Abstract
The heart uses a muscle-specific ribosome in cardiomyocytes, where the ribosomal protein RPL3 is replaced by its paralog RPL3L. Rare biallelic RPL3L mutations cause fatal neonatal dilated cardiomyopathy, yet the mechanisms that link genotype to heart failure are unclear. Despite the recessive inheritance pattern in humans, Rpl3l knockout mice show no overt cardiac phenotype, probably because of compensatory RPL3 upregulation through unknown mechanisms. Here we report four additional cases and propose a unifying pathogenetic model by integrating human genetics, patient tissues and isogenic cell models. Affected individuals typically carry one of two recurrent hotspot missense variants paired with a private allele. Whereas non-hotspot variants phenocopy knockout and allow RPL3 compensation, hotspot variants induce nucleolar protein aggregation, disrupt rRNA processing and block compensation by preserving the role of RPL3L in repressing RPL3 via unproductive splicing. These findings establish combined loss-of-function and gain-of-function mechanisms for RPL3L-associated cardiomyopathy and inform genetic screening, diagnosis and therapeutic development.
Similar content being viewed by others
Main
One of the most fundamental yet least understood adaptations in the heart is that cardiomyocytes use a specialized ribosome to decode genetic information and synthesize proteins1,2. During vertebrate heart development, the ubiquitous ribosomal protein L3 (RPL3)—a gatekeeper of the ribosome decoding center3 and the protein closest to the ribosome RNA-based active site4—is gradually replaced by its paralog, RPL3-like (RPL3L)5. RPL3L shares 78% amino acid identity with RPL3 and eventually becomes the dominant form in adult human left ventricular cardiomyocytes (95%)6, to a lesser extent in skeletal muscle (60%) and it is barely detectable in non-muscle cells7. Interestingly, this switch is reversed during adult muscle hypertrophy after injury or atrophy2,5,8. The physiological function and regulatory mechanism of this highly conserved and tightly controlled ribosomal switching remains poorly understood, highlighting a crucial gap in our understanding of cardiac biology at the most fundamental level.
We9 and others10,11,12,13,14 recently identified a subtype of dilated cardiomyopathy (DCM) associated with biallelic RPL3L mutations, now classified as dilated cardiomyopathy-2D (CMD2D; OMIM no. 619371). Among seven affected families, individuals with mutations in both RPL3L alleles consistently develop severe neonatal DCM, often resulting in fatal heart failure9,10,11,12,13,14. By contrast, heterozygous carriers in the same family remain unaffected. Although the autosomal recessive inheritance pattern suggests a loss-of-function (LOF) of these variants and RPL3L deficiency as the likely cause of CMD2D, Rpl3l-knockout mice generated by several labs display only mild cardiac dysfunction5,8,15,16. Notably, in all Rpl3l-knockout mouse models, there is a consistent upregulation of the ubiquitous paralog RPL3. The molecular mechanism according to which RPL3 is upregulated in Rpl3l-knockout mice remains unclear. Whether a lack of RPL3 compensation in human underlie the severe DCM symptoms remains unexplored.
In this study, we present four additional cases of RPL3L-linked CMD2D and uncover a unifying pathogenetic mechanism through integrated analyses of population genetic data, patient cardiac tissue and isogenic cells expressing various RPL3L variants.
Additional cases highlight hotspot variants as potential drivers
We identified four additional unrelated cases of CMD2D (case reports in the Supplementary Note), increasing the total number of affected families to eleven (see the clinical and molecular characteristics of all patients in Supplementary Table 1). Meta-analyses of all 11 cases highlight two prominent features: compound heterozygosity and hotspot missense variants. Nine of the eleven families harbor two distinct alleles in each case—that is, compound heterozygosity (lines in Fig. 1a). The two exceptions are a homozygous R116H case identified in a consanguineous population12 and a homozygous D308V case reported in this study (because of segmental uniparental disomy; Supplementary Note). Of the 14 variants, 10 are missense, including the previously unreported R58Q. The four remaining variants are putative LOF, including three truncating frameshift variants (T340Nfs*25 previously unreported, threonine to asparagine followed by 25 amino acids) and a splice donor variant (c.1167+1G>A) that causes exon 9 skipping (Δexon9)17. Although variants are distributed across nearly the entire RPL3L protein (Fig. 1a,b) and all affected amino acid positions are deeply conserved across vertebrates (Extended Data Fig. 1), two notable hotspots emerge: D308 (D308N or D308V), observed in six families (including all four cases reported in this study), and G27 (G27D), found in three additional families (Fig. 1a). Collectively, D308N/V and G27D account for 82% (9 out of 11) of affected families.
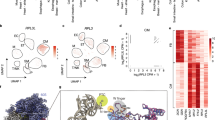
a, Schematic representation of neonatal DCM-associated variants across the RPL3L protein. Each circle denotes a missense variant, while rectangles represent frameshift variants or the splice variant (Δexon9). Variants are color-coded according to family, with lines connecting compound heterozygous variants identified in the same family. Variants identified in the four additional cases are highlighted with a star. The protein domains within RPL3L are labeled. The hotspot variants G27D and D308N/V are emphasized in bold. b, Mapping of the variants (in red) onto the AlphaFold-predicted three-dimensional structure of RPL3L (AF-Q92901-F1). The region encoded by exon 9, deleted in the splice variant (Δexon9) is highlighted in salmon. c, Clustering of RPL3L missense variants based on allele frequency (gnomAD) and predicted pathogenicity (AlphaMissense). Variants identified in the same family are connected by dashed lines. d, Putative human RPL3L knockout in the general population.
Notably, the ten missense variants can be categorized into two groups based on allele frequency and pathogenicity predicted using AlphaMissense18, a deep learning model built on the AlphaFold2 protein structure prediction tool (Fig. 1c). The hotspot variants G27D and D308N/V fall into the group with extremely low allele frequencies (<10−5, Genome Aggregation Database (gnomAD)19) and are predicted to be highly pathogenic. Notably, both G27D and D308N are enriched in admixed American populations (Supplementary Table 2), although they remain exceedingly rare (~10−5). The rarity of these hotspot variants in the general population and their repeated occurrence in a substantial proportion of affected families strongly challenges an LOF mechanism for these variants. Variants in the low-pathogenicity group are more frequent (still very rare, 10−5 to 10−4) and predicted to be less pathogenic (Fig. 1c). Interestingly, all compound heterozygous individuals carry one variant from the high-pathogenicity group and another from the low-pathogenicity group (linked by dashed lines).
Human knockouts of RPL3L exist
Consistent with the absence of a severe cardiac phenotype in Rpl3l-knockout mice, live human knockouts of RPL3L presumably free of severe early-onset DCM have been identified in large-scale genetic studies, including gnomAD19, All of Us20 and the UK Biobank21 (Fig. 1d). One individual carries a premature stop codon at Arg226 in both alleles (R226X), truncating nearly half of the 407-amino-acid protein. In addition, 17 individuals are homozygous for the splice donor variant (c.1167+1G>A). The lack of severe cardiac dysfunction in both putative RPL3L-knockout humans and Rpl3l-knockout mice strongly suggests that RPL3L-linked DCM is unlikely to be caused by simple RPL3L deficiency, despite the autosomal recessive inheritance pattern in affected families.
Ribosome deficiency in patient hearts
To investigate the pathogenetic mechanisms of RPL3L mutations in DCM, we examined the explanted heart from a 5-month-old male patient carrying D308N/T189M9. Bioanalyzer analysis of total RNA revealed a drastic reduction in the 28S rRNA peak compared to control CM heart ventricular tissues or human AC16 cardiomyocyte-like cells (Fig. 2a). The 28S rRNA is the scaffold of the 60S large subunit of the ribosome. Its loss is consistent with the vital roles of RPL3 (and presumably also RPL3L in cardiomyocytes) both as a core structural component of 60S and as a key regulator of 28S rRNA biogenesis and pre-60S ribosome assembly via direct binding to 28S rRNA and its precursors22.
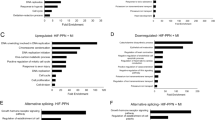
a, Left, schematics depicting the ribosome, including the 60S and 40S subunits scaffolded with 28S and 18S rRNAs, respectively. RPL3/RPL3L is highlighted in red and other ribosomal proteins in gray. Right, bioanalyzer electrophoresis of total RNAs isolated from explanted patient heart ventricular tissue with RPL3L mutations (T189M/D308N), control heart tissues from a 10-year-old female patient with restrictive CM or a 48-year-old male patient with ischemic CM, or AC16 cells. b, Differential gene expression analysis comparing two ventricular regions in the explanted heart and age-matched and sex-matched healthy control heart ventricular tissue (n = 2 technical replicates for D308N/T189M and two biological replicates for controls). Genes encoding 60S and 40S ribosomal proteins are colored red and blue, respectively. RPL3 and RPL3L are highlighted in green and light blue, respectively. c, Top four enriched GO terms in upregulated and downregulated genes. d–g, Cumulative distribution function plots for four significant gene signatures: ribosome (d), apoptosis (e), systolic heart failure (f) and fetal heart myeloid cell (g). Purple/blue indicates genes of interest and black indicates all other expressed genes. The median log2(fold change) and one-sided Kolmogorov–Smirnov test P values are also shown above each plot.
Comparing the transcriptome of the explanted heart to two healthy ventricular tissues with matched sex (male) and age (226 and 127 days versus 5 months of our sample)23, we uncovered widespread alterations in gene expression, including 1,529 significantly upregulated genes and 1,853 significantly downregulated genes (false discovery rate (FDR) < 0.05 and fold change > 1.5; Fig. 2b and Supplementary Table 3). Among the downregulated genes are many ribosomal protein genes (red and blue, Fig. 2b), including RPL3L (green, log2(fold change) = −1.25, FDR = 5.5 × 10−5). Indeed, Gene Ontology (GO) analyses identified ‘cytoplasmic translation’ as the most enriched pathway in downregulated genes (FDR = 4.1 × 10−6; Fig. 2c), in addition to ‘cardiac muscle contraction’ (FDR = 1.7 × 10−2). Interestingly, ‘immune response’ was the most enriched pathway in upregulated genes (FDR = 1.1 × 10−6; Fig. 2c), in line with increased inflammation in failing hearts24. We further quantified the change of 34,837 molecular signatures (gene sets)25 (Supplementary Table 4), which confirmed a global downregulation of the ribosome (P = 3.1 × 10−10; Fig. 2d) and upregulation of apoptosis (P = 1.9 × 10−53; Fig. 2e), systolic heart failure (P = 1.5 × 10−17; Fig. 2f) and fetal heart myeloid cell signatures (log2(fold change) = 2.05; Fig. 2g) in patient tissue.
Surprisingly, unlike the robust compensatory increase of Rpl3 mRNA and protein in all Rpl3l-knockout mice5,8,15,16, RPL3 mRNA is slightly downregulated (by 21%, FDR = 0.60, light blue, Fig. 2b) in patient heart tissues like many other ribosomal protein genes (Fig. 2b,d). When normalized to the average expression of all 60S ribosomal protein genes in the same sample, neither RPL3 nor RPL3L was significantly changed (Extended Data Fig. 2). The lack of significant RPL3 compensation in patient hearts probably contributes to the several cardiac defects seen in human patients and may explain the phenotypic differences between humans and knockout mice.
Isogenic cell line models of RPL3L variants
Both D308N and T189M are expressed at comparable levels in the mRNA-seq data (Extended Data Fig. 2c) and mass spectrometry (MS) data (Extended Data Fig. 2d) in the patient heart. To determine the impact of individual variants, we sought to generate isogenic cell lines expressing various RPL3L variants. Of note, RPL3L is not expressed in human induced pluripotent stem cell-derived cardiomyocytes26 (Extended Data Fig. 3), which is consistent with the notion that induced pluripotent stem cells-cardiomyocytes resemble immature fetal cardiomyocytes27,28, while RPL3L expression is restricted to mature cardiomyocytes. We chose to use human AC16 cells, ventricular cardiomyocyte-like cells commonly used to study cardiac gene expression and function29, including the regulation of ribosomes and translation in cardiac hypertrophy30.
Wild-type AC16 cells naturally express only RPL3, not RPL3L. To switch to RPL3L variants, we introduced via lentivirus a doxycycline (DOX)-inducible short hairpin RNA (shRNA) targeting RPL3 (shRPL3) to deplete RPL3 in an inducible manner (Fig. 3a). A control cell line was also generated integrating a scrambled shRNA (shScramble). The shRPL3 cell line serves as a critical baseline control for the cellular defect and toxicity caused by the LOF of ribosomal proteins. Using shRPL3 cells, we further integrated a second DOX-inducible transgene expressing hemagglutinin (HA)-tagged RPL3L (RPL3L–HA), including the wild-type protein, two hotspot variants (D308N/G27D), two non-hotspot variants (T189M/R161W) and a single-nucleotide polymorphism (SNP) variant encoding P291L, the most common missense variant in RPL3L (allele frequency 4.26%, TOPMed31). The SNP variant is predicted to be benign and is homozygous in at least 715 living individuals.
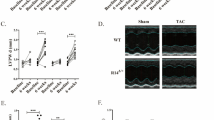
a, Wild-type (WT) AC16 cells were stably integrated with either a control shRNA (shScramble) or a RPL3-targeting shRNA (shRPL3), both inducible by DOX. The shRPL3 cell line was further integrated with HA-tagged RPL3L (or its variants), also expressed from a DOX-inducible promoter. b, Quantification of RPL3 mRNA abundance after 5 days of DOX-induced knockdown (n = 3), normalized to shScramble cells. c, RPL3L and its variants were expressed at a level comparable to that of endogenous RPL3 in shScramble cells. Expression was quantified in transcripts per million using RNA-seq and kallisto (n = 3). d, Bioanalyzer electrophoresis on total RNA from the indicated cell lines was quantified using ImageJ and then overlayed to match 18S peaks. e, Polysome sedimentation traces showing the loss of 60S, 80S and heavy polysomes in shRPL3 and D308N cells, but not WT, SNP or T189M cells (n = 3). f, Cell growth defects in D308N and shRPL3 cells. AC16-derived cells were seeded at equal density and counted every 2 days (n = 4 biological replicates). Data were analyzed using multiple two-sided t-tests with FDR correction for multiple comparisons, comparing shRPL3 to D308N. P = 0.067 at 10 days. The y axis is shown on a logarithmic scale. The percentage values represent the cell density at day 10 relative to the wild-type RPL3L cell line. b,f, Data are expressed as the mean ± s.d. *P < 0.01, **P < 0.001, ***P < 0.001, ****P < 0.00001.
By titrating the DOX concentration, we were able to reduce RPL3 by up to 81% (quantitative PCR with reverse transcription (RT–qPCR); Fig. 3b) while expressing RPL3L (wild-type and variants) at a level comparable to that of endogenous RPL3 in the control shScramble cells (1.67–3.09-fold overexpression, quantified using RNA-seq; Fig. 3c). The incomplete knockdown of RPL3 recapitulates the residual RPL3 expression detected in control and patient hearts (Supplementary Table 3) and in mouse cardiomyocytes5. At the protein level, the abundance of wild-type RPL3L was 61% lower than that of endogenous RPL3 in wild-type AC16 cells (MS; Extended Data Fig. 4), suggesting lower translation efficiency or shorter protein half-life for RPL3L. Therefore, this isogenic system enabled allele-specific switching from RPL3 to RPL3L at the physiological level upon DOX treatment (Fig. 3b,c). Of note, we observed a higher level of RPL3 in cells expressing the two non-hotspot variants (R161W and T189M) compared to other cells (Fig. 3b), despite comparable level of shRNA expression (Extended Data Fig. 5). The mechanism of this variant-specific RPL3 compensation is described below.
Hotspot variants are linked to ribosomal and cellular defects
Using these isogenic cell lines, we found that the 28S rRNA loss observed in the explanted D308N/T189M patient heart tissue (Fig. 2a) was accurately recapitulated in cells expressing the D308N variant (purple, Fig. 3d) but not the T189M variant (green) or wild-type protein (blue). Notably, 28S rRNA loss in D308N cells was much stronger than that of shRPL3 cells (black), despite having similar RPL3 levels (Fig. 3b). Given that the only difference between these two cell lines is the expression of the D308N variant, the stronger 28S rRNA defect indicates toxicity caused by the D308N variant. In addition, the moderate 28S rRNA loss in shRPL3 cells suggests that RPL3/RPL3L deficiency alone is insufficient to fully account for the defects observed in patients (Fig. 2a). We further confirmed that the loss of 28S rRNA scales with D308N expression in a DOX concentration series, with a concomitant accumulation of the 45S pre-rRNA (Extended Data Fig. 6a,b), indicating pre-rRNA processing defects associated with D308N.
Consistent with a defect in ribosome biogenesis, sucrose gradient sedimentation revealed a dramatic reduction in 60S and monosomes and polysomes in cells expressing the hotspot variant D308N (Fig. 3e) or G27D (Extended Data Fig. 6c), compared to wild-type, SNP, T189M or R161W cells (Fig. 3e and Extended Data Fig. 6c). Notably, the loss of ribosomes in D308N-expressing cells was slightly more severe compared to cells not expressing any RPL3L variant (shRPL3) (Fig. 3e), suggesting that the D308N variant does not contribute to ribosome biogenesis.
In line with a severe loss of ribosomes, cells expressing the hotspot variants D308N (Fig. 3f) or G27D (Extended Data Fig. 6d) exhibited a severe growth defect (note that the y axis is on a logarithmic scale). For instance, expressing D308N for 10 days led to a 40-fold reduction in cell counts (compared to wild-type RPL3L cells; Fig. 3f, cyan versus red). Interestingly, cells expressing the D308N variant exhibited a 6.4-fold greater growth defect compared to cells lacking both RPL3 and RPL3L expression (shRPL3) (cyan versus blue, Fig. 3f), again suggesting toxic gain-of-function (GOF) for D308N. This was further confirmed in a DOX concentration series, where even a low level of D308N expression caused a more severe growth defect than shRPL3 (Extended Data Fig. 6e).
Of note, the control cell line expressing RPL3 (shScramble) grew faster than cells expressing wild-type RPL3L (Fig. 3f), mirroring a previous mouse study showing that RPL3L expression is associated with reduced protein synthesis and slower muscle cell growth2. Interestingly, cells expressing the T189M variant of RPL3L grew faster than cells expressing the WT RPL3L (Fig. 3f), which may be attributed to a higher expression of RPL3 in this cell line (Fig. 3b; also see below on RPL3 compensation).
Taken together, we found that the hotspot variants G27D and D308N, but not their coinherited non-hotspot variants T189M and R161W, are responsible for ribosome biogenesis defects and cellular toxicity.
Hotspot variants are sequestered in the nucleus
We then examined the subcellular localization of RPL3L variants using fluorescence imaging of the HA-tag (teal, Fig. 4a). In contrast to the predominantly cytoplasmic localization of the wild-type RPL3L and the common SNP variant expected from a ribosomal protein, the hotspot variants D308N and G27D both exhibited nearly exclusive nuclear localization (Fig. 4a, teal; quantified in Fig. 4b), probably explaining the severe loss of 60S, 80S and polysomes in these cells (Fig. 3e). Notably, compared to all other variants, there was a significant decrease in the cytoplasmic/nuclear ratio of 28S rRNA levels in the two-hotspot-variant D308N/G27D-expressing cells (Fig. 4a, pink; quantified in Fig. 4c), which is consistent with a loss of total 28S rRNA (Fig. 3d) and accumulation of nuclear 45S pre-rRNA (Extended Data Fig. 6b). This 28S rRNA localization defect was not observed in shRPL3 cells, again suggesting that 28S rRNA loss in hotspot-variant-expressing cells is not because of simple RPL3L deficiency, but a toxic GOF of both hotspot variants. The absence of hotspot variants in the cytoplasm further suggests that the increased cellular toxicity observed in hotspot-variant-expressing cells compared to shRPL3 cells (Fig. 3f) is unlikely to arise simply from hotspot variants outcompeting residual RPL3 for ribosome incorporation.
a, Cells were stained for anti-HA antibody (RPL3L and variants) and 28S rRNA fluorescence in situ hybridization (FISH) probes and analyzed using ×60 confocal microscopy. b, Violin plot quantification of cytoplasmic and nuclear abundances of RPL3L–HA signals for the indicated cell lines, measured per cell (n = 26–41 cells). c, As in b but for 28S rRNA. One-way analysis of variance (ANOVA) was used for the statistical analysis. d, Human pre-rRNA processing into mature 18S, 5.8S and 28S rRNAs. e, Overlayed representative bioanalyzer traces of nuclear RNAs isolated from various AC16 cells. Traces are aligned to match the 18S rRNA peaks. Labeled peaks are boxed in d. f, Quantification of 45S normalized to 18S. n = 3. Data are expressed as the mean ± s.d. A one-way ANOVA was used for the statistical analysis. g, Nuclear RNA-seq read count in each region normalized to 18S and then to wild-type RPL3L-expressing cells (n = 2 of the biological replicates of nuclear fractionated RNA from f). NS, not significant. **P < 0.001, ****P < 0.00001.
Similar to the wild-type protein and the SNP variant, the non-hotspot T189M variant was predominantly localized to the cytoplasm (Fig. 4a). T189M-expressing cells showed a normal cytoplasmic/nuclear ratio for 28S rRNAs (Fig. 4c), which is consistent with a lack of defect in polysome trace (Fig. 3e) and cellular growth (Fig. 3f), potentially because of RPL3 compensation (Fig. 3b). Intriguingly, the other non-hotspot R161W variant was sequestered in the nucleus (Fig. 4a,b). However, unlike the hotspot variants and consistent with its lower predicted pathogenicity (Fig. 1c), R161W-expressing cells had a normal cytoplasmic/nuclear 28S rRNA ratio (Fig. 4a,c), which is consistent with a lack of defect in polysome trace (Extended Data Fig. 6a) or cell toxicity (Extended Data Fig. 6b), presumably because of RPL3 compensation in R161W cells (Fig. 3b).
Hotspot variants impair rRNA processing
To further investigate the ribosome biogenesis defect, we analyzed nuclear 45S pre-rRNA processing, the production of mature rRNAs (28S, 18S and 5.8S) via intermediates (Fig. 4d). Using bioanalyzer analysis of nuclear RNAs, we observed in hotspot-variant-expressing cells an increase in peaks corresponding to the 45S and 32S pre-rRNA (Fig. 4e,f), suggesting impaired rRNA processing. The accumulation of pre-rRNA was confirmed using qPCR with primers amplifying the 5′ external transcribed spacer (ETS) and internal transcribed spacer 1 (ITS1) (Extended Data Fig. 7).
We then sequenced the total nuclear RNA in replicates with deep coverage (approximately 20 million reads mapped to 45S). After normalizing to the 18S reads in the same sample and then to the same region in wild-type RPL3L cells, there was an 8–19% decrease of 28S in shRPL3, G27D and D308N cells, compared to wild-type, R161W or T189M cells, which is consistent with a 28S biogenesis defect (Fig. 4g). The smaller decrease of nuclear 28S rRNA compared to the much-pronounced decrease in total RNA (Fig. 3d) suggests additional loss of 28S rRNA in the cytoplasm, presumably because of the lack of RPL3L in the cytosol (Fig. 4a–c). In line with the bioanalyzer and qPCR results, cells expressing the two hotspot variants accumulated the highest level for all three precursor regions in 45S (5’ ETS, ITS1 and ITS2; Fig. 4g). The increased pre-rRNA accumulation observed in hotspot variant cells relative to shRPL3 cells further supports the notion that expression of these recurrent variants contributes additional toxicity.
Hotspot variants bind more strongly to nucleolar proteins
To gain biochemical insights into the toxicity of the hotspot variants, we profiled the nuclear interactomes of RPL3L variants using immunoprecipitation against HA-tag followed by MS in triplicate (Supplementary Table 5). Interestingly, principal component analysis (PCA) of the proteomics data separates the two hotspot variants from the rest in the first principal component (PC1), suggesting global rewiring of the interactome for hotspot variants (Fig. 5a). The second principal component (PC2) separates the three nuclear variants (G27D, D308N and R161W) from the two cytoplasmic variants (T189M and wild type). The close clustering of G27D and D308N, and their separation from R161W despite a shared pattern of nuclear sequestration (Fig. 4a,b), suggests that R161W and the two hotspot variants are retained in the nucleus through distinct mechanisms.
a, PCA based on the abundance of proteins coimmunoprecipitated with each RPL3L variant. n = 3. PC1 and PC2 represent the first and second principal components. b, Enrichment of GRWD1 and C7orf50 in proteins coimmunoprecipitated with the hotspot variants D308N and G27D compared to other variants. Peptide intensities were normalized by the median of all samples and then log2-transformed. The error bars represent the s.d. centered around mean values. n = 3 biological replicates of immunoprecipitated lysates. c, Immunoblotting from cells transfected with the indicated amount of wild-type RPL3L or D308N plasmids in AC16 cells for 24 h. Soluble and insoluble proteins were separated and probed for the indicated antibodies. n = 3 independently performed biological replicates for RPL3L, RPL3 and GAPDH; n = 1 for RPL4 and C7orf50. d, Transient transfection of wild-type RPL3L and the D308N variant at the indicated ratios (total 1.5 µg plasmid) followed by the detection of RPL3L aggregates. Top, representative immunoblots. Bottom, quantification of RPL3L aggregates normalized to transfection with 100% wild-type protein. The expected values assumed no interference between wild-type and D308N protein and were calculated by subtracting the signal intensity of RPL3L bands in wild-type at 100% from D308N, split into 25% segments. n = 3 biological replicates for RPL3L–HA. Data are expressed as the mean ± s.d. e, AC16 cells with 10 ng ml−1 (+) or 100 ng ml−1 (+++) DOX for 96 h were transfected with 1 µg D308N–HA for 24 h. Aggregation of the nucleolar proteins RPL4 and C7orf50 was analyzed by immunoblotting (representative of three replicates). f, Quantification of e. n = 3 biological replicates. Data are expressed as the mean ± s.d. Multiple two-sided t-tests were used to compare low RPL3L and high RPL3L, adjusted for multiple comparisons. RPL4 P = 0.001; C7orf50 P = 0.008. *P < 0.05, **P < 0.001.
Interestingly, GRWD1 (known as Rrb1 in yeast) and C7orf50 (Rbp95), the top two proteins driving the separation of hotspot variants from other variants, both exhibited increased affinity for the hotspot variants, supporting the concept of toxic GOF effects (Fig. 5b). GRWD1 serves as the dedicated chaperone for RPL3 (and also likely RPL3L), binding RPL3 cotranslationally to prevent its aggregation and degradation, ensuring its proper delivery to the assembly site on nucleolar pre-60S particles32. C7orf50 interacts with RPL3 both genetically and physically, binding adjacent to RPL3 on the 28S rRNA during early pre-60S ribosome biogenesis33. Several other 60S biogenesis factors also exhibited enrichment in the nuclear pulldown of RPL3L variants (Extended Data Fig. 8). The sequestration of these essential 60S ribosome biogenesis factors by hotspot variants probably contributes to the 60S biogenesis defect and the toxic GOF associated with these variants.
Hotspot variants promote protein aggregation
One established toxic GOF mechanism according to which an autosomal recessive missense mutation contributes to diseases is via mutant protein aggregation, which can be mitigated by the presence of the wild-type protein34. We found that the D308N protein has a much higher tendency to aggregate compared with wild-type RPL3L (Fig. 5c). In addition, the expression of D308N, but not wild-type RPL3L, causes other nucleolar proteins such as RPL3, RPL4 and C7orf50 to aggregate, but not cytoplasmic GAPDH (Fig. 5c). These observations are consistent with an increased association of hotspot variants with many nucleolar proteins involved in 60S ribosome biogenesis shown in the interactome analysis (Fig. 5b and Extended Data Fig. 8). Note that these experiments were performed in wild-type AC16 cells, which predominantly express RPL3 but not RPL3L, resembling early heart development (thus, it is relevant for this neonatal disease).
We then tested whether D308N aggregation is inhibited in the presence of wild-type RPL3L, as in heterozygous carriers of D308N who showed no severe cardiac defect. We pooled wild-type RPL3L and D308N plasmids (both HA-tagged) at various ratios (0, 25, 50, 75, 100% of D308N) and transfected the pooled plasmids into wild-type AC16 cells (Fig. 5d). Although there was less aggregate with increased wild-type RPL3L (Fig. 5d), this could be explained by a decreased level of D308N in the pool of plasmids (Fig. 5d). Thus, wild-type RPL3L does not prevent the aggregation of D308N itself. However, we found that DOX-inducible expression of RPL3L ameliorated the increased aggregation of RPL4 and C7orf50 caused by D308N (Fig. 5e,f). Thus, the wild-type RPL3L allele in heterozygous carriers of D308N mitigates both ribosome deficiency and the proteotoxicity caused by D308N.
Hotspot variants block posttranscriptional RPL3 compensation
The molecular mechanism underlying RPL3 compensation in Rpl3l-knockout mice remains unclear. As described earlier (Fig. 3b), we observed near-complete compensation of RPL3 mRNA in the non-hotspot-variant-R161W-expressing cells and to a lesser extent in T189M cells, but not in the two hotspot variant cells (D308N and G27D). This was confirmed at the protein level (Fig. 6a). Importantly, there was no significant change in the expression of RPL3-targeting shRNA (Extended Data Fig. 5), ruling out the possibility that RPL3 recovery was because of loss of shRNA-mediated knockdown in cells expressing R161W or T189M. The difference in RPL3 compensation again suggests that the non-hotspot variants R161W and T189M, but not hotspot variants G27D and D308N, are functionally equivalent to RPL3L knockout.
a, Immunoblotting showing compensatory increase of RPL3 protein in R161W and T189M cells. DCM-associated RPL3L variants were expressed for 120 h in DOX and whole-cell lysates were immunoblotted with the indicated antibodies (n = 3 independent experiments). Band intensity was first normalized to GAPDH and then to wild-type for RPL3L and shScramble for RPL3 and RPS6, respectively. b, RPL3 mRNA half-life was enhanced in R161W cells compared to G27D cells. Cells expressing RPL3L variants were incubated with 50 μg ml−1 of the RNA Polymerase II inhibitor α-amanitin for the indicated time points before analysis for RPL3 mRNA relative to GAPDH, normalized to 0 h (n = 3 biological replicates per time point; data are expressed as ± s.e.m.). Data were analyzed using a two-way ANOVA between G27D and R161W conditions (P = 0.015). c, Compensatory RPL3 mRNA increase occurred entirely in the cytoplasm in R161W cells. RNA underwent subcellular fractionation for amplification of RPL3 levels relative to GAPDH. n = 4 biological replicates analyzed using a two-sided t-test. d, A conserved RPL3 NMD isoform was detected in AC16 cells. The top two tracks are RNA-seq coverage and junctions from one shScramble replicate and three replicates of wild-type RPL3L (pooled to match the total junction reads). PhyloP, conservation scores from the alignment of 100 vertebrate genomes. The red asterisk indicates a premature stop codon in the NMD isoform. e, Percentage junction reads supporting the NMD isoform (out of NMD and canonical junctions combined) from whole-cell poly(A) + RNA-seq in cells expressing several RPL3L variants. n = 3 biological replicates of total RNA. Data are expressed as the mean ± s.d. f, RPL3 mRNA level in total RNAs isolated from wild-type AC16 cells expressing DOX-inducible RPL3L (without HA) for 96 h in the absence of shRPL3. n = 3 biological replicates, mean ± s.d., analyzed using multiple unpaired, two-sided t-tests between the no DOX and 1,000 ng ml−1 samples: RPL3 (P = 0.00007), RPL4 (P = 0.054), RPL10A (P = 0.003). g, CLIP–qPCR analyses using lysate from mock-transfected control cells (RPL3L−) or RPL3L–HA-transfected cells (RPL3+), followed by immunoprecipitation (IP) using anti-HA or IgG (mock). Two primer pairs were used for RPL3 intron 3. Data are shown as the mean ± s.d. n = 3 biological replicates. h, Scatter plot showing a negative correlation between the expression of the NMD isoform and RPL3 mRNA in whole-cell RNA-seq. i, Left, representative gel image of semiquantitative RT–qPCR products using primers spanning exons 3 and 4 in nuclear RNAs. Right, quantification in three replicates. n = 3 biological replicates of nuclear RNA. Data are expressed as the mean ± s.d. and were analyzed using a one-way ANOVA followed by Dunnett’s multiple comparisons test (comparison to wild-type RPL3L: shRPL3 P = 0.006; R161W P = 0.004; T189M P = 0.005). j, Percentage of junction reads supporting the canonical and NMD isoform in nuclear total RNA-seq. b,c,f, Data are expressed as the mean ± s.d. **P < 0.001, ***P < 0.0001.
We ruled out transcriptional upregulation as the mechanism of RPL3 compensation because neither RNA polymerase II occupancy at the RPL3 promoter (Extended Data Fig. 9a) nor nascent RNA levels measured using intronic qPCR (Extended Data Fig. 9b) differed significantly between G27D-expressing and R161W-expressing cells.
We then assessed the stability of RPL3 mRNA. Compared to cells expressing the G27D variant, RPL3 mRNA was significantly more stable in cells expressing the R161W variant (Fig. 6b; P < 0.05). Consistent with a posttranscriptional mechanism, the increase in steady-state RPL3 mRNA level was primarily due to increased cytoplasmic RPL3 mRNA, with no significant change in the nuclear fraction (Fig. 6c).
RPL3 compensation via loss of RPL3L-induced unproductive splicing
Nonsense-mediated mRNA decay (NMD) is a major cytoplasmic RNA degradation pathway that targets transcripts containing a poison exon—an alternatively spliced exon introducing a premature termination codon35. RPL3 has a deeply conserved NMD isoform generated by activation of a cryptic 3′ splice site in intron 3, extending exon 4 at the 5′ end to introduce a premature termination codon36 (Fig. 6d). The percentage of the RPL3 NMD isoform increased from 1.4% in control cells to 42% in NMD-deficient HEK 293 cells37 (Extended Data Fig. 9c), indicating that a substantial fraction of RPL3 transcripts is continually produced but rapidly eliminated by NMD. Although overexpression of RPL3 has been shown to promote NMD-mediated auto-inhibition in rat PC12 cells (adrenal medulla tumor)36, we observed no significant change in the RPL3 NMD isoform when RPL3 was overexpressed in C2C12 mouse skeletal muscle cells15 (Extended Data Fig. 9d) or knocked down in AC16 human cardiac muscle cells (shScramble versus shRPL3; Fig. 6e), suggesting that in muscle cells, RPL3 may not strongly regulate its own NMD isoform.
By contrast, expression of RPL3L led to a substantial 12.2-fold increase in the RPL3 NMD isoform compared to RPL3-expressing cells (wild-type RPL3L versus shScramble; Fig. 6e), indicating that RPL3L is a far more potent activator of RPL3 NMD splicing than RPL3 itself. Consistently, reanalysis of three previous mouse Rpl3l-knockout datasets5,8,15 showed that the NMD isoform is on average 7.85-fold more abundant in the hearts of wild-type mice (expressing RPL3L) than in Rpl3l-knockout mice (where RPL3 compensates) (Extended Data Fig. 9d). Supporting these in vivo observations, DOX-induced overexpression of RPL3L in mouse C2C12 cells resulted in a twofold increase of the RPL3 NMD isoform compared to RPL3 overexpression (4.87% versus 2.43%; Extended Data Fig. 9d). Similarly, overexpression of human RPL3L in wild-type AC16 cells (without shRPL3) using a stably integrated, DOX-inducible cassette produced up to a 6.1-fold increase of the RPL3 NMD isoform with increasing levels of RPL3L (Extended Data Fig. 9e). Remarkably, saturating DOX levels (100 ng ml−1) were sufficient to reduce RPL3 mRNA by 50% in the absence of shRPL3 (Fig. 6f). These results in human and mouse, both in vitro and in vivo, demonstrate that RPL3L is the primary activator of the deeply conserved RPL3 NMD isoform in muscle cells.
To test whether the RPL3L protein promotes unproductive splicing of RPL3 transcripts through direct RNA binding, we performed ultraviolet cross-linking and immunoprecipitation (CLIP) in wild-type AC16 cells transfected with RPL3L–HA. Compared with either mock (IgG) immunoprecipitation in RPL3L–HA-expressing cells or RPL3L–HA immunoprecipitation in cells lacking the construct, RPL3L–HA specifically bound intron 3 upstream of the alternative 3′ splice site, but not RPL3 exons or other RNAs (Fig. 6g). These results demonstrate that RPL3L directly binds RPL3 intron 3 to drive unproductive splicing of RPL3 mRNA, thereby revealing a previously unrecognized moonlighting function of RPL3L beyond its role in the ribosome.
Hotspot variants block RPL3 compensation by inducing NMD
Intriguingly, the NMD isoform also increased dramatically in cells expressing the two hotspot variants (G27D and D308N) in addition to wild-type and the SNP variant, but not the two non-hotspot variants (R161W and T189M) (Fig. 6e), suggesting that both the SNP and hotspot variants largely retain the ability to induce RPL3 unproductive splicing, whereas non-hotspot variants have lost it. Notably, there is a remarkable negative correlation between the activation of the NMD isoform and the expression of RPL3 mRNA across RPL3L-variant-expressing cells (Pearson’s correlation coefficient R = −0.94, P = 0.005; Fig. 6h), suggesting that RPL3L-mediated NMD is a major regulator of RPL3 abundance, and that RPL3 compensation in R161W and T189M cells is at least in part due to their inability to induce RPL3 NMD.
To determine the abundance of the NMD isoform before degradation in the cytosol, we examined its abundance in the nucleus. Consistent with the whole-cell mRNA-seq data, we robustly detected an RT–qPCR band corresponding to the NMD isoform in cells expressing wild-type, D308N and G27D, but not shRPL3 cells or cells expressing the non-hotspot variant T189M or R161W (Fig. 6i). Using junction reads from nuclear RNA-seq data, we found that the NMD isoform accounts for approximately 50% of RPL3 transcripts in cells expressing wild-type, D308N and G27D variants, but close to 0% in other cells (Fig. 6j), in line with a 50% loss of RPL3 in the cytosol (Fig. 6f). Thus, approximately 50% of RPL3 mRNAs are repressed by RPL3L and hotspot variants.
Defects in engineered compound heterozygous cells
Patients with RPL3L-linked DCM often carry a combination of a hotspot variant and a non-hotspot variant, such as the D308N/T189M and G27D/R161W pairs we have characterized. Given the opposing effects of the hotspot and non-hotspot variants on ribosome biogenesis and cell growth (Figs. 3–6), we sought to explore the impact of these variants when both are present in the same cell, as is common in most patients with DCM.
To address this, we integrated a DOX-inducible G27D variant into cells expressing the R161W variant, creating a model of compound heterozygosity in the patient with G27D/R161W DCM. Compared to R161W cells, G27D/R161W cells expressed a significantly higher level of the RPL3 NMD isoform (representative gel image in Fig. 7a and quantification of four replicates in Fig. 7b). Using a puromycin incorporation assay to measure protein synthesis rate, we observed a significant reduction in protein synthesis rate in G27D/R161W cells compared to those expressing R161W alone, although the rate was still higher than in cells expressing G27D alone (Fig. 7c). In line with a reduced protein synthesis rate, compound heterozygous cells grew significantly more slowly (Fig. 7d). These results suggest that the ability of hotspot variants to inhibit RPL3 compensation probably contributes to the defect in compound heterozygous patients.
a, Representative gel analysis of RT–qPCR products using the same primers as in Fig. 6h for R161W cells and G27D/R161W compound heterozygous cells with various DOX concentrations. b, Quantifications of a (n = 4 biological replicates, analyzed using multiple two-sided t-tests with FDR correction (two-stage setup method); P = 0.002). c, Cells stably integrated with G27D, R161W or both G27D and R161W were induced with DOX for 144 h. Puromycin was added to a 1 μM final concentration for 30 min before cells were collected for immunoblotting (n = 4 biological replicates). Quantification is shown on the right; significance was determined using an unpaired, two-tailed t-test (P = 0.024). d, Comparing the growth of R161W and G27D/R161W cells. n = 2 biological replicates and n = 4 technical replicates, analyzed using a two-way ANOVA (P = 0.0001). e, A model of RPL3–RPL3L paralog switching and pathogenetic mechanism in neonatal DCM. In wild-type (WT) cardiomyocytes, RPL3L represses RPL3 expression through alternative-splicing-coupled, nonsense-mediated decay (AS-NMD), leading to the predominance of RPL3L ribosomes. In RPL3L-knockout (KO) cells and those expressing non-hotspot variants, this repression is relieved, allowing RPL3 to compensate and produce predominantly RPL3 ribosomes. By contrast, in cells expressing hotspot variants, RPL3 remains repressed while mutant RPL3L is retained in the nucleus, resulting in ribosome deficiency. Hotspot variants further promote nucleolar protein aggregation and rRNA processing defects, driving additional ribosome deficiency and cellular toxicity. Data are expressed as the mean ± s.d. *P < 0.05, ***P < 0.0001.
Discussion
We propose a unifying pathogenetic mechanism according to which diverse mutations in a muscle-specific ribosome drive severe neonatal DCM (Fig. 7e). Central to this model is a deeply conserved, ribosome-independent moonlighting function of RPL3L: it represses RPL3 abundance, thereby enabling RPL3 compensation when RPL3L is lost (Fig. 7e, left and middle). Although RPL3 is transcriptionally active (Extended Data Fig. 9a,b), half of its transcripts are rapidly degraded via NMD triggered by RPL3L-dependent alternative splicing (Fig. 6 and Extended Data Fig. 9). In the absence of functional RPL3L, such as in RPL3L-knockout cells or cells carrying LOF variants, this repression is relieved and RPL3 is upregulated. This mechanism explains the compensatory upregulation of RPL3 in Rpl3l-knockout mice5,8,15,16 and probably contributes both to physiological paralog switching during muscle development1,5,38 and to its reversal during adult muscle hypertrophy2.
Although seemingly wasteful, the constant transcription and degradation of RPL3 mRNAs may confer a regulatory advantage by allowing rapid induction of RPL3 protein in response to change in RPL3L protein levels, without requiring de novo transcriptional activation. Moreover, RPL3 introns harbor four highly conserved small nucleolar RNAs (snoRNAs; SNORD-43, -139, -83A and -83B), whose biogenesis probably depends on RPL3 transcription. Thus, the posttranscriptional regulation of RPL3 by AS-NMD ensures precise and fast control of RPL3 protein abundance while preserving the production of snoRNAs critical for ribosome biogenesis.
Central to the pathogenetic mechanism is the differential effect of hotspot versus non-hotspot variants on RPL3 compensation (Fig. 7e). The two hotspot variants G27D and D308N/V retain the moonlighting function of repressing RPL3 through NMD (Fig. 6e–j) yet lose their primary role as a ribosomal protein by being sequestered in the nucleus and disrupting pre-rRNA processing (Fig. 4). This combination—loss of ribosomal function coupled with blockade of RPL3 compensation—results in ribosome deficiency, as observed in patient hearts (Fig. 2a–d), and probably represents the principal driver of disease. By contrast, non-hotspot variants act more like simple LOF alleles, permitting RPL3 to escape NMD and substitute for RPL3L in ribosomes (Fig. 6e–j).
Beyond blocking RPL3 compensation and causing ribosome deficiency, hotspot variants exert additional toxic effects that cannot be explained by RPL3L/RPL3 loss or ribosome deficiency alone. Compared with shRPL3 cells, which display a similar degree of RPL3 deficiency (Fig. 3b), cells expressing hotspot variants show a more severe loss of 28S rRNA (Fig. 3d and Fig. 4a–c) and 60S ribosomes (Fig. 3e), greater accumulation of pre-rRNAs (Fig. 4e–g), and a stronger growth defect (Fig. 3f). Mechanistically, these defects probably reflect a toxic GOF compared to wild-type RPL3L: hotspot variants exhibit enhanced binding to nucleolar proteins (Fig. 5a,b), an increased tendency to self-aggregate and promote aggregation of other nucleolar proteins (Fig. 5c), and subsequent nuclear retention (Fig. 4a,b). Thus, the combination of exacerbated ribosomal and cellular phenotypes compared to shRPL3 cells, together with the enhanced binding and aggregation properties compared to wild-type RPL3L, indicate that hotspot variants promote disease through mechanisms extending beyond RPL3L deficiency alone.
Taken together, our findings indicate that RPL3L-linked DCM arises from a dual mechanism: a toxic GOF coupled with loss of ribosomal function and inhibition of RPL3 compensation. Our work establishes a framework for interpreting the growing spectrum of RPL3L variants and lays the foundation for therapeutic development. More broadly, given that approximately 80% of Mendelian disease genes have duplicated paralogs39, similar combinatorial mechanisms may underlie a wider spectrum of genetic disorders.
Methods
Ethics statement
The studies, including clinical evaluation and genetic analyses, were reviewed and approved by respective institutional ethics committees from the following participating centers: Children’s Hospital of Philadelphia (Institutional Review Board no. 16-013278) and Kasturba Medical College and Hospital, Manipal, India (IEC: 363/2020). Written informed consent was obtained from the families for clinical exome and genome sequencing and for publication of de-identified information.
Cell culture, drugs and transfection
AC16 cells were cultured at 37 °C, 5% CO2 in low-glucose DMEM (cat. no. 11-885-084, Thermo Fisher Scientific) supplemented with 10% FCS. HEK 293T cells were cultured in DMEM with 10% FCS. For the immunofluorescence assays, 2 × 105 cells were plated in six-well plates and incubated for 24 h. For stable cell generation, EZ-Tet-RPL3-shRNA was cotransfected with CMV-dR8.91 and MD2.G plasmids for 72 h in HEK 293T cells to produce virus. The medium was centrifuged for 5 min at room temperature, filtered using a 0.45-µm Syringe Filter Unit (Thermo Fisher Scientific), and 200 µl viral medium was added to 2 × 105 AC16 cells for 48 h before selection using 200 µg ml−1 hygromycin B until control cells (non-transduced) were dead. For stable generation of pLenti-RPL3L, the above procedure was followed, except that transduction was carried out on EZ-Tet-RPL3-shRNA/AC16 cells and selection was carried out by sorting for BV421+ cells. For G27D/R161W stable cells, pLenti-G27D-HA-mCherry was transduced into R161W-HA-BFP cells and sorted for BFP+ and mCherry+ cells. For the mRNA stability assay, AC16 cells were treated with 50 mg ml−1 α-amanitin for the indicated time points.
Plasmids
To generate plasmid expressing shRNA against RPL3, a reported high-efficiency shRNA sequence for RPL3 was identified (Sigma-Aldrich) and ~70 nt single-stranded oligonucleotides were ordered from Integrated DNA Technologies and annealed by heating to 95 °C and cooled at a rate of 5 °C min−1. Double-stranded oligonucleotides had two 5′ overhangs corresponding to NheI and EcoRI sites, respectively. Double-stranded DNA oligonucleotides were ligated to EZ-Tet-pLKO-Hygro (cat. no. 85972, Addgene)40 using the Quick Ligation Kit (New England Biolabs). The pLenti-RPL3L plasmid was derived from pLentiRNACRISPR_007 (cat. no. 138149, Addgene)41. A gene fragment containing the wild-type human RPL3L sequence with a C-terminal HA-tag was delivered on pUC57 (Genewiz). The PCR products for wild-type RPL3L-HA derived from pUC57, the pLentiRNACRISPR_007 backbone and EBFP (unpublished in-house plasmid) with overlapping ends were assembled using Gibson cloning with the NEBuilder HiFi DNA Assembly Master Mix (New England Biolabs), producing DOX-inducible RPL3L and constitutive EBFP expression. To obtain the RPL3L mutation plasmids, primers (Supplementary Table 6) were designed against pUC57-RPL3L-HA with single-nucleotide substitutions in the 5′ region of the forward primer sequence. Mutagenesis was performed using the Site Directed Mutagenesis Kit (New England Biolabs) as per the manufacturer’s instructions.
Immunofluorescence and FISH
A total of 15,000 cells were seeded on Lab-Tek II chamber slides overnight, washed with PBS and fixed in 300 µl paraformaldehyde for 15 min at room temperature. Fixed cells were washed three times with PBS (3 min each), permeabilized in 0.5% Triton X-100 for 15 min, washed three times and blocked in 3% BSA/PBS for 1 h. Cells were incubated with primary antibody in 3% BSA overnight at 4 °C, washed three times, then incubated with secondary antibody for 1 h at room temperature in the dark. After 3× washing, cells were stained with 1:1,000 4,6-diamidino-2-phenylindole for 5 min, washed three times and imaged on a Nikon Eclipse Ti confocal microscope with ×60 lens. RPL3L was detected with anti-HA (clone 2-2.2.14, Thermo Fisher Scientific); secondary antibodies were goat anti-rabbit Alexa Fluor 488 and goat anti-mouse Alexa Fluor 594 (Invitrogen). For rRNA FISH, cells were fixed as above but permeabilized with 70% ethanol for ≥1 h. Custom 28S rRNA FISH probes (Supplementary Table 6) conjugated to CAL Fluor Red 590 (Stellaris) were incubated with samples for 16 h in the dark, washed with buffers A + B (Stellaris), then incubated with anti-HA antibody as above.
Polysome fractionation
Sucrose gradient density ultracentrifugation separated monosomal and polysomal ribosomal fractions. AC16 cells in 15-cm dishes were lysed with 800 µl ice-cold lysis buffer (20 mM HEPES, 150 mM KCl, 10 mM MgCl2, 0.5% NP-40, 1× EDTA-free PIC, 2.5 mM dithiothreitol, 25 U ml−1 Turbo DNase I, 10 U SuperaseIN, 2 mM Vanadyl Ribonucleoside Complex, 100 µg ml−1 CHX) by scraping. Lysates were triturated 10× with a 23-gauge syringe, incubated at 4 °C for 30 min with rotation, then centrifuged at 14,000g for 7 min. Supernatant (700 µl) was loaded onto a 10–50% sucrose gradient (made in lysis buffer plus 2.5 mM dithiothreitol, 160 U SuperaseIN, 100 µg ml−1 CHX) and ultracentrifuged at 248,000g for 2 h. Gradient formation and fractionation used a Biocomp Gradient Profiler. For protein extraction, 1 volume TCA was added to 4 volumes of sample, incubated 10 min on ice and centrifuged at 18,000g for 5 min at 4 °C. Pellets were washed twice with ice-cold acetone, dried at 95 °C for 5 min, then resuspended in 4× Sample reducing agent (GenScript) diluted 1:1 in radioimmunoprecipitation assay (RIPA) and heated 5 min at 95 °C for SDS–PAGE.
Immunoblotting
Immunoblot analysis was performed after SDS–PAGE using standard protocols. The antibodies used were anti-HA (clone 2-2.2.14, Thermo Fisher Scientific), anti-RPL3 (clone 3365, Sigma-Aldrich), anti-RPL28 (cat. no. 62192, Invitrogen), anti-RPS6 (clone 54D2, Cell Signaling Technology), anti-GAPDH (cat. no. PA1-988, Invitrogen) and anti-Vinculin (clone E1E9V, Cell SignalingTechnology), anti-C7orf50 (cat. no. 20797-1-AP, Proteintech) and anti-RPL4 (cat. no. 11302-1-AP, Proteintech).
RNA extraction and analysis and qPCR
Total RNA was isolated using the NucleoSpin RNA Plus kit (MACHAREY-NAGEL), including use of a genomic DNA removal column. For rRNA analysis, 500 ng RNA was run on a 2100 Bioanalyzer (Agilent Technologies) at the Columbia University’s Molecular Pathology Core. For qPCR, cDNA synthesis was performed using 500 ng RNA with SuperScript IV Reverse Transcriptase (Invitrogen) as per the manufacturers protocol with oligo(dT) primers. Real-time PCR was carried out using the PowerUp SYBR Green Master Mix using 1:3 diluted cDNA. For RT–qPCR, triplicates were performed with 0.5 µM primer and 1 µl cDNA per reaction in the QuantStudio 7 Flex Real-Time PCR System. Ct values obtained were plotted using GAPDH mRNA as the loading control. Quantifications were performed as per the MIQE guidelines. Statistical tests were performed with Prism 9 (GraphPad Software) and two-sided Student’s t-test were used to calculate the P values. Data are expressed as the mean ± s.d. for the number of replicates indicated, with P < 0.01 considered statistically significant. All primer sequences are displayed in Supplementary Table 6.
Nuclear and cytoplasmic RNA extraction
Cells were washed with ice-cold PBS, scraped into 1.7-ml tubes and pelleted at 500g for 5 min at 4 °C. Cytoplasmic RNA was extracted by lysing cells in cytoplasmic lysis buffer (40 mM HEPES-NaOH, pH 7.5, 160 mM KCl, 10 mM MgCl2, 0.5% NP-40, 0.5% glycerol and SuperaseIN) for 30 min at 4 °C with rotation, followed by centrifugation at 2,000g for 2 min to separate the cytoplasmic supernatant. Nuclear pellets were washed in cytoplasmic lysis buffer (without detergent), centrifuged at 500g for 5 min and subjected to a wash with nuclear lysis buffer (10 mM Tris-HCl, pH 8.0, 1.5 mM KCl, 2.5 mM MgCl2, 5% glycerol, 0.5% Triton X-100, 0.5% deoxycholate and 20 U SuperaseIN) for 15 min at 4 °C. Final washes were performed with cytoplasmic lysis buffer (without detergent); nuclei were pelleted at 2,000g for 2 min. Both cytoplasmic and nuclear RNA were extracted as above.
Co-immunoprectipitation
To prepare the magnetic bead–antibody complex, 50 µl of Dynabeads Protein G (Invitrogen) was washed three times with lysis buffer then incubated with 5 µl anti-HA (clone 2-2.2.14) in 400 µl lysis buffer for 1 h at room temperature with constant rotation. Magnetic bead–antibody complex was captured using the Magnetic Separation Rack (New England Biolabs), washed two times with cytoplasmic lysis buffer (20 mM Tris-HCl, pH 7.5, 150 mM KCl, 10% glycerol and 0.1% NP-40) then three times with lysis buffer; 2× 15-cm plates were grown at 80% confluency, cells were washed two times in ice-cold PBS and lysed in 400 µl of ice-cold lysis buffer. The supernatant was added to the magnetic bead–antibody complex and incubated with rotation at 4 °C overnight. Next day, the bead–antibody–antigen complex was washed three times for 10 min at 4 °C with ice-cold lysis buffer. Protein was eluted by incubation with 40 µl pH 3 glycine and heating at 70 °C with shaking for 10 min, then neutralized with 1/10 volume, pH 8.0, Tris.
Proteomics
Explanted frozen patient tissue was homogenized under liquid nitrogen and lysed in RIPA for MS analysis. For global quantitative proteomics, diaPASEF-based proteomics was used. Proteins were reduced and alkylated with 10 mM tris(2-carboxyethyl)phosphine and 40 mM CAA at 45 °C for 15 min, then digested overnight with trypsin/Lys-C (1:50 ratio) at 37 °C. Peptides were acidified with 1% TFA and cleaned using SDB-RPS StageTips before drying. Dried peptides were resuspended in liquid chromatography buffer (3% acetonitrile, 0.1% formic acid); 200 ng per sample was analyzed using diaPASEF on a timsTOF Pro instrument. Peptides were separated over 87 min on a reverse-phase C18 column (25 cm × 75 µm, 1.6 µm) at 300 nl min−1 using a gradient from 2–80% mobile phase B (0.1% formic acid in acetonitrile). The timsTOF Pro acquired data in diaPASEF mode using 32 × 50 Th isolation windows (m/z 400–1,200) with collision energy ramped from 59 eV to 20 eV as a function of mobility. Raw files were searched using DIA-NN with the UniProtKB/Swiss-Prot mouse database in library-free mode with match-between-runs enabled. Carbamidomethylation was the fixed modification; methionine excision, oxidation and N-terminal acetylation were variable modifications. Peptide length was 7–30 amino acids, precursor charge 1–4; FDR was set to 1% at the protein and peptide levels. Custom FASTA files containing mutant D308N peptides were analyzed on MaxQuant42 for peptide abundance relative to wild-type peptide abundance. For measuring RPL3 and RPL3L absolute protein abundances, cells expressing either paralog were cultured for 8 days, pelleted and processed using MS as per the conditions for frozen explanted heart tissue.
Co-immunoprecipitation data analysis
Quantitative analysis of co-immunoprecipitation MS data was performed using the DEP (Differential Enrichment analysis of Proteomics data) R package43. Raw LFQ intensities of proteins with more than one missing value per condition were filtered out. Data were then normalized using variance-stabilizing normalization. Missing values were imputed using random draws from a Gaussian distribution centered in a minimal value (MinProb). Differential enrichment was tested against the RPL3L control group. Significantly enriched proteins were determined by applying an FDR-adjusted P threshold of P < 0.05 and a log2(fold change) cutoff of log2(1.5).
Chromatin immunoprecipitation
Cells were crosslinked with 1% formaldehyde in the culture medium for 15 min at room temperature, followed by quenching with 125 mM glycine for 5 min. Cells were pelleted and washed. The pellet was resuspended in RIPA buffer with protease inhibitor cocktail (Roche), incubated on ice and sonicated to shear chromatin (~1-kb fragments). The lysate was centrifuged at 12,000g for 10 min at 4 °C and the volume was increased with dilution buffer (165 mM NaCl, 0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCl, pH 8.0). Chromatin was incubated with 5 μg of anti-RNA polymerase II (clone F-12, Santa Cruz Biotechnology) overnight at 4 °C. The chromatin-antibody mix was incubated with pre-washed Dynabeads and incubated at room temperature for 45 min. Beads were washed five times in dilution buffer and eluted in genomic DNA extraction kit (MACHAREY-NAGEL) lysis buffer, resuspended in 50 μl of water. PCR was performed using purified DNA and compared to the input samples. Two-sided Student’s t-tests were carried out to calculate the P value.
Nuclear localization calculation
Nuclear localization was calculated as a ratio of the immunofluorescent co-localization signal of anti-HA, with nuclear regions compared to the anti-HA signal with cytoplasmic regions, determined as cellular regions lacking in 4,6-diamidino-2-phenylindole signal. The same calculation was performed for 28S rRNA. All calculations were carried out with ImageJ. The nuclear–cytoplasmic signal ratios for each cell were compiled and underwent a one-way ANOVA with Dunnett correction for multiple comparisons across conditions.
Tissue poly(A)+ RNA-seq
Two small left ventricular tissues from the heart explanted from the individual carrying compound heterozygous D308N/T189M RPL3L alleles were homogenized using a pestle and mortar under liquid nitrogen. Homogenized tissue was submerged in TRIzol and extracted using the Direct-zol kit (Zymo Research). RNA integrity was analyzed using bioanalyzer electrophoresis. Samples underwent poly(A) RNA pulldown and libraries were constructed using Illumina TruSeq chemistry. Libraries were sequenced at the Columbia Sulzberger Genome Center with a NovaSeq 6000 (Illumina). Real-time analysis (Illumina) was used for base calling and bcl2fastq2 (v.2.19) was used for converting BCL to FASTQ format. Gene expression was quantified using kallisto (v.0.46.1) with a customized transcriptome index combining human MANE Select v.1.4 and sequences of transcripts on plasmids used to generate the AC16 cell lines (for example, shRPL3). RNA-seq data for two age-matched and sex-matched control samples were downloaded from the European Nucleotide Archive (samples 5818sTS and 5836sTS23) and analyzed similarly. DESeq2 was used for differential gene expression analysis44. In brief, the non-normalized read counts of each gene (kallisto output) from the four samples were merged into a single table and used as input for DESeq2. Of the 19,344 genes quantified, 13,360 passed a filter for expressed genes (at least ten reads in more than two samples). These expressed genes were used in DESeq2 to identify genes differentially expressed between patient and control samples. GO analysis was performed using DAVID45, comparing upregulated or downregulated genes to all expressed genes using terms in the GOTERM_BP_DIRECT group. For molecular signature (gene set) analysis, the log2(fold change) of genes in each gene set from the Human Molecular Signatures Database (v.2024.1)25,46 was compared to all other expressed genes using a two-sided Kolmogorov–Smirnov test. Only gene sets with a minimum of 50 expressed genes were retained.
AC16 cell line poly(A)+ RNA-seq
Total RNA was extracted as described above from each AC16 cell line in triplicate. Poly(A) + RNA-seq and subsequent gene expression quantification using kallisto were performed as described above for tissue samples.
rRNA analysis using RNA-seq
Nuclear RNA was isolated and sequenced as described above, skipping the poly(A) pulldown. Reads were mapped to a custom bowtie index for 45S rRNA (NR_145819.1). The program bedtools multicov was used to calculate the read count in each region of the 45S, including both mature rRNAs (18S, 28S, 5.8S) and transcribed spacers (5′ ETS, ITS1, ITS2). For each sample, region counts were divided by the total read counts mapped to 45S. These normalized abundances were then further normalized by the same region in the wild-type RPL3L sample.
Analysis of RPL3 NMD isoform using RNA-seq data
Reads were mapped to the human (hg38) or mouse (mm10) genome using STAR47. The resulting BAM files were loaded into the Integrative Genomics Viewer and splice junction plots (sashimi plots) were generated. In addition to the AC16 data, we also analyzed HEK 293 RNA-seq data with and without depleting the essential NMD factors SMG6 and SMG7 (ref. 37) (SRR27241537–SRR27241542), three mouse datasets with Rpl3l knockout5,8,15 and one mouse C2C12 cell line data overexpressing Rpl3 or Rpl3l15.
Isolation of aggregated proteins
To separate soluble and insoluble proteins, cells were pelleted and lysed in RIPA buffer. Lysed cells were sonicated for 5 s and centrifuged for 21,000g for 30 min at 4 °C. Soluble supernatants were saved for downstream analysis. Pellets were washed twice in RIPA at room temperature, then dissolved in urea buffer (20 mM Tris-HCl, pH 7.5, 2% SDS, 8 M urea) and sonicated for 3 min. Soluble and dissolved insoluble fractions were analyzed using SDS–PAGE and immunoblotting.
CLIP
Wild-type RPL3L cells in 15-cm plates were crosslinked using the Stratalinker UV crosslinker 2400, scraped in PBS and centrifuged. For immunoprecipitation, Dynabeads (Thermo Fisher Scientific) were washed three times in lysis buffer, resuspended in lysis buffer with 4 µl anti-HA antibody (cat. no. 71-5500) and incubated with rotation at room temperature for 1 h. Antibody–beads were washed with IP500 buffer (20 mM Tris-HCl, pH 7.5, 150 mM KCl, 10% glycerol (v/v) and 0.1% NP-40 (v/v)) and three times in lysis buffer. Lysates were incubated with antibody–beads overnight at 4 °C, washed twice in high-salt buffer (20 mM HEPES-KOH, pH 7.5, 350 mM KCl, 10 mM MgCl2 and 0.1% (vol/vol) NP-40). RNA on washed beads was eluted in TRIzol and inputs and immunoprecipitation samples were isolated using chloroform extraction.
Statistics and reproducibility
Replicates and statistical tests are described in the figure legends. Sample sizes were not predetermined using statistical methods. No sample size calculations were performed. Sample size was governed by tissue availability. No samples were excluded. Investigators were not blinded to the groups when performing the initial analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data have been deposited in the Gene Expression Omnibus (GEO)48, including the RNA-seq for the patient tissue (accession no. GSE285722) and AC16 cells (accession no. GSE302553). The MS proteomics data for the patient samples and the AC16 RPL3L interactome have been deposited in the ProteomeXchange Consortium via PRIDE49 (PXD066450). The MS proteomics data for AC16 and the wild-type RPL3L cells have also been deposited via PRIDE (PXD066930).
Code availability
The custom code used in this study is available at GitHub (https://github.com/xuebingwu/rpl3l-mutant-paper).
References
Van Raay, T. J., Connors, T. D., Klinger, K. W., Landes, G. M. & Burn, T. C. A novel ribosomal protein L3-like gene (RPL3L) maps to the autosomal dominant polycystic kidney disease gene region. Genomics 37, 172–176 (1996).
Chaillou, T., Zhang, X. & McCarthy, J. J. Expression of muscle-specific ribosomal protein L3-like impairs myotube growth. J. Cell Physiol. 231, 1894–1902 (2016).
Meskauskas, A. & Dinman, J. D. Ribosomal protein L3: gatekeeper to the A site. Mol. Cell 25, 877–888 (2007).
Meskauskas, A., Harger, J. W., Jacobs, K. L. & Dinman, J. D. Decreased peptidyltransferase activity correlates with increased programmed-1 ribosomal frameshifting and viral maintenance defects in the yeast Saccharomyces cerevisiae. RNA 9, 982–992 (2003).
Milenkovic, I. et al. Dynamic interplay between RPL3- and RPL3L-containing ribosomes modulates mitochondrial activity in the mammalian heart. Nucleic Acids Res. 51, 5301–5324 (2023).
Litviňuková, M. et al. Cells of the adult human heart. Nature 588, 466–472 (2020).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Grimes, K. M. et al. Rpl3l gene deletion in mice reduces heart weight over time. Front. Physiol. 14, 1054169 (2023).
Ganapathi, M. et al. Bi-allelic missense disease-causing variants in RPL3L associate neonatal dilated cardiomyopathy with muscle-specific ribosome biogenesis. Hum. Genet. 139, 1443–1454 (2020).
Das, B. B., Gajula, V., Arya, S. & Taylor, M. B. Compound heterozygous missense variants in RPL3L genes associated with severe forms of dilated cardiomyopathy: a case report and literature review. Children 9, 1495 (2022).
Nannapaneni, H., Ghaleb, S., Arya, S., Gajula, V. & Taylor, M. B. Further evidence of autosomal recessive inheritance of RPL3L pathogenic variants with rapidly progressive neonatal dilated cardiomyopathy. J. Cardiovasc. Dev. Dis. 9, 65 (2022).
Al-Hassnan, Z. N. et al. Categorized genetic analysis in childhood-onset cardiomyopathy. Circ. Genom. Precis. Med. 13, 504–514 (2020).
Bajpai, A. K. et al. Exploring the regulation and function of Rpl3l in the development of early-onset dilated cardiomyopathy and congestive heart failure using systems genetics approach. Genes 15, 53 (2023).
Yang, Q. et al. Novel compound heterozygous variants in the RPL3L gene causing dilated cardiomyopathy type-2D: a case report and literature review. BMC Med. Genomics 16, 127 (2023).
Shiraishi, C. et al. RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function. Nat. Commun.14, 2131 (2023).
Brown, L. P. & McCarthy, J. J. Striated Muscle-Specific Ribosomal Protein L3-like: Effect of Knockout on Cardiac Function and Protein Translation. PhD thesis, Univ. Kentucky (2021).
Thorolfsdottir, R. B. et al. Coding variants in RPL3L and MYZAP increase risk of atrial fibrillation. Commun. Biol. 1, 68 (2018).
Cheng, J. et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 381, eadg7492 (2023).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100 (2024).
Bick, A. G. et al. Genomic data in the All of Us Research Program. Nature 627, 340–346 (2024).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Dorner, K., Ruggeri, C., Zemp, I. & Kutay, U. Ribosome biogenesis factors—from names to functions. EMBO J. 42, e112699 (2023).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Adamo, L., Rocha-Resende, C., Prabhu, S. D. & Mann, D. L. Reappraising the role of inflammation in heart failure. Nat. Rev. Cardiol. 17, 269–285 (2020).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Pozo, M. R., Meredith, G. W. & Entcheva, E. Human iPSC-cardiomyocytes as an experimental model to study epigenetic modifiers of electrophysiology. Cells 11, 200 (2022).
Guo, Y. & Pu, W. T. Cardiomyocyte maturation: new phase in development. Circ. Res. 126, 1086–1106 (2020).
Thomas, D., Cunningham, N. J., Shenoy, S. & Wu, J. C. Human-induced pluripotent stem cells in cardiovascular research: current approaches in cardiac differentiation, maturation strategies, and scalable production. Cardiovasc. Res. 18, 20–36 (2022).
Davidson, M. M. et al. Novel cell lines derived from adult human ventricular cardiomyocytes. J. Mol. Cell. Cardiol. 39, 133–147 (2005).
Hedaya, O. M. et al. Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy. Nat. Commun. 14, 6166 (2023).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Pillet, B. et al. Dedicated chaperones coordinate co-translational regulation of ribosomal protein production with ribosome assembly to preserve proteostasis. eLife 11, e74255 (2022).
Bhutada, P. et al. Rbp95 binds to 25S rRNA helix H95 and cooperates with the Npa1 complex during early pre-60S particle maturation. Nucleic Acids Res. 50, 10053–10077 (2022).
Di Fede, G. et al. A recessive mutation in the APP gene with dominant-negative effect on amyloidogenesis. Science 323, 1473–1477 (2009).
Fair, B. et al. Global impact of unproductive splicing on human gene expression. Nat. Genet. 56, 1851–1861 (2024).
Cuccurese, M., Russo, G., Russo, A. & Pietropaolo, C. Alternative splicing and nonsense-mediated mRNA decay regulate mammalian ribosomal gene expression. Nucleic Acids Res. 33, 5965–5977 (2005).
Britto-Borges, T., Gehring, N. H., Boehm, V. & Dieterich, C. NMDtxDB: data-driven identification and annotation of human NMD target transcripts. RNA 30, 1277–1291 (2024).
Burn, T. C. et al. Generation of a transcriptional map for a 700-kb region surrounding the polycystic kidney disease type 1 (PKD1) and tuberous sclerosis type 2 (TSC2) disease genes on human chromosome 16p3.3. Genome Res. 6, 525–537 (1996).
Dickerson, J. E. & Robertson, D. L. On the origins of Mendelian disease genes in man: the impact of gene duplication. Mol. Biol. Evol. 29, 61–69 (2012).
Frank, S. B., Schulz, V. V. & Miranti, C. K. A streamlined method for the design and cloning of shRNAs into an optimized Dox-inducible lentiviral vector. BMC Biotechnol. 17, 24 (2017).
Wessels, H.-H. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat. Biotechnol. 38, 722–727 (2020).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Zhang, X. et al. Proteome-wide identification of ubiquitin interactions using UbIA-MS. Nat. Protoc. 13, 530–550 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995 (2013).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
We thank members of the Wu lab (especially L. Xing) and the Cardiometabolic Genomics Program at Columbia University Irving Medical Center for discussion and comments on the manuscript. This work is supported by National Institutes of Health (NIH)/National Heart, Lung, and Blood Institute grant 1R01HL171664-01 and Pershing Square Foundation MIND Prize (X.W.) and a postdoctoral fellowship from the American Heart Association (M.R.M.). This research was funded in part through the NIH/National Cancer Institute Cancer Center Support grant P30CA013696 and used the Genomics and High Throughput Screening Shared Resource, the Molecular Pathology Shared Resource and the CCTI Flow Cytometry Core. The CCTI Flow Cytometry Core is supported in part by the Office of the Director, NIH under award nos. S10OD030282 and S10OD020056. Biospecimens and the data used for this research were obtained from the Columbia University Biobank, which is partially supported by the Columbia University’s Clinical and Translational Science Award funded through grant no. UL1TR001873. The study of case 4 was supported by the DBT/Wellcome Trust India Alliance grant titled ‘Center for Rare Disease Diagnosis, Research and Training’ (grant IA/CRC/20/1/600002) to A.S.
Author information
Authors and Affiliations
Contributions
X.W. and M.R.M. conceived the functional study and wrote the manuscript with input from all authors. M.R.M. performed most of the experiments. E.R.R. assisted in the protein aggregation experiments. M.G. coordinated the case report sample collection and provided critical feedback. T.M.L. and J.M.F. identified and characterized the D308N/T340Nfs*26 case. M.V.P., A.B. and P.J. identified and characterized the D308N/R58Q case. A.L.R., R.C.A. and D.N. identified and characterized the D308N/c.1167+1G>A case. A.A., A.S. and S.S.N. identified and characterized the D308V/D308V case. R.K.S. generated the proteomics data. Y.Y. assisted with imaging. E.J.G. contributed the control patient heart samples. W.K.C. contributed the D308N/T189M tissue sample. W.K.C., M.P.R. and F.Y. edited the manuscript and provided critical feedback.
Corresponding authors
Ethics declarations
Competing interests
X.W. is a member of the scientific advisory board for Epitor Therapeutics. W.K.C. serves on the board of directors at Prime Medicine. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Eva Maria Novoa, Roddy Walsh, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Conservation of RPL3L protein sequence across eleven vertebrate species.
RPL3L protein sequences were downloaded from NCBI. Multiple sequence alignment was performed using Clustal Omega/EMBL-EBI. Positions affected by DCM missense variants are highlighted.
Extended Data Fig. 2 Expression of RPL3 and RPL3L in explanted patient hearts.
a, RPL3 expression (mRNA-seq) normalized to average 60S genes. N = 2 technical replicates for D308N/T189M, 2 biological replicates for controls. Error bars signify s.d centered on mean value. Unpaired two-sided t-test. b, same as a but for RPL3L. c, RNA-seq read coverage (BAM file visualized in IGV genome browser) showing that both D308N and T189M variants are equally expressed. d, Intensity of the peptide containing the D308 residue in the T189M variant and the peptide containing the N308 residue in the D308N variant normalized to the total intensity in RPL3L. The T189 site was not detected as it lacks nearby trypsin sites. Mean value of N = 2 technical replicates ± s.d. Unpaired two-sided t-test. ns, not significant.
Extended Data Fig. 3 iPSC cardiomyocytes (iPSC-CMs) do not express RPL3L.
Data was analyzed from RNA-seq performed by Pozo et al26 comparing iPSC-CMs to fetal and adult human heart.
Extended Data Fig. 4 Comparing RPL3 and RPL3L protein abundance.
Mass spectrometry was performed using lysate from wild type AC16 cells (expressing RPL3) and AC16 cells expressing shRPL3 and RPL3L-WT. Total peptide abundance (intensity) was calculated for each cell line, separating peptides that are unique to each protein from those that are ambiguous. N = 3 biological replicates. Data shown as mean ± s.d.
Extended Data Fig. 5 Expression of the RPL3-targeting shRNA in RPL3L variant-expressing cell lines.
TPM, tags per million (N = 3 biological replicates).
Extended Data Fig. 6 Effect of variants on rRNA processing, translation, and cell growth.
a, 28S/18S rRNA ratio in D308N cells under various dox concentrations. N = 3 biological replicates. b, same as a for 45S/18S rRNA ratios. c, Polysome fractionation for G27D and R161W, as in Fig. 3e. d, Cell growth defect for G27D and R161W, as in Fig. 3f. N = 3 biological replicates with 2 technical replicates. e, Cell growth defect for D308N and shRPL3 cells under various dox concentration. N = 2 biological replicates with 2 technical replicates. Data are mean ± s.d. ns, not significant; * P < 0.05; ** P < 0.001.; *** P < 0.0001; **** P < 00001.
Extended Data Fig. 7 Quantification of pre-rRNA.
qRT-PCR of 5’ ETS and ITS1 on the 45S pre-rRNA using nuclear RNAs isolated from indicated cell lines (N = 3 biological replicates, except RPL3L-WT which is 2). Data shown as mean ± s.d. P value as shown. Analyzed by two-sided t-test.
Extended Data Fig. 8 Enrichment of 60S ribosome biogenesis factors in proteins co-immunoprecipitated with nuclear RPL3L variants.
Peptide intensities are normalized by the median of all samples and then log2-transformed. Error bars represent standard deviation. N = 3 biological replicates of immunoprecipitated lysates as in Fig. 5b.
Extended Data Fig. 9 Mechanisms of RPL3 compensation.
a, No difference in transcription activity in RPL3 promoter assayed by RNA Pol II ChIP-qPCR. Multiple primer pairs were used for each region. An intergenic region upstream of RPL3L was used as a negative control of non-transcribed region (N = 3 biological replicates of immunoprecipitated chromatin). Data shown as mean ± s.d. Statistical significance determined by two-sided t-test between G27D and R161W. b, R161W does not induce higher RPL3 intronic RNA as a proxy of transcription levels. Total RNA was isolated and cDNA was synthesized using random primers. Primers targeting intron 3 (not overlapping with the NMD exon) and intron 9 of RPL3 were used for amplification (N = 3 biological replicates). Analyzed by two-sided t-test between G27D and R161W. Error bars represent standard deviation. ns, not significant. c, RNA-seq coverage and junction reads in control HEK293 cells and SMG6-KD/SMG7-KO cells from a previous study37. N = 3 biological replicates from that study. Data shown as mean ± s.d. d, RNA-seq coverage and junction reads in mouse hearts (WT and KO) and C2C12 cells (OE, overexpression) from three previous studies: Grimes et al.8, Milenkovic et al.5, and Shiraishi et al.15 Replicates were pooled. e, Wild type AC16 cells expressing dox-inducible RPL3L (without HA) for 96 h were analyzed with RT-PCR for RPL3 NMD exon activation in the absence of shRPL3. The numbers below the gel are relative band intensity (NMD/Canonical) normalized to lane 2 (No Dox). Related to Fig. 6f.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note
Supplementary Tables 1–6 (download XLSX )
Supplementary Table 1: Clinical and molecular characteristics of all patients with RPL3L-associated dilated cardiomyopathy; Supplementary Table 2: Enrichment of some RPL3L variants in admixed American populations. Allele counts and frequencies of some RPL3L variants in various population genetic databases; Supplementary Table 3: Differentially expressed genes in D308N/T189M patient hearts (DESeq2); Supplementary Table 4: Gene set analysis; Supplementary Table 5: AC16 interactome; Supplementary Table 6: Oligonucleotides used.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Full-length, unprocessed gels or immunoblots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Full-length, unprocessed gels or immunoblots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download PDF )
Full-length, unprocessed gels or immunoblots.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed agarose gels.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Murphy, M.R., Ganapathi, M., Rotlevi, E.R. et al. Pathogenetic mechanisms of muscle-specific ribosomes in dilated cardiomyopathy. Nat Cardiovasc Res 5, 51–66 (2026). https://doi.org/10.1038/s44161-025-00761-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-025-00761-8