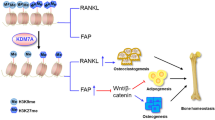
Abstract
Histone methylation plays a critical role in chromatin accessibility and transcription regulation, with implications for various disorders, including osteoporosis. Jumonji C domain-containing protein 7 (JMJD7) demethylates arginine (R) residues on histones, influencing tissue metabolism and integrity. However, its role in bone tissue remains uncharacterized. We have discovered JMJD7 loss in human osteoporotic bone biopsies, and neonatal osteoblast-specific Jmjd7 knockout mice exhibit delayed cranial suture closure and premature mortality. Adult female Jmjd7 knockout mice, but not males, develop a smaller stature with hallmark features of osteoporosis and visceral adiposity. Forced Jmjd7 expression mitigates estrogen deficiency-induced bone loss. Jmjd7 deletion alters the transcriptomic landscape and promotes the H3R2me1-enriched epigenome, particularly affecting cellular energy metabolism and suppressing osteogenic differentiation of bone marrow mesenchymal cells. Runx2 is, among others, a functional epigenomic target of Jmjd7. Mechanistically, Jmjd7 loss disrupts energy production by shifting towards anaerobic glycolysis at the expense of mitochondrial oxidative phosphorylation. This metabolic shift is mediated through inhibition of complex I activity and reduced production of isocitrate dehydrogenase (Idh) and its intermediate α-ketoglutarate (α-KG). Notably, α-KG supplementation reverses H3R2me1-dependent transcriptional repression and mitigates post-translational arginine methylation and ubiquitination of Idh and Runx2, counteracting Jmjd7 deletion-induced loss of mineralized matrix synthesis. Furthermore, α-KG supplementation improves osteogenic differentiation and bone formation, attenuating both Jmjd7 loss- and estrogen deficiency-induced osteoporosis. Taken together, Jmjd7 is indispensable for bone integrity, and its loss accelerates osteoporosis through epigenetic repression of α-KG production, affecting mitochondrial energy metabolism and Runx2 signaling. This study reveals a novel anabolic function of Jmjd7 in maintaining bone mass homeostasis and emphasizes the essential role of its cofactor α-KG in promoting bone health.
Similar content being viewed by others
Introduction
Bone homeostasis is finely tuned through a dynamic balance between bone acquisition by osteoblasts and remodeling by osteoclasts. Osteoblasts produce mineralized extracellular matrices to support bone anabolism and secrete key osteoclastogenic cytokines that regulate osteoclast formation and bone remodeling [1]. Disruption of this equilibrium between bone-forming and bone-resorbing activities leads to altered skeletal microarchitecture [2], ultimately resulting in osteoporosis characterized by low bone mass, deteriorated trabecular microstructure, and reduced biomechanical strength [3].
Mitochondrial energy metabolism plays a pivotal role in osteogenic differentiation and bone formation. For example, β-oxidation of long-chain fatty acids is required for parathyroid hormone-mediated osteogenic capacity and bone mineral acquisition [4]. Mitochondrial oxidative phosphorylation in periosteal progenitor cells is indispensable for proper cortical bone development [5]. The tricarboxylic acid (TCA) intermediate α-ketoglutarate (α-KG) is necessary for glutamine metabolism during osteogenic differentiation of skeletal stem cells [6] and mediates serine metabolism required for osteoclast formation from bone-marrow macrophage precursor cells [7]. As a co-factor, α-KG influences numerous epigenetic pathways contributing to tissue metabolism in both physiological and pathological contexts [8].
Epigenetic modifications, particularly post-translational methylation of lysine (K) or arginine (R) residues in histones, affect chromatin accessibility and gene transcription [9]. Emerging evidence reveals a strong correlation between histone methylation and osteoporotic disorders. For example, H3K9 demethylase Kdm7a knockout results in high bone mass and counteracts ovariectomy-induced osteoporosis [10]. Deletion of H3K4 methyltransferase Ash1l in osteoclasts accelerates bone loss [11], while H3K36 methyltransferase NSD2 mediates melatonin-induced promotion of osteogenic differentiation of human bone-marrow mesenchymal cells [12].
Histone arginine methyltransferases also play significant roles in bone metabolism. Prmt3 promotes H4R3 methylation and osteogenic activity of mesenchymal progenitor cells, and its silencing accelerates bone mass loss [13]. Carm1 knockout induces low bone mass and delayed regeneration capacity after tibia fracture [14]. Prmt5 catalyzes the symmetric dimethylation of H3R2, H4R3, and H3R8, and its knockdown promotes osteogenic activity of mesenchymal progenitor cells [15]. Prmt1 interference represses osteoclastogenic cytokine Rankl-mediated osteoclast formation and reverses ovariectomy-induced bone mass loss [16].
Histone demethylases of the Jumonji C domain-containing protein family (JMJD) remove methyl groups from arginine residues in histones, utilizing α-KG and Fe2+ as co-factors [17], thereby controlling tissue integrity. JMJD5 and JMJD7 preferentially demethylate arginine residues on histones H2, H3, and H4 [18]. While the role of Jmjd6 in histone arginine methylation and glutamate metabolism has been studied in tissue remodeling [19, 20] and tumor growth [21], the specific function of histone arginine demethylases in bone metabolism remains largely unexplored.
In this study, we utilized osteoblast-specific Jmjd7 knockout mice to investigate the role of Jmjd7 in bone metabolism. We aimed to characterize the effects of Jmjd7 loss on bone tissue and metabolic programming in osteogenic cells, revealing new insights into the epigenetic regulation of bone homeostasis.
Results
Jmjd7 loss induces cranial suture defect and neonatal mortality
To investigate the impact of Jmjd7 loss on bone integrity, we generated osteoblast-specific conditional Jmjd7 knockout mice (Jmjd7KO). This was achieved by crossing mice carrying loxP-flanked exons 2-8 of Jmjd7 (Jmjd7loxP) with mice carrying Cre recombinase under the control of the 2.3-kb collagen 1a1 promoter (Col1a1Cre), resulting in tamoxifen-inducible Col1a1Cre-Jmjd7loxP offspring. At three days of age, Col1a1Cre-Jmjd7loxP mice received a single dose of 75 mg/kg tamoxifen to induce Jmjd7KO mice, while tamoxifen-injected Jmjd7loxP littermates served as wild-type controls. Genotyping and verification of bone-specific Jmjd7 deletion were conducted in Jmjd7KO mice (Supplementary Fig. S1).
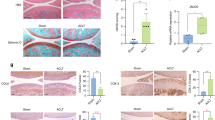
Successful deletion of Jmjd7 was verified by the absence of Jmjd7 mRNA expression and protein levels (Fig. 1a) in osteoblasts of Jmjd7KO mice. Notably, this deletion led to a reduction in histone demethylase activity and an increase in H3R2me1 and H3K27me3 levels (Fig. 1b), indicating alterations in histone methylation patterns. Micro-CT radiography revealed that Jmjd7KO animals exhibited signs of delayed cranial suture closure. Furthermore, these knockout mice exhibited a significantly shortened lifespan, with the majority dying around 3 weeks of age, in sharp contrast to wild-type littermates, which survived throughout the duration of the study (Fig. 1c).
Decreased Jmjd7 mRNA expression and protein levels (a) and histone demethylase activity, but increased H3R2me1 and H3K27me3 levels (b) in Jmjd7KO mice. Micro-CT images showing delayed cranial suture fusion in neonatal Jmjd7KO mice (c); scale bars, 6 mm (top view), 9 mm (lateral view). Analysis of bone microstructure by μCT (d); similar Tb.BMD, BV/TV, Tb.N, and SMI in 12-week-old male Jmjd7KO mice as compared to wild-type mice (sagittal and axial reconstructions, scale bar, 300 μm). Reduced body size and body weight (e; scale bar, 25 mm) with a sparser bone microarchitecture (scale bar, 300 μm) (f) as well as decreases in Tb.BMD, BV/TV, Tb.N, and Tb.Th (g) in 12-week-old female Jmjd7KO mice. Marked differences in visceral (yellow) and subcutaneous (blue) fat deposition (scale bar, 2.5 cm) (h) and reduced expression of the sex-dimorphism regulator Utx (i) in 12-week-old female Jmjd7KO mice. Lower lumbar spine BMD (j) and diminished JMJD7 expression in osteoblasts (k; scale bar, 30 μm) of human osteoporotic bone. JMJD7 levels positively correlated with lumbar spine BMD (l). Data are shown as mean ± SEM and analyzed using Mann-Whitney test. Asterisks (*) denote statistically significant differences (P < 0.05). WT wild-type, cKO conditional Jmjd7 knockout mice, Ctrl control, Op osteoporosis.
Jmjd7 deletion represses bone mass in female mice
To investigate the effect of Jmjd7 loss on bone mass in adult mice, we induced Jmjd7 knockout in six-week-old Col1a1Cre-Jmjd7loxP and Jmjd7loxP (wild-type, WT) mice via tamoxifen injections at 75 mg/kg/day for five consecutive days. Jmjd7 deletion was confirmed one week after the final injection, and the animals were subsequently monitored until 12 weeks of age. In male mice, analysis of trabecular bone microarchitecture revealed no significant differences between Jmjd7KO and WT mice. Key parameters including trabecular bone mineral density (Tb.BMD), trabecular volume (BV/TV), trabecular number (Tb.N), and structure model index (SMI) were comparable between the two groups (Fig. 1d). Interestingly, female Jmjd7KO mice exhibited distinct phenotype changes. These animals developed a smaller stature and maintained lower body weights compared to female WT mice throughout the study period (Fig. 1e). Micro-CT analysis of bone microstructure in female Jmjd7KO mice revealed a significantly sparser trabecular bone network (Fig. 1f), characterized by marked decreases in Tb.BMD, BV/TV, Tb.N, and trabecular thickness (Tb.Th) (Fig. 1g). Furthermore, female Jmjd7KO mice displayed pronounced adiposity, with substantially larger volumes of subcutaneous and abdominal fat compared to sex- and age-matched WT mice (Fig. 1h).
Serum estrogen, estradiol, and testosterone levels varied according to sex; however, these sex hormone levels were comparable between sex-matched Jmjd7KO and WT mice (Supplementary Fig. S2). The X-linked H3K4 demethylase Kdm5c [22] and the H3K27 demethylase Utx [23] are involved in sex-specific tissue metabolism. Utx also plays a crucial role maintaining in bone integrity [24] and regulating adipose tissue metabolism [25]. Our data revealed a correlation between sex-dependent bone loss in Jmjd7KO mice and Utx expression. Specifically, female Jmjd7KO mice, but not males, exhibited decreased Utx expression in bone tissue as compared to their sex-matched WT counterparts. In contrast, Kdm5c expression was similar between female and male Jmjd7KO and WT mice (Fig. 1i).
Human osteoporotic bones are JMJD7-depleted
To investigate the potential role of JMJD7 in human osteoporosis, we analyzed bone specimens obtained from patients undergoing spinal surgery, comparing those with and without osteoporosis. Bone mineral density measurements of L2-L5 lumbar spine revealed significantly lower values in patients with a history of osteoporosis compared to control subjects (Fig. 1j). Immunohistochemical analysis of osteoblasts adjacent to osteoporotic bone demonstrated relatively low JMJD7 protein levels (Fig. 1k). Importantly, JMJD7 levels in osteoblasts showed a positive correlation with spine BMD. Receiver operating characteristics (ROC) curve analysis indicated that JMJD7 levels could effectively distinguish between osteoporotic and non-osteoporotic bone specimens (Fig. 1l).
Jmjd7 regulated osteoporosis development in ovariectomized mice
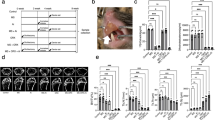
To investigate the role of Jmjd7 in osteoporosis development, we utilized a model of estrogen deficiency-induced osteoporosis. Six-week-old female Jmjd7loxP and Col1a1Cre-Jmjd7loxP mice were administered tamoxifen at 75 mg/kg/day for five consecutive days to induce Jmjd7 knockout, which was confirmed at eight weeks of age. Bilateral ovariectomy was performed on both Jmjd7KO and WT mice 12 weeks of age (Fig. 2a). At the end of the experiment, ovariectomized Jmjd7KO mice exhibited a sustained loss of Jmjd7 expression compared to ovariectomized WT mice (Fig. 2a). Micro-CT radiography revealed that ovariectomy induced a sparse bone microstructure in WT mice. However, this effect was significantly exacerbated in Jmjd7KO mice, which exhibited a more pronounced loss of trabecular bone network compared to ovariectomized WT mice (Fig. 2b). Quantitative analysis confirmed that ovariectomized Jmjd7KO mice experienced greater reductions in Tb.BMD, BV/TV, Tb.Th, and Tb.N and displayed greater enhancement in Tb.Sp and SMI (Fig. 2c). Interestingly, the impact on cortical bone parameters, including cortical BMD (Ct.BMD) and thickness (Ct.Th), was similar between the two groups (Fig. 2d). Notably, ovariectomized Jmjd7KO mice developed kyphosis-like deformities (Fig. 2e) characterized by significantly larger Cobb angles of the thoracic spine compared to ovariectomized WT mice (Fig. 2f).
Schematic study design and confirmation of persistent Jmjd7 loss in sham and OVX Jmjd7KO mice (a). Micro-CT images showing severe trabecular bone loss in OVX Jmjd7KO mice (b; scale bar, 350 μm). Significant reductions in Tb.BMD, Tb.Th, BV/TV, and Tb.N and increases in Tb.Sp and SMI in OVX Jmjd7KO and mice (c). No significant changes in Ct.BMD and Ct.Th between groups (d). OVX Jmjd7KO mice displayed kyphotic spinal curvature (scale bar of left panels, 3 cm; scale bar of right panels, 1 cm) (e). Increased thoracic Cobb angles in OVX Jmjd7KO vs. OVX WT mice (f). Lentiviral Jmjd7 overexpression in OVX mice restored high Jmjd7 and reduced H3R2me1 levels in osteoblasts (g). Forced Jmjd7 expression preserved trabecular bone structure and mitigated estrogen deficiency-induced loss of Tb.BMD and BV/TV (h; scale bar, 300 μm). Data are shown as mean ± SEM from 5 to 7 mice analyzed by ANOVA test and Bonferroni post hoc test. Similarly, Jmjd7 lentivirus treatment enhanced trabecular bone network and attenuated Jmjd7KO-induced bone loss (i; scale bar, 300 μm). Data are shown as mean ± SEM from 3 or 4 mice analyzed by Mann-Whitney test. Asterisks (*) denote statistically significant differences (P < 0.05).
To investigate the effect of Jmjd7 on estrogen deficiency-induced bone loss, we administered lentivirus encoding Jmjd7 to 12-week-old ovariectomized mice via tail vein injection one week postoperatively. IVIS imaging confirmed successful transduction in bone tissue one week after systemic injection, as evidenced by GFP fluorescence and expression of reporter eGFP and Jmjd7 in bone-marrow mesenchymal cells (Supplementary Fig. S3). Forced Jmjd7 expression inhibited H3R2me1 levels in osteoblasts (Fig. 2g). Skeleton in Jmjd7-treated ovariectomized mice exhibited relatively more interwoven trabeculae microarchitecture compared to mock-treated ovariectomized mice eight weeks after treatment. Gain of Jmjd7 function mitigated estrogen deficiency-induced loss of BMD and BV/TV (Fig. 2h), suggesting that Jmjd7 function in osteoblasts played an important role in keeping bone integrity during osteoporosis. Furthermore, lentivirus Jmjd7-treated 12-week-old female Jmjd7KO mice exhibited more spacious trabecular bone network with higher Tb.BMD and BV/TV than mock-treated Jmjd7KO mice eight weeks after treatment (Fig. 2i).
Jmjd7 deletion dysregulates bone formation and osteoclastic resorption
Histomorphometric analyses of bone tissue in Jmjd7KO mice revealed features indicative of low dynamic bone formation, characterized by decreased mineral apposition rate (MAR) and bone formation rate (BFR/BS) (Fig. 3a). Static bone histomorphology assessment demonstrated that Jmjd7 loss resulted in significantly reduced trabecular bone volume (Fig. 3b), fewer osteocalcin-immunostained osteoblasts (Ob.N/mm), and an increased number of TRAP-stained osteoclasts (Oc.N/mm) (Fig. 3c, d).
Reduced calcein double-label fluorescence indicates decreased mineral apposition rate (MAR) and bone formation rate (BFR/BS) in Jmjd7KO vs. WT mice (a, scale bar, 20 μm). Von Kossa staining revealed reduced trabecular bone area and BV/TV in Jmjd7KO mice (b, scale bar, 2 mm). Representative images show hematoxylin/eosin (H&E) staining, osteocalcin immunostaining, and TRAP histochemical staining of bone tissue (c, scale bar, 75 μm). Jmjd7 loss decreased osteoblast numbers (Ob.N) and increased osteoclast numbers (Oc.N) (d). Downregulation of osteogenic genes post-Jmjd7 deletion (e). Reduced mineralized matrix formation in Jmjd7KO mice (f; scale bar, 5 mm). Elevated Tnfsf11 expression in BMSCs from Jmjd7KO mice (g) and increased marrow adipocytes (h; scale bar, 20 μm). Enhanced TRAP-positive osteoclast formation and osteoclast marker expression of bone-marrow macrophage progenitors (i; scale bar, 20 μm). Elevated Utx protein (j) and increased osteogenic marker expression and mineralized matrix synthesis in Utx-treated female Jmjd7KO osteoblasts (k; scale bar, 5 mm). Data are presented as mean ± SEM from 4 to 6 mice analyzed by Mann-Whitney test and ANOVA test with Bonferroni post hoc test. Asterisks (*) denote statistically significant differences (P < 0.05).
In vitro studies further corroborated these findings. Bone-marrow mesenchymal cells from Jmjd7KO mice exhibited impaired osteogenic differentiation capacity compared to WT counterparts. This was evidenced by reduced expression of key osteogenic markers, including Runx2, osterix (Sp7), bone alkaline phosphatase (Alpl), collagen 1 (Col1a1), and osteocalcin (Bglap) (Fig. 3e), as well as diminished von Kossa-stained mineralized matrix formation (Fig. 3f). Conversely, Jmjd7KO bone-marrow stromal cells displayed significantly higher expression of crucial osteoclastogenic cytokine Tnfsf11 (Fig. 3g) and enhanced differentiation into Nile red-stained adipocytes (Fig. 3h) compared to WT cells.
Bone-marrow macrophages from Jmjd7KO mice exhibited increased osteoclastogenic capacity. This was characterized by increased TRAP-stained osteoclast formation (Fig. 3i) and markedly elevated expression of key osteoclast markers - nuclear factor of activated T cells 1 (Nfatc1), cathepsin K (Ctsk), matrix metalloproteinase 9 (Mmp9), and carbonic anhydrase 2 (Car2) (Fig. 3i).
Moreover, female Jmjd7KO bone-marrow mesenchymal cells displayed reduced expression of estrogen receptor-α, Utx, osteogenic markers and the osteogenic factors Igf2 and Wnt3a, while showing increased expression of the bone formation inhibiting factor sclerostin, as compared to male Jmjd7KO mice. In contrast, male Jmjd7KO mice exhibited bone regulator expression similar to their wild-type counterparts (Supplementary Fig. S2).
Given that female Jmjd7KO mice exhibited reduced X-linked Utx expression, we examined its contribution to osteogenic activity. Lentiviral Utx overexpression (Fig. 3j) restored osteogenic marker expression and rescued von Kossa-stained mineralized matrix biosynthesis in bone-marrow stromal cells from female Jmjd7KO mice (Fig. 3k).
Jmjd7 deletion alters gene expression profiles, and energy metabolism in osteoblasts
To investigate the effects of Jmjd7 loss on osteoblastic activity, we performed RNA sequencing analysis on calvaria osteoblasts from eight-week-old WT and Jmjd7KO mice, confirming reduced Jmjd7 expression in the KO cells (Supplementary Fig. S3). Principal component analysis revealed distinct clustering of gene expression profiles between Jmjd7KO and WT osteoblasts (Fig. 4a). Volcano plot (Fig. 4b) and heatmap (Fig. 4c) analyses identified 519 differentially expressed genes, which were associated with plenty of biological processes. Gene ontology pathway analysis revealed downregulation of processes related to osteoblast differentiation, bone mineralization, chromatin remodeling, and DNA demethylation (Fig. 4d). Conversely, processes involved in glycolytic, ATP metabolism, fatty acid biosynthesis, and inflammatory cytokine production were upregulated (Fig. 4d). KEGG pathway analysis further demonstrated that these differentially expressed genes contributed to multiple metabolic activities, including fatty acid, pyruvate, carbon metabolism, glycolysis, and oxidative phosphorylation (Fig. 4e). Additionally, gene enrichment set analysis uncovered that Jmjd7 knockout led to suppression of mitochondrial bioenergetics (Fig. 4f).
Principal component analysis showing distinct clustering of gene expression profiles (a). Volcano plot depicting 519 differentially expressed genes (b), with corresponding heatmap of changes (c). Gene ontology analysis reveals enrichment in osteoblast differentiation, bone mineralization, chromatin remodeling, glycolysis, and cytokine signaling (d). KEGG pathway analysis indicates involvement in carbon metabolism, fatty acid metabolism, and OXPHOS (e). GESA demonstrates reduced mitochondrial metabolic activity in Jmjd7KO osteoblasts (f). RNA sequencing was performed on osteoblasts from three Jmjd7KO and WT mice.
Loss of Jmjd7 function represses mitochondrial energy metabolism
Our analyses revealed significant alterations in energy metabolism in Jmjd7KO osteoblasts. Jmjd7KO osteoblasts exhibited enhanced glycolytic activity, as evidenced by elevated extracellular acidification rates (Fig. 5a), including glycolysis, glycolytic capacity, and glycolytic reserve (Fig. 5b). This was further corroborated by increased lactate production (Fig. 5c). Conversely, mitochondrial function was markedly impaired in Jmjd7KO osteoblasts. We observed (1) reduced mitochondrial respiration capacity (Fig. 5d), (2) decreased basal, maximal, spare, and ATP-linked oxygen consumption rates (Fig. 5e), (3) downregulation of key enzymes involved in mitochondrial oxidative phosphorylation (OXPHOS) (Fig. 5f), and (4) diminished complex I activity (Fig. 5g). The loss of Jmjd7 function resulted in a significant reduction in total ATP production, primarily due to suppressed mitochondrial ATP synthesis, partially offset by a modest increase in glycolysis-derived ATP (Fig. 5h).
Increased extracellular acidification rates (ECAR) (a), enhanced glycolytic parameters (b) and lactate production (c), but decreased oxygen consumption rate (OCR) in Jmjd7KO osteoblasts (d). Lower basal, maximal, spare, and ATP-linked respirations (e) and reduced key OXPHOS enzymes levels in Jmjd7KO compared to WT (f). Decreased complex I activity (g) and ATP production (h). Altered TCA enzyme activities of Pdh, Kgdh, Sdh, and Idh in Jmjd7KO osteoblasts (i). Reduced nuclear and cytosolic Idh protein levels (j) and decreased α-KG production (k). Molecular weight rulers are indicated in kDa on right. Data are presented as mean ± SEM from 5 to 7 mice analyzed by Mann-Whitney test. Asterisks (*) denote statistically significant differences (P < 0.05).
Jmjd7 deletion further caused differential effects on key TCA cycle enzymes, including (1) increased pyruvate dehydrogenase activity and (2) decreased activities of succinate dehydrogenase, ketoglutarate dehydrogenase, and isocitrate dehydrogenase (Idh) (Fig. 5i). Notably, Idh, a crucial enzyme that converts isocitrate into α-ketoglutarate (α-KG), was significantly affected. α-KG serves as an indispensable co-factor, working in concert with Fe2+ to facilitate the removal of methyl groups in histones by the JMJD family [17]. Our data demonstrated that Jmjd7 loss resulted in reduced Idh levels in both cytosolic and nuclear fractions (Fig. 5j). Consequently, α-KG production in Jmjd7KO osteoblasts was substantially lower than in WT osteoblasts (Fig. 5k).
Jmjd7 regulates transcription and post-translational modification of Idh and Runx2
To characterize the H3R2me1-enriched epigenome that may impair osteoblast anabolism, we performed chromatin immunoprecipitation sequencing (ChIP-seq) on WT and Jmjd7KO osteoblasts (Supplementary Fig. S4). Meta-profile analysis identified a pronounced enrichment of H3R2me1 around 5 kb upstream and downstream of transcription start sites in Jmjd7KO osteoblasts (Fig. 6a). A Venn diagram showed 1,801 and 986 genes enriched for H3R2me1 in WT and Jmjd7KO cells, respectively (Fig. 6b). Gene ontology (GO) analysis indicated involvement in processes such as cellular respiration, glucose metabolism, fatty acid metabolism, osteoclast differentiation, IGF receptor signaling, and bone formation (Fig. 6c). Furthermore, KEGG pathway analysis highlighted pathways related to oxidative phosphorylation, PTH production, Wnt signaling, cellular senescence, and carbohydrate metabolism (Fig. 6d). HOMER motif enrichment analysis identified transcription factors associated with H3R2me1-enriched regions that play key roles in cartilage and bone metabolism (Fig. 6e), notably targeting Osr2 [26] and the osteogenic transcription factor Runx2. Loss of Jmjd7 resulted in increased H3R2me1 enrichment at Runx2-associated regulatory regions (Fig. 6e), suggesting epigenetic repression of this key transcription factor.
Genome-wide H3R2me1 metaprofiles ± 5 kb around TSS (a) and Venn diagram showing overlap enriched genes (b). GO and KEGG analyses link altered gene expression to bone formation, metabolism, and demethylation (c, d). HOMER motif analysis shows H3R2me1-enriched transcription factors, including Osr2, and Runx2 (e). Increased H3R2me1 and H3K27me3 levels upon Jmjd7 deletion (f). Enhanced H3R2me1/H3K27me3 enrichment at Idh and Runx2 promoters upon Jmjd7 loss (g) correlates with suppressed mRNA expression (h). Jmjd7 deletion enhanced arginine methylation and ubiquitination of Idh and Runx2, respectively (i, j), while α-KG stabilizes Idh and Runx2 proteins, increases Bglap and Col1a1 mRNA, and promotes mineralized matrix production in Jmjd7KO osteoblasts (k). Volcano plot showing differentially expressed genes (e.g., Igf1, Bmp2, and Wnt3a) between vehicle- and α-KG-treated Jmjd7KO osteoblasts (l). KEGG pathway of α-KG-induced gene expression profiles highlighting intracellular pathway such as Igf binding, osteoblast differentiation, Bmp signaling, collagen matrix formation, and angiogenesis (m). JMJD7-IN-1 and DMOG repressed histone arginine methylase activity (n) and enhanced H3R2me1 levels (o) and impaired Col1a1 and Bglap expression (p) in lentivirus Jmjd7-treated osteoblasts. ChIP-seq and RNA sequencing were performed from 3 independent experiments. Data are presented as mean ± SEM from 4 or 5 independent experiments analyzed by ANOVA test and Bonferroni post hoc test. Asterisks (*) denote statistically significant differences (P < 0.05).
We investigated how Jmjd7 loss led to Idh reduction, inducing α-KG underproduction and osteoblast dysfunction. Epigenetic analyses demonstrated increased H3R2 mono-methylation and H3K27 trimethylation in Jmjd7KO osteoblasts (Fig. 6f). Chromatin immunoprecipitation studies revealed increased enrichment of H3R2me1 and H3K27me3 marks at the promoter regions of Idh and Runx2 (Fig. 6g), which correlated with reduced mRNA expression of these genes (Fig. 6h). Notably, α-KG supplementation reversed these epigenetic alterations and restored gene expression, accompanied by upregulation of Jmjd5 and Jmjd6. Pharmacological inhibition of Jmjd6 activity with WL12 attenuated the α-KG-induced reduction of H3R2 mono-methylation and osteogenic marker expression in Jmjd7KO osteoblasts (Supplementary Fig. S5).
In line with the association between non-histone protein arginine methylation and the osteogenic capacity of mesenchymal progenitor cells [27], further investigation uncovered that Jmjd7 deletion resulted in decreased protein levels of Idh and Runx2. This decrease was accompanied by increased methyl-arginine and ubiquitin modifications detected in their immunoprecipitated complexes (Fig. 6i, j). Here, α-KG supplementation mitigated these effects (Supplementary Fig. S5), improving the expression of osteogenic markers and enhancing extracellular matrix mineralization (Fig. 6k).
Transcriptomic analyses of vehicle-treated and α-KG-supplemented Jmjd7KO osteoblasts revealed significant alterations, with 564 upregulated and 471 downregulated genes (Fig. 6l). GO analyses further identified enrichment in pathways related to IGF binding, osteoblast differentiation, BMP signaling, extracellular matrix collagen composition, and angiogenesis (Fig. 6m).
Additionally, inhibition of Jmjd7 activity by JMJD7-IN-1 and blockage of 2-oxyglutarate-dependant dioxygenase activity by DMOG decreased histone arginine demethylase activity (Fig. 6n) and increased H3R2me1 levels. These changes were accompanied by reduced protein levels of Runx2 and Idh (Fig. 6o) and lower expression of Col1a1 and Bglap (Fig. 6p) in lentivirus-mediated Jmjd7-treated osteoblasts.
α-KG supplementation attenuates osteoporosis progression in Jmjd7KO mice
Given the observed improvement in mineralized matrix production in Jmjd7KO osteoblasts following α-KG supplementation, we investigated its potential to mitigate bone mass loss in Jmjd7KO mice. Experimental cohorts were administered 0.25% or 0.75% α-KG in drinking water (pH 7.2) ad libitum. α-KG-supplemented mice exhibited significantly elevated serum α-KG concentrations three days after treatment initiation, while returned to vehicle-treated levels three days after the supplementation ceased. Metabolic analyses revealed that α-KG-supplemented mice had lower overall energy expenditure, while their feed and water intake remained comparable to those of vehicle-treated control mice (Supplementary Fig. S6).
Treating Jmjd7KO mice with α-KG for 12 weeks did not affect feed and water intake, body weight or serum biochemistry parameters, including glucose, GOT, cholesterol, BUN, and creatinine, relative to controls (Supplementary Fig. S7). Supplementation with α-KG significantly ameliorated Jmjd7 deletion-induced alterations in serum α-KG and bone turnover markers, reducing the loss of osteocalcin (bone formation marker) and attenuating the elevation of CTX-1 (bone resorption marker) (Fig. 7a). Furthermore, α-KG-supplemented Jmjd7KO mice exhibited improved trabecular bone microarchitecture (Fig. 7b), with attenuated reductions in BMD, BV/TV, Tb.Th, and Tb.N compared to vehicle-fed Jmjd7KO mice (Fig. 7c).
α-KG supplementation improved serum α-KG, osteocalcin and CTX-1 levels (a). Micro-CT images demonstrating marked preservation of trabecular bone structure in α-KG-fed mice (b; scale bar, 400 μm). Enhanced Tb.BMD, BV/TV, Tb.Th, and Tb.N (c). Calcein labeling, H&E, and TRAP histochemical staining demonstrated improved bone tissue quality (d; scale bar, 30 μm), while α-KG supplementation reversed decreased BFR/BS and suppressed Oc.N (e), enhanced mineralized matrix synthesis by bone-marrow mesenchymal cells (f; scale bar, 5 mm), and reduced osteoclast numbers from bone-marrow macrophage precursor cells (g; scale bar, 20 μm). Data are presented as mean ± SEM from 5 to 7 animals analyzed by ANOVA test with Bonferroni post hoc test. Asterisks (*) denote statistically significant differences (P < 0.05).
Histomorphometric analysis demonstrated that α-KG supplementation reversed impairments in calcein-labelled bone mineral apposition and mitigated the increase in TRAP-stained osteoclasts (Fig. 7d). This resulted in attenuation of BFR/BS loss and normalization of Oc.N within the skeletal microenvironment (Fig. 7e).
Ex vivo analyses corroborated these findings, showing that α-KG preserved the osteogenic capacity of bone-marrow mesenchymal cells, as evidenced by enhanced mineralized matrix synthesis and expression of osteogenic markers (Fig. 7f). Additionally, α-KG supplementation reduced osteoclast formation from bone-marrow macrophage precursor cells in Jmjd7KO mice (Fig. 7g).
α-KG attenuates estrogen deficiency-induced bone remodeling
The promising effects of α-KG supplementation on bone integrity in Jmjd7KO mice prompted us to investigate its potential in mitigating estrogen deficiency-induced bone deterioration. Ovariectomized mice were administered 0.75% α-KG in drinking water (pH 7.2) ad libitum for 8 weeks. Feed and water intake, body weight and serum biochemical parameters were similar in α-KG- and vehicle-treated ovariectomized mice (Supplementary Fig. S7). Interestingly, α-KG supplementation reduced Jmjd7 loss in bone tissue (Fig. 8a) and serum α-KG loss and CTX-1 overproduction (Fig. 8b) in ovariectomized mice. α-KG-treated ovariectomized mice exhibited improved trabecular bone microstructure (Fig. 8c) with higher Tb.BMD, BV/TV, Tb.Th, Tb.N (Fig. 8d). Additionally, Ct.BMD and Ct.Th were increased compared to vehicle-treated ovariectomized mice (Fig. 8e).
α-KG supplementation restored Jmjd7 levels (a) and elevated serum α-KG while normalizing CTX-1 in OVX mice (b). Micro-CT reveals reduced trabecular bone loss (c; scale bar, 400 μm); α-KG-treatment enhanced Tb.BMD, BV/TV, Tb.Th, Tb.N (d), Ct.BMD and Ct.Th (e). Robust calcein-labeled bone mineralization and reduced TRAP-positive osteoclasts in α-KG-fed mice (f; scale bar, 50 μm). Elevated bone formation rate and reduced osteoclast overburden (g). Enhanced mineralized matrix production and osteogenic gene expression in bone-marrow stromal cells (h; scale bar, 5 mm). Inhibited osteoclast formation and osteoclastogenic gene expression in bone-marrow macrophages (i; scale bar, 20 μm). Data are presented as mean ± SEM from 5 to 7 animals and analyzed by ANOVA test with Bonferroni post hoc test. Asterisks (*) denote statistically significant differences (P < 0.05).
α-KG supplementation ameliorated the estrogen deficiency-induced loss of bone mineral acquisition and attenuated osteoclast formation (Fig. 8f). This was evidenced by improvements in BFR/BS and a reduction in Oc.N (Fig. 8g). Ex vivo studies showed that α-KG treatment reversed the underproduction of mineralized matrix and loss of osteogenic marker expression in bone-marrow mesenchymal cells from ovariectomized mice (Fig. 8h). Concurrently, α-KG supplementation reduced osteoclast formation and suppressed expression of key osteoclast markers in these animals (Fig. 8i).
Discussion
Epigenetic regulation, particularly histone methylation, has been increasingly recognized as a critical factor in bone homeostasis and the osteoporotic skeleton. Previous studies have suggested that histone arginine methylation by protein arginine methyltransferases enhances osteogenic differentiation of mesenchymal stem cells and bone formation [13, 14]. Our study builds upon this foundation, revealing for the first time the loss of JMJD7, a histone arginine demethylase, in human osteoporotic bone biopsies. To further investigate this finding, we generated an osteoblast-specific Jmjd7 knockout mouse model, uncovering novel anabolic actions of methyl H3R2 eraser, in synergy with the TCA cycle intermediate α-KG, on osteoblastic activity and bone mass homeostasis.
Our data demonstrate the essential role of Jmjd7 in osteoblasts for proper bone development. Neonatal deletion of this epigenetic histone modifier resulted in a prominent feature of delayed closure around the cranial suture. This observation aligns with other studies highlighting the importance of protein arginine methyltransferases (Prmts) in cranial and limb integrity. For example, cranial neural crest-specific Prmt1 knockout mice exhibit cranial bone underdevelopment [28], while mice deficient in Prmt5 specifically in cartilage showed reduced bone volume in long bones compared to wild-type mice [29].
Intriguingly, Jmjd7 loss caused a sex-specific inhibition of bone integrity in adult mice, with osteoporosis-like features occurring solely in female Jmjd7KO animals. This female-dependent bone deterioration presumably results from the loss of X-linked H3K27me3 demethylase Utx, a key regulator for sex dimorphism [24, 30]. Notably, Utx loss in cranial progenitor cells led to cranial bone underdevelopment predominantly in female animals [31]. The observed increase in osteogenic marker expression in lentivirus Utx-treated female Jmjd7KO bone-marrow stromal cells corroborated Utx’s role in female-dependent bone loss. Furthermore, the female Jmjd7KO mice exhibited a plethora of dysregulated bone gain and remodeling reactions, including low mineral acquisition, reduced osteoblast numbers, impaired osteogenic differentiation capacity, and increased osteoclast formation. These factors collectively explain the presence of low bone mass and porous microstructure in these animals.
In addition to skeletal abnormalities, Jmjd7KO mice developed high adiposity in both visceral and subcutaneous compartments. This observation is consistent with growing evidence showing that histone arginine hypermethylation enhances adipocyte formation [32] and promotes osteoblast dysfunction [33]. Furthermore, Utx interference promotes the differentiation of mesenchymal stem cell toward adipocytes [34]. Osteoclastogenic cytokine Rankl (encoded by Tnfsf11) has been shown to promote whole-body energy metabolism and fat accumulation [35]. Given Utx loss and high Tnfsf11 expression, and increased adipocyte formation capacity in the Jmjd7KO mouse bone microenvironment, the bone-fat crosstalk in the osteoporotic skeleton may contribute to the increased body adiposity observed in these knockout mice. Consequently, these combined bone catabolic effects culminated in a severe osteoporosis condition with kyphosis-like features upon estrogen deficiency.
To elucidate the molecular mechanisms underlying the suppression of bone mass loss, we characterized the transcriptomic landscape changes between Jmjd7KO and WT osteoblasts. GO and KEGG pathway analyses revealed major gene expression changes enriched in various cellular and metabolic activities, including osteoblast differentiation, chromatin remodeling, glycolysis, and mitochondrial energy metabolism. In vitro models uncovered a bioenergetic programming shift in Jmjd7KO osteoblasts from mitochondrial OXPHOS toward compensatory anaerobic glycolysis.
While glycolysis typically accounts for energy production during nitric oxide-dependent osteogenic lineage specification of bone-marrow stromal cells [36] and Lgr4-mediated anabolic activity of preosteoblasts [37], the mitochondrial TCA cycle coupled with glycolysis is necessary to fuel efficient osteoblast differentiation [38]. Moreover, inhibition of glycolysis has been shown to enhance mitochondrial OXPHOS for osteoblastic activity, mitigating age-induced bone loss [39]. Conversely, impaired mitochondrial OXPHOS blocks osteogenic differentiation of skeletal progenitor cells [40]. Together, these findings suggest that the bioenergetic programming shift caused by Jmjd7 deletion leads to insufficient overall ATP production, which may fail to meet the energy demands required for osteoblast anabolism.
α-KG emerged as a functional molecule in the Jmjd7 regulation of mitochondrial energetics and osteoblastic activity. This TCA cycle intermediate, produced by the key enzyme IDH [41], is also required by the JMJD family histone demethylases to remove methylated groups from histones [42]. To demonstrate an α-KG dependent anabolic activity of Jmjd7KO osteoblasts, we investigated osteogenic transcription factors Runx2 and Osterix (Sp7), along with crucial bone proteins alpha-1 type I collagen (encoded by Col1a1), osteocalcin (encoded by Bglap), and von Kossa-stained extracellular matrix mineral deposits. These transcription factors regulate phenotypes of osteogenic cell lineages [43, 44], and cranial bone underdevelopment is observed in mice deficient in Runx2 [45], which may explain the delayed cranial suture closure seen in neonatal Jmjd7KO mice.
Furthermore, α-KG promoted transcriptomic changes, including upregulation of Igf1, Bmp2, and Wnt3a in Jmjd7KO osteoblasts. Of note, α-KG supplementation confirmed protective effects on the synthesis of mineralized extracellular matrices, consistent with reduced osteoporosis development in both Jmjd7KO mice and ovariectomized mice. The beneficial effects of α-KG on bone integrity align with previous studies demonstrating reduced bone loss in aged mice upon α-KG supplementation [46, 47]. We identified that Jmjd7 removed the methyl group from H3R2 via 2-oxoglutarate-dependent dioxygenase activity. α-KG supplementation repressed H3R2 methylation in Jmjd7KO osteoblasts by promoting Jmjd6. Previous studies demonstrate that competitive inhibition of α-KG-dependent dioxygenase represses vitamin C-induced histone demethylation critical for osteogenic differentiation [48]. Additionally, Jmjd6 catalyzes the demethylation of H3R2 [49]. Our data thus provide the first evidence of Jmjd7’s regulatory actions on α-KG and H3R2 in bone tissue metabolism.
We characterized H3R2me1-enriched epigenomic profiles that contributed to a range of biological activities, including cell respiration, osteoclast differentiation, and bone formation, Wnt signaling pathway, and OXPHOS. The osteogenic transcription factor Runx2 was, among the prominent targets. These findings further corroborated the features of energy dysmetabolism and loss of osteogenic capacity in Jmjd7KO osteoblasts. We identified two intracellular pathways through which Jmjd7 controls of Idh and Runx2 function. Jmjd7 loss led to Idh and Runx2 transcription repression by promoting methyl H3R2 enrichment at their respective promoter regions and increased arginine methylation of these two proteins, ultimately targeting them for ubiquitination. To the best of our knowledge, no prior studies have examined the effect of histone arginine demethylase on Runx2 stability in bone tissue. However, loss of protein arginine methyltransferase function has been reported to inhibit Runx2 signaling during osteogenic differentiation of bone-marrow mesenchymal cells [50] and calcified matrix production in vascular cells [51]. Additionally, protein arginine methyltransferases, alongside methylated arginine-containing proteins, are upregulated during Rankl-induced osteoclast differentiation of bone-marrow macrophage precursor cells [16] and adipocyte formation from myogenic progenitor cells [52].
In summary, our investigations shed light on novel bone regulatory actions of Jmjd7, which, at least in part, controls histone and non-histone arginine methylation critical for mineralized matrix synthesis in osteoblasts. Loss of Jmjd7 function in osteoblasts appears to promote methylated H3R2-dependent transcriptional repression and arginine methylation of Idh and Runx2, exerting deleterious effects on bone formation by impairing TCA cycle intermediate α-KG-dependent mitochondrial energy metabolism, while enhancing anaerobic glycolysis. This study further highlights the potential for developing osteoporosis prevention strategies by manipulating Jmjd7 function or α-KG action.
Materials and Methods
Human bone specimens
Protocols for clinical leftover specimen use were reviewed and approved by the Institutional Review Board, Chang Gung Medical Foundation (Affidavit #201701826B0 and #202002225B0). With informed consent before surgery, patients (n = 13; 11 females and 2 males; age 71.6 ± 3 years) with history of osteoporosis who required lumbar spine decompression, vertebroplasty, and fixation and the discectomy for spine fracture, degenerative spinal stenosis, and vertebral disc herniation were enrolled in the osteoporosis group (Op). Patients (n = 13; 7 females and 6 males; age 49.2 ± 3.2 years) without a history of osteoporosis who required spine surgery were enrolled in the control group (Ctrl). Bone mineral density of L2-L5 spine was quantified using dual energy X-ray absorptiometry, with a T-score < -2.5 of lumbar spine used to diagnose osteoporosis. Leftover bone specimens were obtained from osteotomized tissue during surgery.
Osteoblast-specific Jmjd7KO mice
Animal protocols were reviewed and approved by the IACUC of Kaohsiung Chang Gung Memorial Hospital (IACUC Affidavit #2020121201). All animals were group housed in a specific-pathogen free animal facility with sterile drinking water and chow diet provided ad libitum. C57BL/6 N mice carrying a loxP-flanked Jmjd7 construct spanning exons 2-8 (ENSMUSE00001270566–ENSMUSE00000344742; strain C57BL/6N-Jmjd7tm1aNarl/Narl; Jmjd7loxP; RMRC13335; National Center for Biomodels, Taiwan) were crossed with C57BL/6 mice expressing tamoxifen-inducible Cre recombinase under the control of a 2.3-kb collagen 1a1 promoter (JAX stock #016241; B6.Cg-Tg(Col1a1-cre/ERT2)1Crm/J; The Jackson Laboratory; Bar Harbor, ME) to generate Col1a1Cre-Jmjd7loxP mice. Genotyping was performed by PCR using primers for Cre and loxP sites (Supplementary Table 1). Neonatal experiments: To assess the role of Jmjd7 in bone integrity, three-day-old Col1a1Cre-Jmjd7loxP mice and their wild-type littermate controls carrying the Jmjd7loxP allele were injected with a single dose of tamoxifen (75 mg/kg; Aldrich-Sigma, St Louis, MO) and monitored until three weeks of age. Adult animals: In a separate set of experiments, six-week-old Col1a1Cre-Jmjd7loxP mice and littermate controls received daily tamoxifen injections (75 mg/kg) for five consecutive days, with genotyping confirmed one week after treatment. Animals were subsequently observed till 12 weeks of age. Sex- and age-matched animals were randomly grouped; and in vivo studies were performed by investigators blinded to animal genotypes and treatment conditions.
Ovariectomized mice
Six-week-old female Jmjd7loxp and Col1a1Cre-Jmjd7loxP mice were administered tamoxifen (75 mg tamoxifen/kg/day) for 5 consecutive days, and Jmjd7 deletion was confirmed at eight weeks of age. Twelve-week-old female C57BL/6 mice, Jmjd7KO mice, and WT mice were subjected to bilateral ovariectomy or sham surgery under isoflurane anesthesia. Eight weeks postoperatively, mice were euthanatized, and peripheral blood was collected via an intracardiac puncture, followed by harvesting of bone specimens.
In vivo lentivirus Jmjd7 treatment
Full-length Jmjd7 cDNA sequence was inserted into pMIF-cGFP-Zeo vectors. Packing vectors (pPACKF1; System Biosciences, Mountain View, CA) were co-transfected into 293 T cells to produce lentivirus suspension using Lipofectamine® 3000 (Therno Fisher Scientific, Waltham, MA). After ultracentrifugation and titration into 1010 infectious units/ml, lentivirus Jmjd7 or empty vector particles as mock were injected into 12-weeks-old ovariectomized C57BL/6 mice (109 infectious units/mice) via tail veins 1 week upon removing ovaries, as previously described [53]. Eight weeks postoperatively, mice were euthanatized, and bone tissue were dissected. In separate experiments, 12-week-old female Jmjd7KO and WT mice were tail vein injected lentivirus Jmjd7 or empty vector particles (109 infectious units/mice). Eight weeks after injection, animals were euthanatized, and bone tissue were dissected. Lentiviral tissue tropism was assessed by analyzing eGFP fluorescence in the lung, liver, kidney and bone of anesthetized mice using an IVIS Lumina II In Vivo Imaging System (PerkinElmer Inc, Waltham, MA) one week after injection. Fluorescence intensity was quantified following the manufacturer’s instructions.
In vivo α-KG supplementation
In sterile drinking water, α-ketoglutarate (α-KG; 75890; Aldrich-Sigma, St. Louis, MO) was mixed to prepare 0.25% and 0.75% α-KG solution (w/v; pH 7.2). One week after tamoxifen injection, female Jmjd7KO and WT mice were provided with α-KG solution and chow diet ad libitum for 12 weeks. Separately, ovariectomized C57BL/6 mice received 0.75% α-KG solution and chow diet ad libitum for eight weeks one week postoperatively. Animals’ metabolic profiles were investigated using Phenomaster (TSE System, Homburg, Germany), according to the manufacturer’s instruction. Mice were single housed and provided with feed and water ad libitum in metabolic cages of a controlled climate chamber (23 °C, 12 h dark/12 h light cycle) for 72 h. Oxygen consumption, carbon dioxide production, respiratory exchange ratio, heat production, feed and water intake were monitored and normalized with body weight using an in-house High Throughput Physiological Signaling Calculation Unit.
μCT analysis
Upon anesthesia, whole body scans were performed using μCT imaging platform (1176 Skyscane, Bruker, Billerica, MA) to capture radiographs with a pixel size of 35 × 35 × 35 μm. Image reconstruction and contouring were performed using SKYSCAN CT-Analysis software. Calibration and binarization of muscle and adipose tissue images were achieved with a Micro-CT Mouse Phantom (QRM-707137, Freiburg, Germany). The binarization thresholds of subcutaneous fat and visceral adipose were set between 75 and 95. Volumes of subcutaneous fat, visceral adipose tissue, and total tissue were calculated using the in-house software, as previously described [54]. For bone mass quantification and microstructural analysis, long bones and the spine were scanned to capture radiographs of 9 × 9 × 9 μm pixel size [53]. Specimen density was quantified using a calcium hydroxyapatite phantom (QRM-70134). Measurements included trabecular bone mineral density (Tb.BMD), volume (BV/TV), thickness (Tb.Th), number (Tb.N), separation (Tb.Sp), structure model index (SMI), cortical BMD (Ct.BMD), and thickness (Ct.Th). An orthopedic surgeon, blinded to the experiment, evaluated the bone images. The thoracic spine curve was quantified using Cobb angle, as previously described [55].
Histomorphometry and immunohistochemistry
Animals were injected 25 mg/kg fluorescence calcein 9 days and 3 days before the end of the experiment. Tibia specimens were embedded in methyl acryl resin and sectioned into 4 μm thickness. Calcein labeling was quantified using fluorescence microscope and image analysis software (Zeiss) to calculate mineral apposition rate (MAR; μm/day) and bone formation rate (BFR/BS; μm3/μm2/day). To evaluate trabecular volume (BV/TV, %) and osteoclast number (Oc.N/mm), respectively, von Koss stain and tartrate-resistant acid phosphatase (TRAP) histochemical staining kits were used. In some experiments, gross histology was assessed using hematoxylin and eosin staining. Osteoblasts were identified in tissue sections using immunohistochemistry with an osteocalcin antibody (ab93876; Abcam, Cambridge, UK) and Super SensitiveTM IHC Detection Systems (BioLaboratories; Fremont, CA). Osteoblast number (Ob.N/mm) was then calculated. Four random fields and 3 random sections of each specimen were selected. Histological analysis was performed by a pathologist blinded to the experiments.
Osteogenic, adipogenic, and osteoclast differentiation
Bone-marrow stromal cells were isolated from femurs [56] and incubated in 24-well plates at a density of 2 × 105 cells/well in osteogenic (A1007201; Thermal Fisher Scientific) or adipogenic medium (A100701) supplemented with 10% fetal bovine serum, for 15 and 21 days, respectively. In some experiments, bone-marrow mesenchymal cells were transfected lentivirus Utx and empty vector (108 infectious units/ml) for 48 hours and cultured in osteogenic medium. Mineralized matrices were characterized using von Kossa staining, while cytoplasmic oil droplets were stained using fluorescence Nile red. To quantify osteoclast formation, bone-marrow macrophages were isolated [56] and incubated in 48-well plates at a density of 5×104 cells/well in osteoclast culture medium (KT-647; Kamiya BioMedical Co, Seattle, WA) for 7 days. Osteoclasts were stained using TRAP Staining kits (KT-008l). The areas of mineralized matrices and the number of Nile red-stained adipocytes, and TRAP-stained osteoclasts in 3 random fields of per well of 3 different wells per mouse were counted.
RT-PCR
Following reverse transcription (RT) of 1 μg of total RNA, the resulting cDNAs were mixed with gene of interest-specific primers (as detailed in Supplementary Table 1) and TaqMan™ Universal PCR Master Mix. PCR amplification was performed using an ABI StepOne Plus 96-Well Real-Time PCR Detection System (Thermo Fisher Scientific). Computed cycle threshold (Ct) values of each gene were normalized to 18S rRNA expression levels. Changes in mRNA expression levels were calculated using 2ΔΔCt method, where ΔCt represents the difference between the Ct vales of designated gene and 18S (Ctgene-Ct18S) and ΔΔCt represents the ΔCt values of CtJmjd7KO and CtWT.
Calvaria osteoblast isolation
Six-week-old female Col1a1Cre-Jmjd7loxP mice were administered tamoxifen at 75 mg/kg/day for 5 consecutive days. Age- and sex-matched tamoxifen-treated Jmjd7loxP mice served as WT controls. At eight weeks of age, both Jmjd7KO mice and WT mice were euthanatized via overdose of anesthetics. Osteoblasts were isolated from calvarial bone using 0.2% collagenase and 0.25% trypsin (Gibco™; Thermo Fisher Scientific, Waltham, MA) as described previously [57]. Cells were cultured in DMEM supplemented with 10% fetal bovine serum until reaching confluence. In some experiments, Jmjd7KO and WT osteoblasts at a density of 106 cells/well (6-well plates) were transfected lentivirus Jmjd7 and empty vector (108 infectious units/ml) for 48 hours and followed by incubating in osteogenic medium with 30 μM JMJD7-IN-1 (MBS5765205; MyBiosource, San Diego, CA), 200 nM DMOG (D3695; Sigma-Aldrich, St Louis, MO), 4 μM WL12 (T9939; TargetMol Chemical Inc, Boston MA) and 2 mM α-ketoglutarate (pH 7.2).
RNA-sequencing
Total RNA was isolated from 107 calvaria osteoblasts from Jmjd7KO and WT mice using Collibri Stranded RNA Library Prep Kits (ThermoFisher Scientific). Jmjd7 deletion was confirmed using RT-PCR (Supplementary S3). RNA sequencing library was performed using an Illumina NovaSeq platform (Illumina Inc., San Diego, CA). Raw sequencing data were trimmed, and quality-controlled using Trimmoatic software (http://www.usadellab.org/cms/?page=trimmomatic) [58]. The processed reads were aligned to the reference genome using HISAT2 program. Differentially expressed gene (DEG) analysis was conducted using DESeq2. Genes with a fold change \(\ge\) 2 and adjusted P < 0.05 were considered DEG [58]. Gene ontology and KEGG pathway analyses of DGE were performed using the Cluster Profiler, following the manufacturer’s instruction (www.toolsbiotech.com).
ChIP-sequencing
H3R2me1 in 5 × 106 osteoblasts was immunoprecipitated using either an H3R2me1 monoclonal antibody (ab176844; Abcam, Cambridge, UK) or control IgG antibody (ab172730), utilizing the MilliporeSigma™ Magna ChIP™ A/G Chromatin Immunoprecipitation Kits (1710085; Thermo Fisher Scientific). Following chromatin extraction from the immunocomplexes, DNA libraries were sequenced using a 2 × 150 bp paired-end configuration on the Illumina HiSeq/Novaseq platform (Illumina, San Diego. CA). Quality control was performed using Cutadapt V1.3 and FastQC V0.11.4, retaining datasets with over 2 × 107 reads (Supplementary Fig. S4). Sequencing reads were trimmed (average length < 150 bp), aligned to the reference genome using Bowtie2 V2.2.6, ensuring more than 2 × 107 total mapped reads. Peak calling and characterization were conducted using MACS2 V2. Fold change of read per million (RPM) was calculated using R package (V1.16). Heatmaps illustrating signal enrichment ± 5 kb around TSS were generated with Deeptools V3.3.0 and ChIPseeker V3.11. Genome-wide gene expression profiles and functional ontology analyses of the aligned gene sets were performed using GOSeq V1.34.1. Additionally, KEGG pathway enrichment was analyzed through bioinformatics tools (http://en.wikipedia.org/wiki/KEGG).
ChIP-PCR
The enrichment of Runx2 and Idh in 2 ng DNA samples isolated from H3R2me1 immunocomplexes was quantified using SYBR® Green-conjugated primers specific for Runx2, Idh, and Gadph promoters (see Supplementary Table 1 for details) to determine Ct values. The relative enrichment of H3R2me1 at these promoters was calculated using the equation 2-ΔCt, where ΔCt resembled for the difference between Ctgene and CtGadph [59].
Seahorse influx analysis
Extracellular acidification rate (ECAR) of 105 osteoblasts was quantified using Agilent Seahorse XF Glycolysis Stress Test Kits (102601-100, Agilent, Santa Clara, CA) with a Seahorse XFe Analyzer. The assay involved sequential additions of glucose, oligomycin and 2-deoxyglucose and Seahorse XFe Analyzer. Glycolysis, glycolytic capacity and glycolytic reserve were calculated using Seahorse XF Report Generator, according to the manufacturer’s instructions. Mitochondrial oxygen consumption rate (OCR) was measured using Agilent Seahorse XFp Cell Mito Stress Test Kits (103015-100). The assay involved sequential additions of oligomycin, FCCP, and a combination of antimycin and rotenone. Basal respiration, maximal respiration, spare respiration capacity, ATP-linked respiration, and proton leak in osteoblasts, were calculated from the OCR measurements. ATP production from glycolysis and mitochondrial respiration was measured using Seahorse XF Real-Time ATP Rate Assay Kits (103592-100), according to the manufacturer’s instructions.
Mitochondrial complex activity
Mitochondria were extracted from 2 × 107 osteoblasts using Mitochondria Isolation Kits (ab110170; Abcam, Cambridge, UK). The activities of mitochondrial complexes I (ab109721, Abcam), II + III (ab109908), and IV (ab109911) were quantified using 50 μg protein/20 μl mitochondrial isolates.
ELISA and enzymatic activity assay
Osteocalcin (ab28536; Abcam, Cambridge, UK), CTX-1 (MBS9901663; MyBiosource, San Diego, CA), estrogen (ab285291), estradiol (MBS261250), and testosterone (ab285350) levels in serum were measured using specific ELISA kits. The production of α-KG in serum and mitochondrial isolates was quantified using the Alpha Ketoglutarate Assay Kit (ab83431). Additionally, the activities of Pdh (ab109902), Idh (ab102528), Kgdh (ab185440), and Sdh (ab228560) were quantified using their respective kits. Histone demethylase activity was measured using Histone Demethylase Fluorescent Activity Kit (EIAHDMF, Invitrogen™, Thermo Fisher Scientific).
Immunoblotting
Jmjd7 (SAB3500097l Sigma-Aldrich), H3R2me1, H3K27me3 (ab6002), Idh (ab172964), Runx2 (ab76956), monomethylated arginine (ab414), ubiquitin (ab134953), Hdac1 (ab7028), and actin (ab6276) in osteoblasts were analyzed using specific antibodies and Pierce™ Fast Western Blot Kits (Thermo Fisher Scientific). Mitochondrial oxidative phosphorylation (OXPHOS) in mitochondrial isolates was assessed using the Total OXPHOS Rodent WB Antibody Cocktail (ab110413).
Statistical analysis
Distribution of data was analyzed using Kolmogorov-Smirnov test. Data of clinical specimens and WT mice and Jmjd7KO mice were analyzed using Mann-Whitney test. Investigations of WT mice and Jmjd7KO with or without ovariectomy and α-KG supplementation were analyzed using ANOVA test followed by Bonferroni post hoc test. P value < 0.05 was considered significantly different.
Data availability
RNA sequencing readouts are deposited in GEO database (accession number GSE305876). Data related to laboratory animal experiments and in vitro osteoblast models are available on request from the corresponding author. Clinical data are not available due to General Data Protection Regulations.
References
Stegen S, Carmeliet G. Metabolic regulation of skeletal cell fate and function. Nat Rev Endocrinol. 2024;20:399–413.
Compston JE, McClung MR, Leslie WD. Osteoporosis. Lancet. 2019;393:364–76.
Walker MD, Shane E. Postmenopausal osteoporosis. N Engl J Med. 2023;389:1979–91.
Alekos NS, Kushwaha P, Kim SP, Li Z, Abood A, Dirckx N, et al. Mitochondrial beta-oxidation of adipose-derived fatty acids by osteoblasts fuels parathyroid hormone-induced bone formation. JCI Insight. 2023;8:e165604.
Khan MP, Sabini E, Beigel K, Lanzolla G, Laslow B, Wang D, et al. HIF1 activation safeguards cortical bone formation against impaired oxidative phosphorylation. JCI Insight. 2024;9:e182330.
Yu Y, Newman H, Shen L, Sharma D, Hu G, Mirando AJ, et al. Glutamine metabolism regulates proliferation and lineage allocation in skeletal stem cells. Cell Metab. 2019;29:966–78.
Stegen S, Moermans K, Stockmans I, Thienpont B, Carmeliet G. The serine synthesis pathway drives osteoclast differentiation through epigenetic regulation of NFATc1 expression. Nat Metab. 2024;6:141–52.
Zhao J, Yao K, Yu H, Zhang L, Xu Y, Chen L, et al. Metabolic remodelling during early mouse embryo development. Nat Metab. 2021;3:1372–84.
Rossmann MP, Hoi K, Chan V, Abraham BJ, Yang S, Mullahoo J, et al. Cell-specific transcriptional control of mitochondrial metabolism by TIF1γ drives erythropoiesis. Science. 2021;372:716–21.
Shan L, Yang X, Liao X, Yang Z, Zhou J, Li X, et al. Histone demethylase KDM7A regulates bone homeostasis through balancing osteoblast and osteoclast differentiation. Cell Death Dis. 2024;15:136.
Zhao X, Lin S, Ren H, Sun S, Zheng L, Chen LF, et al. The histone methyltransferase ASH1L protects against bone loss by inhibiting osteoclastogenesis. Cell Death Differ. 2024;31:605–17.
Xie Y, Han N, Li F, Wang L, Liu G, Hu M, et al. Melatonin enhances osteoblastogenesis of senescent bone marrow stromal cells through NSD2-mediated chromatin remodelling. Clin Transl Med. 2022;12:e746.
Min Z, Xiaomeng L, Zheng L, Yangge D, Xuejiao L, Longwei L, et al. Asymmetrical methyltransferase PRMT3 regulates human mesenchymal stem cell osteogenesis via miR-3648. Cell Death Dis. 2019;10:581.
Li JY, Wang TT, Ma L, Zheng LL. CARM1 deficiency inhibits osteoblastic differentiation of bone marrow mesenchymal stem cells and delays osteogenesis in mice. Biochim Biophys Acta Mol Cell Res. 2023;1870:119544.
Kota SK, Roening C, Patel N, Kota SB, Baron R. PRMT5 inhibition promotes osteogenic differentiation of mesenchymal stromal cells and represses basal interferon stimulated gene expression. Bone. 2018;117:37–46.
Choi JH, Jang AR, Kim DI, Park MJ, Lim SK, Kim MS, et al. PRMT1 mediates RANKL-induced osteoclastogenesis and contributes to bone loss in ovariectomized mice. Exp Mol Med. 2018;50:1–15.
Manni W, Jianxin X, Weiqi H, Siyuan C, Huashan S. JMJD family proteins in cancer and inflammation. Signal Transduct Target Ther. 2022;7:304.
Liu H, Wang C, Lee S, Deng Y, Wither M, Oh S, et al. Clipping of arginine-methylated histone tails by JMJD5 and JMJD7. Proc Natl Acad Sci USA. 2017;114:E7717–E7726.
Zhao J, Adams A, Roberts B, O’Neil M, Vittal A, Schmitt T, et al. Protein arginine methyl transferase 1- and Jumonji C domain-containing protein 6-dependent arginine methylation regulate hepatocyte nuclear factor 4 alpha expression and hepatocyte proliferation in mice. Hepatology. 2018;67:1109–26.
Guo Z, Hu YH, Feng GS, Valenzuela Ripoll C, Li ZZ, Cai SD, et al. JMJD6 protects against isoproterenol-induced cardiac hypertrophy via inhibition of NF-κB activation by demethylating R149 of the p65 subunit. Acta Pharm Sin. 2023;44:1777–89.
Jablonowski CM, Quarni W, Singh S, Tan H, Bostanthirige DH, Jin H, et al. Metabolic reprogramming of cancer cells by JMJD6-mediated pre-mRNA splicing associated with therapeutic response to splicing inhibitor. Elife. 2024;12:RP90993.
Cui X, Hou L, Yan B, Liu J, Zhang C, Sui P, et al. Sexual dimorphism in the mouse bone marrow niche regulates hematopoietic engraftment via sex-specific KDM5c/CXCL12 signaling. J Clin Invest. 2025;135:e182125.
Cheng MI, Li JH, Riggan L, Chen B, Tafti RY, Chin S, et al. The X-linked epigenetic regulator UTX controls NK cell-intrinsic sex differences. Nat Immunol. 2023;24:780–91.
Gao CW, Lin W, Riddle RC, Kushwaha P, Boukas L, Björnsson HT, et al. A mouse model of Weaver syndrome displays overgrowth and excess osteogenesis reversible with KDM6A/6B inhibition. JCI Insight. 2024;9:e173392.
Li F, Jing J, Movahed M, Cui X, Cao Q, Wu R, et al. Epigenetic interaction between UTX and DNMT1 regulates diet-induced myogenic remodeling in brown fat. Nat Commun. 2021;12:6838.
Shao R, Suo J, Zhang Z, Kong M, Ma Y, Wen Y, et al. H3K36 methyltransferase NSD1 protects against osteoarthritis through regulating chondrocyte differentiation and cartilage homeostasis. Cell Death Differ. 2024;31:106–18.
Wu J, Chen X, Sehgal P, Zhang T, Jackson-Weaver O, Gou Y, et al. Arginine methylation of R81 in Smad6 confines BMP-induced SMAD1 signaling. J Biol Chem. 2021;296:100496.
Gou Y, Li J, Wu J, Gupta R, Cho I, Ho TV, et al. Prmt1 regulates craniofacial bone formation upstream of Msx1. Mech Dev. 2018;152:13–20.
Ramachandran J, Liu Z, Gray RS, Vokes SA. PRMT5 is necessary to form distinct cartilage identities in the knee and long bone. Dev Biol. 2019;456:154–63.
Clotet-Freixas S, Zaslaver O, Kotlyar M, Pastrello C, Quaile AT, McEvoy CM, et al. Sex differences in kidney metabolism may reflect sex-dependent outcomes in human diabetic kidney disease. Sci Transl Med. 2024;16:eabm2090.
Pribadi C, Cakouros D, Camp E, Anderson P, Gronthos S. KDM6A-mediated regulation of cranial frontal bone suture fusion in mice is sex dependent. Stem Cells Dev. 2023;32:398–409.
Zhu Q, Wang D, Liang F, Tong X, Liang Z, Wang X, et al. Protein arginine methyltransferase PRMT1 promotes adipogenesis by modulating transcription factors C/EBPbeta and PPARgamma. J Biol Chem. 2022;298:102309.
Dashti P, Lewallen EA, Gordon JAR, Montecino MA, van Leeuwen JPTM, Stein GS, et al. Protein arginine methyltransferases PRMT1, PRMT4/CARM1 and PRMT5 have distinct functions in control of osteoblast differentiation. Bone Rep. 2023;19:101704.
Hemming S, Cakouros D, Isenmann S, Cooper L, Menicanin D, Zannettino A, et al. EZH2 and KDM6A act as an epigenetic switch to regulate mesenchymal stem cell lineage specification. Stem Cells. 2014;32:802–15.
Mori K, Mizokami A, Sano T, Mukai S, Hiura F, Ayukawa Y, et al. RANKL elevation activates the NIK/NF-kappaB pathway, inducing obesity in ovariectomized mice. J Endocrinol. 2022;254:27–36.
Jin Z, Kho J, Dawson B, Jiang MM, Chen Y, Ali S, et al. Nitric oxide modulates bone anabolism through regulation of osteoblast glycolysis and differentiation. J Clin Invest. 2021;131:e138935.
Yang YY, Zhou YM, Xu JZ, Sun LH, Tao B, Wang WQ, et al. Lgr4 promotes aerobic glycolysis and differentiation in osteoblasts via the canonical Wnt/beta-catenin pathway. J Bone Min Res. 2021;36:1605–20.
Lee WC, Ji X, Nissim I, Long F. Malic enzyme couples mitochondria with aerobic glycolysis in osteoblasts. Cell Rep. 2020;32:108108.
Hollenberg AM, Smith CO, Shum LC, Awad H, Eliseev RA. Lactate dehydrogenase inhibition with oxamate exerts bone anabolic effect. J Bone Min Res. 2020;35:2432–43.
Lin C, Yang Q, Guo D, Xie J, Yang YS, Chaugule S, et al. Impaired mitochondrial oxidative metabolism in skeletal progenitor cells leads to musculoskeletal disintegration. Nat Commun. 2022;13:6869.
Kang HS, Lee JH, Oh KJ, Lee EW, Han BS, Park KY, et al. IDH1-dependent alpha-KG regulates brown fat differentiation and function by modulating histone methylation. Metabolism. 2020;105:154173.
Li X, Wu F, Günther S, Looso M, Kuenne C, Zhang T, et al. Inhibition of fatty acid oxidation enables heart regeneration in adult mice. Nature. 2023;622:619–26.
Lui JC, Raimann A, Hojo H, Dong L, Roschger P, Kikani B, et al. A neomorphic variant in SP7 alters sequence specificity and causes a high-turnover bone disorder. Nat Commun. 2022;13:700.
Qin X, Jiang Q, Nagano K, Moriishi T, Miyazaki T, Komori H, et al. Runx2 is essential for the transdifferentiation of chondrocytes into osteoblasts. PLoS Genet. 2020;16:e1009169.
Jin R, Zhang H, Lin C, Guo J, Zou W, Chen Z, et al. Inhibition of miR338 rescues cleidocranial dysplasia in Runx2 mutant mice partially via the Hif1a-Vegfa axis. Exp Mol Med. 2023;55:69–80.
Wang Y, Deng P, Liu Y, Wu Y, Chen Y, Guo Y, et al. Alpha-ketoglutarate ameliorates age-related osteoporosis via regulating histone methylations. Nat Commun. 2020;11:5596.
Cheng C, Xing Z, Hu Q, Kong N, Liao C, Xu S, et al. A bone-targeting near-infrared luminescence nanocarrier facilitates alpha-ketoglutarate efficacy enhancement for osteoporosis therapy. Acta Biomater. 2024;173:442–56.
Thaler R, Khani F, Sturmlechner I, Dehghani SS, Denbeigh JM, Zhou X, et al. Vitamin C epigenetically controls osteogenesis and bone mineralization. Nat Commun. 2022;13:5883.
Chang B, Chen Y, Zhao Y, Bruick RK. JMJD6 is a histone arginine demethylase. Science. 2007;318:444–7.
Li Z, Wang P, Li J, Xie Z, Cen S, Li M, et al. The N(6)-methyladenosine demethylase ALKBH5 negatively regulates the osteogenic differentiation of mesenchymal stem cells through PRMT6. Cell Death Dis. 2021;12:578.
Zhou G, Zhang C, Peng H, Su X, Huang Q, Zhao Z, et al. PRMT3 methylates HIF-1α to enhance the vascular calcification induced by chronic kidney disease. Mol Med. 2024;30:8.
Kim KH, Oprescu SN, Snyder MM, Kim A, Jia Z, Yue F, et al. PRMT5 mediates FoxO1 methylation and subcellular localization to regulate lipophagy in myogenic progenitors. Cell Rep. 2023;42:113329.
Lian WS, Ko JY, Chen YS, Ke HJ, Hsieh CK, Kuo CW, et al. MicroRNA-29a represses osteoclast formation and protects against osteoporosis by regulating PCAF-mediated RANKL and CXCL12. Cell Death Dis. 2019;10:705.
Lin YH, Lian WS, Wu RW, Chen YS, Wu SL, Ko JY, et al. Trimethylamine-N-oxide accelerates osteoporosis by PERK activation of ATF5 unfolding. Cell Mol Life Sci. 2025;82:13.
Hey HWD, Lam WMR, Chan CX, Zhuo WH, Crombie EM, Tan TC, et al. Paraspinal myopathy-induced intervertebral disc degeneration and thoracolumbar kyphosis in TSC1mKO mice model-a preliminary study. Spine J. 2022;22:483–94.
Tang L, Wu M, Lu S, Zhang H, Shen Y, Shen C, et al. Fgf9 negatively regulates bone mass by inhibiting osteogenesis and promoting osteoclastogenesis via MAPK and PI3K/AKT signaling. J Bone Min Res. 2021;36:779–91.
Kim I, Park S, Kim J, Park SY, Seo J, Roh S. Treatment with Lactobacillus paracasei L30 extract induces osteogenic differentiation of human bone marrow mesenchymal stem cells in vitro. Biomed Pharmacother. 2025;184:117913.
Sewe SO, Silva G, Sicat P, Seal SE, Visendi P. Trimming and validation of Illumina short reads using Trimmomatic, trinity assembly, and assessment of RNA-Seq data. Methods Mol Biol. 2022;2443:211–32.
Yi SJ, Jang YJ, Kim HJ, Lee K, Lee H, Kim Y, et al. The KDM4B-CCAR1-MED1 axis is a critical regulator of osteoclast differentiation and bone homeostasis. Bone Res. 2021;9:27.
Acknowledgements
This work is supported by [NHRI-EX114-11029SI] and [NHRI-EX114-11437SI] from National Health Research Institute; [NSTC110-2314-B-182A-018-MY3] form National Science and Technology Council; [CMRPG8M1281-3] from Chang Gung Memorial Hospital, Taiwan and BMTFR [13XP5206], Germany. Authors also thank Core Laboratory for Gnotobiotic Medicine and Center for Laboratory Animals, Kaohsiung Chang Gung Memorial Hospital, Taiwan, and Taiwan Animal Consortium, for facility support.
Funding
This work was supported by [NHRI-EX114-11029SI] and [NHRI-EX114-11437SI] from the National Health Research Institute; [NSTC110-2314-B-182A-018-MY3] from National Science and Technology Council; and [CMRPG8M1281-3] from Chang Gung Memorial Hospital, Taiwan, as well as the BMFTR [13XP5206], Germany.
Author information
Authors and Affiliations
Contributions
Study conception and design: R-WW, W-SL, Y-HL, Y-SC, S-YW, S-LW, M-YL, HJ, and F-SW; cell culture analysis: Y-SC; RNA-seq, ChIP-seq, bioinformation analysis, & ChIP-PCR: W-SL and Y-SC; in vivo & μCT analysis: Y-SC and S-YW; histology & immunoblotting: S-LW and M-YL; osteoporosis diagnosis & clinical specimens collection: R-WW; Analysis and interpretation of data: R-WW, W-SL, Y-HL, HJ, and F-SW; Article drafting and revision: R-WW, W-SL, Y-HL, HJ, and F-SW.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
All methods involving human bone specimens were conducted in accordance with the Declaration of Helsinki and relevant national guidelines and regulations in Taiwan. The protocols were reviewed and approved by the Institutional Review Board of Chang Gung Medical Foundation (IRB Affidavit #201701826B0 and #202002225B0), and written informed consent was obtained from all patients. Animal experiments described were performed in compliance with the Guidelines for the Care and Use of Laboratory Animals, Taiwan. Animal protocols were reviewed and approved by the Institutional Animal Care and Use Committee of Kaohsiung Chang Gung Memorial Hospital (IACUC Affidavit #2020121201).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, RW., Lian, WS., Lin, YH. et al. Histone arginine demethylase JMJD7 promotes bone formation through regulating α-Ketoglutarate metabolism. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01727-4
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01727-4