Abstract
The gut microbiota and its metabolites critically regulate immune cell phenotype, function and energy metabolism. We screened a collection of gut microbiota-related metabolites to identify modulators of mitochondrial metabolism in T cells. Here we show that indole-3-propionic acid (IPA) stimulates mitochondrial respiration of CD4+ T cells by increasing fatty acid oxidation (FAO) and amino acid oxidation (AAO), while inhibiting glycolytic capacity. IPA also impacts CD4+ T cell behaviour by inhibiting their differentiation to type 1 and type 17 helper T cell phenotypes. Mechanistically, the metabolic and immune effects of IPA are mediated by peroxisome proliferator-activated receptor-β/δ. The administration of IPA rescues mitochondria respiration in mice with gut bacteria depletion or colitis by enhancing FAO and AAO in colonic CD4+ T cells. Adoptive transfer experiments show that IPA acts on CD4+ T cells to exert its protective effect against inflammation. Collectively, our study reveals that the anti-inflammatory effects of IPA are mediated by metabolic reprogramming of CD4+ T cells toward the enhancement of mitochondrial respiration.
Similar content being viewed by others
Main
Host–microbiota interactions are now recognized as a significant factor in the maintenance of health and in the onset of several diseases, including inflammatory bowel disease (IBD), Crohn’s disease (CD) and ulcerative colitis (UC), which are chronic, relapsing inflammatory disorders of the gastrointestinal tract that disrupt patients’ health and quality of life1,2.
Classical host–microbe interaction concepts rely on the recognition of conserved microbial motifs by innate immunity sensors or on the effect of microbial molecules on a host cell receptor3,4. The gut microbiota plays a critical role in maintaining mucosal immunity, immune cell maturation, differentiation and functions5. Moreover, energy metabolism plays a crucial role in mounting the appropriate immune cell response. It has been demonstrated that gut microbiota influences immune cell metabolism, particularly through the impact of its metabolites such as short-chain fatty acids (SCFAs), bile acids (BAs) and tryptophan-derivedmetabolites6,7. These metabolites can alter the physiology and function of host cells by modulating major energy metabolic pathways, including glycolysis, the tricarboxylic acid (TCA) cycle and fatty acid (FA) and amino acid (AA) metabolism.
The gut bacteria have a special relationship with host metabolism, notably via the mitochondria, due to their common origin. Significant functional, morphological and molecular overlaps exist between gut bacteria and mitochondria8, and emerging evidence implicates bidirectional communication between these entities to regulate the fate and function of each other9. At the same time, mitochondria are responsible for numerous essential cellular functions. Notably, the integrity of mitochondrial function in intestinal epithelial systems plays a critical role in maintaining the homeostasis of intestinal health10. Several studies also implicate mitochondrial dysfunction in IBD pathogenesis11,12.
In this study, we focused on the effect of the gut microbiota on T cells, which are crucial players in intestinal health and disease, including IBD13,14. Phenotypic screening of microbiota-derived metabolites for their effects on T cells’ energy metabolism led to the identification of indole-3-propionic acid (IPA), which we discovered can increase mitochondrial respiration in CD4+ T cells by enhancing FA oxidation (FAO) and AA oxidation (AAO) both in vitro and in vivo. The rewiring of T cell energy metabolism, which was dependent on PPAR β/δ signalling yet independent of aryl hydrocarbon receptor (AhR) signalling, underlies the reprogramming of CD4+ T cell’s differentiation and anti-inflammatory effects. IPA reduces the differentiation of CD4+ T cells toward type 1 helper T (TH1) and TH17 cells without interfering with regulatory T (Treg) cell activity. This effect occurs following a relatively short exposure and does not require the continuous presence of IPA. IPA exhibited protective effects in a colitis mouse model, which aligns with human data from IBD cohorts.
Results
IPA reprograms the energy metabolism of human T cells
Based on a literature search, we selected 85 metabolites produced or transformed by the gut microbiota to be subjected to phenotypic screening of human T cell to identify molecules able to enhance energy metabolism. These include notably BAs, SCFAs, AAs, organic acids and tryptophan derivatives15,16 (Supplementary Table 1 and Fig. 1a).
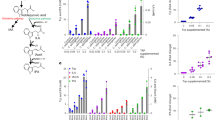
a, Schematic diagram showing the metabolic screening process from Jurkat T cell line to human PBMCs. Created with BioRender.com. b, Schematic diagram showing the protocol of human PBMCs treated with IPA in basal and CD3/CD28 Dynabeads activation state. Created with BioRender.com. c,d, t-SNE (t-distributed stochastic neighbour embedding) analysis of PBMC subsets and puromycin-ATP synthesis signal in control (Con, c) and CD3/CD28 groups (d). e, Metabolic profile of distinct subsets in PBMCs, including CD4+, CD8+, DN+ T cells, B cells and HLA-DR+ antigen-presenting cells induced by IPA (10, 100 and 1,000 μM) in basal conditions. n = 7 donors per group. f, Bar chart indicating CD3/CD28 activation highly increased total ATP synthesis in lymphocyte subsets in PBMCs. n = 6 donor per group. g, Metabolic profile of distinct lymphocyte subsets in PBMCs, including CD4+, CD8+ and DN+ T cells induced by IPA (10, 100 and 1,000 μM) in a CD3/CD28 activation state. n = 3–6 donors per group. Each point represents an independent donor with value below 0 or above 100 are excluded from the analysis. Data represent mean ± s.e.m. analysed by one-way analysis of variance (ANOVA) with Tukey’s correction for multiple comparisons (e,g) and mean ± s.e.m. analysed by two-tailed Student’s t-test (f). NS, not significant.
We monitored the impact screened metabolites had on ATP levels of Jurkat human T cell line cultured in the presence or absence of 2-deoxy-D-glucose (2-DG; a glucose analogue that inhibits glycolysis) and oligomycin (that inhibits the mitochondrial- ATP synthase) to gain insights into the pathways influenced by each metabolite. IPA particularly piqued our interest because it increased ATP levels in basal conditions in a manner dependent both upon mitochondrial respiration and, to a lesser extent, glycolysis (Extended Data Fig. 1a–c). We then evaluated the effect of IPA on the energy metabolism of different primary cell subsets of human peripheral blood mononuclear cells (PBMCs) using SCENITH17 (single-cell energetic metabolism by profiling translation inhibition), a technology that profiles energy metabolism in heterogeneous samples with single-cell resolution (Fig. 1b). Using CD3, CD4, CD8, CD19 and HLA-DR markers, five phenotypically distinct clusters were generated through dimensionality reduction analysis. Compared with the control (Con) setting, CD3/CD28 activation, allowing for efficient activation of T cells, profoundly modified energy metabolism without altering the immune cell population composition (Fig. 1c,d). In basal conditions, IPA increased mitochondrial dependence of CD4+ T cells, double-negative (DN) T cells (CD3+CD4−CD8−), CD8+ T cells, CD19+ B cells and HLA-DR+ antigen-presenting cells by increasing the utilization of glucose, pointing to a ubiquitous response. On the contrary, IPA decreased glycolytic capacity, FAO and AAO capacity (Fig. 1e). In fact, following CD3/CD28 activation, the total ATP content was dramatically increased in CD4+, CD8+, DN T cells and B cells, meaning that cellular energy metabolism was strongly activated (Fig. 1c,d,f). As previously shown18, CD3/CD28 activation of T cells led to a massive increase of glycolytic capacity and, conversely, to a decreased mitochondrial dependence. Treatment of activated T cells with IPA led to a metabolic shift away from CD3/CD28-dependent aerobic glycolysis and towards enhanced mitochondrial respiration. In line, FAO and AAO capacity were significantly increased by IPA treatment in the CD3/CD28 activated cells compared with the CD3/CD28-activated control cells that were not incubated with IPA (Fig. 1g).
To further investigate whether IPA directly alter the energy metabolism state of CD4+ T cells, independently of the cell activation status, we included CD69 as an early T cell activation marker in the SCENITH panel. Regardless of baseline or CD3/CD28 activation status, different doses of IPA (from 1 μM to 1,000 μM) did not change CD69 mean fluorescence intensity (MFI), nor the proportion of CD69+ cells in CD4+ T cells in human PBMCs (Supplementary Fig. 1a). When the analysis was restricted to CD69+CD4+ T cells, IPA could still increase mitochondrial dependence through boosting FAO and/or AAO, which is consistent with what we observed previously for total CD4+ T cells (Extended Data Fig. 2a). We also further subdivided CD4+ helper T (TH) lymphocytes, focusing more on the role of IPA on changing the proportion and energy metabolism of already established TH memory subsets, including TH1, TH17 and TH1/TH17. IPA did not change the proportion of TH1, TH1/TH17 or TH17 memory cell proportion in total CD4+ T cells in both basal and CD3/CD28 activation state (Supplementary Fig. 1b). Regarding the SCENITH energy metabolism part, in accordance with what we observed for total CD4+ T cells, IPA increased mitochondrial dependence in TH1 and TH17 memory TH cells (Extended Data Fig. 2b,c). IPA also had similar but weaker effects in CXCR3+CCR6+ TH1/TH17 cell subsets (Supplementary Fig. 1c). Taken together, these results demonstrate that IPA boosts human primary T cells energy metabolism by increasing mitochondrial respiration in both basal and CD3/CD28 activation conditions.
IPA alters the metabolic profile of mouse CD4+ T cells
To explore the mechanisms involved in the effects of IPA, we first analysed primary CD4+ T cells from mouse spleen using SCENITH (Extended Data Fig. 3a). CD3/CD28 activation increased total ATP synthesis in CD4+ and CD8+ T cells as expected (Extended Data Fig. 3b,c). In activated CD4+ T cells, IPA significantly increased mitochondrial respiration and, conversely, reduced glycolytic capacity. This effect was achieved by increasing the FAO and AAO capacity and reducing glucose dependence (Extended Data Fig. 3d). However, IPA did not alter the metabolic profile of CD8+ T cells (Extended Data Fig. 3e). Taken together, these results show that IPA promotes a metabolic shift to boost the energy metabolism of primary mouse and human CD4+ T cells alike (Extended Data Fig. 3f).
IPA boosts mitochondrial activity of mouse CD4+ T cells through FAO and AAO, while inhibiting glycolysis capacity
The mouse model being relevant for studying the effects of IPA on CD4+ T cells, we employed purified primary mouse spleen CD4+ T cells and assessed their metabolic activity using the Seahorse Flux Analyzer, which concomitantly measures cellular oxygen consumption (OCR) and extracellular acidification rate. As expected, CD3/CD28 activation strongly increased glycoATP (ATP generated through glycolysis) and, to a lesser extent, mitoATP (ATP produced through oxidative phosphorylation; OxPhos) (Fig. 2a). The ATP rate index, the ratio of mitoATP on glycoATP, significantly decreased with CD3/CD28 activation, confirming a switch from mitochondrial respiration to aerobic glycolysis.
a, GlycoATP and MitoATP production of mouse CD4+ T cells induced by IPA in basal and CD3/CD28 activation status in 6 h. n = 6. b, ATP rate index of CD4+ T cells treated with and without IPA in basal and CD3/CD28 activation state at 6 h. n = 6 except for Con and IPA1000 group with n = 5 (outliers are removed for analysis). c, Energetic map generated by ATP rate assay. n = 6 except for Con and IPA1000 group with n = 5. d, Seahorse Mitostress assay analysis of mouse CD4+ T cells cultured with and without CD3/CD28 in the presence of DMSO or IPA (from 1 μM to 1,000 μM) at 6 h. e, Basal and maximal respiration measurement in the Mitostress assay. For d,e, n = 8 for Con, all doses of IPA in basal conditions and CD3/CD28+ IPA100, CD3/CD28+ IPA1000. n = 6 for the remaining three. f, Maximal respiration generated from Seahorse substrate oxidation assay of CD4+ T cells cultured with and without CD3/CD28 in the presence of DMSO or IPA (100 and 1,000 μM) in control (Con), etomoxir, BPTES and UK5099 groups. The data are shown at 6 h. n = 3 for Con and CD3/CD28 group, n = 4 for IPA-treated groups in basal and CD3/CD28 activation state. g, Seahorse glycolytic rate assay analysis of mouse CD4+ T cells cultured with and without CD3/CD28 in the presence of DMSO or IPA (from 1 μM to 1,000 μM)) at 6 h. n = 8. h, Basal and compensatory glycolysis measurement in glycolytic rate assay above. n = 8. i, Lactate production of CD4+ T cells stimulated with/without CD3/CD28 in the presence of DMSO or IPA (from 1 μM to 1,000 μM)) at 6 h. n = 6 except n = 5 for IPA1000 and CD3/CD8 group. j–l, Representative images (j) and mitochondrial and cristae parameter analysis (k,l) of CD4+ T cells cultured for 24 h in basal and CD3/CD28 activation state in the presence of DMSO or IPA (1,000 μM). Scale bar, 500 nm. For mitochondrial number, n = 7–11 (n = 11 for Con, n = 10 for IPA and CD3/CDC28+ IPA, n = 7 for CD3/CD28 group); for the other parameters, 20–40 mitochondria were analysed (n = 35 for Con, n = 23 for IPA, n = 30 for CD3/CD28, n = 22 for CD3/CD28+ IPA). Data represent mean ± s.e.m. and were analysed by one-way ANOVA with Tukey’s correction for multiple comparisons (e,f,h,i,k,l), mean ± s.e.m. analysed by two-way ANOVA with Tukey’s correction for multiple comparisons (a) and mean ± s.e.m. analysed by two-tailed Student’s t-test (b).
At steady state, IPA increased OCR without significant changes in the ATP rate index. However, in the CD3/CD28 activation context, IPA increased OCR as well as ATP rate index, showing an activation of mitochondrial respiration (Fig. 2b). The energetic map further illustrates the overall effect of IPA on the energy metabolism of CD4+ T cells (Fig. 2c). In Mitostress assay, different doses of IPA from 1 μM to 1,000 μM increased mitochondrial basal and maximal respiration (Fig. 2d,e). Mitochondrial mass, as assessed by MitoTracker Green, was moderately increased by 1,000 μM IPA at steady state but not in the CD3/CD28 activation context (Extended Data Fig. 4a). MitoTracker Deep Red (MTDR) was used to evaluate the mitochondrial membrane potential (ΔΨm) that results from the generation of a proton motive force. CD3/CD28 activation led to increased MTDR MFI and this effect was significantly increased by IPA, confirming that IPA boosts ΔΨm (Extended Data Fig. 4b). This result was also confirmed after normalizing ΔΨm by mitochondrial mass (Mito Deep Red:Mito Green ratio; Extended Data Fig. 4c). We then measured cytoplasmic reactive oxygen species (ROS) and mitochondrial ROS (mitoROS), which are OxPhos byproducts. Both cytoplasmic ROS and mitoROS were decreased by IPA in CD3/CD28 activation context at the 48-h and 72-h time points (Supplementary Fig. 2a,b). The analysis of mitoROS to cytoplasmic ROS ratio showed that ROS were predominantly generated in the mitochondria at baseline, whereas ROS mainly originated from the cytoplasm in the context of CD3/CD28 activation. IPA reduced ROS production in both cytoplasm and mitochondria with a more pronounced effect in cytoplasm (Supplementary Fig. 2a–c). Taken together, these results showed that the beneficial impact of IPA on mitochondrial respiration does not come at the potentially damaging expense increasing mitochondrial or cytoplasm ROS levels.
Seahorse substrate oxidation assay was then used to determine which metabolic substrate route into mitochondria IPA was able to manipulate. UK5099 (an inhibitor of the mitochondrial pyruvate carrier MPC), Etomoxir (an inhibitor of CPT1b and FAO) and BPTES (a glutaminase inhibitor), were used to specifically block mitochondrial pyruvate utilization, FAO and glutaminolysis, respectively (Extended Data Fig. 4d). The effect of IPA in boosting maximal respiration was lost in the presence of Etomoxir and BPTES but not in the presence of UK5099 (Fig. 2f and Extended Data Fig. 4e). These results were consistent with those obtained with SCENITH in mixed cell systems (Extended Data Fig. 3d,f) and confirmed that IPA-dependent effects on mitochondrial respiration are mediated by FAO and AAO. In addition, IPA led to increased uptake of lipids, as demonstrated by experiments with BODIPY 493/503 and BODIPY FL C12, which are fluorescent dyes that stain neutral lipids and synthetic precursors for a wide variety of fluorescent phospholipids, respectively (Supplementary Fig. 2d,e). Of note, the expression of CD36, a scavenger receptor facilitating lipid transport within the cells, also increased in activated cells treated with IPA (Supplementary Fig. 2f). To further demonstrate the critical role of lipids in IPA’s effect on boosting mitochondrial respiration, we performed a new Seahorse Mitostress assay under conditions of exogenous lipid deprivation and replenishment with the long-chain FA palmitic acid (PA). As shown in Extended Data Fig. 4f,g, PA replenishment was able to restore the IPA effect partially. Furthermore, the effect of IPA was also lost in the presence of CD36 neutralizing antibody, demonstrating that the effect of IPA in changing cellular energy metabolism is dependent on the CD36 lipid transporter (Extended Data Fig. 4h,i).
Different doses of IPA exhibited a significant effect in reducing basal and compensatory glycolysis of CD4+ T cells (Fig. g,h). Consistent with what we observed in the Seahorse glycolytic rate assay, lactate production in the cell culture supernatant was inhibited, mainly by high doses of IPA (Fig. 2i). Thus, these results demonstrate IPA increases CD4+ T cell mitochondrial respiration by promoting FAO and AAO. In parallel, IPA also inhibits aerobic glycolysis.
IPA increases mitochondrial respiration in a Cpt1a-dependent, but OPA1-independent manner
The ultrastructure of the mitochondrial inner membrane (MIM) is a critical regulator of mitochondrial respiration19. Cristae, which are MIM invaginations, harbour the majority of OxPhos machinery required to generate ATP. In T cells, modulation of mitochondrial structure via the ablation of the MIM shaping protein OPA1 changes T cell energy metabolism and immune functions20. Given the positive effects of IPA on energy metabolism of human and mouse T cells, we sought to assess mitochondrial structure using transmission electron microscopy (TEM). Although IPA did not change the number and length of mitochondria, it significantly increased the width and total area of mitochondria in activated CD4+ T cells. Notably, we observed effects on the organization of cristae, which are the MIM invaginations; IPA treatment led to an increase in cristae density, cristae score, which evaluates cristae abundance and form21, and total cristae area (Fig. 2j–l). These results show that IPA affects the distribution and density of mitochondrial cristae without affecting the mitochondria number and length, which explains how IPA stimulates the production of energy through the electron transport chain (ETC)22.
Not limited to human or mouse T cells, we found that IPA had a broad effect on boosting mitochondrial respiration in HeLa cells and mouse embryonic fibroblasts (MEFs) (Extended Data Fig. 5a,b). To determine whether OPA1 was required to potentiate the effects of IPA on mitochondrial metabolism, we used MEF lacking OPA1 (Optic atrophy 1, Opa1KO), in which we previously showed manifest defects in cristae maintenance, OxPhos activity, mitochondrial membrane potential and respiration23. Notably, IPA was able to suppress defects in mitochondrial respiration in Opa1KO MEFs and enhance respiration in wild-type (WT) MEFs (Extended Data Fig. 5b). Moreover, defects in mitochondrial membrane potential and mitochondrial mass observed in Opa1KO MEF cells were rescued by IPA (Extended Data Fig. 5c). Taken together, these results demonstrate that IPA enhances mitochondrial respiration independently of OPA1.
To investigate whether IPA has a direct effect on mitochondria, we further tested IPA on isolated cardiac mitochondria and assessed mitochondrial respiration by high-resolution respirometry24 in the presence of various substrates of the TCA and inhibitors of the OxPhos system. Preincubation of IPA did not increase oxygen consumption rates under any of the substrate conditions that are known to promote phosphorylation-dependent respiration (Extended Data Fig. 5d). Respirometry studies also excluded that enhanced mitochondrial respiration by IPA is results from the uncoupling of the proton motive force across the MIM as CCCP-induced uncoupling after IPA addition was just as effective in maximizing mitochondrial respiration (Extended Data Fig. 5e). These results indicate that IPA is not a mitochondrial uncoupler and requires intact cellular structure and the participation of other signals to exert its effect of increasing mitochondrial respiration.
As recent data suggest potential off-target effects of Etomoxir25, although at much higher dose than the one we used, we applied siRNA in MEFs as an alternative approach to evaluate the involvement of FAO in the effects of IPA (Extended Data Fig. 5f). siRNA targeting the key enzyme Cpt1a significantly reduced this gene expression to about 20% of the level in the Con and mock-treated groups (Extended Data Fig. 5g). Seahorse experiments showed that the effects of IPA on increasing mitochondrial respiration were lost when Cpt1a was knocked down (Extended Data Fig. 5h,i). These results support the fact that the effect of IPA in promoting mitochondrial respiration is dependent on the FAO pathway.
IPA increases mitochondrial OxPhos via PPARβ/δ and independently of AhR signalling pathway
IPA is known to be an AhR agonist6. However, as shown using the AhR reporter system and in accordance with previous studies26, IPA has a weak ability to activate AhR compared with other microbiota-derived indoles such as indole acetic acid (IAA), oxindole and indoxyl-3-sulphate (I3S) (Extended Data Fig. 6a,b). 6-Formylindolo[3,2-b]carbazole (Ficz), which is a highly potent ligand of AhR did not change the energy metabolism of WT and AhR−/− CD4+ T cells (Extended Data Fig. 6c,d). Moreover, IPA was able to increase mitochondrial basal and maximal respiration of AhR−/− CD4+ T cells (Extended Data Fig. 6e,f). These results show that AhR is not involved in the effects of IPA on CD4+ T cell energy metabolism.
To get deeper insights into the mechanisms underpinning IPA-dependent metabolic reprogramming, we performed proteomic analyses on activated CD4+ T cells with or without IPA treatment. Gene set enrichment analysis (GSEA) showed that glycolysis (P = 0.04) and the MYC pathway (P = 0.0696) were upregulated in the CD3/CD28 group (Extended Data Fig. 6g). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed that the PPAR signalling pathway was significantly upregulated by IPA (Fig. 3a). Differential abundance analysis showed several enriched proteins in the IPA treatment setting, including long-chain-fatty acid-CoA ligase (ACSBG1), ubiquitin-40S ribosomal protein S27a (RPS27a) and serine/threonine-protein kinase pim-1 (PIM1), all of which are involved in the PPAR signalling pathway (Fig. 3b,c).
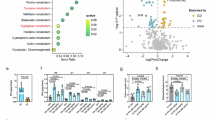
a, Proteomics KEGG enrichment analysis. b,c, Volcano plot (b) and circle heatmap (c) of proteomics showing differentially expressed proteins between CD3 (CD3/CD28) and CIPA2 (CD3/CD28 + IPA1000) group. Red boxes indicate highly expressed proteins. d, PPAR transactivation assay indicating that IPA of 1,000 μM activates PPARβ/δ nuclear receptor. n = 4. e–g, Seahorse Mitostress assay analysis of mouse CD4+ T cells cultured with and without CD3/CD28 in the presence of DMSO or IPA (100 and 1,000 μM) at 6 h in the basal condition (e), in the presence of GSK3787 (f) and GW6471 (g). n = 6 for each except n = 4 for IPA100 and IPA1000 (e,g); n = 6 for Con and CD3/CD28, n = 4 for IPA100, IPA1000 and n = 8 for CD3/CD28+ IPA100, CD3/CD28+ IPA1000 (f). h, Basal and maximal respiration measurement of Mitostress assay above in basal condition and in the presence of GSK3787, GW6471. n = 4–8. The exact n value is consistent with the above description of e–g. A two-tailed Welch t-test was used for the analysis of the proteomics data and a P value of 0.05 was applied to determine differentially expressed proteins. Data represent mean ± s.e.m. analysed by one-way ANOVA with Tukey’s correction for multiple comparisons (d) and mean ± s.e.m. analysed by two-way ANOVA with Fisher LSD test (h).
PPAR signalling has been implicated in enhanced mitochondrial and metabolic activity in various in vitro and in vivo paradigms27. To determine whether IPA activates PPAR signalling, we used a HEK293 reporter cell line and monitored PPAR transactivation. This effect disappears when dominant-negative PPARα or PPARβ/δ were transiently expressed instead of PPARγ, thus excluding the role of the latter (Fig. 3d). GW501516, a selective PPARδ agonist, increased basal and maximal respiration of activated CD4+ T cells (Extended Data Fig. 6h,i), an effect that was blocked by the specific PPARβ/δ inhibitor GSK3787. Similar results were observed with PPARα agonist GW7647 and the selective PPARα inhibitor GW6471 (Extended Data Fig. 6j,k). These results suggest that activation of PPARα or PPARβ/δ itself can promote mitochondrial respiration in CD4+ T cells. The effect of IPA on increasing CD4+ T cell basal and maximal respiration was abolished in the presence of GSK3787 but not of GW6471, meaning that IPA acts on PPARβ/δ to exert its influence on energy metabolism (Fig. 3e–h). Taken together, these results demonstrate that the effects of IPA on CD4+ T cell energy metabolism are dependent on PPARβ/δ but not on PPARα, PPARγ or AhR signalling pathways.
IPA inhibits TH1 and TH17 cell differentiation without altering Treg cells of mouse CD4+ T cells
We next explored the effects of IPA on the immunophenotypic fate of mouse CD4+ T cells. Total CD4+ or naive CD4+ T cells isolated from WT C57BL/6J mouse splenocytes were submitted to different cytokine environments to induce differentiation toward TH1, TH17 and Treg cell phenotypes in the presence or absence of IPA (Extended Data Fig. 7a). For total CD4+ T cells, as shown by flow cytometry analyses, IPA did not affect cell viability under TH1 cell differentiation conditions but increased the percentage of live cells to a certain extent (Fig. 4a). Additionally, IPA dramatically decreased IFNγ+ TH1 percentage and IFNγ MFI in both total CD4+ T cells and IFNγ+ TH1 populations, with a dose-dependent effect (Fig. 4b–d). IPA also decreased the percentage of IFNγ+ T-bet+ TH1 cells and T-bet MFI (Extended Data Fig. 7b,c). In the TH17 cell differentiation settings, IPA decreased IL-17A+ TH17 cell percentage and IL-17A MFI without any effect on cell viability (Fig. 4e–h). IPA also decreased the percentage of IL-17A+ RORγt+ TH17 cells and RORγt MFI (Extended Data Fig. 7d,e). Consistent with results in total CD4+ T cells, IPA reduced T-bet MFI in TH1 cells and RORγt MFI in TH17 cells, respectively (Extended Data Fig. 7f). However, in Treg cell differentiation conditions, IPA did not show clear effect on CD4+FOXP3+ percentage or on FOXP3 MFI (Fig. 4i–l). Taken together, these results demonstrate that IPA directs CD4+ T cells toward an anti-inflammatory phenotype by decreasing TH1 and TH17 cell differentiation without affecting the Treg cell phenotype.
a, Flow cytometry analysis of cell viability under TH1 cell differentiation conditions. n = 3. b, Representative flow cytometry plots and quantitative comparison of IFNγ+TH1 percentage of CD4+ T cells stained intracellularly for IFNγ and IL-4 in Con, IPA100 and IPA1000 groups. n = 3. c, Representative IFNγ expression histograms and quantitative comparison of IFNγ MFI of CD4+ T cells in Con, IPA100 and IPA1000 groups. n = 3. d, Quantitative comparison of IFNγ MFI of IFNγ+TH1 cells in Con, IPA100 and IPA1000 groups. n = 3. e, Flow cytometry analysis of cell viability under TH17 differentiation conditions. n = 3. f, Representative flow cytometry plots and quantitative comparison of IL-17A+TH17 percentage of CD4+ T cells stained intracellularly for IL-17A and IFNγ in Con, IPA100 and IPA1000 groups. n = 3. g, Representative IL-17A expression histograms and quantitative comparison of IL-17A MFI of CD4+ T cells in Con, IPA100 and IPA1000 groups. n = 3. h, Quantitative comparison of IL-17A MFI of IL-17A+TH17 cells in Con, IPA100 and IPA1000 groups. n = 3. Flow cytometry and its quantification of CD4+ T cells stained intracellularly for IL-17A and IFNγ. n = 3. i, Flow cytometry analysis of cell viability under Treg differentiation conditions. n = 3. j–l, Flow cytometry (j) and its quantification (k,l) of CD4+ T cells stained intracellularly for FOXP3. n = 3. m,n, Flow cytometry (m) and its quantification (n) of naive CD4+ T cells stained intracellularly for IFNγ and IL-4. n = 4. o,p, Flow cytometry (n) and its quantification (p) of naive CD4+ T cells stained intracellularly for IL-17A and IFNγ. n = 3. q,r, Flow cytometry and its quantification of naive CD4+ T cells proliferation stained with CFSE. n = 5. Data represent mean ± s.e.m. analysed by one-way ANOVA with Tukey’s correction for multiple comparisons (a–i,k,l,n,p) and mean ± s.e.m. analysed by two-way ANOVA with Tukey’s correction for multiple comparisons (r).
We then evaluated the effect of IPA on naive CD4+ T cell differentiation. All doses of IPA (from 1 μM to 1,000 μM) exhibited significant inhibitory effects on TH17 cell differentiation, while only highest doses (100 and 1,000 μM) showed significant effects on TH1 differentiation (Fig. 4m–p). IPA at 1,000 μM was able to significantly inhibit naive CD4+ T cell proliferation in basal (no cytokine environment), TH1 and TH17 cell differentiation conditions (Fig. 4q,r). In conclusion, these data show that IPA can inhibit TH1 and TH17 cell differentiation, with a much stronger effect for TH17 cells, and only the highest dose can inhibit naive CD4+ T cell proliferation.
To determine whether IPA can exhibit its ‘reprogramming’ effects after a short exposure or if permanent exposure is required, we performed a ‘washout’ experiment (Extended Data Fig. 7g). As shown previously, under usual settings, IPA exhibits an inhibitory effect on the differentiation of TH1 and TH17 cells. Of note, the effects of IPA were still visible after 24 and 48 h of washout (Extended Data Fig. 7h,i). These results suggest that the effect of IPA on regulating TH1 and TH17 cell reprogramming is a long-lasting phenomenon that does not require continuous exposure to IPA. In addition, IPA also has the ability to hinder the differentiation of naive CD4+ T cells into TH1 and TH17 cells during the final restimulation, which provided us with some hints that for differentiated TH1 and TH17 cells, IPA was still able to inhibit their proportion when subjected to secondary antigenic strikes (Extended Data Fig. 7j,k).
IPA promotes CD4+ T cell differentiation toward anti-inflammatory direction through metabolic reprogramming and PPARβ/δ signalling pathway
To investigate whether the effect of IPA on altering CD4+ T cell phenotype is dependent on energy metabolism rewiring, CD4+ T cells differentiation was performed with or without distinct metabolic pathway inhibitors (Extended Data Fig. 8a,b). In the basal condition, IPA exerts a sustained and stable inhibitory effect on TH1 and TH17 cells (Fig. 5a,f,g,l). As shown in Extended Data Fig. 8c,d, oligomycin, when used at 3 nM concentration and higher, induced a significant drop in the proportion of TH1 cells without negatively impacting cell viability. Furthermore, the effect of IPA on decreasing TH1 cell percentage was abolished in the presence of oligomycin at the same doses (Fig. 5b and Extended Data Fig. 8d). Therefore, the effect of IPA on reducing the TH1 cell phenotype is mediated by the induction of mitochondrial ATP synthesis. Etomoxir and TOFA (a competitive inhibitor of acetyl-CoA carboxylase blocking endogenous lipid synthesis), but not BPTES, decreased the effect of IPA on reducing TH1 cell percentage and IFNγ MFI (Fig. 5c–f and Extended Data Fig. 8g), indicating that IPA requires FAO and endogenous FA synthesis to modulate T cell fate. Oligomycin at a concentration of 0.1 nM and above can affect the differentiation percentage of TH17 cells (Extended Data Fig. 8e). The effects of IPA on TH17 cell differentiation were also inhibited by oligomycin (Fig. 5h and Extended Data Fig. 8f). Moreover, BPTES, but not etomoxir and TOFA, blocked the inhibitory effect of IPA on TH17 cell differentiation (Fig. 5i–l and Extended Data Fig. 8h). Taken together, these results show that the effects of IPA on TH1 and TH17 cell differentiation are mediated by the modulation of mitochondria energy metabolism with a dominant role of lipid oxidation for TH1 cell and of glutaminolysis for TH17 cells.
a–f, Flow cytometry and its quantification of CD4+ T cells stained intracellularly for IFNγ and IL-4. CD4+ T cells were cultured in TH1 cell differentiation condition treated with DMSO or IPA (100 and 1,000 μM) in baseline and in the presence of specific metabolic pathway inhibitors including oligomycin (Oligo), etomoxir (Eto), TOFA and BPTES. n = 3. g–l, Flow cytometry and its quantification of CD4+ T cells stained intracellularly for IL-17A and IFNγ. CD4+ T cells were cultured in TH17 cell differentiation condition treated with DMSO or IPA (100 and 1,000 μM) in baseline and in the presence of specific metabolic pathway inhibitors as mentioned above. n = 3. Data represent mean ± s.e.m. analysed by two-way ANOVA with Tukey’s correction for multiple comparisons (f,l).
We then evaluated whether the effect of IPA on CD4+ T cell differentiation was dependent on PPARβ/δ, as it is for the metabolic reprogramming. As shown in Extended Data Fig. 8i,j, PPARδ agonist GW501516 decreased TH1 cell proportion and IFNγ MFI in a dose-dependent manner. And IPA lost its ability to reduce the proportion of TH1 cells in the presence of the specific PPARβ/δ inhibitor GSK3787 (Extended Data Fig. 8k). A similar effect was observed for TH17 cells (Extended Data Fig. 8l–n). Taken together, these results demonstrate that IPA activates the PPARβ/δ signalling pathway, triggering the reprogramming of CD4+ T cell energy metabolism and ultimately causing CD4+ T cells to differentiate into an anti-inflammatory direction.
IPA shifts energy metabolism of CD4+ T cell towards mitochondrial respiration in vivo
We next examined whether IPA influences the metabolic profile of CD4+ T cells in vivo using SCENITH. IPA and many other metabolites are continuously produced by the gut microbiota, obfuscating the effects that these gut bacteria have on host cell energy metabolism and function. Therefore, we orally administered a cocktail of nonabsorbable antimicrobial agents (ABX) to adult conventional mice for 7 days to deplete their gut microbiota, together with IPA or a vehicle solution. This allowed evaluating the effect of IPA in mice with a mature immune system but without the interference of endogenously produced metabolites (Fig. 6a). Moreover, the use of nonabsorbable ABX allowed us to avoid the deleterious effects on mitochondrial metabolism that cell-penetrant antibiotics designed to inhibit bacterial ribosomes could have on mitochondrial translation in host cells28,29. As expected, ABX treatment massively depleted the gut microbial content without any effect of IPA (Fig. 6b). The total number of cells isolated from the small intestine and colon lamina propria (LP) did not change in ABX- and IPA-treated mice (Extended Data Fig. 9a). However, in the ABX-treated condition, IPA increased the proportion of colonic CD4+ T cells, while decreasing the proportion of CD8+ T cells. This phenomenon was not observed in the small intestine (Fig. 6c,d and Extended Data Fig. 9b).
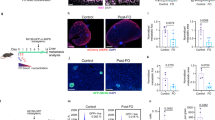
a, Schematic diagram showing the protocol of ABX treatment and SCENITH of small intestine and colon LP. IPA (200 mg kg−1) or corresponding vehicle control were administered to mice daily by oral gavage. Created with BioRender.com. b, Total faecal bacteria amount of WT, ABX and ABX + IPA group. n = 5. c,d, The proportion of each cell population including CD4+, CD8+, DN and CD4+CD8+ double positive cell subsets under the TCRαβ+ cell gate in colonic LP of WT, ABX and ABX + IPA mice. n = 7 for WT and ABX + IPA, n = 9 for ABX. (e) ATP synthesis in WT, ABX and ABX + IPA group of CD4+ and CD8+ T cell subsets in colon LP. n = 16 for WT, n = 15 for ABX, n = 12 for ABX + IPA. f, Metabolic profile of CD4+ T cells of colonic LP in WT, ABX and ABX + IPA group. n = 13 for WT, n = 12 for ABX and ABX + IPA groups. g, Metabolic profile of CD8+ T cells of colonic LP in WT, ABX and ABX + IPA group. n = 13 for WT, n = 12 for ABX and ABX + IPA groups. Data represent mean ± s.e.m. analysed by one-way ANOVA with Tukey’s correction for multiple comparison (b,d,e) and mean ± s.e.m. analysed by two-way ANOVA with Tukey’s correction for multiple comparisons (f,g).
Moreover, compared with untreated mice, colon LP CD4+, CD8+ and γδ T cells from ABX-treated mice exhibited a decrease in total ATP synthesis. IPA administration rescued the decrease ATP synthesis induced by ABX treatment, mainly for CD4+ T cells. No significant changes were observed in cells isolated from the small intestine, suggesting a lower impact of the gut microbiota on immune cells’ energy metabolism in this compartment (Fig. 6e and Extended Data Fig. 9c,d).
In colon CD4+ T cells and, to a lesser extent in CD8+ T cells, ABX treatment shifted the energy metabolic profile by decreasing mitochondrial dependence and increasing glycolytic capacity and glucose dependence. IPA administration reversed the ABX effects on the energy metabolism of colonic CD4+ T cells, mostly by increasing FAO and AAO. More moderate trends were also observed for colonic CD8+ T cells (Fig. 6f,g). ABX also induced some changes in colonic B and γδ T cells with only minor effects of IPA (Extended Data Fig. 9e,f). Taken together, these results demonstrate that gut microbiota depletion alters the energy metabolism of several immune cells in colon LP and that IPA administration counteracts these alterations by boosting FAO, AAO and, eventually, mitochondrial activity.
IPA rewires the energy metabolism and phenotype of colonic CD4+ T cells during inflammation and protects from colitis
We then sought to interrogate the clinical relevance of our results in the context of IBD. CD and UC are two types of IBD and are characterized by chronic intestinal inflammation. We took advantage of a longitudinal cohort of patients with CD or UC who were sampled before initiating biologic therapy and then 2 and 14 weeks later (Fig. 7a and Extended Data Fig. 10a). Notably, serum IPA concentrations were higher in healthy controls (HCs) compared with patients with CD or UC (Fig. 7b). Linear models were conducted to evaluate the dynamic changes between responders and nonresponders (NRs) over time. Patients inactive at week 14 were considered responders. A modest divergence in serum IPA levels between responders and NRs to biologic therapy appeared progressively as expected, which reached statistical significance at week 26 post-therapy induction (P = 0.043 and Fig. 7c). This result shows that the concentration of IPA differs between the responders and NRs during the treatment process over time. Significant negative correlations were observed between serum IPA and C-reactive protein (CRP) (a systemic marker of inflammation) levels, clinical activity scores in UC (partial Mayo score) and CD (CDAI) (Fig. 7d). All the data above suggest the importance of IPA for the progress of IBD treatment.
a, Schematic diagram showing healthy HCs and IBD patient cohort collection. Created with BioRender.com. b, IPA concentration in serum of healthy donors (Control) and patients with CD or UC. c, Dynamic changes of IPA concentration in responders and NRs over 26 weeks. n = 54 for CD (NR/R = 39/15), n = 53 for UC (NR/R = 37/16). Linear models were applied for analysis without multiple corrections. d, Correlation analysis between serum IPA and disease activity indicators (CRP, partial Mayo and CDAI). n = 179 for HC, n = 77 for CD and n = 80 for UC (b,d). Linear mixed coefficient models were used for analysis with multiple hypothesis correction using Benjamini–Hochberg adjustment. e,f, Body weight curve and DAI scoring during the whole process of the DSS model in WT mice. n = 10 for Con and DSS+ IPA group; n = 8 for DSS group. The P values in the graph show the comparison between groups DSS + IPA and DSS. g–i, Haematoxylin and eosin (H&E) staining (g), colon length (h), histology score of colon tissue at D12 in Con, DSS and DSS + IPA groups (i). n = 9 for colon length comparison. For H&E histology, four samples were randomly selected. Scale basr, 200 μm. j, Metabolic profile of colonic CD4+ T cells in Con, DSS and DSS + IPA groups at D12. n = 5–9 per group. n = 7 for Con, n = 9 for DSS, n = 6 for DSS+ IPA. k,l, Flow cytometry (k) and its quantification (l) of colonic CD4+ T cells stained intracellularly for IL-17A and IFNγ. n = 5 for Con, n = 6 for DSS and DSS+ IPA group. m, MFI analysis of IL-17A and IFNγ in colonic CD4+ T cells. n = 5 for Con, n = 6 for DSS and DSS + IPA group. Data represent mean ± s.e.m. analysed by two-way ANOVA with Tukey’s correction for multiple comparisons (e,f,j) and mean ± s.e.m. analysed by one-way ANOVA with Tukey’s correction for multiple comparisons (h,i,l,m).
In accordance with previous studies30,31, IPA oral gavage improved the inflammation severity in dextran sulphate sodium (DSS)-induced colitis in mice, as demonstrated by a lower weight loss, disease activity index (DAI), colon length shortening and histological score (Fig. 7e–i and Extended Data Fig. 10b). Intestinal inflammation was associated with a shift in colon LP CD4+ T cells energy metabolism with decreased mitochondrial dependence and increased glycolytic capacity, meaning that aerobic glycolysis was strongly induced. Similarly to our in vitro findings, IPA reversed these changes in mitochondrial metabolism via increasing FAO and/or AAO (Fig. 7j). IPA also compensated for some alterations observed in γδT cells (Extended Data Fig. 10c). In colitis settings, IPA also impacted CD4+ T cells phenotype by decreasing IL-17A+ TH17 and IFNγ+ TH1 cell proportions as well as IL-17A and IFNγ MFI (Fig. 7k–m). IPA did not significantly impact Treg cell proportions or FoxP3 MFI values in vivo (Extended Data Fig. 10d,e). Taken together, these results show that the protective role of IPA in DSS-induced colitis is associated with the rewiring of CD4+ T cell energy metabolism toward mitochondrial respiration and an anti-inflammatory phenotype.
Anti-inflammatory effect of IPA in colitis model in vivo are mediated by CD4+ T cells
To further evaluate whether the protective effect of IPA in colitis is dependent on the adaptive immune system, we used Rag2−/− mice that lack a functional adaptive immune system and submitted them to DSS colitis with or without IPA treatment. In this setting, IPA was no longer able to exhibit its protective effects (Extended Data Fig. 10f–j), indicating that a functional adaptive immune system is essential for the beneficial effects of IPA. To test whether CD4+ T cells were required in the protective effects of IPA, we adoptively transferred CD4+ T cells from WT mice submitted to colitis with or without IPA treatment to Rag2−/− mice submitted to DSS-induced colitis (Fig. 8a). The mice transferred with IPA-treated CD4+ T cells (IPA CD4+ T→Rag2) exhibited decreased colitis severity compared with their counterparts that received nontreated CD4+ T cells (no-IPA CD4+ T→Rag2) (Fig. 8b–f). Similarly, CD4+ T cells obtained from microbiota-depleted WT mice treated with IPA in a non-inflammatory context (Fig. 8g) also exhibited protective effects when adoptively transferred to Rag2−/− mice (Fig. 8h–l). Taken together, these findings demonstrate that the protective effect of IPA on colitis is mediated by CD4+ T cells.
a, Schematic diagram showing the protocol of adoptive transfer model. Created with BioRender.com. b,c, Body weight curve (c) and DAI scoring (d) of Rag2−/− recipient mice that received either no-IPA-treated CD4+ T cells or IPA-treated CD4+ T cells during the whole adoptive transferred process. n = 8 for no-IPA and n = 6 for IPA group. d–f, H&E staining (d), colon length (e) and histology score (f) of colon tissue at D13 in no-IPA CD4+ T→Rag2 and IPA CD4+ T→Rag2 mice. n = 8 for no-IPA and n = 6 for IPA group for colon length comparison. For H&E histology, n = 3 for no-IPA and n = 4 for IPA group were randomly selected. Scale bars, 200 μm. g, Schematic diagram showing the protocol of adoptive transfer model. Created with BioRender.com. p.o., oral administration. h,i, Body weight curve (h) and DAI scoring (i) of Rag2−/− recipient mice that received either ABX CD4+ T cells or ABX + IPA CD4+ T cells during the whole adoptive transferred process. n = 9 for ABX and n = 7 for ABX + IPA group. j–l, H&E staining (j), colon length (k) and histology score (l) of colon tissue at D12 in ABX CD4+ T→Rag2 and ABX + IPA CD4+ T→Rag2 mice. n = 9 for ABX and n = 7 for ABX + IPA group for colon length comparison. For H&E histology, five samples were randomly selected. Scale bars, 200 μm. All data (except a,d,g,j) represent mean ± s.e.m. analysed by two-tailed unpaired Student’s t-test.
Discussion
In the past decades, tremendous progress has been made in the field of immunometabolism, revealing the crucial role of intracellular metabolism in regulating immune cell function32. In this context, it is essential to uncover the factors that modulate cellular energy metabolism. The gut microbiota has been shown to massively impact immune system maturation and functions. Given the intimate relatedness between mitochondria and bacteria, it has been logically suspected to be involved33,34. Indeed, published studies have already demonstrated some effects of the gut microbiota on intestinal epithelial and immune cell energy metabolism, but the available data are globally scarce7. Moreover, an altered gut microbiota composition and diversity have been involved in many human diseases characterized by an inappropriate immune response35,36. It therefore suggests that some alterations in the gut microbiota composition and functions may contribute to altered immune cell energy metabolism and thus to pathogenesis. This is particularly the case for IBD.
A growing body of literature describes the concept that metabolism is a rheostat for intestinal inflammation, which seems to be closely related to ‘metabolic inflammation’ established in obesity, metabolic dysfunction-associated fatty liver disease and related diseases37,38,39. Schulz-Kuhnt et al. showed that modulation of the expression of ATP citrate lyase (ACLY), which controls the immunometabolism in mucosal T cell and colitis model, could serve as a new strategy to improve the resolution of gut inflammation40. However, the relationships between immunometabolism, microbiome and inflammation remain largely unknown.
Multiple studies have shown mitochondrial plasticity, which modulate cellular metabolic changes is integral to T cells. Long-lived memory T (TM) cells after infection or cancer engage FAO to fuel the TCA cycle41. Naive CD4+ T cell activation triggers a unique mitochondrial proteome remodelling where enhanced one-carbon metabolism takes place, which is pivotal for its sustained activation and survival42. However, few studies have directly identified intervention methods to enhance mitochondrial respiration in T cells. In our study, we screened a collection of microbiome-derived metabolites for their ability to boost energy metabolism in T cells and discovered that IPA yielded a strong and rapid ability to increase mitochondrial respiration and OxPhos via boosting the utilization of FAO and AAO metabolism of CD4+ T cells, directing these cells to adopt an anti-inflammatory phenotype both in vitro and in vivo. Additionally, IPA also promoted mitochondrial respiration in already established TH1 cells as well as TH17 memory CD4+ T cells. The effect of IPA on boosting CD4+ T cell mitochondrial respiration was attenuated by either serum deprivation or CD36 inhibition. PA replenishment partially compensated for the consequences of serum deprivation, demonstrating the involvement of exogenous FA transport in the effects of IPA. Moreover, IPA remodelled mitochondrial cristae area and density in a PPARβ/δ-dependent and OPA1-independent manner. Buck et al. have shown that OPA1 is required for TM cell generation, which have fused mitochondria and increased OxPhos over glycolysis20. However, the similar modulations of mitochondrial shape and function by IPA do not seem to require OPA1, at least in MEFs, which may suggest that there are multiple, nonredundant ways where mitochondrial and metabolic reprogramming can take place. Of note, IPA is a fast-acting metabolite, leading to increased respiration and membrane potential in a few hours of treatment, which is more rapid than other agonists and chemical agents that enhance mitochondrial structure and function43. Previous studies have shown the effects of IPA on parenchymal cells, and lung epithelial cells, reducing their mitochondrial membrane potential and mitoROS production44. However, in mouse MEFs and human HeLa cells we tested, we did not observe negative effects on mitochondrial membrane potential and respiration. In cardiomyocytes, IPA enhanced mitochondrial respiration at early time points, but long-term treatment had the opposite effect45. These studies showed that IPA affects immune and parenchymal cell metabolism differently.
Various indoles, which are derived from the metabolism of tryptophan by the gut microbiota, including IAA, indole acrylic acid, indole lactic acid (ILA) and IPA, are AhR agonists and can cooperate with other gut microbiota-derived metabolites, such as SCFAs and BAs to relieve epithelial stress, regulate intestinal immunity and have a beneficial effect on intestinal inflammation, colorectal cancer and many other diseases30,31,46,47,48. IPA is a relatively weak AhR agonist compared with other indole derivatives, and indeed, its effect on CD4+ T cell energy metabolism does not depend on the AhR signalling pathway. Instead, the effects are mediated by PPAR. PPARs, including PPARα, PPARβ/δ and PPARγ are nuclear transcription factors that can be activated by FAs. Among them, PPARβ/δ plays a key role in regulating glucose and lipid metabolism and maintaining mitochondria function27. Wang et al. have shown that the CD36–PPARβ axis can precisely regulate the fitness and biogenesis of mitochondria, thus reprogramming Treg cells to adapt to the tumour microenvironment49. Our proteomics analysis pointed toward the PPAR signalling pathway and functional experiments confirmed the involvement of PPARβ/δ in both metabolic and immune effects of IPA. Indeed, previous structural analysis showed that IPA can be used as a building scaffold for PPAR agonist synthesis50.
We showed that the IPA level is decreased in patients with IBD and negatively correlates with disease activity. As previously shown, we observed that IPA exhibits protective effects in mouse model of colitis30,31. We unravelled that this protection is mediated by effects on CD4+ T cell energy metabolism. We showed that the effects of IPA on energy metabolism underlie profound changes in CD4+ T cell immunophenotype toward an anti-inflammatory direction. Among the different helper T cell subsets, different subsets have distinct energy metabolism preferences. Treg cells dominantly depend on mitochondria respiration, FAO and glutaminolysis, whereas TH1, TH2 and TH17 cells mainly rely on aerobic glycolysis and glutaminolysis for their growth, proliferation and functions18. This is in line with the effects of IPA on cellular energy metabolism, promoting mitochondrial respiration and inhibiting aerobic glycolysis, leading eventually to TH1 and TH17 cell inhibition. The effect of IPA on TH1 cell phenotype was mostly driven by FA metabolism, whereas it was dependent on glutaminolysis for TH17 cells. For Treg cells, as mitochondrial respiration is already largely activated under baseline conditions, it is likely that IPA cannot induce an additional increase in the polarized phenotype of Treg cells. Our study included CD4+ T cells from healthy humans and mouse spleen and intestinal LP. However, we did not study CD4+ T cells from human patients with IBD, which is one of the limitations of our work.
In conclusion, our study reveals that the anti-inflammatory effects of IPA are mediated by metabolic reprogramming of CD4+ T cells via favouring mitochondrial metabolism and ultimately resulting in protection in intestinal inflammation contexts. Besides providing promising therapeutic avenues in IBD, these results highlight the intricacy between gut microbiota functions and immune cell metabolism and phenotype, with potential consequences and applications in multiple human diseases.
Methods
Our research complies with all relevant ethical regulations. The research protocol followed the World Medical Association Declaration of Helsinki. Approval for human studies was obtained from the local ethics committee (for human blood, Comité de Protection des Personnes Ile-de-France IV, IRB 00003835 Suivitheque study; registration number 2012/05NICB; for the IBD cohort, ethics committee of the medical faculty of Kiel University; number AZ D489/14, A 124/14 and AZ 156/03-2/13). Animal experiments were carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the French Ministry of Agriculture and the European Union. All animal experiments were subjected to and approved by the Institutional Ethics Committee and the French Ministry of Research (APAFIS no. 37837-202206300029683 v.3 and no. 47879-2024030216059527 v.3).
Cell lines and in vitro primary cell culture
Human Jurkat cells (clone E6-1, TIB-152, ATCC) were cultured in RPMI 1640 GlutaMAX supplement medium with 10% fetal bovine serum (FBS) (Gibco, 10437028), 1% penicillin–streptomycin solution (Gibco, 15140122) and 1× MEM Non-Essential Amino Acids Solution (Gibco, 11140035). WT and Opa1KO MEF cells were described previously23 and were provided and validated by the laboratory of T.W. MEFs were cultured in DMEM, 1× high D-glucose (4.5 g l−1) + GlutaMAX + sodium pyruvate + Phenol Red (Thermo Fisher, 11594446) with 5% FBS (Thermo Fisher, 11573397) and 1% penicillin–streptomycin solution (Sigma p4333). HeLa cells (CCL-2, ATCC) were cultured in DMEM 1× high D-glucose (4.5 g l−1)+ 10% FBS+ 1% penicillin–streptomycin solution and pyruvate. HepG2-Lucia (hpgl-ahr, Invivogen) was maintained according to the manufacturer’s protocol. All the cell lines were kept in 37 °C, 5% CO2 humidified incubator. All the cell lines were kept in 37 °C, 5% CO2 humidified incubator.
Human PBMCs from whole blood were isolated by using 50-ml SepMate PBMC Isolation Tubes with an insert for density gradient centrifugation (Stem Cell, 85450). Pre-warmed density gradient centrifugation solution Histopaque (Sigma, 10771) has been added to the tube in advance. Blood was collected into EDTA-coated vacutainer tubes (BD Biosciences) and diluted 1:1 with washing buffer whose recipe is PBS with 1% FBS and 2 mM EDTA (Thermo Fisher, 15575020). Then, the diluted blood was gently pipetted onto the SepMate PBMC Isolation Tubes. The tubes were centrifuged at 1,200g for 10 min at room temperature. After centrifugation, the top liquid above the insert were poured off, followed by washing and then lysing the red blood cells (BioLegend, 420301). PBMCs with high viability were obtained and cultured in RPMI 1640 GlutaMAX Supplement medium with 10% FBS, 1% penicillin–streptomycin solution, 1 mM sodium pyruvate (Gibco, 11360070), 1× MEM Non-Essential Amino Acids Solution and 50 μM β-mercaptoethanol, at 37 °C with 5% CO2.
Mice were killed according to institutional ethical guidelines. The spleens were aseptically removed and transferred to a sterile Petri dish containing culture medium. Spleens were gently smashed by using a sterile syringe and passed through a 70-µm cell strainer (Corning, 352350) placed over a 50-ml conical tube to mechanically dissociate the tissue. After centrifugation, washing and red blood cell lysis was carried out. A single cell suspension of spleen was obtained and cultured in RPMI 1640 GlutaMAX plus 10% FBS, 1 mM sodium pyruvate, 1 × MEM Non-Essential Amino Acids and 50 μM β-mercaptoethanol for future study.
IBD cohort
The cohort was 179 healthy controls, 77 patients with CD and 80 patients with UC with observations at baseline, 2–14 weeks following the initialization of biologic therapy. Patients were recruited between 27 August 2014 and 11 January 2021 by the Department of Internal Medicine I at the University Medical Center Schleswig–Holstein (Campus Kiel, Germany), and the healthy reference population was recruited in 2016 at the University Medical Center Schleswig–Holstein (Campus Kiel, Germany). None of these participants had received any antibiotics or other medication 2 months before inclusion. All enrolled patients with IBD or healthy controls gave informed consent and wrote informed consent forms. Participants were enrolled regardless of sex; no restrictions were placed on the proportion of male or female participants. Individuals with diabetes were excluded from the analysis. CD patients with CRP ≥ 5 and/or CDAI > 150 and/or HBI ≥ 5 were considered active, and patients with UC with CRP ≥ 5 and/or CAI ≥ 5 and/or partial Mayo ≥ 2 were considered active. For the longitudinal analysis, only patients active at baseline were included (n = 138); patients inactive at week 14 were considered as responders. Targeted metabolomics of serum samples was performed using the Biocrates MxP Quant kit. Linear mixed models were used to assess the relationship between log-transformed IPA and the three disease activity indicators (CRP, CDAI and partial Mayo), as well as between the healthy reference population and UC/CD. Detailed information has been described elsewhere51.
Animals
WT and other genetic knockout mice were maintained on a C57BL/6J background. All the mice used in DSS colitis or adoptive transfer moder were female and 10–12 weeks old. For ABX-treatment experiment or in vitro splenocyte isolation, no significant sex differences were found in the reported experiments. Mice were kept in a specific-pathogen-free environment and provided with ad libitum access to adequate food (LASQCdiet Rod16, LASvendi) and water. They were maintained on a 12-h light–dark cycle and the temperature was constant. No statistical methods were used to pre-determine sample sizes but our sample sizes are similar to those reported in previous publications52. All experimental animals were randomly divided into groups. Data collection and analysis were performed blind to the conditions of the experiments. The animal facility was granted approval (D-75-12-01) given by the French Administration. All experiments were conducted according to the European Communities Council Directive (2010/63/UE) for the care and use of animals for experimental procedures and complied with the regulations of the French Ethics Committee in Animal Experiment «Charles Darwin» registered by the French authority under CE005. All procedures were approved by this committee and all efforts were made to minimize suffering.
Method details
ATP measurement
CellTiter-Glo Luminescent Cell Viability Assay (Promega, G7573) was used to measure ATP levels of Jurkat cells following the manual instructions. A standard curve of ATP was performed for ATP quantification. After 8 h of treatment, Jurkat cells were collected and incubated in CellTiter-Glo luminescence ATP reconstituted reagent for 10 min and luminescence was measured by a multimode SpectraMax.
Protein quantification
Protein amounts were measured by using BC Assay: Protein assay kit (Interchim, FT-40840A) following the manufacturer’ instructions.
SCENITH
For in vitro human PBMCs or mouse splenocytes, 0.2 million cells seeded in a 96-well plate were pre-treated with or without different dosages of IPA (Sigma, 220027) including 1, 10, 100 and 1,000 μM for 2 h, then stimulated with or without CD3/CD28 T-Activator Dynabeads (Thermo Fisher, 11132D for human; Thermo Fisher, 11453D for mouse), respectively for 4 h. After metabolite treatment and CD3/CD28 activation, cells were collected and treated during 15 min with control, 2-DG, oligomycin and the sequential combination of the two inhibitors (2-DG + oligomycin). Puro (final concentration 10 μg ml−1) was added in the last 15–45 min of the metabolic inhibitor treatment. After puromycin incubation, cells were washed in cold washing buffer and then Fc receptor blockade, cell live–dead staining, surface staining and anti-puro monoclonal antibody with Alexa Fluor 488 staining were performed in sequence. Finally, cell samples were collected and analysed by flow cytometry.
Total and naive CD4+ T cell isolation and differentiation
Mouse CD4+ T cells were isolated and purified using a mouse CD4+ T cell isolation kit (Miltenyi, 130-104-454) according to the manufacturer’s instructions. For CD4+ T cell differentiation, 0.1 million cells were plated in a 96-well plate and activated by CD3/CD28 Dynabeads and differentiated into various TH cell subsets using the following cytokine environment. For TH1 cells, 1 ng ml−1 mIL-2 (Miltenyi, 130-120-331) and 10 ng ml−1 mIL-12 (Miltenyi, 130-096-707). For TH17 cells, 20 ng ml−1 mIL-6 (Miltenyi, 130-096-682), 10 ng ml−1 IL-23 (Miltenyi, 130-096-676), 10 ng ml−1 mIL-1β (Miltenyi, 130-101-681) and 2 ng ml−1 hTGF-β (Miltenyi, 130-095-067). For Treg cells, 20 ng ml−1 hIL-2 (R&D systems, 202-IL-010) and 10 ng ml−1 hTGF-β were used. For naive CD4+ T cell differentiation, a naive CD4+ T Cell Isolation kit (Miltenyi, 130-104-453) was used. Then, 0.05 million cells were plated in a 96-well plate, activated by CD3/CD28 Dynabeads and differentiated into TH1 and TH17 cell phenotypes. The cytokine environment is described as follows. For TH1 cells, 0.5 ng ml−1 mIL-2 (Miltenyi, 130-120-331) and 0.01 ng ml−1 mIL-12 (Miltenyi, 130-096-707). For TH17 cells, 20 ng ml−1 mIL-6 (Miltenyi, 130-096-682), 10 ng ml−1 IL-23 (Miltenyi, 130-096-676), 10 ng ml−1 mIL-1β (Miltenyi, 130-101-681) and 2 ng ml−1 hTGF-β (Miltenyi, 130-095-067). Cells were treated with or without different doses of IPA (from 1 μM to 1,000 μM). For purely cellular phenotyping experiments, after 3 days of treatment, differentiated CD4+ T or naive CD4+ T cells were restimulated with 50 ng ml−1 phorbol 12-myristate 13-acetate (PMA; Sigma, P8139) and 1 μM ionomycin (InvivoGen, 56092-82-1) in the presence of 5 μg ml−1 Brefeldin A (Sigma, B7651) in the last 5 h. Finally, the samples were collected and a phenotype test was performed using cytokine and TF factor markers.
For the IPA ‘washout’ experiment, all naive CD4+ T cells were incubated with IPA for 24 h in TH1 or TH17 cell differentiation conditions. This would be considered the T0 time point. Next, various doses of IPA (from 1 μM to 1,000 μM) persisted in the culture medium for one group, whereas it was completely washed out for the other group. We observed the TH1 and TH17 cell differentiation phenotype 24 h (T24) and 48 h (T48) after IPA was washed out. In addition, to study whether IPA could still exert its role on hindering TH1 and TH17 cell differentiation of naive CD4+ T cells in the acute phase, IPA (from 1 μM to 1,000 μM) was added only during the last 5 h of restimulation by PMA/ionomycin.
For energy metabolism and PPARβ/δ signalling pathway intervention experiments, various doses of oligomycin from 1 μM to 0.1 nM, 1 μM etomoxir, 1 μM TOFA, 3 μM BPTES or 10 μM GSK3787 were added into the culture system with metabolite IPA together in different differentiation cytokine conditions. For GW501516, 5 μM and 50 μM were tested as positive controls for the PPARδ nuclear receptor. On day 2, the supernatant was discarded and fresh metabolites and corresponding inhibitors were re-added. After 3 days of treatment, cell samples were collected and analysed for TH1 and TH17 cell phenotypes by flow cytometry. All flow cytometry analyses were performed on Cytoflex (Beckman) and data were analysed with FlowJo software.
Naive CD4+ T cell proliferation
Naive CD4+ T cells were isolated and purified from mouse splenocytes by using a Miltenyi naive CD4+ T cell isolation kit (Miltenyi, 130-104-453). Purified cells were stimulated with CD3/CD28 Dynabeads under varying experimental conditions, including no cytokine and TH1 or TH17 cell differentiation conditions in the presence or absence of IPA at specified concentrations (from 1 μM to 1,000 μM). Cytokine compositions for TH1 and TH17 cell differentiation were maintained as described above. Cell proliferation was assessed using the CellTrace CFSE Cell Proliferation kit (Thermo Fisher Scientific, C34554) following the manufacturer’s protocol. After 5 days of culture, cells were collected and analysed by flow cytometry to quantify proliferation.
Metabolic flux analysis (Seahorse)
WT or AhR−/− mouse CD4+ T cells (0.2 million cells per well, plated in poly-L-lysine pre-coated plate to immobilize the cells immediately before the experiment) were incubated with or without IPA in basal or CD3/CD28 activation state. Then, 2 × 104 of WT and OPA1KO MEFs were seeded in a 96-well plate and treated with or without IPA for 6 h. The OCR and extracellular acidification rate were determined using a Seahorse XF96 Extracellular Flux Analyzer (Seahorse Bioscience) following the protocols recommended by the manufacturer. For the real-time ATP rate assay, mitoATP and glycoATP were calculated with a sequential injection of 1.5 μM oligonucleotide and 0.5 μM rotenone or antimycin A (Agilent, 103592-100). For the Mitostress assay, OCR was tested by sequential injection of 1.5 μM oligomycin, 1 μM FCCP (carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone) and 0.5 μM rotenone and antimycin A for CD4+ T cells (Agilent, 103015-100). For the glycolytic rate assay, a Seahorse XF Glycolytic Rate Assay kit (Agilent Technologies, 103344-100) was used according to the manufacturer instructions. However, for MEFs, 2 μM FCCP was used in the experiment. The mitochondrial energy metabolism substrates were detected by Seahorse XF Substrate Oxidation Stress Test kits (Agilent, 103672-100, 103673-100, 103674-100). Data were analysed by wave software (Agilent).
For PA replenishment experiments, CD4+ T cells were stimulated with or without CD3/CD28 Dynabeads in the presence or absence of various doses of IPA (from 1 μM to 1,000 μM). We set up different serum conditions including 10% FBS, 0 FBS, bovine serum albumin (BSA) vehicle control (8.3%) and PA (200 μM) replenishment. A Seahorse Mitostress assay was performed after 6 h incubation as described above. For CD36 lipid transporter inhibition experiment, CD4+ T cells were pre-incubated with CD36 neutralizing antibody or its isotype control at the same concentration 10 μg ml−1 for 30 min. Then, the cells were stimulated with or without CD3/CD28 Dynabeads in the presence or absence of various doses of IPA (from 1 μM to 1,000 μM) for 6 h. Finally, a Seahorse Mitostress assay was performed as described above.
For the PPAR signalling pathway intervention experiments, 1 μM GSK3787 or 5 μM GW6471 was added into the culture system with IPA together to treat the cells for 6 h.Then, 5 μM GW501516 or 1 μM GW7647 was tested for PPARδ and PPARα positive control, respectively. Ficz at 10 ng ml−1 was used as a positive control for the AhR signalling pathway. Finally, cells were collected and a Seahorse Mitostress assay was performed according to the manual.
Lactate production measurement
Total CD4+ T cells were isolated from mice and cultured under different experimental conditions. Cells were either stimulated with CD3/CD28 Dynabeads or left unstimulated, in the presence or absence of increasing concentrations of IPA (1 μM to 1,000 μM). Following a 6-h incubation period, cell culture supernatants were collected and analysed for lactate concentration using a commercially available assay kit (Promega, J5021), following the manufacturer’s protocol.
MitoTracker Green/Deep Red and ROS production
For mitochondria membrane potential and mitochondrial mass detection, CD4+ T cells were cultured with or without IPA in basal or CD3/CD28 activation state for 24 h. Cells were collected and incubated with MitoTracker Green FM (1:6,000) and MitoTracker Deep Red (1:5,000) in FACS buffer for 20 min at room temperature (Thermo Fisher Scientific). The cell treatment time was extended to 48 and 72 h for the detection of ROS production in cytoplasm and mitochondria. Finally, the cells were incubated with 5 μM mitoSOX (Thermo Fisher) and 10 μM DCFDA (Sigma) for 20 min at 4 °C and assayed by flow cytometry.
Lipid fluorescent analogues
Consistent with the MitoTracker experiment, CD4+ T cells were cultured with or without IPA in basal or CD3/CD28 activation state for 24 h. Cells were collected and incubated with 10 μM BODIPY 493/503 or 5 μM BODIPY FL C12 in FACS buffer for 20 min at 4 °C (Thermo Fisher Scientific) and assayed by flow cytometry.
Electron microscopy
For TEM, mouse CD4+ T cells were chemically fixed with 2.5% glutaraldehyde in culture medium for 1 h at room temperature, washed in 1× PHEM buffer (60 mM PIPES, 25 mM HEPES, 10 mM EGTA, 2 mM MgCl2, pH 7.3) and centrifuged at 9,500g. Cells were postfixed in a mix of 1% osmium tetroxide and 1.5% potassium ferrocyanide for 1 h, washed in 1× PHEM, incubated in 1% tannic acid for 30 min. After another wash in 1× PHEM, cells were incubated in 1% osmium tetroxide for 1 h, washed in water and dehydrated in ethanol in an increasing series from 25% to 100%. All samples were then incubated in propylene oxide and infiltrated in an Epon resin/propylene oxide mix v/v53. Samples were then embedded in pure Epon resin (EMS), centrifuged at 13,680 rcf, followed by polymerization for 48 h at 60 °C. Ultrathin sections (70 nm) were cut with a Leica Ultracut UC7 microtome, and stained with 4% uranyl acetate, followed by 3% lead citrate. TEM images were captured with a Tecnai Spirit 120 kV TEM equipped with a bottom-mounted Eagle 4k × 4k camera (FEI). The methods of analysing mitochondrial number, global shape and cristae parameter were indicated previously21.
Mitochondrial fluorescence imaging
The mitochondria morphology images of WT or Opa1KO MEFs were acquired using the Operetta CLS High-Content Analysis systems (PerkinElmer)23. Mitochondria membrane potential was stained by using tetramethylrhodamine ethyl ester perchlorate (TMRE), and mitochondria mass was represented by genetically encoded mitochondrially targeted YFP (mitoYFP).
Mitochondria isolation
Cardiac mitochondria were isolated freshly. In brief, ventricles were separated from the atria and non-myocardial tissues, cut into small pieces and homogenized manually in an ice-cold 2-ml homogenizer using IB buffer (275 mM sucrose, 20 mM Tris and 1 mM EGTA–KOH, pH 7.2) with trypsin–EDTA (0.05%). To inhibit trypsin activity, BSA fatty acid-free (0.25 mg ml−1) and protease inhibitor cocktail (Roche) were added. Cardiac homogenates were then centrifuged at a lower speed (1,000g, 10 min, 4 °C) to remove nuclei and debris, followed by a higher-speed centrifugation (3,200g, 15 min, 4 °C) to isolate the crude mitochondrial fraction. Finally, the crude mitochondrial pellet was resuspended in IB buffer, and the protein concentration was measured using a Bradford assay.
High-resolution respirometry
Oxygen consumption was measured in cardiac mitochondria using high-resolution respirometry (O2k-Fluorespirometer). For this procedure, cardiac mitochondria were freshly isolated from adult WT mice (8 weeks old) as detailed above. Specifically, 50 µg of cardiac mitochondria were resuspended in Mir05 buffer (3 mM MgCl2-6H2O, 60 mM lactobionic acid, 20 mM taurine, 10 mM KH2PO4, 20 mM HEPES–KOH, 110 mM sucrose, 0.5 mM EGTA–KOH and 1 g l−1 BSA). Two protocols were performed. The first protocol evaluated whether IPA act directly on the isolated mitochondria. Before oxygen evaluation analysis, mitochondria were incubated with IPA (100 µM) or vehicle solution (DMSO) for 4 h under gentle agitation at 4 °C. The mitochondrial respiration capacity was determined through the sequential addition of PGM (10 mM pyruvate, 5 mM glutamate and 5 mM malate; state 2) in the presence of 1 mM ADP (state 3) to evaluate complex I-driven respiration; rotenone (0.5 µM) and ascorbate (2 mM), N,N,N′,N′-tetramethyl-p-phenylenediamine (TMPD; 0.5 mM) and carbonyl cyanide m-chlorophenyl hydrazone (CCCP; 2 µM) for complex IV-driven respiration. The second protocol evaluated whether IPA acts as an uncoupling drug. Mitochondrial respiration capacity was determined through the sequential addition of PGM (10 mM pyruvate, 5 mM glutamate and 5 mM malate; state 2) in the presence of 1 mM ADP (state 3) to evaluate complex I-driven respiration; oligomycin (1 µM) to block ATP synthase (complex V) followed by IPA (100 µM) or vehicle solution (DMSO); to serve as an experimental control, CCCP (2 µM) was used (a known mitochondrial uncoupling drug) and ascorbate (2 mM), TMPD (0.5 mM) and CCCP (2 µM) for complex IV-driven respiration.
Cpt1a knockdown by using siRNA approach in MEF cells
A total of 100,000 MEFs were seeded in antibiotic-free medium 24 h before transfection to achieve 70–90% confluency at the time of transfection. For each well of a six-well plate, Cpt1a siRNA (final concentration 20 nM, siGENOME Mouse Cpt1a siRNA, SMARTPool, Horizon Discovery) and Lipofectamine RNAiMAX (7.5 µl per well, Thermo Fisher) were separately diluted in Opti-MEM Reduced-Serum Medium (Thermo Fisher). Both tubes were inverted by hand for 2–3 s. Then, the diluted siRNA was combined and incubated with the diluted Lipofectamine RNAiMAX reagent for 10 min at room temperature to form siRNA–lipid complexes. The mixture was added dropwise to cells, followed by gentle swirling. Cell culture medium should be refreshed after 6 h transfection at 37 °C in a 5% CO2 humidified atmosphere to avoid cytotoxicity. Control transfections included non-targeting siRNA (negative control, Sigma) and transfection reagent-only (mock control). Finally, the cells were collected after 48 h for Cpt1a expression detection and a further Seahorse Mitostress experiment.
RNA extraction, reverse transcription and qPCR
A total 0.5 million MEFs were collected and RNA were extracted by using QIAGEN RNeasy Micro kit (QIAGEN, 74004) according to the manufacturer’s instructions. Finally, RNA concentration and purity were assessed by Nanodrop (A260/A280 ratio ≥ 1.8). Reverse transcription was carried out using an Applied Biosystems High-Capacity cDNA Reverse Transcription kit (Thermo Fisher) and the cycle as followed: step 1, 25 °C for 10 min; step 2, 37 °C for 120 min; and step 3, 85 °C for 5 min, then the cDNA samples were held in 4 °C. Cpt1a and housekeeping gene RPLP0 were quantified using specific primers. Please refer to the supplementary material for the specific sequences.
Cpt1a: forward: 5′-TGGCATCATCACTGGTGTGTT-3′,
Reverse: 5′-GTCTAGGGTCCGATTGATCTTTG-3′.
RPLP0: forward: 5′-AGATTCGGGATATGCTGTTGGC-3′,
Reverse: 5′-TCGGGTCCTAGACCAGTGTTC-3′.
Amplifications were performed with the following conditions: denaturation step 10 min at 95 °C; 40 cycles of 95 °C denaturation for 15 s; and 55 °C extension for 1 min followed by a standard melting curve programme.
Proteomics sequencing and data analysis
Materials
MS-grade acetonitrile (ACN), MS-grade H2O, ammonium bicarbonate (NH4HCO3), MS-grade formic acid (FA), Tris(2-carboxyethyl) phosphine hydrochloride (TCEP-HCl) and S-methyl methanethiosulphonate (MMTS) were from Thermo Fisher Scientific. Sequencing-grade trypsin/Lys C mix was from Promega. Ammonium bicarbonate (NH4HCO3) was from Sigma-Aldrich.
Samples preparation before LC–MS/MS analysis
A 6× volume of cold acetone (−20 °C) was added to a sample volume containing 10 µg of protein extracts. Vortexed tubes were incubated overnight at −20 °C then centrifuged for 10 min at 11,000 rpm and 4 °C. The supernatant was removed, then the protein pellets were dissolved in urea 8 M NH4HCO3 25 mM buffer.
Samples were then reduced with TCEP-HCl 10 mM and alkylated with MMTS 20 mM. After a 16-fold dilution in NH4HCO3, samples were digested overnight at 37 °C by a mixture of trypsin/Lys C (1:20 enzyme:substrate ratio). The digested peptides were loaded and desalted on Evotips Pure, provided by Evosep one (Odense) according to the manufacturer’s procedure.
LC–MS/MS acquisition
Samples were analysed on a timsTOF Pro 2 mass spectrometer (Bruker Daltonics) coupled to an Evosep one system (Evosep) operating with the 30SPD method developed by the manufacturer. In brief, the method is based on a 44-min gradient and a total cycle time of 48 min with a C18 analytical column (0.15 × 150 mm, 1.9-µm beads, ref EV-1106) equilibrated at 40 °C and operated at a flow rate of 500 nl min−1. H2O/0.1% FA was used as solvent A and ACN/ 0.1% FA as solvent B. The timsTOF Pro 2 was operated with a DIA-PASEF method comprising 12 pydiAID frames with three mass windows per frame resulting in a cycle time of 0.975 s as described in Bruker application note LC–MS 218.
Data analysis
MS raw files were processed using Spectronaut 18 (Biognosys). Data were searched against the SwissProt reviewed Mus musculus (11_2023, 17199 entries) database. Specific tryptic cleavages were selected and a maximum of two missed cleavages were allowed. The following post-translational modifications were considered for identification: acetyl (protein N-term), oxidation (M), as variables and MMTS (C) as fixed. The maximum number of variable modifications was set to 2. Identifications were filtered based on a 1% precursor and protein q-value cutoff threshold. The protein LFQ method was set to automatic and the quantity was set at the MS2 level with a cross-run normalization applied. Multivariate statistics on protein measurements were performed using Qlucore Omics Explorer v.3.9 (Qlucore AB). A positive threshold value of 1 was set to allow a log2 transformation of abundance data for normalization that is all abundance data values below the threshold are replaced by 1 before transformation. The transformed data were finally used for statistical analysis, GSEA and KEGG enrichment analysis using Majorbio Cloud54. The differentially expressed proteins between the two groups were analysed using the Welch’s t-test, with P = 0.05 as the cutoff value for statistical analysis. Among them, the high expression protein was upregulated by more than 1.2 times, and the low expression protein was downregulated by 0.83 times.
PPAR transactivation assay
Transactivation of PPARβ following IPA treatment was measured using a luciferase-based assay as described previously55. In brief, HEK293 cells (CRL-1573, ATCC) were seeded in 12-well plates at a density of 5 × 104 cells per well and grown to 50–70% confluence. Cells were co-transfected with 500 ng of a dominant-negative vector targeting PPARα, or PPARγ or PPARβ/δ, along with 500 ng of the PPRE-Luc reporter plasmid (kindly provided by H. Fahmi) and 100 ng of the pRL-TK plasmid (Promega) for normalization. The transfection process was performed using DharmaFECT kb, following the manufacturer’s instructions. After 24 h, the cells were stimulated with IPA at the indicated concentrations for 48 h. Luciferase activity (both firefly and Renilla) was then measured according to the Promega protocol for dual-luciferase assays.
AhR activity measurement
The AhR activity was measured using HepG2-Lucia AhR reporter cells (InvivoGen, France). Following the manufacturer’s protocol, cells were plated in a 96-well format, stimulated for 24 h and subsequently assessed for luciferase activity. Measurements were performed on a luminometer with Quanti-Luc reagent (InvivoGen). The metabolites indole, indoxyl-3-sulphate (I3S) (I3875), indole-3 aldehyde (I3A) (129445), hydroxyindole (Ind-OH) (H31859), IPA (220027), indole-3-lactic acid (ILA) (I5508), IAA (I3750), oxindole (O9808), tryptamine (193747), tryptophol (T90301) and Ficz (SML1489) were all purchased from Sigma-Aldrich. The results were normalized by luciferase activity from each sample and cytotoxicity measurements (CytoTox 96 Non-radioactive Cytotoxicity Assay, Promega).
ABX-treated mouse model
Mice were treated with a cocktail of antibiotics including neomycin (1 g l−1, Sigma), colistin (850,000 IU l−1, Sigma), gentamycin (35 mg l−1, Sigma), vancomycin (500 mg l−1, Sigma) and amphotericin-B (10 mg l−1, Sigma) in drinking water for 7 days to deplete gut microbiota. IPA was dissolved in a mixed solution with 10% DMSO (Sigma), 40% PEG300 (MCE, HY-Y0873), 5% Tween-80 (Sigma, P1754) and 45% saline. IPA (200 mg kg−1) and the corresponding vehicle control were administered to mice daily by oral gavage.
Faecal bacteria DNA extraction and qPCR
Faecal genomic DNA was extracted from the stool samples using the Gordon DNA extraction method56. Bacterial quantification was performed by qPCR using SYBR Green PCR Master Mix (Applied Biosystems) in a StepOnePlus apparatus (Applied Biosystems). Each reaction was carried out in duplicate in a final volume of 20 µl with 0.2 µM each primer and 5 µl of diluted DNA. Total bacteria were quantified using specific primers (Supplementary material provides detailed sequences): Uni 16SR 5′-ATTACCGCGGCTGCTGGC-3′ and Uni 16SF 5′-ACTCCTACGGGAGGCAGCAGT-3′. Amplifications were performed with the following conditions: denaturation step 10 min at 95 °C; 40 cycles of 95 °C denaturation for 15 s; and 55 °C extension for 1 min followed by a standard melting curve programme.
Mouse DSS model
To induce colitis in WT mice, mice were administered drinking water supplemented with 2% DSS (MP Biomedicals) for 7 days and were then allowed to recover by drinking normal water for the following 5 days. Then, 3% DSS was used in the Rag2−/− colitis experiment. Different percentages of DSS were used in the experiments, depending on the product batch and the mice genotype, to avoid reaching a predefined limit point (where weight loss exceeds 20% of the initial weight). For all the DSS and adoptive transfer model, we all used female mice aged 10–12 weeks to avoid excessive inflammation and immune rejection between males and females during the experiment. IPA (200 mg kg−1) and the corresponding vehicle control were administered to mice daily by oral gavage. For the DSS model, mice were weighed and scored daily for disease severity and colon tissue was obtained on day 12 to perform haematoxylin and eosin (H&E) staining.
Isolation of lamina propria immune cells
For the isolation of LP immune cells, the small intestine or colon was cut out and opened longitudinally. Peyer’s patches were removed simultaneously. Tissues were washed with cold 1× PBS to remove the faecal and then were cut into 0.5–1-cm long pieces cross-sectionally and then mixed with 5 ml prewarmed HBSS 1× with 5% FBS, EDTA (5 mM, Thermo Fisher), 1% penicillin–streptomycin, 10 mM HEPES and dithiothreitol (DTT; 0.145 mg ml−1, Sigma-Aldrich) in the shaker at 37 °C for 20 min. The supernatants containing epithelial cells and intraepithelial lymphocytes (IELs) were discarded and pellets were washed with cold PBS 1×. The tissue pieces were then transferred to a new tube and digested with collagenase type VIII (0.5 mg ml−1, Sigma-Aldrich) supplemented with DNase I (1 mg ml−1, Sigma-Aldrich) for 30 min. All the contents were passed through a 100-μm cell strainer. LPLs were obtained using 40/80 Percoll centrifugation (Sigma-Aldrich). Cells were centrifuged briefly and suspended in complete RPMI containing 10% FBS, 1% HEPES (Gibco), 100 U ml−1 penicillin, 100 μg ml−1 streptomycin (Gibco) and 50 μM 2-mercaptoethanol (Gibco). For the ABX-treated mouse model, 0.5–1 million cells were plated in 96-well plates for the following SCENITH experiments. For the DSS colitis model, LP immune cells were used for energy metabolism detection by SCENITH and CD4+ T cell immunophenotype exploration. Cells were stimulated with 50 ng ml−1 PMA (Sigma, P8139) and 1 μM ionomycin (InvivoGen, 56092-82-1) in the presence of Brefeldin A (Sigma, B7651) for 4 h to determine the cytokine production by TH1 and TH17 cells.
Rag2 −/− adoptive transfer model
For donor cell preparation, mesenteric lymph nodes were collected from DSS mice treated with or without IPA or ABX-depleted mice administered with or without IPA. Donor mice were administrated with IPA (200 mg kg−1) or corresponding vehicle control daily by oral gavage. Then, CD4+ T cells were isolated and purified by using mouse CD4+ T cell isolation kit. One million CD4+ T cells were injected intraperitoneally into Rag2−/− recipient mice submitted to DSS colitis on day 0 and day 7. All the donor and recipient mice are aged between 10 and 12 weeks. Mice were weighed and scored daily for disease severity, and colon tissue was obtained on day 12 to perform H&E staining.
Histology
The histological scoring on H&E-stained slides was evaluated by three main categories correlating with inflammation, including immune cell infiltration, mucosal damage and changes in mucosal architecture. The specific scoring criteria have been described previously57.
Statistical analysis
All data were represented as mean ± s.e.m. and GraphPad Prism v.10.0 (GraphPad Software) was used for statistical analysis. Data distribution was assumed to be normal but this was not formally tested. Data from two groups were compared using a Student’s t-test. Data from more than two groups were compared using a one-way analysis of variance (ANOVA) or two-way ANOVA as described in the figure caption. The n values referring to replicate numbers are also noted. A P value less than 0.05 was considered statistically significant and specific P values are shown in each figure.
For the data obtained from human subjects, linear mixed models were used to assess the relationship between log-transformed IPA and the three disease activity indicators (CRP, CDAI and partial Mayo), as well as between the healthy reference population and UC/CD. Multiple hypothesis correction was accomplished using Benjamini–Hochberg adjustment for the comparison between controls, UC and CD. CRP values were not available for the healthy reference population. To assess differences between responders and non-responders over the 26-week period following biologic therapy induction, we performed linear models at each time point. Although linear mixed-effects models with group × time interactions were initially considered, these models attenuated later group differences due to minimal early divergence and partial pooling across time points. Visual inspection of the data indicated that group separation emerged progressively, in line with our previous hypothesis. We therefore adopted time-point-specific models to better capture the temporal dynamics of the response. Given that these analyses reflect a single, biologically driven hypothesis regarding group divergence over time, we did not apply corrections for multiple comparisons. Results from both linear and mixed-effects models are included in the Supplementary Information for comparison. Linear mixed models included an interaction term between time in weeks (coded as an ordered factor) and response. Contrast analysis was applied to assess the pairwise differences between responders and non-responders at each time point. Patient IDs were included as random intercepts, and P values were calculated using Satterthwaite’s method at a nominal significance value of 5%. Sex assigned at birth and age were included as covariates in all models. Residual versus fitted plots were used to check for signs of non-linearity, unequal variance, or outliers. Models were built using R v.4.3.2, linear mixed models were built using lmerTest (v.3.1-3), linear models with stats (v.4.4.2) and contrast analysis was performed using emmeans (v.1.10.6).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Any additional information required to reanalyse the data reported in this paper is available from the lead contact upon request. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD057537. Source data are provided with this paper.
Code availability
R code for the analysis of IPA concentrations in the IBD cohort is provided in the Supplementary Materials.
References
Glassner, K. L., Abraham, B. P. & Quigley, E. M. M. The microbiome and inflammatory bowel disease. J. Allergy Clin. Immunol. 145, 16–27 (2020).
Lee, M. & Chang, E. B. Inflammatory bowel diseases (IBD) and the microbiome-searching the crime scene for clues. Gastroenterology 160, 524–537 (2021).
Hayward, J. A., Mathur, A., Ngo, C. & Man, S. M. Cytosolic recognition of microbes and pathogens: inflammasomes in action. Microbiol Mol. Biol. Rev. 82, e00015–e00018 (2018).
Krautkramer, K. A., Fan, J. & Backhed, F. Gut microbial metabolites as multi-kingdom intermediates. Nat. Rev. Microbiol. 19, 77–94 (2021).
Shi, N., Li, N., Duan, X. & Niu, H. Interaction between the gut microbiome and mucosal immune system. Mil. Med. Res. 4, 14 (2017).
Agus, A., Planchais, J. & Sokol, H. Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724 (2018).
Michaudel, C. & Sokol, H. The gut microbiota at the service of immunometabolism. Cell Metab. 32, 514–523 (2020).
Roger, A. J., Munoz-Gomez, S. A. & Kamikawa, R. The origin and diversification of mitochondria. Curr. Biol. 27, R1177–R1192 (2017).
Yardeni, T. et al. Host mitochondria influence gut microbiome diversity: a role for ROS. Sci. Signal 12, eaaw3159 (2019).
Moschandrea, C. et al. Mitochondrial dysfunction abrogates dietary lipid processing in enterocytes. Nature 625, 385–392 (2024).
Ho, G. T. & Theiss, A. L. Mitochondria and inflammatory bowel diseases: toward a stratified therapeutic intervention. Annu Rev. Physiol. 84, 435–459 (2022).
Lee, H., Jeon, J. H. & Kim, E. S. Mitochondrial dysfunctions in T cells: focus on inflammatory bowel disease. Front. Immunol. 14, 1219422 (2023).
Martini, G. R. et al. Selection of cross-reactive T cells by commensal and food-derived yeasts drives cytotoxic T(H)1 cell responses in Crohn’s disease. Nat. Med. 29, 2602–2614 (2023).
Gomez-Bris, R. et al. CD4 T-Cell Subsets and the Pathophysiology of Inflammatory Bowel Disease. Int. J. Mol. Sci. 24, 2696 (2023).
Matsumoto, M. et al. Impact of intestinal microbiota on intestinal luminal metabolome. Sci. Rep. 2, 233 (2012).
Zeng, X. et al. Gut bacterial nutrient preferences quantified in vivo. Cell 185, 3441–3456 e3419 (2022).
Arguello, R. J. et al. SCENITH: a flow cytometry-based method to functionally profile energy metabolism with single-cell resolution. Cell Metab. 32, 1063–1075 e1067 (2020).
Hamaidi, I. & Kim, S. Sirtuins are crucial regulators of T cell metabolism and functions. Exp. Mol. Med. 54, 207–215 (2022).
Wai, T. Is mitochondrial morphology important for cellular physiology?. Trends Endocrinol. Metab. 35, 854–871 (2024).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Lam, J. et al. A universal approach to analyzing transmission electron microscopy with ImageJ. Cells 10, 2177 (2021).
Kondadi, A. K., Anand, R. & Reichert, A. S. Cristae membrane dynamics – a paradigm change. Trends Cell Biol. 30, 923–936 (2020).
Cretin, E. et al. High-throughput screening identifies suppressors of mitochondrial fragmentation in OPA1 fibroblasts. EMBO Mol. Med. 13, e13579 (2021).
Donnarumma, E. et al. Mitochondrial fission process 1 controls inner membrane integrity and protects against heart failure. Nat. Commun. 13, 6634 (2022).
Choi, J. et al. Etomoxir repurposed as a promiscuous fatty acid mimetic chemoproteomic probe. iScience 27, 110642 (2024).
Modoux, M. et al. Butyrate acts through HDAC inhibition to enhance aryl hydrocarbon receptor activation by gut microbiota-derived ligands. Gut Microbes 14, 2105637 (2022).
Li, Y. et al. Peroxisome proliferator-activated receptors: a key link between lipid metabolism and cancer progression. Clin. Nutr. 43, 332–345 (2024).
Moullan, N. et al. Tetracyclines disturb mitochondrial function across eukaryotic models: a call for caution in biomedical research. Cell Rep. 10, 1681–1691 (2015).
Richter, U., Lahtinen, T., Marttinen, P., Suomi, F. & Battersby, B. J. Quality control of mitochondrial protein synthesis is required for membrane integrity and cell fitness. J. Cell Biol. 211, 373–389 (2015).
Flannigan, K. L. et al. The pregnane X receptor and indole-3-propionic acid shape the intestinal mesenchyme to restrain inflammation and fibrosis. Cell Mol. Gastroenterol. Hepatol. 15, 765–795 (2023).
Wang, G. et al. Microbiota-derived indoles alleviate intestinal inflammation and modulate microbiome by microbial cross-feeding. Microbiome 12, 59 (2024).
Lercher, A., Baazim, H. & Bergthaler, A. Systemic immunometabolism: challenges and opportunities. Immunity 53, 496–509 (2020).
Boguszewska, K., Szewczuk, M., Kazmierczak-Baranska, J. & Karwowski, B. T. The similarities between human mitochondria and bacteria in the context of structure, genome, and base excision repair system. Molecules 25, 2857 (2020).
Lubin, J. B. et al. Arresting microbiome development limits immune system maturation and resistance to infection in mice. Cell Host Microbe 31, 554–570 e557 (2023).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Ni, J., Wu, G. D., Albenberg, L. & Tomov, V. T. Gut microbiota and IBD: causation or correlation?. Nat. Rev. Gastroenterol. Hepatol. 14, 573–584 (2017).
Adolph, T. E. et al. The metabolic nature of inflammatory bowel diseases. Nat. Rev. Gastroenterol. Hepatol. 19, 753–767 (2022).
Tilg, H., Zmora, N., Adolph, T. E. & Elinav, E. The intestinal microbiota fuelling metabolic inflammation. Nat. Rev. Immunol. 20, 40–54 (2020).
Yki-Jarvinen, H., Luukkonen, P. K., Hodson, L. & Moore, J. B. Dietary carbohydrates and fats in nonalcoholic fatty liver disease. Nat. Rev. Gastroenterol. Hepatol. 18, 770–786 (2021).
Schulz-Kuhnt, A. et al. ATP citrate lyase (ACLY)-dependent immunometabolism in mucosal T cells drives experimental colitis in vivo. Gut 73, 601–612 (2024).
Corrado, M. et al. Dynamic cardiolipin synthesis is required for CD8(+) T cell immunity. Cell Metab. 32, 981–995 e987 (2020).
Ron-Harel, N. et al. Mitochondrial biogenesis and proteome remodeling promote one-carbon metabolism for T cell activation. Cell Metab. 24, 104–117 (2016).
Miret-Casals, L. et al. Identification of new activators of mitochondrial fusion reveals a link between mitochondrial morphology and pyrimidine metabolism. Cell Chem. Biol. 25, 268–278 e264 (2018).
Perdijk, O. et al. Antibiotic-driven dysbiosis in early life disrupts indole-3-propionic acid production and exacerbates allergic airway inflammation in adulthood. Immunity 57, 1939–1954 e1937 (2024).
Gesper, M. et al. Gut-derived metabolite indole-3-propionic acid modulates mitochondrial function in cardiomyocytes and alters cardiac function. Front. Med. 8, 648259 (2021).
Han, J. X. et al. Microbiota-derived tryptophan catabolites mediate the chemopreventive effects of statins on colorectal cancer. Nat. Microbiol 8, 919–933 (2023).
Cui, W. et al. Gut microbial metabolite facilitates colorectal cancer development via ferroptosis inhibition. Nat. Cell Biol. 26, 124–137 (2024).
Jia, D. et al. Microbial metabolite enhances immunotherapy efficacy by modulating T cell stemness in pan-cancer. Cell 187, 1651–1665 e1621 (2024).
Wang, H. et al. CD36-mediated metabolic adaptation supports regulatory T cell survival and function in tumors. Nat. Immunol. 21, 298–308 (2020).
Kuhn, B. et al. Structure-based design of indole propionic acids as novel PPARα/γ co-agonists. Bioorg. Med. Chem. Lett. 16, 4016–4020 (2006).
Harris, D. M. M. et al. Tryptophan degradation as a systems phenomenon in inflammation – an analysis across 13 chronic inflammatory diseases. eBioMedicine 102, 105056 (2024).
Michaudel, C. et al. Rewiring the altered tryptophan metabolism as a novel therapeutic strategy in inflammatory bowel diseases. Gut 72, 1296–1307 (2023).
Scandola, C., Lanza, F., Gachet, C. & Eckly, A. In situ exploration of murine megakaryopoiesis using transmission electron microscopy. J. Vis. Exp. https://doi.org/10.3791/62494 (2021).
Han, C. et al. Majorbio Cloud 2024: update single-cell and multiomics workflows. Imeta 3, e217 (2024).
Moulin, D. et al. Rosiglitazone induces interleukin-1 receptor antagonist in interleukin-1β-stimulated rat synovial fibroblasts via a peroxisome proliferator-activated receptor β/δ-dependent mechanism. Arthritis Rheum. 52, 759–769 (2005).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Sokol, H. et al. Card9 mediates intestinal epithelial cell restitution, T-helper 17 responses, and control of bacterial infection in mice. Gastroenterology 145, 591–601 e593 (2013).
Acknowledgements
The authors express their sincere gratitude to all colleagues who contributed to this work. The authors thank the China Scholarship Council for the financial support to the first author Q.L. We are grateful for support for equipment from the French Government Programme Investissements d’Avenir France BioImaging (no. ANR-10-INSB-04-01) and the French government (Agence Nationale de la Recherche) Investissement d’Avenir programme, Laboratoire d’Excellence) ‘Integrative Biology of Emerging Infectious Diseases’ (ANR-10-LABX-62-IBEID). The authors thank the animal facility care staff of the Saint-Antoine Hospital (UMR S938, PHEA) for their technical support, and thank S. Truong and N. Bernard (Saint-Antoine Hospital, APHP, France) for their clinical help. This project received funding from the European Research Council (ERC; ENERGISED, ERC-2021-COG-101043802) and MSD-AVENIR to H.S. and the EKFS Clinician Scientist Professorship (2020_EKFS.11), the BMBF (eMED Juniorverbund ‘Try-IBD’ 01ZX1915A) and the DFG RU5042 to K.A. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Q.L. and H.S. conceptualized and designed the study and performed the data analysis. Q.L. performed all the experiments unless otherwise indicated. R.O.F., V.P., L. Creusot, A.H.A., S.A., Z.H., Y.H., L. Chollet, C.D., N.R., M.-L.M. and T.W. provided technical assistance for the in vitro and in vivo experiments. M.A.C.-R. and Y.Z. conducted mitochondrial isolation, high-resolution respirometry detection and mitochondrial fluorescence imaging experiment. D.M.M.H., S.S. and K.A. provided human healthy and IBD patient clinical data and conducted the relevant analysis. F.L., S.E.S. and H.G. provided the knockout mice. C.S. performed electron microscopy experiments. T.L. helped to write the ethics project and provided technical support for animal experiments. G.C. conducted proteomic sequencing. R.J.A. provided SCENITH reagent kits and related technical support. J.B., M.D.A.-P. and M.C.B. helped with flow cytometry panel design and data analysis for human PBMC TH1/TH17 cell memory subsets. D.M. performed PPAR transactivation assays in HEK293 cells. L.B. and N.R. performed DNA extraction and qPCR bacterial quantification from mouse faeces. Q.L., R.O.F., M.L.M., T.W. and H.S. discussed the experiments and results. Q.L. and H.S. wrote the paper. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
H.S. reports lecture fees, board membership or consultancy from Carenity, AbbVie, Astellas, Danone, Ferring, Mayoly Spindler, MSD, Novartis, Roche, Tillots, Enterome, BiomX, Takeda and Biocodex, has stocks from Enterome, and is co-founder of Exeliom Biosciences. K.A. reports lecture fees, board membership or consultancy from AbbVie, Ferring, Johnson & Johnson, Takeda and Lilly. All other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Chaoran Li, Nicholas Jones and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Metabolite screening heatmap.
(A-C) Metabolite screening heatmap showing the mean ratio values of various doses of IPA-treated groups and control (Con) group in basal, 2-DG and oligomycin conditions. After 8 h incubation, ATP levels and protein amount were measured in Jurkat cells.
Extended Data Fig. 2 Effects of IPA on early activated CD4+ T cells and memory TH cell subsets.
(A) SCENITH readouts including mitochondrial dependence, glycolytic capacity, glucose dependence and FAO and/or AAO capacity of CD69+ CD4 + T cells in Con, CD3/CD28 activation and CD3/CD28 plus various doses of IPA (1, 10, 100, 1000 μM) conditions at 6 h time point, n = 6. (B-C) SCENITH readouts including mitochondrial dependence, glycolytic capacity, glucose dependence and FAO and/or AAO capacity of TH1 and TH17 Th memory subsets in Con, CD3/CD28 activation and CD3/CD28 plus various doses of IPA (1, 10, 100, 1000 μM) conditions at 6 h time point, n = 7 for each (outliers below 0 or above 100 are removed during analysis). Panel (A-C) represent mean ± SEM were analysed by two-tailed paired t-test based on different donors.
Extended Data Fig. 3 IPA alters the metabolic profile of mouse splenic CD4 + T cells.
(A) Schematic diagram showing the protocol of mouse splenocyte isolation and SCENITH experiment procedure. Created with BioRender.com. (B) UMAP analysis of splenocyte subsets in control (Con) and CD3/CD28 groups. (C) ATP synthesis generated by SCENITH in mouse CD4+ and CD8 + T cells that were cultured with or without CD3/CD28 activation in the presence of DMSO or IPA (10, 100, 1000 μM) for 6 h. n = 6. (D, E) Metabolic profile of CD4+and CD8 + T cells that were cultured in the same conditions above. n = 5. (F) Mechanism diagram showing the effect of IPA on altering the energy metabolism. (C-E) represent mean ± SEM analysed by one-way ANOVA with Tukey’s correction for multiple comparisons.
Extended Data Fig. 4 IPA Increases Mitochondrial Respiration via CD36 and Exogenous Lipids.
(A-C) MitoTraker Green, MitoTracker Deep Red and the ratio of the two markers of mouse CD4 + T cells cultured with/without CD3/CD28 in the presence of DMSO or IPA (100, 1000 μM) at 24 h time point. n = 3. (D) Schematic diagram showing the function of specific metabolic pathway inhibitors: UK5099 (mitochondrial pyruvate transportation), Etomoxir (FAO) and BPTES (glutanimonlysis). Created with BioRender.com. (E) Seahorse substrate oxidation assay of mouse CD4 + T cells cultured with/without CD3/CD28 activation in the presence of DMSO or IPA (100, 1000 μM) in the presence or absence of etomoxir, BPTES and UK5099 in 6 h. n = 3 for Con and CD3/CD28 group, n = 4 for IPA-treated groups in basal and CD3/CD28 activation state. (F-G) Seahorse Mitostress assay of CD4 + T cell in different serum conditions including 10% FBS, 0 FBS, BSA vehicle control and PA replenishment. The cells were stimulated with/without CD3/CD28 in the presence of DMSO or IPA (from 1 μM to 1000 μM) at 6 h. n = 3 for Con and n = 4 for the rest groups in different FBS conditions in Panel (F). n = 6 for Con and n = 8 for the rest groups in different FBS conditions in Panel (G). (H-I) Seahorse Mitostress assay of CD4 + T cell under IgG2a isotype control and CD36 neutralizing antibody conditions. The cells were stimulated with/without CD3/CD28 in the presence of DMSO or IPA (from 1 μM to 1000 μM) at 6 h. n = 6 for Con and n = 8 for the rest groups in LgG2a and CD36 Ab conditions. Panel (G), (I) represent mean ± SEM, were analysed by two-way ANOVA with Tukey’s correction for multiple comparisons. Panel (A-C) represent mean ± SEM, were analysed by one-way ANOVA with Tukey’s correction for multiple comparisons.
Extended Data Fig. 5 IPA increases mitochondrial respiration in a Cpt1a dependent, but OPA1-independent manner.
(A) Seahorse Mitostress assay curve and related OCR parameter analysis showing the effect of IPA on HELA cells cultures with DMSO or IPA (10, 100 μM) in 6 h. n = 6 for Con group, n = 8 for the other 3 IPA-treated groups. (B) Seahorse Mitostress assay curve and related OCR parameter analysis showing the effect of IPA on WT or Opa1KO MEF cells cultured with DMSO or IPA (10, 100 μM) in 6 h. n = 5 for WT-Con, n = 6 for Opa1KO-Con, n = 8 for the other groups. (C) Mitochondria images and Mitomass, mitochondria membrane potential analysis of WT or Opa1KO MEF cells cultured with DMSO or 10 μM IPA in 24 h. n = 8. (D, E) Figure showing the effect of IPA on isolated cardiac mitochondria. n = 3. (F) Cpt1a siRNA workflow in MEF cells. Created with BioRender.com. (G) Cpt1a expression in mRNA level in Con, siRNA Cpt1a, mock and non-target negative control groups. n = 3. (H, I) Seahorse Mitostress assay analysis of MEF cells cultured in the presence of DMSO or IPA (10, 100 μM) in WT and siRNA Cpt1a groups at 6 h. n = 6. All the panels except immunofluorescence images in panel (C) and siRNA workflow in panel (F) represent mean ± SEM. (A) were analysed by two-way ANOVA with Tukey’s correction for multiple comparisons. (B), (G) and (I) were analysed by one-way ANOVA with Tukey’s correction for multiple comparisons. Panel (D), (E) were analysed by multiple unpaired t-test.
Extended Data Fig. 6 The effect of IPA on CD4 + T cell energy metabolism is mediated by the PPARβ/δ nuclear receptor, not the canonical AhR signalling pathway.
(A) Schematic diagram showing the principle of HepG2-Lucia AhR reporter cell line. Created with BioRender.com. (B) Heatmap showing AhR activity induced by different Tryptophan metabolites. Ficz: 10 ng/mL. n = 3. (C) Seahorse Mitostress assay curve of WT and AhR−/− CD4 + T cells in basal and CD3/CD28 activation state in the presence of DMSO and Ficz (10 ng/mL). (D) Basal and maximal respiration analysis of the condition above. For Panel (C) and (D), n = 6 for Con and CD3/CD28 group; n = 4 for Ficz and CD3/CD28+Ficz group. (E) Seahorse Mitostress assay curve of WT and AhR−/− CD4 + T cells in basal and CD3/CD28 activation state in the presence of DMSO and IPA (100, 1000 μM). (F) Basal and maximal respiration analysis of the condition above. For Panel (E) and (F), n = 6 for each except n = 4 for IPA100 and IPA1000 groups. (G) GSEA of proteomics showing that Glycolysis and MYC pathway were enriched in CD3/CD28 groups compared with CD3/CD28 + IPA1000 group. Label CD3: CD3/CD28 group; CIPA2: CD3/CD28 + IPA1000 group. (H) Seahorse Mitostress assay of CD4 + T cells cultured with/without CD3/CD28 activation in the presence of DMSO and GW501516 (5 μM). The curves showed basal condition and in the presence of the inhibitor GSK3787 in sequence in 6 h. (I) Basal and maximal respiration of CD4 + T cells generated from Mitostress assay treated with/without GW501516 in basal and CD3/CD28 activation state in the absence/presence of GSK3787 inhibitor. For Panel (H) and (I): n = 6 for Con and CD3/CD28 group; n = 4 for the other two. (J) Seahorse Mitostress assay of CD4 + T cells cultured with/without CD3/CD28 activation in the presence of DMSO and GW7647 (1 μM). The urves showed basal condition and in the presence of the inhibitor GW6471 in sequence in 6 h. (K) Basal and maximal respiration of CD4 + T cells generated from Mitostress assay treated with/without GW7647 in basal and CD3/CD28 activation state in the absence/presence of GW6471 inhibitor. For Panel (J) and (K): n = 6 for Con and CD3/CD28 group; n = 4 for the other two. All the panels except (A), (B) and (G) represent mean ± SEM. (D), (F), (I) and (K) were analysed by two-way ANOVA with multiple comparisons.
Extended Data Fig. 7 IPA induces a durable inhibitory effect on TH1 and TH17 cell reprogramming that persists upon secondary challenge.
(A) Schematic diagram showing the protocol of mouse CD4 + T cell differentiation. Created with BioRender.com. (B-C) Flow cytometry and its quantification of CD4 + T cells stained intracellularly for T-bet and IFN-γ. n = 3. (D-E) Flow cytometry and its quantification of CD4 + T cells stained intracellularly for RORγt and IFN-γ to detect TH17 phenotype. n = 3. (F) Flow cytometry and its quantification of T-bet in TH1 and RORγt in TH17 populations respectively. n = 6 for each for T-bet marker; n = 6 for Con and n = 3 for IPA100 and IPA1000 group for RORγt marker. (G) Schematic diagram of IPA ‘washout’ experiment. Created with BioRender.com. (H, I) Flow cytometry quantitative analysis of TH1 and TH17 proportion in T24 and T48 time points in two circumstances: IPA persisted in the culture medium during the whole process while for the other group, IPA was washed out after 24 h of incubation. n = 3. (J) Flow cytometry and its quantification of naïve CD4 + T cells stained intracellularly for IFN-γ and IL-4. In one group, IPA was applied continuously throughout the entire process, while in the other group, IPA was only added during the last 5 h of restimulation. n = 3 for Con and n = 4 for the other groups. (K) Flow cytometry and its quantification of naïve CD4 + T cells stained intracellularly for IL-17A and IFN-γ. In one group, IPA was applied continuously throughout the entire process, while in the other group, IPA was only added during the last 5 h of restimulation. n = 3 for each in the whole process condition; n = 4 for each upon restimulation. All the panels except panel (A) and (G) represent mean ± SEM. Data in panel (B-F) were analysed by one-way ANOVA with Tukey’s correction for multiple comparisons. Panel (H-K) were analysed by two-way ANOVA with Tukey’s correction for multiple comparisons.
Extended Data Fig. 8 IPA changes the phenotype of mouse CD4 + T cells through rewiring the energy metabolism and PPARβ/δ nuclear receptor.
(A-B) Schematic diagram showing the intervention methods of different metabolic pathways and the experimental procedure. Panel (A) and (B) were Created with BioRender.com. (C) IFN-γ + TH1 percentage of CD4 + T cells in the presence of various doses of oligomycin from 1 μM to 0.1 nM. n = 3. (D) Figure showing the effect of IPA of 100, 1000 μM on TH1 phenotype in baseline and in the presence of various doses of oligomycin from 1 μM to 0.1 nM. n = 3. (E) IL-17A + TH17 percentage of CD4 + T cells in the presence of various doses of oligomycin from 10 nM to 0.1 nM. n = 3. (F) Figure showing the effect of IPA of 100, 1000 μM on TH17 phenotype in baseline and in the presence of various doses of oligomycin from 10 nM to 0.1 nM. n = 3. (G) Flow cytometry analysis of IFN-γ MFI. CD4 + T cells were cultured in TH1 differentiation condition with DMSO or IPA (100, 1000 μM) in baseline and in the presence of specific metabolic pathway inhibitors including oligomycin, etomoxir, TOFA and BPTES. n = 3. (H) Flow cytometry analysis of IL-17A MFI. CD4 + T cells were cultured in TH17 differentiation condition with DMSO or IPA (100, 1000 μM) in baseline and in the presence of specific metabolic pathway inhibitors as mentioned above. n = 3. (I-J) Flow cytometry and its quantification of CD4 + T cells stained intracellularly IFN-γ and IL-4 to detect TH1. CD4 + T cells were cultured in TH1 differentiation condition with DMSO and GW501516 (5, 50 μM). n = 3. (K) Flow cytometry data showing the effect of IPA on IFN-γ + TH1 percentage in the absence/presence of GSK3787. n = 6 and n = 3 for Con group in basal and GSK3787 conditions respectively; n = 3 for IPA100 and IPA1000 in the presence/absence of GSK3787. (L-M) Flow cytometry and its quantification of CD4 + T cells stained intracellularly IL-17A and IFN-γ to detect TH17. CD4 + T cells were cultured in TH17 differentiation condition with DMSO and GW501516 (5, 50 μM). n = 6 for Con and n = 3 for other two doses of GW501516 groups. (N) Flow cytometry data showing the effect of IPA on IL-17A + TH17 percentage in the absence/presence of GSK3787. n = 6 for Con group in the presence/absence of GSK3787; n = 3 and n = 6 for the two doses of IPA in the presence/absence of GSK3787 respectively. Panel (C), (E), (J), (M) represent mean ± SEM analysed by one-way ANOVA with Tukey’s correction for multiple comparisons. Panel (D), (F), (G), (H), (K), (N) represent mean ± SEM analysed by two-way ANOVA with Tukey’s correction for multiple comparisons.
Extended Data Fig. 9 The regulatory effect of IPA on small intestinal lamina propria cells is less pronounced than that of colon.
(A) The total cell number of LP of small intestine and colon in WT, ABX and ABX + IPA mice. n = 7 for WT and n = 9 for ABX and ABX + IPA groups. (B) The proportion of each cell population including CD4 + , CD8 + , DN, DP subsets under the TCRαβ+ cell gate of LP in small intestine in WT, ABX and ABX + IPA mice. n = 8 for WT, n = 10 for ABX and n = 9 for ABX + IPA. (C) ATP synthesis in WT, ABX and ABX + IPA mice of γδ T cell and B cell subsets in colonic LP. For γδ T cell: n = 8 for WT, n = 10 for ABX and n = 9 for ABX + IPA. For B cell: n = 13 for WT, n = 15 for ABX and ABX + IPA groups. (D) ATP synthesis in WT, ABX and ABX + IPA group of CD4 + , CD8 + , γδ T cell and B cell subsets in LP of small intestine. For CD4 + , CD8 + T cell and B cell: n = 13 for WT, n = 16 for ABX and ABX + IPA groups. For γδ T cell: n = 7 for WT, n = 10 for ABX and n = 9 for ABX + IPA groups. (E) Metabolic profile of B cell of LP in small intestine and colon in WT, ABX and ABX + IPA mice. For colon: n = 13 for WT, n = 12 for ABX and ABX + IPA groups. For SI: n = 10 for WT, n = 12 for ABX and ABX + IPA groups. (F) Metabolic profile of γδ T cell of LP in small intestine and colon in WT, ABX and ABX + IPA mice. For colon: n = 7 for WT and ABX + IPA groups, n = 9 for ABX groups. For SI: n = 3 for WT, n = 6 for ABX and ABX + IPA groups. (A–D) represent mean ± SEM analysed by one-way ANOVA with Tukey’s correction for multiple comparisons. (E) and (F) represent mean ± SEM analysed by two-way ANOVA with Tukey’s correction for multiple comparisons. Results for the colon are solid columns, and for the small intestine are striped columns with diagonal lines.
Extended Data Fig. 10 The protective effect of IPA against DSS colitis is dependent on CD4 + T cell TH1 and TH17 metabolic reprogramming without altering Treg phenotype in vivo.
(A) Table showing the clinical data of healthy controls and IBD patient cohort. (B) Schematic diagram showing the protocol of DSS colitis experiment treated with or without IPA. IPA (200 mg/kg) or corresponding vehicle control were administered to the mice daily by oral gavage. Created with BioRender.com. (C) Metabolic profile of colonic CD8 + T cell, B cell and γδ T cell of control (Con), DSS and DSS + IPA mice. n = 7 for Con, n = 9 for DSS, n = 6 for DSS + IPA. (D-E) Flow cytometry and its quantification of colonic CD4 + T cells stained intracellularly for FOXP3. n = 5 for Con, n = 6 for DSS and DSS + IPA groups. (F-G) Body weight curve and DAI scoring during the whole process of DSS model in Rag2-/- mice. n = 5 for Con, n = 6 for DSS and DSS + IPA groups. (H-J) Colon length, histology score and H & E staining of colon tissue at D7 in Con, DSS and DSS + IPA Rag2-/- mice. n = 5 for Con and DSS; n = 6 for DSS + IPA groups for colon length comparison. For H & E histology, n = 4 for Con and DSS; n = 3 for DSS + IPA groups were randomly selected. Scale bar: 200 μm. Panel (E), (H), (I) represent mean ± SEM analysed by one-way ANOVA with Tukey’s correction for multiple comparisons. Panel (C), (F), (G) represent mean ± SEM analysed by two-way ANOVA with Tukey’s correction for multiple comparisons.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1 showing all the metabolites involved in metabolite screening in Jurkat cells. Supplementary Figs. 1 and 2, FACS gating strategy; R code availability; oligonucleotide sequences.
Supplementary Data (download XLSX )
Supplementary source data of Supplementary Figs. 1 and 2.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Q., de Oliveira Formiga, R., Puchois, V. et al. Microbial metabolite indole-3-propionic acid drives mitochondrial respiration in CD4+ T cells to confer protection against intestinal inflammation. Nat Metab 7, 2510–2530 (2025). https://doi.org/10.1038/s42255-025-01396-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01396-6
This article is cited by
-
L-kynurenine reshapes immune microenvironment to alleviate methamphetamine-induced chronic lung injury through gut-lung axis
Microbiome (2026)
-
The gut microbiome in solid-organ and haematopoietic-stem-cell transplantation
Nature Reviews Microbiology (2026)
-
IPA brews metabolic balance in gut immunity
Nature Metabolism (2025)