fos in human B- and T-cell NFAT (nuclear factor of activated T cells): an acidic region is required for in vitro assembly. Mol. Cell Biol. 14, 6886–6895 (1994).
Jain, J. et al. Normal peripheral T-cell function in c-Fos-deficient mice. Mol. Cell Biol. 14, 1566–1574 (1994).
Pollard, J. M. et al. Pregnancy dedifferentiates memory CD8+ T cells into hypofunctional cells with exhaustion-enriched programs. JCI Insight 9, https://doi.org/10.1172/jci.insight.176381 (2024).
Norton, M. T., Fortner, K. A., Oppenheimer, K. H. & Bonney, E. A. Evidence that CD8 T-cell homeostasis and function remain intact during murine pregnancy. Immunology 131, 426–437 (2010).
Xu, X. et al. The altered immunological status of children conceived by assisted reproductive technology. Reprod. Biol. Endocrinol. 19, 171 (2021).
Hu, C. et al. Supraphysiological estradiol promotes human T follicular helper cell differentiation and favours humoural immunity during in vitro fertilization. J. Cell Mol. Med. 25, 6524–6534 (2021).
Gao, F. et al. Single-cell profiling of the peripheral blood immune landscape during mid- and late-stage pregnancy. Physiol. Genomics 56, 855–868 (2024).
Orvieto, R. et al. C-reactive protein levels in patients undergoing controlled ovarian hyperstimulation for IVF cycle. Hum. Reprod. 19, 357–359 (2004).
Kollmann, Z., Schneider, S., Fux, M., Bersinger, N. A. & von Wolff, M. Gonadotrophin stimulation in IVF alters the immune cell profile in follicular fluid and the cytokine concentrations in follicular fluid and serum. Hum. Reprod. 32, 820–831 (2017).
Ye, H. Y. et al. Serum granulosa cell-derived TNF-α promotes inflammation and apoptosis of renal tubular cells and PCOS-related kidney injury through NF-κB signaling. Acta Pharm. Sin. 44, 2432–2444 (2023).
Pellicer, A. et al. The pathogenesis of ovarian hyperstimulation syndrome: in vivo studies investigating the role of interleukin-1beta, interleukin-6, and vascular endothelial growth factor. Fertil. Steril. 71, 482–489 (1999).
Orta-Mascaró, M. et al. CD6 modulates thymocyte selection and peripheral T cell homeostasis. J. Exp. Med. 213, 1387–1397 (2016).
Nagy, B. et al. Key to life: physiological role and clinical implications of progesterone. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms222011039 (2021).
Abramov, Y. et al. Plasma inflammatory cytokines correlate to the ovarian hyperstimulation syndrome. Hum. Reprod. 11, 1381–1386 (1996).
Baumgarten, S. C. et al. Evaluation of the maternal systemic immune system during frozen euploid embryo transfer according to cycle outcome. J. Reprod. Immunol. 164, 104261 (2024).
Zhao, Y. H. et al. Genomic expression profiles of blood and placenta reveal significant immune-related pathways and categories in Chinese women with gestational diabetes mellitus. Diabet. Med. 28, 237–246 (2011).
Lekva, T., Norwitz, E. R., Aukrust, P. & Ueland, T. Impact of systemic inflammation on the progression of gestational diabetes mellitus. Curr. Diab. Rep. 16, 26 (2016).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Grabski, I. N., Street, K. & Irizarry, R. A. Significance analysis for clustering with single-cell RNA-sequencing data. Nat. Methods 20, 1196–1202 (2023).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Guo, X. et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 24, 978–985 (2018).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913.e1819 (2021).
Mercado-Evans, V. et al. Gestational diabetes augments group B Streptococcus infection by disrupting maternal immunity and the vaginal microbiota. Nat. Commun. 15, 1035 (2024).
Wedekind, L. & Belkacemi, L. Altered cytokine network in gestational diabetes mellitus affects maternal insulin and placental-fetal development. J. Diab. Complications 30, 1393–1400 (2016).
Joshi, N. P., Madiwale, S. D., Sundrani, D. P. & Joshi, S. R. Fatty acids, inflammation and angiogenesis in women with gestational diabetes mellitus. Biochimie 212, 31–40 (2023).
Zhao, J. H. et al. Genetics of circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat. Immunol. 24, 1540–1551 (2023).
Zhang, F. et al. Adaptive immune responses to SARS-CoV-2 infection in severe versus mild individuals. Signal Transduct. Target Ther. 5, 156 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. https://doi.org/10.1038/s41596-024-01045-4 (2024).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Zhu, H. GDM_scRNAseq_ART: analysis code for ART-associated gestational diabetes single-cell transcriptomics. Zenodo, https://doi.org/10.5281/zenodo.18195363 (2026).
Acknowledgements
This research was supported by the Beijing Zhongwei Joint Funds of the Zhejiang Provincial Natural Science Foundation of China (project no. LBY24H040001 and project no. LBY24H040002), Ningbo Youth Science and Technology Innovation Leaders Project (project no. 2023QL058), Ningbo Public Welfare Technology Application Research Program (project no. 2023S113), Ningbo Key Research and Development Project (project no. 2024Z223 and project no. 2024Z225).
Author information
Authors and Affiliations
Contributions
J.X., L.D.J., N.L.S.T., and H.Z. conceived the study and designed the experiments. H.Z., J.C., Q.Y.X., L.Q.X., D.H., X.C., and J.H.H. performed the experiments, conducted analysis, and interpretation of the data. T.C., P.H.W., Y.Q.L., W.Y., B.Q.L., D.D.G., M.W.Y., and Y.Y.W. collected samples and clinical information from patients. H.Z., J.C., J.X., L.D.J., and N.L.S.T. composed, reviewed, and revised the manuscript. J.X. and L.D.J. contributed data and provided resources for the study. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Nishel M. Shah and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Eirini Trompouki and Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, H., Cai, J., Xu, Q. et al. Single-cell and proteomic profiling unveil aberrant immune phenotypes in gestational diabetes following assisted reproductive technology. Commun Biol 9, 335 (2026). https://doi.org/10.1038/s42003-026-09615-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09615-w