Abstract
Metabolomic indices summarizing diet-related metabolic responses are instrumental for examining and replicating diet–disease associations. Here we aim to identify metabolomic signatures characterizing the amounts and types of dietary carbohydrate and assess their associations with type 2 diabetes (T2D) risk. Nutritional metabolomics indices were developed using data from 1,196 healthy participants in the Lifestyle Validation Study with 7-day diet records (7DDRs). Elastic net regression within cross-validation was used to derive metabolomic indices of total carbohydrates and primary food sources. Replication was conducted using feeding menu data among 153 women from the Nutrition and Physical Activity Assessment Study. Associations with incident T2D were examined using multivariable Cox regression in 11,454 participants from the Nurses’ Health Study, Nurses’ Health Study II and Health Professionals Follow-up Study. Metabolites positively associated with total carbohydrates and added sugars mainly included glycerolipids (diacylglycerols and triglycerides), whereas glycerophospholipids (phosphatidylethanolamines and phosphatidylcholines) were inversely associated. Whole grains were linked to betaine, 3-indolepropionic acid (IPA) and hippuric acid; vegetables and legumes to IPA, N-acetylornithine and pipecolic acid; and fruits to proline-betaine and IPA. Identified metabolomic signatures showed significant correlations with a 7-day diet record-assessed diet in the Lifestyle Validation Study (Pearson r 0.33–0.65). In the Nutrition and Physical Activity Assessment Study, the metabolomic index of total carbohydrates was also significantly correlated with intake (r = 0.40). Signatures for total carbohydrates, added sugars, refined grains and potatoes were associated with higher T2D risk (HR per s.d. (95% confidence interval): 1.07 (1.02–1.12), 1.09 (1.03–1.14), 1.12 (1.07–1.18) and 1.36 (1.29–1.43)), whereas whole grain, vegetable, fruit and legume signatures were inversely associated (HR per s.d. (95% confidence interval): 0.73 (0.70–0.77), 0.95 (0.90–0.99), 0.88 (0.83–0.92) and 0.93 (0.88–0.97)). The metabolomic signatures of carbohydrate sources were differentially associated with T2D risk, highlighting the utility of blood metabolomics to objectively capture dietary carbohydrates and support dietary guidelines emphasizing fruits, vegetables and whole grains for diabetes prevention.
Similar content being viewed by others
Main
Carbohydrate intake plays a pivotal role in human health, influencing various metabolic processes and disease outcomes. Evidence unequivocally demonstrates that carbohydrates from different food sources may have different health consequences, highlighting the importance of carbohydrate quality1,2. In general, diets rich in whole grains, legumes, fruits and vegetables are associated with lower risks of obesity, type 2 diabetes (T2D) and cardiovascular disease in epidemiological studies3,4. On the other hand, diets high in readily absorbable sugars or starch, typically low in nutrient density and high in glycaemic load, including added sugars, refined grains and fried potatoes, show the opposite associations1,2,5. There was a suggestion of heterogeneity from these analyses that may be partially related to measurement errors associated with the recall-based dietary assessment tools used in these studies4,5. It is critical to use objective biomarkers that are free of the same measurement errors of the recall-based methods to replicate these important associations. However, valid biomarkers are scarce for quantifying the intake of total carbohydrates or its primary sources, with perhaps the exception of plasma alkylresorcinol levels for specific whole-grain foods (for example, rye)6 and certain lipid metabolites, such as triacylglycerol and lysophosphatidylcholine, which may reflect total carbohydrate intake7.
The human metabolome consists of a repertoire of small molecular-weight compounds with both endogenous and exogeneous origins and may serve as an alternate avenue for identifying objective markers that reflect diet. In comparison to the traditional nutrient biomarkers, metabolomics signatures often comprise multiple metabolites that, through certain combinations, jointly ‘register’ the responses to dietary exposures. Some previous studies have explored the utility of leveraging metabolomics data to measure diet. For example, a recent analysis used metabolomics data to develop macronutrient intake calibration equations for the evaluations of associations between calibrated macronutrient intake with disease outcomes7. Moreover, several studies have demonstrated that diets with a lower glycaemic index and glycaemic load are linked to favourable metabolomic profiles, characterized by shifts in triacylglycerol levels and reduced concentrations of markers associated with inflammation and oxidative stress8,9. Legume consumption was linked to a distinctive metabolomic signature of amino acids, cortisol and lipid species10. Fibre intake may be reflected by a group of metabolites, such as indolepropionate (3-indolepropionic acid, IPA), 3-phenylpropionate and methylguanidine11,12. Despite these advances, one limitation shared by previous studies is that the diet was often assessed using food frequency questionnaires (FFQs) or other recall-based instruments10,11, except for a feeding study7,12. As such, the measurement errors of these instruments may be propagated into the metabolomic signatures. In addition, few large prospective studies have thoroughly examined the associations of metabolites and built metabolomics signatures reflecting different amounts and types of carbohydrate intake and subsequent risk of incident T2D.
To fill the knowledge gaps, this current study aims to first identify metabolomics signatures of the amounts and types of carbohydrate assessed using 7-day diet records (7DDRs) and then to seek replications of the signatures in the independent feeding study7 that prepared controlled meals for study participants. Lastly, the current analysis examines the signatures in relation to T2D risk in three prospective cohort studies of US men and women.
Results
Characteristics of the study participants
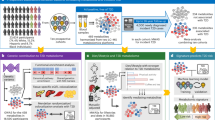
The current study consists of three main components (Fig. 1): (1) the discovery phase: a metabolomics analysis that examined relationships between dietary carbohydrate intake and plasma metabolites in the Lifestyle Validation Study (LVS); (2) the external replication phase: replicated the metabolomics signatures that reflect total carbohydrate intake and subtypes of dietary carbohydrate in the Nutrition and Physical Activity Assessment Study Feeding Study (NPAAS-FS) within the Women’s Health Initiative (WHI) cohort; (3) cohort analyses that investigated the prospective associations between carbohydrate metabolomic signatures and the risk of developing T2D in the Nurses’ Health Study (NHS), Nurses’ Health Study II (NHSII) and Health Professionals Follow-up Study (HPFS).
a, In the discovery phase, participants are enroled from the LVS. A total of 1,196 participants who had 7DDR data for carbohydrate intake and metabolomics data with missing rate <30% were included. Elastic net regression was used to select metabolites and build metabolite profile scores correlated to total carbohydrate and carbohydrate intake from different dietary sources. b, In the external replication phase, participants were from the NPAAS-FS. A total of 153 women completed the 14-day feeding study and attended two clinic visits. We mapped metabolites from the LVS with this metabolomics data from the NPAAS-FS. A correlation analysis was employed to examine the relationships between the metabolomic profile scores and carbohydrate intake in replication phase of NPAAS-FS. c, For the cohort analysis of metabolite profiles with incident T2D, participants were selected from NHS, NHSII and HPFS. Blood samples were collected from NHS participants during 1989–1990, from NHSII participants during 1996–1999 and from HPFS participants during 1993–1995. We used Cox regression models to evaluated the prospective associations of carbohydrate associated metabolites and metabolite profile scores with incident T2D over follow-up. All icons adapted from Icons8 (https://icons8.com/icons).
The metabolomics analysis of LVS involved 1,196 participants, of whom 62.1% were women, with a mean age of 66.4 years. The average percentage of calories from carbohydrates was 45.7% (Extended Data Table 1). The NPAAS-FS study involved 153 women, with a mean age of 74.9 years. The cohort analysis consisted of 11,454 participants, with an average age of 52.6 years. Compared with participants who did not develop T2D during the follow-up period, those who did were more likely to have hypertension, hyperlipidemia and a higher body mass index (BMI) at the time of blood draw. In addition, they were less likely to engage in physical activity, had a lower Alternative Healthy Eating Index (AHEI) and drank less alcohol (Extended Data Table 2)
Carbohydrate consumption and individual metabolites
Plasma metabolites associated with the intake of carbohydrates in the LVS are shown in Fig. 2a,b. A total of 68 metabolites were found to be significantly associated with total carbohydrate consumption. Of these, the metabolites positively associated with total carbohydrate consumption included glycerolipids, such as diacylglycerol (DAG(34:3)) and triglycerides (TAG(50:4), TAG(52:6) and TAG(52:7)), as well as IPA, N-acetyltryptophan and proline-betaine. Conversely, metabolites inversely associated included glycerophospholipids, such as phosphatidylethanolamine (PE(P-36:1)/PE(O-36:2) and PE(P-38:4)/PE(O-38:5)) and phosphatidylcholine (PC(P-34:4)/PC(O-34:5)), and sphingolipids, such as sphingomyelin (SM(d18:1/20:0)), carnitine (C20, C26) and 2-aminobutyric acid.
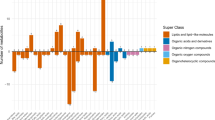
a, Volcano plot showing the associations between total carbohydrate consumption and plasma metabolites by using multivariate linear regressions in LVS (n = 1,196). The models were adjusted by cohort, age, BMI, ancestry, smoking, physical activity, alcohol drinking, AHEI, total calorie intake and per cent of calories from protein. Two-sided statistical tests were employed, with Bonferroni correction applied for multiple comparisons for analysing individual metabolites (P < 0.05/293). The metabolites above the dotted line are significant. b, A polar plot illustrating significant associations from the multivariate linear regressions. The data are represented as −log10(P) values for the associations. The bars pointing to outside the circle indicates positive and pointing inwards indicates negative. c, Selected coefficients for the metabolites from elastic net regression models for total carbohydrate consumption. Negative coefficients are oriented to the left, whereas positive coefficients are on the right. d, Triad method of total carbohydrate intake assessment in the LVS. e, The relationship between quintiles of total carbohydrate intake and predicted metabolomic indices in the three-pooled cohort study (n = 11,454). A correlation analysis was conducted between total carbohydrate intake and predicted metabolomic indices. The x axis represents the quintiles of total carbohydrate intake accessed in cohort study (percentage of calories from carbohydrates). The y axis denotes the metabolomic indices. In the box plots, the centre line represents the median, the bounds of the box represent the 25th and 75th percentiles, the whiskers extend to the smallest and largest values within 1.5× the interquartile range from the lower and upper quartiles, and the points beyond the whiskers denote the outliers. Two-sided statistical tests were employed. Met scores, carbohydrate-related metabolomic indices; rQR, correlation between FFQ and 7DDR; rRM, correlation between 7DDR and metabolomic indices; rQM, correlation between FFQ and metabolomic indices; rQT, validity coefficient of the FFQ; rRT, validity coefficient of the 7DDR; rMT, validity coefficient of metabolomic indices.
The associations between carbohydrate types and plasma metabolites are shown in Fig. 3. Similar to total carbohydrate consumption, positive correlates with added sugar consumption, including TAGs, whereas metabolites that were inversely associated mainly included PEs and PCs. However, for whole-grain consumption, positively associated metabolites included betaine, IPA and hippuric acid. Inversely associated metabolites included PEs and PCs. In particular, metabolites positively associated with fruit consumption included proline-betaine and IPA. For vegetable consumption, positively associated metabolites included IPA, N-acetylornithine and pipecolic acid. Regarding legume consumption, positively associated metabolites were similar to those associated with vegetables, particularly pipecolic acid, N-acetylornithine and IPA. For potato consumption, positively associated metabolites included triacylglycerol (TG) (52:5) and allantoin, whereas hippuric acid was inversely associated with potato intake. Additional statistically significant metabolites were detected for the intake of fried potatoes in comparison with baked potatoes, including C12 carnitine, vitamin A and several lipids (Supplementary Fig. 1).
a–g, Volcano plots showing the associations between added sugar (a), whole grain (b), refined grain (c), fruit (d), vegetable (E), legume (f), potato intake (g) and plasma metabolites by using multivariate linear regressions in the LVS (n = 1,196). The models were adjusted by for cohort, age, BMI, ancestry, smoking, physical activity, alcohol drinking and AHEI, excluding whole grains when analysing whole grains, similar for vegetables, fruits and legumes) and total calorie intake. The variables were accessed at the blood draw. Two-sided statistical tests were employed, with Bonferroni correction applied for multiple comparisons for analysing individual metabolites (P < 0.05/293). The metabolites above the dotted line are significant.
Metabolomic indices of carbohydrate consumption
A total of 36 metabolites for total carbohydrate consumption were selected to build metabolomic signatures (Fig. 2c). The selected metabolites for total carbohydrate consumption included IPA, glycerophospholipids and carboxylic acids and derivatives such as 2-aminobutyric acid. The metabolomic signatures of added sugar, whole grain, refined grain, fruit, vegetable, potato and legume consumption were shown in Extended Data Figs. 1 and 2 and Supplementary Figs. 2–8, respectively. The metabolomic indices based on the selected metabolites were significantly correlated with each corresponding carbohydrate food consumption derived from the 7DDRs in the LVS (Pearson r ranging from 0.33 (potatoes) to 0.65 (total carbohydrate)). Specifically, in the training set, we observed a Pearson r of 0.63 for total carbohydrate, 0.45 for added sugar, 0.30 for whole grains, 0.17 for refined grains, 0.50 for fruits, 0.43 for vegetables, 0.30 for potatoes and 0.29 for legumes. In the testing set, the corresponding Pearson r was 0.60 for total carbohydrate, 0.46 for added sugar, 0.25 for whole grains, 0.33 for refined grains, 0.50 for fruits, 0.41 for vegetables, 0.20 for potatoes and 0.28 for legumes (all P values <0.05) (Table 1). The identified metabolomic signatures showed significant correlations with the ‘true’ intake (Pearson r ranging from 0.47 (potatoes) to 0.69 (total carbohydrate)), using the triad method (Fig. 2d and Extended Data Fig. 3).
In the replication phase using NPAAS-FS, the metabolomics signatures were also significantly correlated with carbohydrate consumption estimated from the feeding menu data, with Pearson r of 0.40 for total carbohydrate, 0.21 for added sugar, 0.27 for whole grains, 0.21 for refined grains, 0.07 for fruits, 0.16 for vegetables and 0.28 for potatoes (Table 1 and Fig. 4). We further calculated partial correlation coefficients with adjustment for ancestry, age at blood draw, BMI at blood draw, smoking status, alcohol intake, physical activity and AHEI, which did not materially change our results.
a–f, The correlation coefficients between metabolomic indices and intake of total carbohydrate (a), whole grain (b), added sugar (c), refined grain (d), vegetable (e) and potato (f) in NPAAS-FS within the WHI (n = 153). The solid line represents the fitted regression line, and the shaded area indicates the 95% confidence interval.
In the pooled sample from the three cohorts (n = 11,454), the metabolomic indices were significantly associated with carbohydrate consumption (P < 0.001) (Fig. 2e). Pearson correlations with the corresponding carbohydrate intake derived from the FFQs were 0.38 for total carbohydrates, 0.24 for added sugar, 0.16 for whole grains, 0.09 for refined grains, 0.27 for fruits, 0.18 for vegetables, 0.07 for potatoes and 0.11 for legumes (Table S1). In the sensitivity analysis using least absolute shrinkage and selection operator (LASSO), the selected metabolites and the identified metabolomic signatures were similar to those selected by elastic net regression (Supplementary Table 2). In the analyses of metabolite selection robustness using stability selection with elastic net regression, the main metabolites identified in the original analysis were also selected as robust features (Supplementary Fig. 9).
Associations of metabolomic indices with the risk of T2D
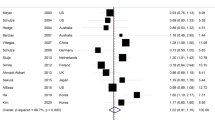
Table 2 presents prospective associations between the identified metabolomic signatures of total and subtypes of carbohydrate consumption and the risk of developing T2D. After adjusting for lifestyle and dietary risk factors, the HRs (95% confidence interval (CIs)) for T2D (per s.d. increase) were 1.07 (1.02–1.12; P = 0.01) for the total carbohydrate signature, 1.09 (1.03–1.14; P = 0.001) for added sugar, 1.12 (1.07–1.18; P < 0.001) for refined grains, 1.36 (1.29–1.43; P < 0.001) for potatoes, 0.73 (0.70–0.77; P < 0.001) for whole grains, 0.95 (0.90–0.99; P < 0.001) for fruits, 0.88 (0.83–0.92; P < 0.001) for vegetables and 0.93 (0.88–0.97; P < 0.001) for legumes. When total potatoes were further divided into fried and boiled/baked potatoes, the increased risk of T2D by potato intake was mainly driven by fried potatoes. The risk of T2D was significantly higher for fried potatoes (HR of 1.49, 95% CI 1.41–1.57; P < 0.001), whereas there was no significant association for baked potatoes (HR of 1.01, 95% CI 0.96–1.06; P = 0.73). In these models, we adjusted for BMI at early adulthood instead of BMI at blood draw, as concurrent BMI may function as an intermediate outcome in the pathway between carbohydrate intake and T2D risk (Supplementary Table 3). We additionally adjusted for the respective dietary carbohydrate variables (for example, whole grain or added sugar intake) in the analyses. The results remained largely unchanged after this adjustment, indicating that the associations were independent of self-reported dietary intake (Supplementary Table 4). The metabolite set enrichment analysis revealed that groups of triacylglycerols (with ≤56 carbons and ≤3 double bonds) and diacylglycerols were positively associated with T2D, whereas groups of cholesterol esters and plasmalogens showed significant inverse associations with T2D (Extended Data Fig. 4). In the sensitivity analysis of the T2D case–control study, after adjusting for potential confounders, the metabolite profile scores showed significant associations with T2D risk: consumptions of added sugar (HR of 1.21) and potatoes (HR of 1.84) was associated with an increased T2D risk, whereas whole grains (HR of 0.70), fruits (HR of 0.79), vegetables (HR of 0.85) and legumes (HR of 0.71) were linked to a lower risk (all P < 0.05) (Extended Data Table 3). Total carbohydrate signature was not significantly associated with T2D risk in this sample.
Discussion
The current analysis demonstrated that individual food sources of selected carbohydrates might have different associations with the blood metabolome. Metabolites of amino acids or phytochemicals, such as IPA, hippuric acid, proline-betaine, pipecolic acid and N-acetylornithine were among top metabolites associated with the intake of whole grains, fruits, vegetables and legumes. By contrast, lipid species, such as TAGs, were more likely among top predictors of intakes of added sugar or fried potatoes. Integrating dietary and metabolomics data from two independent studies, we also identified and replicated objective metabolomic signatures of carbohydrate intake. The objective metabolomic indices showed differential associations with T2D risk that were consistent with those observed in cohort studies that used FFQs to evaluate carbohydrate intakes4,5. Overall, the current study demonstrated the utility of metabolomics in developing objective indices of selected carbohydrate intake, which facilitate the validation of dietary assessment instruments and corroboration of associations with disease outcomes.
Previous research has identified certain metabolites related to carbohydrate or carbohydrate-related dietary consumption, and some of these metabolites are consistent with our findings. Metabolites, such as IPA, methylguanidine and 3-phenylpropionate, have been linked to dietary fibre intake11,12. Whole-grain intake was associated with pipecolic acid, betaine, tetradecanedioic acid and glucuronidated alkylresorcinols in cohort studies13. An intervention study supported that increasing whole-grain intake led to significant alterations in metabolite profiles. These changes included elevated levels of alkylresorcinols as well as several microbial metabolites, such as pipecolic acid betaine, sinapyl alcohol and enterolactone14. Furthermore, a feeding study found that a low-glycaemic whole-grain diet increased beneficial metabolites such as inositol, citrulline and glutamine, while reducing markers such as melatonin, trimethylamine and branched-chain amino acids associated with inflammation and insulin resistance9. Metabolites such as proline-betaine and hippuric acid have been linked to fruits and vegetables consumption in observational studies15,16. Our study contributed to this research by comprehensively identifying plasma metabolites associated with total carbohydrates and distinguishing between those related to high- and low-quality carbohydrate types.
Notably, total carbohydrate intake was associated with the highest diversity of metabolites, underscoring the complexity of carbohydrate composition and metabolism. Carbohydrates from whole grains, fruits, vegetables and legumes exhibit distinct metabolic profiles compared with those from added sugars, refined grains and potatoes. Our study identified several individual metabolites specifically related to fibre-rich carbohydrate foods, including whole grains, vegetables, fruits and legumes. One such metabolite, IPA, is produced in the gut through complex interactions among fibre intake, the gut microbiota fermentation of fibre and tryptophan consumption17. IPA influences metabolic pathways associated with glucose and lipid metabolism and also has anti-inflammatory and antioxidant properties18. Moreover, betaine, naturally found in whole grains, is absorbed by the small intestine and enters the bloodstream, where it functions as an osmolyte and serves as a methyl donor in the methionine cycle19. Betaine has been shown to play a role in reducing insulin resistance20. Proline-betaine is a metabolite commonly found in certain fruits, especially citrus fruits such as oranges and grapefruits21. Moreover, hippuric acid is a key metabolite that arises from the metabolism of dietary polyphenols22, which are abundant in a variety of plant-based foods, including whole grains, fruits and vegetables. In addition, N-acetylornithine is a positive marker of vegetables and legumes. N-acetylornithine is a derivative of the amino acid ornithine. A systematic review found that both N-acetylornithine and pipecolic acid may serve as biomarkers of legume intake23. Pipecolic acid is produced through the microbial metabolism of lysine, an amino acid abundant in vegetables and legumes, and serves as a non-specific biomarker of bean consumption24.
Research on the metabolomic signatures of simple sugars or refined carbohydrates in population-based studies is currently limited. A previous feeding study found that dietary patterns high in refined grains and added sugars is linked to increased levels of amino acid metabolites, purine metabolism and altered insulin response, suggesting a potential role in proinflammatory and insulin-related metabolic changes9. In our study, the metabolites related to more heathy carbohydrate foods were largely distinct from those responsive to added sugar, refined grains or fried potatoes, which are more likely to increase metabolites involved in fat storage and sphingolipid signalling and have a more pronounced adverse impact on glucose and fatty acid synthesis. We observed that the intake of added sugar might influence the levels of glycerolipids, especially TAGs with 46–50 carbons or ≤3 double bonds. This finding aligns with previous studies that have reported an association between lipids with lower carbon numbers and fewer double bonds and an increased risk of diabetes25,26. High total carbohydrate consumption, driven by low-quality carbohydrate, leads to elevated blood glucose levels and increased insulin secretion27, which promotes the conversion of excess glucose into fatty acids through de novo lipogenesis28. On the other hand, PEs, PCs and other glycerophospholipids were negative response markers to the low-quality carbohydrates. Changes in phospholipid metabolism might play a role in the insulin resistance and pathological process of metabolic disorders29,30.
Our research built metabolomic signatures for diverse carbohydrate sources, offering a deeper understanding of their unique metabolic functions. Notably, while some of the top metabolites are part of these indices, the algorithm also incorporated other metabolites that were not initially statistically significant. This may underscore the complex metabolic interactions that may contribute to the enhanced predictive power of our indices. The indices, including those for total carbohydrate and its main dietary sources, were associated with the diet with strength comparable to the established dietary markers31. They were also significantly associated with diet in the independent replication cohort, especially for the index of total carbohydrates. It is worth noting that in these two cohorts, different methods were used to assess diet, including 7DDRs, prepared meal data and FFQs. By employing diverse dietary assessment methods, we can effectively cross-examine the robustness of our findings, as the measurement errors associated with these methods are independent of each other. This methodological rigour significantly promoted validity of the replication process. However, the trade-off of this approach lies in the heterogeneity of dietary assessments and potential metabolomic platform-dependent differences between the studies. For example, the intake levels and variability and spectrum of carbohydrate foods in the LVS 7DDR data and NPAAS-FS menu data differed, and a proportion of metabolites identified in the LVS were missing from the NPAAS-FS metabolomics data, probably due to differences in analytical platforms across laboratories. Collectively, this heterogeneity may explain the seemingly attenuated correlation coefficients between carbohydrate intake variables and their signatures in the replication cohort, in comparison with the estimated correlations between the same signatures and ‘true’ intake in the LVS.
A growing body of evidence from prospective studies, primarily utilizing FFQs, has investigated the relationship between carbohydrate intake and T2D risk1,2,3,5. Although prior research has explored these associations using individual food components or overall carbohydrate intake, our study utilized metabolomic indices and signatures to offer a more comprehensive understanding of these relationships. Our research successfully replicated existing findings based on recall-based methods and also provided deeper insights into the metabolic pathways that connect carbohydrate quality and sources with T2D risk.
Our study has several strengths, including the prospective design of metabolomic indices with risk of developing T2D, the large sample size, the long follow-up, standardized diagnosis, evaluation of multiple carbohydrate subtypes and detailed covariable information. However, the limitations of this study must be addressed. First, the current analysis only included known metabolites, thereby excluding any unnamed or unidentified compounds. Second, we used blood samples for metabolomics analysis at only one timepoint. As the human metabolome may vary over time, repeated measurements may be still needed to achieve a more stable estimate of the metabolomics signatures. Third, metabolites were excluded from the current analysis if they showed unacceptable stability in our pilot study that examined the impact of delayed processing of samples during self-administered blood collection32. It is thus likely that some important lipid response markers were not captured by the current indices, and this may also underlie relative weaker correlations for refined grains and potatoes. Fourth, the metabolomic indices may largely capture the biological responses following carbohydrate intake, and it remains to be elucidated regarding the role of genetics, the human gut microbiome and other factors that may influence the responses. Lastly, although we used objective metabolomic indices and adjusted for a multitude of covariates, we still cannot exclude the role of confounding in our observations (such as food preparation methods, genetic factors, imperfectly measured physical activity and so on).
In conclusion, our study identified a panel of plasma metabolites associated with both total and individual types of carbohydrate intake, including some glycerolipids, glycerophospholipids and other metabolites such as betaine, IPA, hippuric acid, proline-betaine, N-acetylornithine and pipecolic acid. The metabolomic indices of carbohydrate consumption from different dietary sources were differentially associated with the risk of T2D. These findings highlighted the potential utility of metabolomics data to reflect intake of carbohydrates and to examine diet–disease associations objectively. Future studies should further evaluate the performance of these metabolite signatures by validating their associations with carbohydrate intake and T2D risk in independent cohorts with different demographic characteristics. Moreover, harmonization of metabolomics platforms and standardized dietary assessment methods would help to strengthen the generalizability and translational potential of these findings.
Methods
Study design
In the discovery phase, participants are enroled from the Men’s LVS (MLVS) and Women’s LVS (WLVS), the goal of which was to validate self-reported diet and lifestyle through the use of 7DDRs and objective biomarkers33. MLVS was conducted in 2011–2013 within the HPFS cohort and the Harvard Pilgrim Health Care cohort. The WLVS was conducted in 2010–2012 among selected participants from the NHS and NHSII. All LVS participants (including MLVS and WLVS) were free of a history of chronic diseases as per study protocol. In all, 1,196 LVS participants who completed 7DDR assessments and had existing metabolomics data were included in the current analyses.
In the external replication phase, participants were from the NPAAS-FS involving 153 participants of the WHI cohort. The NPAAS-FS was conducted in 2011–2013. The study targeted postmenopausal women who were free from major medical conditions. This 2-week long feeding study provided participants meals that were prepared according to each participant’ habitual diet assessed using a 4-day diet records as a starting point for individualizing diet specifications. A total of 153 women completed the feeding study and attended two clinic visits34. Blood samples were collected after a 2-week controlled feeding period designed to mimic participants’ usual diets, ensuring stable biomarker concentrations and retainment of the intake variations34.
For the cohort analysis of metabolomic profiles with incident T2D, participants were from NHS, NHSII and HPFS cohorts. In brief, blood samples were collected from 32,826 NHS participants during 1989–1990, 29,611 NHSII participants during 1996–1999 and 18,225 HPFS participants during 1993–1995. Metabolomic data were generated from multiple individual studies within these cohorts, which collectively provided data for the third component of the current analyses. Participants with existing metabolomics data were excluded if they had a daily energy intake below 500 kcal for women or 800 kcal for men or above 3,500 kcal for women and 4,000 kcal for men, if they were lost to follow-up after blood collection, or reported a history of cancer, cardiovascular disease or T2D at the time of blood draw. Ultimately, 11,454 participants were included from the pooled cohort (Extended Data Fig. 5). Of note, these participants did not include the LVS participants.
The study protocol has been approved by the Human Subjects Committees of the Harvard T.H. Chan School of Public Health and Brigham and Women’s Hospital. In the WHI study, participants provided written informed consent for the overall WHI programme and the NPAAS-FS substudy. Study protocols were approved by the Institutional Review Board at the Fred Hutchinson Cancer Research Center and all participating clinical centres.
Dietary assessment
We used two sets of 7DDRs data collected during LVS examinations to represent their habitual diet. Participants were provided with detailed instructions for completing their 7DDRs. Participants weighed their food before and after eating and submitted recipes for homemade dishes and labels from commercial products. Nutrition records were analysed using the Nutrition Data System for Research software at the Nutrition Coordinating Center, University of Minnesota, yielding data on over 150 nutrients and dietary constituents35,36. Total carbohydrate intake was expressed as percentage of calories. The intakes of added sugar and carbohydrates from whole grains, refined grains, vegetables, fruits, potatoes and legumes were adjusted for total energy intake using the residual method and then expressed as grams per day. The food contributors for these carbohydrate variables were summarized in Extended Data Table 4. We further categorized potatoes into baked/boiled/mashed potatoes versus fried potatoes.
In addition, participants from the LVS also completed a validated FFQ37. Participants in the NHS, HPFS and NHSII cohorts completed similar FFQs quadrennially since 1984, 1986 and 1991, respectively. Averaged nutrient intake was calculated based on the most recent FFQ cycle before blood collection (1990 for NHS, 1994 for HPFS and 1999 for NHSII). Total and types of carbohydrate intake were calculated by multiplying the frequency of food consumption by the nutrient content based on the Harvard University Food Composition Database and then summing these values. All carbohydrate variables were adjusted for total energy intake.
In the NPAAS-FS, participants’ intake of total and individual types of carbohydrate was derived from menus for preparing the controlled meals. To calculate intake for each food or food group, menu items were converted into standard servings per day using the Nutrition Data System for Research serving sizes. The food intake variables were then calculated by averaging the intake over the 14-day feeding period (mean servings per day)38. In this current study, types of carbohydrate intake included added sugars, whole grains, refined grains, vegetables, whole fruits and potatoes (in grams per day). Similar to the LVS, the total and types of carbohydrate intake were adjusted for total energy intake using the residual method.
Metabolomics measurement
In the LVS, plasma metabolomics profiling was conducted using high-throughput liquid chromatography–mass spectrometry techniques at the Broad Institute of MIT and Harvard (Cambridge, MA)32. Hydrophilic interaction liquid chromatography (HILIC) with positive ionization mode detection (HILIC-pos) was used to separate polar metabolites, HILIC with negative ionization mode detection (HILIC-neg) was utilized for negative ones and C8 chromatography with positive ionization mode detection (C8-pos) was used for polar and non-polar lipids in positive ion mode. Only named metabolites were analysed, excluding features with missing rates >75% or mean coefficient of variation <30%. We also excluded metabolites that did not pass our pilot study investigating the effects of delayed sample processing during blood collection (intraclass correlation coefficient <0.4)32. Two drug metabolites were further excluded (acetaminophen and α-hydroxymetoprolol). Metabolites were then natural log-transformed and scaled to z-scores. Missing data for each metabolite were imputed by replacing missing values with the half of minimum valid value. Finally, we included a total of 293 known metabolites in the metabolomics analysis during the discovery phase (Extended Data Fig. 6).
Serum metabolomics profiles for NPAAS-FS participants were derived using the Metabolon platform (Metabolon), which employs Q-exactive ultrahigh-performance liquid chromatography tandem mass spectrometry38. For the Metabolon method, the sample analysis and data processing, including peak alignment and compound identification, have been detailed in prior publications39. In addition, lipidomics profiling was conducted in Dr Daniel Raftery’s lab at the Northwest Metabolomics Research Center at the University of Washington using the Sciex QTRAP 5500 Lipidyzer platform, which incorporates the SelexION differential mobility spectrometry method that targeted 1,070 lipids in 13 major lipid classes40. The measurements of metabolomics and lipidomics are complementary in the spectrum of metabolites that each method emphasizes. We mapped the metabolites from the three labs between LVS and NPAAS-FS by the HMDB numbers, metabolite names, or synonyms of these metabolites using the Human Metabolome Database or Lipid Maps. Lipid metabolites in the LVS could be mapped by summing species with the same number of carbons and double bonds in NPAAS-FS (Supplementary Table 5).
Ascertainment of T2D
For the cohort analysis, self-reported T2D cases were confirmed via a supplemental questionnaire if at least one of the following criteria from the American Diabetes Association was met: (1) presence of one or more classic symptoms (for example, excessive thirst, frequent urination, weight loss, hunger, itching or coma) along with fasting plasma glucose (PG) ≥126 mg dl−1 (7.0 mmol l−1) or random PG ≥200 mg dl−1 (11.1 mmol l−1); (2) at least two elevated PG levels on separate occasions (fasting PG ≥140 mg dl−1, random PG ≥200 mg dl−1 or PG ≥200 mg dl−1 at 2 h during an oral glucose tolerance test) without accompanying symptoms; or (3) use of hypoglycaemic medication (either insulin or oral hypoglycaemic agents). Before 1998, a fasting PG level of ≥7.8 mmol l−1 (140 mg dl−1) was used for diagnosing diabetes based on National Diabetes Data Group criteria41. Beginning in 2010, HbA1c ≥6.5% was included in the diagnostic criteria42. The validity of the supplementary questionnaire was examined in two prior studies conducted within the NHS and HPFS cohorts. These studies utilized blinded medical record reviews, which confirmed T2D diagnoses in 98% and 97% of participants, respectively43.
Covariates
In the LVS, demographics, lifestyles and medical conditions were assessed from self-reported questionnaires completed at the blood draw, including sex, age, ancestry, body weight, smoking status, physical activity and alcohol drinking. BMI is calculated by dividing a person’s weight in kilograms by the square of their height in metres. The AHEI was derived and cumulatively averaged based on FFQ assessments since baseline (1986 in NHS, 1991 in NHSII and 1986 in HPFS) through 2010.
In the cohort analysis, information was obtained from self-reported biennial questionnaires until blood draw, including age, ancestry, family history of diabetes, BMI at early adulthood (age 18 in the NHS, NHSII or 21 in the HPFS), history of hypertension, history of high cholesterol, fasting status, smoking status, alcohol drinking, physical activity, total calories intake, per cent of calories from protein and AHEI.
Statistical analyses
In the LVS, we first explored metabolites that were associated with carbohydrate intake using multivariate linear regression models, with Bonferroni correction. To build metabolomic indices of carbohydrate intake, we applied elastic net regression to select relevant metabolites from all measured metabolites and constructed metabolomics scores for total carbohydrate intake as well as carbohydrate intake from different dietary sources. Individuals were randomized to either a training set or a testing set in a 7-to-3 fashion. The elastic net regression with a tenfold cross-validation, and leave-one-out approach was performed by using the R cv.glmnet function, with an α of 0.5 to indicate an equal mix of LASSO and Ridge regularization and the optimal lambda value based on the mean square error (‘lambda.min’) to minimize the cross-validation prediction error rate (R package ‘glmnet’)44,45. We then constructed metabolomic indices using the β coefficients estimated from the trained model in both training and testing sets. The performance of the metabolomic indices was evaluated using Pearson correlation coefficients between the indices and carbohydrate intake. Correlations between ‘true’ intake and intakes measured using 7DDRs, FFQ and the metabolomic indices were assessed using the triad method in LVS46,47. In light of the lack of well-accepted quantitative criteria for the performance of dietary biomarkers, we considered r ≥ 0.30 as the evidence of a successful development of the indices. Of note, correlations between diet and many established nutrient biomarkers, such as long-chain n − 3 fatty acids and trans fatty acids, and dietary intake were in the range of 0.30 and above48. We built the same carbohydrate metabolomic indices in the NPAAS-FS using available metabolites and calculated correlation coefficients to quantify the replication performance. Considering the heterogeneity between the discovery cohort and replication cohort in terms of dietary assessments and metabolomic profiling, we used statistical significance at 0.05 as the criterion for determining whether the replication was acceptable. The metabolomic indices were calculated using the following formula:
where Mi represents the level or concentration of the ith metabolite, and βi represents the coefficient associated with the ith metabolite.
In the cohort analysis, we constructed the same indices based on available metabolites. Correlations were calculated to examine the relationships between the indices and carbohydrate intake assessed using FFQs. We used Cox regression models to evaluate prospective associations of the indices with incident T2D during follow-up. Person-time was calculated from the blood collection date until the diagnosis of T2D, death, loss to follow-up or end of the study period (June 2020 in the three cohorts), whichever came first. We built two models with model 1 adjusted for study cohorts and age at blood draw. Model 2 was further adjusted for ancestry (white or others), fasting status (fasting or non-fasting), family history of diabetes (yes or no), smoking status (never smoking or smoking), alcohol drinking (quintiles of continuous), BMI at early adulthood (<25, 25–29.9 or ≥30 kg m−2), physical activity (quintiles of continuous), hypertension (yes or no), high cholesterol (yes or no), total calorie intake (quintiles) and AHEI (quintiles). In addition, several sensitivity analyses were conducted. First, as current BMI can be a potential mediator, mediation analyses using bootstrapping with 500 resamples were employed to explore indirect effects. Second, the selection of metabolites and their coefficients were determined using the LASSO regression with lambda yielding the minimum mean square error value49,50. Third, to assess the robustness of metabolite selection, we performed stability selection by repeatedly fitting elastic net regression models to random subsamples of the data with cross-validated penalty parameters. Metabolites selected in at least 80% of 100 subsampling iterations were considered robust and reproducible. Fourth, we additionally adjusted for the respective dietary carbohydrate variables (for example, whole grain or added sugar intake) to examine whether the metabolomic indices are associated with T2D independently of self-reported diet. Fifth, to gain a better understanding of the underlying biological processes, we organized metabolites into groups and performed metabolite set enrichment analysis to identify those specifically associated with T2D. Last, we examined associations between the indices and the diabetes risk in a nested case–control study of T2D within NHS, which included 1,456 participants (778 diabetes cases and 778 healthy controls) who were free of diabetes at blood draw in 1989–1990 with T2D risk ascertained through 2008. Conditional logistic regression models were used to investigate associations of interest. Two-sided statistical tests (P < 0.05) were employed, with Bonferroni correction applied for multiple comparisons when analysing individual metabolites. All statistical analyses were performed using R version 4.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Because of participant confidentiality and privacy concerns, data are available upon written request. According to standard controlled access procedure, applications to use the NHS, NHSII and HPFS resources will be reviewed by our External Collaborators Committee for scientific aims, evaluation of the fit of the data for the proposed methodology and verification that the proposed use meets the guidelines of the Ethics and Governance Framework and the consent that was provided by the participants. Investigators wishing to use the NHS, NHII and HPFS data are asked to submit a brief description of the proposed project. Further information including the procedures to obtain and access data from the NHS, NHSII and HPFS is described at https://www.nurseshealthstudy.org/researchers (contact: nhsaccess@channing.harvard.edu) and https://hsph.harvard.edu/research/health-professionals/resources/for-external-collaborators/. Researchers are granted access to the WHI datasets for their manuscript proposals that have been reviewed and approved by the WHI Publications and Presentations (P&P) Committee. Investigators wishing to use the WHI datasets are asked to submit a brief description of the proposed project. Further information including the procedures to obtain and access data is described at https://www.whi.org/ (contact: p&p@whi.org).
Code availability
The analysis programmes are publicly available via GitHub at https://github.com/Xiaowen-bot/carb.
References
Ludwig, D. S., Hu, F. B., Tappy, L. & Brand-Miller, J. Dietary carbohydrates: role of quality and quantity in chronic disease. Brit. Med. J. 361, k2340 (2018).
Kelly, R. K. et al. Associations between types and sources of dietary carbohydrates and cardiovascular disease risk: a prospective cohort study of UK Biobank participants. BMC Med. 21, 34 (2023).
Del Carmen Fernandez-Figares Jimenez, M. Plant foods, healthy plant-based diets, and type 2 diabetes: a review of the evidence. Nutr. Rev. 82, 929–948 (2024).
Aune, D. et al. Fruit and vegetable intake and the risk of cardiovascular disease, total cancer and all-cause mortality-a systematic review and dose-response meta-analysis of prospective studies. Int. J. Epidemiol. 46, 1029–1056 (2017).
Schwingshackl, L. et al. Food groups and risk of type 2 diabetes mellitus: a systematic review and meta-analysis of prospective studies. Eur. J. Epidemiol. 32, 363–375 (2017).
Landberg, R., Kamal-Eldin, A., Andersson, A., Vessby, B. & Åman, P. Alkylresorcinols as biomarkers of whole-grain wheat and rye intake: plasma concentration and intake estimated from dietary records1. Am. J. Clin. Nutr. 87, 832–838 (2008).
Prentice, R. L. et al. Biomarker-calibrated macronutrient intake and chronic disease risk among postmenopausal women. J. Nutr. 151, 2330–2341 (2021).
Bullo, M. et al. Plasma metabolomic profiles of glycemic index, glycemic load, and carbohydrate quality index in the PREDIMED study. J. Nutr. 151, 50–58 (2021).
Navarro, S. L. et al. Plasma metabolomics profiles suggest beneficial effects of a low-glycemic load dietary pattern on inflammation and energy metabolism. Am. J. Clin. Nutr. 110, 984–992 (2019).
Margara-Escudero, H. J. et al. Plasma metabolite profile of legume consumption and future risk of type 2 diabetes and cardiovascular disease. Cardiovasc. Diabetol. 23, 38 (2024).
Wang, Z. et al. Gut microbiota and blood metabolites related to fiber intake and type 2 diabetes. Circ. Res. 134, 842–854 (2024).
Prentice, R. L. et al. Biomarkers for components of dietary protein and carbohydrate with application to chronic disease risk in postmenopausal women. J. Nutr. 152, 1107–1117 (2022).
Noerman, S., Virtanen, J. K., Lehtonen, M., Brunius, C. & Hanhineva, K. Serum metabolites associated with wholegrain consumption using nontargeted metabolic profiling: a discovery and reproducibility study. Eur. J. Nutr. 62, 713–726 (2023).
Koistinen, V. M. et al. Metabolic changes in response to varying whole-grain wheat and rye intake. NPJ Sci. Food 8, 8 (2024).
Oude Griep, L. M. et al. A metabolome wide association study of fruit and vegetable consumption and associations with cardiovascular disease risk factors: The International Study of Macro-/Micronutrients and Blood Pressure (INTERMAP) Study. J. Nutr. 155, 122–131 (2025).
Krupp, D., Doberstein, N., Shi, L. & Remer, T. Hippuric acid in 24-hour urine collections is a potential biomarker for fruit and vegetable consumption in healthy children and adolescents. J. Nutr. 142, 1314–1320 (2012).
Hu, Y. et al. Interplay between diet, circulating indolepropionate concentrations and cardiometabolic health in US populations. Gut 72, 2260–2271 (2023).
Konopelski, P. & Mogilnicka, I. Biological effects of indole-3-propionic acid, a gut microbiota-derived metabolite, and its precursor tryptophan in mammals’ health and disease. Int. J. Mol. Sci. 23, 1222 (2022).
Filipcev, B., Kojic, J., Krulj, J., Bodroza-Solarov, M. & Ilic, N. Betaine in cereal grains and grain-based products. Foods 7, 49 (2018).
Kim, D. H. et al. Effect of betaine on hepatic insulin resistance through FOXO1-induced NLRP3 inflammasome. J. Nutr. Biochem. 45, 104–114 (2017).
Heinzmann, S. S. et al. Metabolic profiling strategy for discovery of nutritional biomarkers: proline betaine as a marker of citrus consumption. Am. J. Clin. Nutr. 92, 436–443 (2010).
Clarke, E. D. et al. The relationship between dietary polyphenol intakes and urinary polyphenol concentrations in adults prescribed a high vegetable and fruit diet. Nutrients 12, 3431 (2020).
Sri Harsha, P. S. C. et al. Biomarkers of legume intake in human intervention and observational studies: a systematic review. Genes Nutr. 13, 25 (2018).
Perera, T. et al. Identification and monitoring of metabolite markers of dry bean consumption in parallel human and mouse studies. Mol. Nutr. Food Res. 59, 795–806 (2015).
Rhee, E. P. et al. Lipid profiling identifies a triacylglycerol signature of insulin resistance and improves diabetes prediction in humans. J. Clin. Invest. 121, 1402–1411 (2011).
Taya, N. et al. Change in fatty acid composition of plasma triglyceride caused by a 2 week comprehensive risk management for diabetes: a prospective observational study of type 2 diabetes patients with supercritical fluid chromatography/mass spectrometry-based semi-target lipidomic analysis. J. Diabetes Investig. 14, 102–110 (2023).
Ludwig, D. S. & Ebbeling, C. B. The carbohydrate-insulin model of obesity: beyond ‘calories in, calories out’. JAMA Intern. Med. 178, 1098–1103 (2018).
Ameer, F., Scandiuzzi, L., Hasnain, S., Kalbacher, H. & Zaidi, N. De novo lipogenesis in health and disease. Metabolism 63, 895–902 (2014).
Chang, W., Hatch, G. M., Wang, Y., Yu, F. & Wang, M. The relationship between phospholipids and insulin resistance: from clinical to experimental studies. J. Cell. Mol. Med. 23, 702–710 (2019).
Chen, S. et al. Plasma glycerophospholipid profile, erythrocyte n–3 PUFAs, and metabolic syndrome incidence: a prospective study in Chinese men and women. Am. J. Clin. Nutr. 114, 143–153 (2021).
Landberg, R. et al. Dietary biomarkers-an update on their validity and applicability in epidemiological studies. Nutr. Rev. 82, 1260–1280 (2024).
Townsend, M. K. et al. Reproducibility of metabolomic profiles among men and women in 2 large cohort studies. Clin. Chem. 59, 1657–1667 (2013).
Yuan, C. et al. Relative validity of nutrient intakes assessed by questionnaire, 24-hour recalls, and diet records as compared with urinary recovery and plasma concentration biomarkers: findings for women. Am. J. Epidemiol. 187, 1051–1063 (2018).
Lampe, J. W. et al. Dietary biomarker evaluation in a controlled feeding study in women from the Women’s Health Initiative cohort. Am. J. Clin. Nutr. 105, 466–475 (2017).
Salvini, S. et al. Food-based validation of a dietary questionnaire: the effects of week-to-week variation in food consumption. Int. J. Epidemiol. 18, 858–867 (1989).
Al-Shaar, L. et al. Reproducibility and validity of a semiquantitative food frequency questionnaire in men assessed by multiple methods. Am. J. Epidemiol. 190, 1122–1132 (2021).
Gu, X. et al. Validity and reproducibility of FFQ in measuring food and food group intakes. Curr. Dev. Nutr. 6, 765 (2022).
Playdon, M. C. et al. Measuring diet by metabolomics: a 14-d controlled feeding study of weighed food intake. Am. J. Clin. Nutr. 119, 511–526 (2024).
Zarrouk, E. et al. High-resolution mass spectrometry: theoretical and technological aspects. Toxicologie Analytique et Clinique 34, 3–18 (2022).
Prentice, R. L. et al. Metabolomics biomarkers for fatty acid intake and biomarker-calibrated fatty acid associations with chronic disease risk in postmenopausal women. J. Nutr. 153, 2663–2677 (2023).
Report of the Expert Committee on the Diagnosis and Classification of Diabetes Mellitus. Diabetes Care 20, 1183–1197 (1997).
Association, A.D. Standards of medical care in diabetes—2010. Diabetes Care 33, S11–S61 (2010).
Hu, F. B. et al. Physical activity and television watching in relation to risk for type 2 diabetes mellitus in men. Arch. Intern. Med. 161, 1542–1548 (2001).
Li, J. et al. The Mediterranean diet, plasma metabolome, and cardiovascular disease risk. Eur. Heart J. 41, 2645–2656 (2020).
Wang, F. et al. Plasma metabolomic profiles associated with mortality and longevity in a prospective analysis of 13,512 individuals. Nat. Commun. 14, 5744 (2023).
Kaaks, R. J. Biochemical markers as additional measurements in studies of the accuracy of dietary questionnaire measurements: conceptual issues. Am. J. Clin. Nutr. 65, 1232S–1239S (1997).
Fraser, G. E. & Shavlik, D. J. Correlations between estimated and true dietary intakes. Ann. Epidemiol. 14, 287–295 (2004).
Sun, Q., Ma, J., Campos, H., Hankinson, S. E. & Hu, F. B. Comparison between plasma and erythrocyte fatty acid content as biomarkers of fatty acid intake in US women. Am. J. Clin. Nutr. 86, 74–81 (2007).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Ottosson, F. et al. Plasma metabolome predicts aortic stiffness and future risk of coronary artery disease and mortality after 23 years of follow-up in the general population. J. Am. Heart Assoc. 13, e033442 (2024).
Acknowledgements
We thank the participants and staff of the NHS, NHSII and HPFS for their valuable contributions. The authors also acknowledge the support by the NUS-Harvard Chan WHI to the Nurses’ Health Studies.
We thank the following investigators in the WHI programme—Program Office (National Heart, Lung, and Blood Institute, Bethesda, MD, USA): J. Rossouw, J. Reis and C. Price; Clinical Coordinating Center (Fred Hutchinson Cancer Research Center, Seattle, WA, USA): G. Anderson, R. Prentice, A. LaCroix, and C. Kooperberg; Investigators and Academic Centers (Brigham and Women’s Hospital, Harvard Medical School, Boston, MA, USA): J. E. Manson; (MedStar Health Research Institute/Howard University, Washington, DC, USA) B. V. Howard; (Stanford Prevention Research Center, Stanford, CA, USA) M. L. Stefanick; (University of Arizona, Tucson/Phoenix, AZ, MD, USA) C. A. Thomson; (University at Buffalo, Buffalo, NY, USA) J. Wactawski-Wende; (Wake Forest University School of Medicine, Winston-Salem, NC, USA) S. Shumaker; (University of Massachusetts) B. Silver; (Wake Forest University) M. Vitolins; (University of Alabama at Birmingham) G. Wells; (University at Buffalo) A. Millen; (University of Florida, Gainesville/Jacksonville Florida) M. Limacher; (The Ohio State University) Electra Paskett WHI Memory Study: (Wake Forest University School of Medicine, Winston-Salem, NC) M. Espeland.
The study was funded by the National Institutes of Health (grant nos. UM1 CA186107, U01 CA176726, U01 CA167552, P01 CA87969, R01 HL034594, R01 HL035464, R01 HL60712, R01 DK120870, R01 DK126698, R01 DK119268, U2C DK129670, DK119268, R01 ES022981, R01 ES036206, R21 AG070375, P30 DK035816, P30 CA015704 and R01 CA119171) and the Novo Nordisk Foundation (grant no. NNF24SA0092022).
The WHI programme is funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, US Department of Health and Human Services through contracts 75N92021D00001, 75N92021D00002, 75N92021D00003, 75N92021D00004 and 75N92021D00005.
The funders had no role in the study design; in the collection, analysis and interpretation of data; preparation, review or approval of the manuscript; and decision to submit the manuscript for publication. All authors confirm the independence of researchers from funders.
Author information
Authors and Affiliations
Contributions
Q.S. and X.W. conceived and designed the study. X.W. performed the statistical analysis and data interpretation and drafted the manuscript. All authors conducted the technique review and edited the manuscript. Q.S., X.W. and P.X. are responsible for the later revision of the code and manuscript. X.W. and Q.S. are guarantors of this work and, as such, had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Health thanks Christopher Papandreou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ben Johnson, in collaboration with the Nature Health team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Selected coefficients for the metabolites from elastic regression for added sugar consumption.
Negative coefficients are oriented to the left, while positive coefficients are on the right.
Extended Data Fig. 2 Selected coefficients for the metabolites from elastic regression for whole grain consumption.
Negative coefficients are oriented to the left, while positive coefficients are on the right.
Extended Data Fig. 3 Triad Method assessment of dietary carbohydrate intake.
Triad Method of (A) added sugar intake; (B) refined grain intake (C) whole grain intake; (D) vegetable intake; (E) fruit intake; (F) potato intake; (G) legume intake assessment in Lifestyle Validation Study. Abbreviations: FFQ, food frequency questionnaire; 7DDR, 7-day dietary records; Met scores, carbohydrate related metabolomic indices; rQR, correlation between FFQ and 7DDR; rRM, correlation between 7DDR and metabolomic indices; rQM, correlation between FFQ and metabolomic indices; rQT, validity coefficient of the FFQ; rRT, validity coefficient of the 7DDR; rMT, validity coefficient of metabolomic indices.
Extended Data Fig. 4 Metabolite groups and type 2 diabetes risk.
Association between knowledge-based metabolite groups and risk of type 2 diabetes. Metabolite Set Enrichment Analysis was used to estimate enrichment scores (NES) based on estimates from multivariable Cox models.
Extended Data Fig. 5 Participants selection of the cohort study.
Participants selection of the Nurses’ Health Study (NHS), NHSII, and Health Professionals Follow-Up Study (HPFS).
Extended Data Fig. 6 Metabolites in the discovery phase.
Frequency of included metabolites in the discovery phase from Lifestyle Validation Study (n = 293).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–9 and Tables 1–5.
Supplementary Data 1 (download XLSX )
Source data for Supplementary Figs. 1–9.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Xia, P., Wang, F. et al. Metabolomic signatures of dietary carbohydrates and differential association with type 2 diabetes. Nat. Health 1, 145–157 (2026). https://doi.org/10.1038/s44360-025-00023-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44360-025-00023-8