Abstract
The hypothalamus regulates feeding and metabolic balance in response to metabolic cues. Here we report that extracellular vesicles (EVs) are secreted from the mediobasal hypothalamus in a diurnal manner that is influenced by daily feeding. Sox2-positive tanycytes have a critical role in maintaining the diurnal pattern of hypothalamic EV release. Inhibition of tanycyte EV release leads to a loss of feeding diurnality, weight control and blood glucose homoeostasis, whereas supplementation with tanycytic EVs confers metabolic benefits. We show that a subset of tanycytic EVs carries surface prepro-insulin (ppIns), which mediates recognition and uptake by insulin-receptor-positive hypothalamic neurons. These EVs are loaded with mTORC components, including Rictor in a low-phosphorylation state, and support hypothalamic neuronal signalling. Both ppIns and Rictor are important for the EV-mediated preservation of feeding rhythmicity and resistance to diet-induced metabolic dysfunction. Collectively, these findings identify tanycyte-derived EVs as regulators of feeding diurnality through insulin precursor-directed targeting and delivery of mTORC components to hypothalamic neurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Source data supporting the findings in this study are provided with this paper. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with dataset identifier PXD072591. All other materials and information generated in this study are available upon request. This study did not involve generating any original code. Source data are provided with this paper.
References
Jordan, S. D., Konner, A. C. & Bruning, J. C. Sensing the fuels: glucose and lipid signaling in the CNS controlling energy homeostasis. Cell. Mol. Life Sci. 67, 3255–3273 (2010).
Lam, T. K. Neuronal regulation of homeostasis by nutrient sensing. Nat. Med. 16, 392–395 (2010).
Yue, J. T. & Lam, T. K. Lipid sensing and insulin resistance in the brain. Cell Metab. 15, 646–655 (2012).
Zeltser, L. M., Seeley, R. J. & Tschop, M. H. Synaptic plasticity in neuronal circuits regulating energy balance. Nat. Neurosci. 15, 1336–1342 (2012).
Cota, D. et al. Hypothalamic mTOR signaling regulates food intake. Science 312, 927–930 (2006).
Bruning, J. C. & Fenselau, H. Integrative neurocircuits that control metabolism and food intake. Science 381, eabl7398 (2023).
Myers, M. G. Jr., Affinati, A. H., Richardson, N. & Schwartz, M. W. Central nervous system regulation of organismal energy and glucose homeostasis. Nat. Metab. 3, 737–750 (2021).
Cone, R. D. Anatomy and regulation of the central melanocortin system. Nat. Neurosci. 8, 571–578 (2005).
Herrera Moro Chao, D. et al. Hypothalamic astrocytes control systemic glucose metabolism and energy balance. Cell Metab. 34, 1532–1547.e6 (2022).
Zhang, Y., Reichel, J. M., Han, C., Zuniga-Hertz, J. P. & Cai, D. Astrocytic process plasticity and IKKβ /NF-κB in central control of blood glucose, blood pressure, and body weight. Cell Metab. 25, 1091–1102.e4 (2017).
Kim, J. G. et al. Leptin signaling in astrocytes regulates hypothalamic neuronal circuits and feeding. Nat. Neurosci. 17, 908–910 (2014).
Garcia-Caceres, C. et al. Astrocytic insulin signaling couples brain glucose uptake with nutrient availability. Cell 166, 867–880 (2016).
Garcia-Caceres, C. et al. Role of astrocytes, microglia, and tanycytes in brain control of systemic metabolism. Nat. Neurosci. 22, 7–14 (2019).
Ardanaz, C. G. et al. Astrocytic GLUT1 reduction paradoxically improves central and peripheral glucose homeostasis. Sci. Adv. 10, eadp1115 (2024).
Varela, L. et al. Astrocytic lipid metabolism determines susceptibility to diet-induced obesity. Sci. Adv. 7, eabj2814 (2021).
Zhang, Y. L. et al. Hypothalamic stem cells control ageing speed partly through exosomal miRNAs. Nature 548, 52–57 (2017).
Li, J. X., Tang, Y. Z. & Cai, D. S. IKKβ/NF-κB disrupts adult hypothalamic neural stem cells to mediate a neurodegenerative mechanism of dietary obesity and pre-diabetes. Nat. Cell Biol. 14, 999–1012 (2012).
Xiao, Y. Z. et al. Reducing hypothalamic stem cell senescence protects against aging-associated physiological decline. Cell. Metab. 31, 534–548.e5 (2020).
Yu, B., Ikhlas, S., Ruan, C., Zhong, X. & Cai, D. Innate and adaptive immunity of murine neural stem cell-derived piRNA exosomes/microvesicles against pseudotyped SARS-CoV-2 and HIV-based lentivirus. iScience 23, 101806 (2020).
Ikhlas, S., Usman, A., Kim, D. & Cai, D. Exosomes/microvesicles target SARS-CoV-2 via innate and RNA-induced immunity with PIWI-piRNA system. Life Sci. Alliance https://doi.org/10.26508/lsa.202101240 (2021).
Jung, H. G. et al. Control of aging-associated neurodegeneration via hypothalamic extracellular vesicles containing parathymosin. Cell Rep. 44, 116561 (2025).
Crescitelli, R., Lasser, C. & Lotvall, J. Isolation and characterization of extracellular vesicle subpopulations from tissues. Nat. Protoc. 16, 1548–1580 (2021).
Gomes, P. A. et al. A novel isolation method for spontaneously released extracellular vesicles from brain tissue and its implications for stress-driven brain pathology. Cell Commun. Signal. 21, 35 (2023).
Lee, J. C., Ray, R. M. & Scott, T. A. Prospects and challenges of tissue-derived extracellular vesicles. Mol. Ther. 32, 2950–2978 (2024).
Terlecki-Zaniewicz, L. et al. Extracellular vesicles in human skin: cross-talk from senescent fibroblasts to keratinocytes by miRNAs. J. Invest. Dermatol. 139, 2425–2436.e5 (2019).
Stangler, L. A. et al. Microdialysis and microperfusion electrodes in neurologic disease monitoring. Fluids Barriers CNS 18, 52 (2021).
Kano, M. et al. Tanycyte-like cells derived from mouse embryonic stem culture show hypothalamic neural stem/progenitor cell functions. Endocrinology 160, 1701–1718 (2019).
Hendrickson, M. L., Zutshi, I., Wield, A. & Kalil, R. E. Nestin expression and in vivo proliferative potential of tanycytes and ependymal cells lining the walls of the third ventricle in the adult rat brain. Eur. J. Neurosci. 47, 284–293 (2018).
Yoo, S. et al. Control of neurogenic competence in mammalian hypothalamic tanycytes. Sci. Adv. 7, eabg3777 (2021).
Robins, S. C. et al. α-Tanycytes of the adult hypothalamic third ventricle include distinct populations of FGF-responsive neural progenitors. Nat. Commun. 4, 2049 (2013).
Balland, E. et al. Hypothalamic tanycytes are an ERK-gated conduit for leptin into the brain. Cell Metab. 19, 293–301 (2014).
Guillebaud, F. et al. Glial endozepines reverse high-fat diet-induced obesity by enhancing hypothalamic response to peripheral leptin. Mol. Neurobiol. 57, 3307–3333 (2020).
Harrison, L. et al. Fluorescent blood–brain barrier tracing shows intact leptin transport in obese mice. Int. J. Obes. (Lond.) 43, 1305–1318 (2019).
Smith, A., MacAulay, B., Scheufen, J., Hudak, A. & Abizaid, A. Chronic social defeat stress increases brain permeability to ghrelin in male mice. eNeuro https://doi.org/10.1523/ENEURO.0093-24.2024 (2024)
Gomez, I. M. et al. Hypothalamic tanycytes internalizeghrelin from the cerebrospinal fluid: molecular mechanisms and functional implications. Mol. Metab. 90, 102046 (2024).
Nguyen, L. T. & Dodd, G. T. Hypothalamic neuronal-glial crosstalk in metabolic disease. npj Metab. Health. Dis. 2, 27 (2024).
Porniece Kumar, M. et al. Insulin signalling in tanycytes gates hypothalamic insulin uptake and regulation of AgRP neuron activity. Nat. Metab. 3, 1662–1679 (2021).
Tang, Y., Zuniga-Hertz, J. P., Han, C., Yu, B. & Cai, D. Multifaceted secretion of htNSC-derived hypothalamic islets induces survival and antidiabetic effect via peripheral implantation in mice. Elife https://doi.org/10.7554/eLife.52580 (2020).
Cai, D., Dhe-Paganon, S., Melendez, P. A., Lee, J. & Shoelson, S. E. Two new substrates in insulin signaling, IRS5/DOK4 and IRS6/DOK5. J. Biol. Chem. 278, 25323–25330 (2003).
Dalvi, P. S., Nazarians-Armavil, A., Tung, S. & Belsham, D. D. Immortalized neurons for the study of hypothalamic function. Am. J. Physiol. Regul. Integr. Comp. Physiol. 300, R1030–R1052 (2011).
Nazarians-Armavil, A., Chalmers, J. A., Lee, C. B., Ye, W. & Belsham, D. D. Cellular insulin resistance disrupts hypothalamic mHypoA-POMC/GFP neuronal signaling pathways. J. Endocrinol. 220, 13–24 (2014).
Nazarians-Armavil, A., Menchella, J. A. & Belsham, D. D. Cellular insulin resistance disrupts leptin-mediated control of neuronal signaling and transcription. Mol. Endocrinol. 27, 990–1003 (2013).
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Sjostedt, E. et al. An atlas of the protein-coding genes in the human, pig, and mouse brain. Science https://doi.org/10.1126/science.aay5947 (2020).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. https://doi.org/10.1126/sciadv.abh2169 (2021).
Hornbeck, P. V. et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 43, D512–D520 (2015).
Panwar, V. et al. Multifaceted role of mTOR (mammalian target of rapamycin) signaling pathway in human health and disease. Signal Transduct. Target. Ther. 8, 375 (2023).
Alessi, D. R., Pearce, L. R. & Garcia-Martinez, J. M. New insights into mTOR signaling: mTORC2 and beyond. Sci. Signal. 2, pe27 (2009).
Battaglioni, S., Benjamin, D., Walchli, M., Maier, T. & Hall, M. N. mTOR substrate phosphorylation in growth control. Cell 185, 1814–1836 (2022).
Dibble, C. C., Asara, J. M. & Manning, B. D. Characterization of Rictor phosphorylation sites reveals direct regulation of mTOR complex 2 by S6K1. Mol. Cell. Biol. 29, 5657–5670 (2009).
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor–mTOR complex. Science 307, 1098–1101 (2005).
Wang, J. D., Shao, Y., Liu, D., Liu, N. Y. & Zhu, D. Y. Rictor/mTORC2 involves mitochondrial function in ES cells derived cardiomyocytes via mitochondrial Connexin 43. Acta Pharmacol. Sin. 42, 1790–1797 (2021).
Shiota, C., Woo, J. T., Lindner, J., Shelton, K. D. & Magnuson, M. A. Multiallelic disruption of the rictor gene in mice reveals that mTOR complex 2 is essential for fetal growth and viability. Dev. Cell 11, 583–589 (2006).
Hsu, P. P. et al. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science 332, 1317–1322 (2011).
Sarbassov, D. D. et al. Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr. Biol. 14, 1296–1302 (2004).
Julien, L. A., Carriere, A., Moreau, J. & Roux, P. P. mTORC1-activated S6K1 phosphorylates Rictor on threonine 1135 and regulates mTORC2 signaling. Mol. Cell. Biol. 30, 908–921 (2010).
Parker, B. L. et al. Targeted phosphoproteomics of insulin signaling using data-independent acquisition mass spectrometry. Sci. Signal. 8, rs6 (2015).
Fu, W. & Hall, M. N. Regulation of mTORC2 signaling. Genes (Basel) https://doi.org/10.3390/genes11091045 (2020).
Liu, P. et al. Sin1 phosphorylation impairs mTORC2 complex integrity and inhibits downstream Akt signalling to suppress tumorigenesis. Nat. Cell Biol. 15, 1340–1350 (2013).
Guertin, D. A. et al. Ablation in mic of the mTORC components raptor, rictor, or mLST8 reveals that mTORC2 is required for signaling to Akt-FOXO and PKCα, but not S6K1. Dev. Cell 11, 859–871 (2006).
Treins, C., Warne, P. H., Magnuson, M. A., Pende, M. & Downward, J. Rictor is a novel target of p70 S6 kinase-1. Oncogene 29, 1003–1016 (2010).
Kocalis, H. E. et al. Rictor/mTORC2 facilitates central regulation of energy and glucose homeostasis. Mol. Metab. 3, 394–407 (2014).
Saucisse, N. et al. Functional heterogeneity of POMC neurons relies on mTORC1 signaling. Cell Rep. 37, 109800 (2021).
Chellappa, K. et al. Hypothalamic mTORC2 is essential for metabolic health and longevity. Aging Cell 18, e13014 (2019).
Wrobel, L. et al. mTORC2 Assembly is regulated by USP9X-mediated deubiquitination of RICTOR. Cell Rep. 33, 108564 (2020).
Oh, W. J. & Jacinto, E. mTOR complex 2 signaling and functions. Cell Cycle 10, 2305–2316 (2011).
Rhea, E. M., Rask-Madsen, C. & Banks, W. A. Insulin transport across the blood–brain barrier can occur independently of the insulin receptor. J. Physiol. 596, 4753–4765 (2018).
Kuwabara, T. et al. Insulin biosynthesis in neuronal progenitors derived from adult hippocampus and the olfactory bulb. EMBO Mol. Med. 3, 742–754 (2011).
Mehran, A. E. et al. Hyperinsulinemia drives diet-induced obesity independently of brain insulin production. Cell Metab. 16, 723–737 (2012).
Mazucanti, C. H. et al. Release of insulin produced by the choroid plexis is regulated by serotonergic signaling. JCI Insight 4, https://doi.org/10.1172/jci.insight.131682 (2019).
Hills, C. E. & Brunskill, N. J. C-Peptide and its intracellular signaling. Rev. Diabet. Stud. 6, 138–147 (2009).
Hills, C. E. & Brunskill, N. J. Cellular and physiological effects of C-peptide. Clin. Sci. (Lond.) 116, 565–574 (2009).
Liu, M. et al. Impaired cleavage of preproinsulin signal peptide linked to autosomal-dominant diabetes. Diabetes 61, 828–837 (2012).
Unnerstale, S. & Maler, L. pH-Dependent Interaction between C-peptide and phospholipid bicelles. J. Biophys. 2012, 185907 (2012).
Prevot, V., Cornea, A., Mungenast, A., Smiley, G. & Ojeda, S. R. Activation of erbB-1 signaling in tanycytes of the median eminence stimulates transforming growth factor β1 release via prostaglandin E2 production and induces cell plasticity. J. Neurosci. 23, 10622–10632 (2003).
Lhomme, T. et al. Tanycytic networks mediate energy balance by feeding lactate to glucose-insensitive POMC neurons. J. Clin. Invest. 131, e140521 (2021).
Kim, M. S. et al. Rapid linkage of innate immunological signals to adaptive immunity by the brain–fat axis. Nat. Immunol. 16, 525–533 (2015).
Zhang, G. et al. Hypothalamic programming of systemic ageing involving IKK-β, NF-κB and GnRH. Nature 497, 211–216 (2013).
Zhang, G. et al. Neuropeptide exocytosis involving synaptotagmin-4 and oxytocin in hypothalamic programming of body weight and energy balance. Neuron 69, 523–535 (2011).
Zhang, Y. et al. Hypothalamic stem cells control ageing speed partly through exosomal miRNAs. Nature 548, 52–57 (2017).
Welsh, J. A. et al. MIFlowCyt-EV: a framework for standardized reporting of extracellular vesicle flow cytometry experiments. J. Extracell. Vesicles 9, 1713526 (2020).
van der Vlist, E. J., Nolte-‘t Hoen, E. N., Stoorvogel, W., Arkesteijn, G. J. & Wauben, M. H. Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat. Protoc. 7, 1311–1326 (2012).
O’Donoghue, L. & Smolenski, A. Analysis of protein phosphorylation using Phos-tag gels. J. Proteomics 259, 104558 (2022).
Seo, G. & Wang, W. Characterization of ATG8-family protein phosphorylation by Phos-tag gel for autophagy study. STAR Protoc 5, 102872 (2024).
Kinoshita-Kikuta, E. et al. Dynamic equilibrium of protein phosphorylation by kinases and phosphatases visualized by Phos-tag SDS–PAGE. Kinases Phosphatases 2, 224–239 (2024).
Acknowledgements
We thank the Animal Physiology Core, the Histology Core, the Flow Cytometry Core, the Proteomics Core and the Analytical Imaging Core (with equipment support from S10OD034397-01 and P30CA013330 for confocal microscope) of the Albert Einstein College of Medicine. We also thank the Extracellular Vesicle Core of the University of Pennsylvania for service, and thank Brookhaven National Laboratory for assistance with the nano-particle tracking instrument. We thank S. Sidoli for assisting with proteomics and thank other members of the Cai laboratory for general assistance. This research was supported through Einstein institutional resources and partly through a Milky Way Research Foundation award and National Institutes of Health grants DK121435, AG031774 and HL147477 R01AG031774 (all supports to D.C.).
Author information
Authors and Affiliations
Contributions
Y.C. performed the majority of experiments, including animal models, animal phenotyping, therapeutics, histology, various EV assays, EV characterization and signalling studies, contributed to data analysis, generated figures and assisted with manuscript writing. M.W.K. contributed to the generation of the EV models and EV characterization, assisted with signalling studies and contributed to discussions. G.G. contributed to the evaluation of EV models and discussions. D.C. conceived the hypothesis, constructed the project, designed research strategies and approaches, guided and supervised experimentation, instructed and finalized data analysis, interpreted the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt, in collaboration with the Nature Metabolism team
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quantitative comparison of EVs among brain regions.
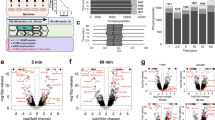
Various brain regions were dissected from adult male C57BL/6 mice during the light/rest phase to release EVs under ex vivo condition for 6 hours, and the released EVs into the medium were profiled for EVs via NTA. Curves, representative NTA; bar graphs, total EVs of each biological sample and each group. Statistics: one-way ANOVA and Tukey post-hoc (right); n = 4 mice per group; data reflect mean ± SEM.
Extended Data Fig. 2 Validation of Rab27a KD model efficiency.
(a–c) Tanycyte immunostaining in Rab27a KD and control (Con) in Figure were quantified for Rab27a in the middle part of mediobasal hypothalamus (a), Sox2-positive cells along the 3V (b), and Nestin in the middle part of mediobasal hypothalamus (c). (d) Mediobasal hypothalamic sections (containing 3V tanycytes) from Rab27a KD and Con were immunostained for cleaved caspase 3. Scale bars, 100 μm. Statistics: two-tailed unpaired t-test; n = 3 mice per group (a, b), n = 4 mice per group (c); data reflect mean ± SEM
Extended Data Fig. 3 Leptin-induced STAT3 signaling in Rab27a KD model.
Overnight-fasted Rab27a KD mice and matched control (Con) received a systemic administration of leptin at a dose of 2 mg/kg body weight or vehicle via intraperitoneal injection. At 45 minutes later, animals were transcardially perfused, and brain sections across the mediobasal hypothalamus were generated and immunostained for phosphorylated STAT3 (pSTAT3). DAPI revealed nuclei of all cells. Scale bar, 100 μm. Immunostaining signals were quantified and analyzed in the bar graph. Statistics: two-way ANOVA with Tukey post-hoc; n = 3 mice per group; data reflect mean ± SEM.
Extended Data Fig. 4 Additional metabolic assessments for male Rab27a KD model.
(a) Male Rab27a KD mice and matched Con mice maintained on normal chow were assessed for energy expenditure (EE) without fasting in metabolic cages. EE values were shown per animal without body weight adjustment. (b–d) After the development of feeding disorder, weight gain, and hyperglycemia under chow feeding, male Rab27a KD subjected to a 10-day rescue experiment in which animals received daily intranasal administration of tanycytic EVs (1 µg) vs. vehicle. Same treatment also applied to Con group for additional information. These animals during the treatment were monitored for food intake (b) and body weight (c) and followed by fasting blood glucose (d) at the end of this treatment. Statistics: two-tailed unpaired t-test (a); two-way repeated measures ANOVA/Tukey post-hoc (b, c), and one-way ANOVA/Tukey post-hoc (d). n = 6 mice per group (a–d); data reflect mean ± SEM.
Extended Data Fig. 5 Composition assessment of Rab27a KD EVs versus control EVs.
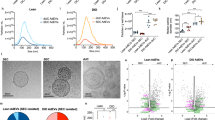
EVs from tanycytes with Rab27a KD vs. matched control (Con) were purified and analyzed by Western blot (a) and proteomics (b). Proteomic datasets were analyzed for protein-level differences based on uncorrected p values and corrected false discovery rate (FDR) q values. The threshold for differential expression levels was set at a 2-fold change based on standard variation criteria. Statistics: two-tailed unpaired t-test; n = 3 biological samples per group.
Extended Data Fig. 6 Additional tanycyte profiling in female Rab27a KD model.
Mediobasal hypothalamus containing tanycytes along the 3V from Rab27a KD and control (Con) were sectioned and immunostained for additional evaluations including tanycyte morphology through Nestin immunostaining (a), apoptotic biomarker through immunostaining of cleaved caspase 3 (b), and transcytosis of Alexa 594-labelled leptin (2 mg/kg body weight) following a systemic administration through an intraperitoneal injection (c). DAPI staining (blue) was merged to show the background of sections. Scale bar, 100 μm.
Extended Data Fig. 7 Additional metabolic assessments for female Rab27a KD model.
(a) Female Rab27a KD mice and matched control (Con) maintained on normal chow were assessed for energy expenditure (EE) without fasting. EE values were calculated per animal without body weight adjustment. (b–d) After the development of feeding disorder, weight gain, and hyperglycemia under chow feeding, female Rab27a KD and Con underwent a 10-day treatment in which animals received daily intranasal administration of tanycytic EVs (1 µg) vs. vehicle. Metabolic phenotypes were monitored for food intake (b) and body weight (c) followed by fasting blood glucose measurement (d). Statistics: two-tailed unpaired t-test (a); two-way repeated measures ANOVA/Tukey post-hoc (b, c), and one-way ANOVA with Tukey post-hoc (d); n = 6 mice per group (a–d); data represent mean ± SEM.
Extended Data Fig. 8 Full-length mRNA of ppIns in tanycyte model.
Full-length mRNA encoding ppIns was amplified by RT-PCR from the in vitro tanycyte model of this project and was then sequenced by Sanger’s sequencing.
Extended Data Fig. 9 Proteomics of EVs deficient of ppIns or Rictor versus control.
EVs from tanycytes with ppIns KD samples vs. matched shRNA-based Control (Con) samples, or Rictor KD samples vs. matched gRNA-based Con samples (the same control in Extended Data 5), were subjected to proteomics (a, b) and measurement of total small RNAs (c). Statistical analysis of protein-level differences was then analyzed based on uncorrected p values and corrected FDR q values (a, b). The threshold for differential expression levels was set at a 2-fold change based on standard variation criteria. Statistics: two-tailed unpaired t-test; n = 3 biological samples per group (a–c); data in bar graphs reflect mean ± SEM.
Extended Data Fig. 10 Conceptual diagram of this study.
a. Tanycytes secrete EVs in a diurnal manner, with EV release rising toward the onset of the rest/sleep phase over the 24-hour cycle. This increase reflects the cumulative effects of multiple meals during the 12-hour active phase. EV levels remain elevated during the early and middle portions of the rest/sleep phase and are further amplified by circadian clock mechanisms, before declining as the vesicles are gradually taken up and cleared. b. A subset of EVs released by these tanycytes display ppIns on their surface, enabling selective recognition and uptake by IR-positive neurons in the adjacent mediobasal hypothalamus. Through this mechanism, these EVs deliver cargos such as mTORC component(s) to target neurons, thereby sustaining mTORC-dependent satiety signaling during the rest/sleep phase, when anorexigenic hormones such as insulin are low. Taken together, in contrast to hormones such as insulin, which act over short time windows (2–3 hours), this EV-based mechanism provides a 12-hour mode of regulating satiety, offering a new and critical perspective on the control of body weight and metabolic balance. Graphics in this figure were generated through BioRender software with institution purchased license.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7 and Uncropped scans 1.
Supplementary Data 1 (download XLSX )
Source data for supplementary figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 6 (download PDF )
Uncropped western blot gel images for Fig. 5.
Source Data Fig. 7 (download XLSX )
Statistical source data for Fig. 6.
Source Data Fig. 8 (download PDF )
Uncropped western blot gel images for Fig. 6.
Source Data Fig. 9 (download XLSX )
Statistical source data for Fig. 7.
Source Data Fig. 10 (download PDF )
Uncropped western blot gel images for Fig. 5.
Source Data Fig. 11 (download XLSX )
Statistical source data for Fig. 8.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 5 (download PDF )
Uncropped western blot gel images for Extended Data Fig. 5.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data for Extended Data Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Choi, Y., Kim, M.W., Go, G. et al. Metabolic regulation by tanycyte-derived extracellular vesicles through insulin precursor-mediated neuronal recognition and mTORC component delivery. Nat Metab 8, 666–684 (2026). https://doi.org/10.1038/s42255-026-01474-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-026-01474-3