Abstract
Morphogens play a critical role in coordinating stress adaptation and aging across tissues, yet their involvement in neuronal mitochondrial stress responses and systemic effects remains unclear. In this study, we reveal that the transforming growth factor beta (TGF-β) DAF-7 is pivotal in mediating the intestinal mitochondrial unfolded protein response (UPRmt) in Caenorhabditis elegans under neuronal mitochondrial stress. Two ASI sensory neurons produce DAF-7, which targets DAF-1/TGF-β receptors on RIM interneurons to orchestrate a systemic UPRmt response. Remarkably, inducing mitochondrial stress specifically in ASI neurons activates intestinal UPRmt, extends lifespan, enhances pathogen resistance, and reduces both brood size and body fat levels. Furthermore, dopamine positively regulates this UPRmt activation, while GABA acts as a systemic suppressor. This study uncovers the intricate mechanisms of systemic mitochondrial stress regulation, emphasizing the vital role of TGF-β in metabolic adaptations that are crucial for organismal fitness and aging during neuronal mitochondrial stress.
Similar content being viewed by others
Introduction
The nervous system serves as a central regulatory hub, detecting both environmental and internal stress stimuli, orchestrating a spectrum of physiological processes, and influencing non-autonomous functions such as foraging behavior, immune responses, body fat storage, systemic stress adaptions, and aging1,2. Disruptions in mitochondrial function significantly affect neuronal activities and can extend their impact to peripheral tissues, yet the mechanisms by which neuronal circuits transmit mitochondrial stress signals to these tissues remain largely unclear.
In Caenorhabditis elegans, various neuronal-specific stressors have been identified that elicit cell non-autonomous responses in peripheral tissues. For instance, knocking down the mitochondrial electron transport chain (ETC) component cco-1 in neurons activates the mitochondrial unfolded protein response (UPRmt) in the intestine, extending lifespan3. Similarly, neuronal expression of YFP with a polyglutamine tract (Q40::YFP), disruption of the mitochondrial dynamic regulator FZO-1, or induction of mitochondrial ROS production within neurons can have cell non-autonomous effects in peripheral tissues4,5,6,7. Additionally, exposure to pathogens bacterial source odorant, 2,3-pentanedione, can induce cell non-autonomous the UPRmt in a serotonin dependent manner8. Interestingly, this neuronal coordination of the mitochondrial systemic stress response is not limited to C. elegans. For example, in mice, heterodeficiency of the mitoribosomal protein Crif1 in hypothalamic proopiomelanocortin (POMC) neurons activate the mitochondrial stress response in distal adipose tissues, protecting obese mice and improving glucose metabolism9. Similarly, genetic inactivation of OPA1, a mitochondrial cristae-remodeling protein, in POMC neurons reduces white adipose tissue (WAT) lipolysis, leading to obesity10.
Recent studies have identified secreted “mitokine” signals released from neurons experiencing mitochondrial dysfunction that elicit cell non-autonomous effects. These “mitokine” signals include the morphogen Wnt/EGL-20, neurotransmitters like serotonin, and various neuropeptides4,5,11,12. The recognition of Wnt as a “mitokine” signal has prompted speculation about the broader role of morphogens in inter-tissue stress communication.
Wnt and transforming growth factor beta (TGF-β) are crucial morphogens with conserved functions in metazoan. TGF-β extensively cross-talks with the Wnt pathway in various biological contexts, including epithelial-mesenchymal transition, bone formation and remodeling, enterocyte differentiation, cardiac fibrogenesis, and colorectal cancer13,14,15,16,17. In C. elegans, neuronal TGF-β signaling regulates development18, food intake19, pathogen avoidance20,21, fat metabolism22, post-prandial quiescence23, and food-leaving behavior24. While Wnt signaling has been implicated in inter-tissue mitochondrial stress communication, the role of TGF-β signaling in orchestrating cell non-autonomous mitochondrial stress responses and influencing systemic metabolic states remains elusive.
To further understand how morphogens participate in the neuronal coordination of systemic mitochondrial stress response, we demonstrate that DAF-7, a TGF-β ligand, secreted from ASI sensory neurons, acts on DAF-1, a TGF-β receptor expressed on RIM interneurons, to coordinate the systemic UPRmt activation. Combining genetic and cell biology approaches, we found that ASI-specific mitochondrial stress is sufficient to induce UPRmt activation in the intestine, leading to extended lifespan and increased pathogen resistance. Notably, this ASI-RIM-mediated UPRmt activation is positively regulated by dopamine but negatively regulated by GABA. Our study establishes a direct link between neuronal mitochondrial stress and systemic stress adaption and longevity via the TGF-β/DAF-7 signaling pathway, highlighting the critical role of neuronal coordination in systemic metabolic adaptations and aging for organismal fitness.
Results
TGF-β/DAF-7 is required for the induction of the cell non-autonomous UPRmt
To investigate the involvement of TGF-β signaling in cell non-autonomous UPRmt activation, we utilized a UPRmt reporter strain (hsp-6p::gfp) in C. elegans lacking TGF-β ligands, under neuronal mitochondrial stress induced by either neuronal cco-1 knockdown or neuronal Q40::YFP expression3,5. C. elegans features two TGF-β superfamily signaling pathways, each comprising distinct ligands, receptors, and Smads. Specifically, daf-7, the homolog of human GDF11, encodes the ligand of the TGF-β Dauer pathway18, while dbl-1, the homolog of human BMP5, encodes the ligand for the TGF-β Sma/Mab pathway25.
We found that the mutation of daf-7 significantly suppressed the induction of hsp-6p::gfp expression in animals with neuronal cco-1 knockdown or neuronal Q40::YFP expression (Fig. 1a, b and Supplementary Fig. 1a–d). Moreover, overexpression of daf-7p::daf-7 partially restored hsp-6p::gfp expression in daf-7(e1372) mutants (Fig. 1a, b and Supplementary Fig. 1c, d).
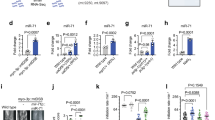
a Representative photomicrographs showing the bright-field images of hsp-6p::gfp; and the fluorescence images of hsp-6p::gfp in wild type background, neuronal cco-1 knockdown background, neuronal cco-1 knockdown and daf-7 mutation background, neuronal cco-1 knockdown, daf-7 mutation and daf-7p::daf-7(cDNA) overexpressing background. The posterior region of the intestine where hsp-6p::gfp expressed is highlighted. Scale bar, 100 μm. b Fluorescence intensities of hsp-6p::gfp in animals shown in (a) were quantified using ImageJ. n = 30 worms. c Relative mRNA levels of hsp-6 and hsp-60 in control and daf-7 mutation with or without neuronal cco-1 knockdown background. n = 6 biological replicates. d Representative fluorescence photomicrographs of hsp-6p::gfp in neuronal cco-1 knockdown background with the expression of daf-7 dsRNA in indicated sub-type neurons. Scale bar, 100 μm. e Fluorescence intensities of hsp-6p::gfp in animals shown in (d) were quantified using ImageJ. n = 30 worms. f Representative fluorescence images of daf-7p::gfp (FK181) and str-3p::mCherry reporters. Scale bar, 25 μm. g Representative fluorescence photomicrographs of daf-7p::gfp with or without the expression of rgef-1p::cco-1 hairpin(HP). Scale bar, 25 μm. h Fluorescence intensities of daf-7p::gfp in ASI neurons of animals shown in (f) were quantified using ImageJ. n = 12 neurons. i Relative mRNA level of daf-7 in control and rgef-1p::cco-1HP animals. n = 4 biological replicates. p values were annotated via two-tailed unpaired t test in (h) and (i), via ordinary one-way ANOVA in (b) and (e), via two-way ANOVA in (c). Error bars, SEM. Source data are provided as a Source Data file. See also Supplementary Fig. 1.
Consistent with the reporter results, the daf-7 mutation significantly suppressed endogenous hsp-6 and hsp-60 mRNA levels in animals with neuronal cco-1 knockdown or neuronal Q40::YFP expression compared with wild-type animals (Fig. 1c and Supplementary Fig. 1e). Additionally, previous studies have shown that the cell non-autonomous UPRmt can also be activated by neuronal Wnt/EGL-20 expression11, ADL-neuronal Gas (gsa-1) or Gaq (egl-30) activation12, and the transgenerational effect of neuronal mitochondrial stress (TGW)26. Our findings indicate that the cell non-autonomous UPRmt induced by these systems is dependent on daf-7 (Supplementary Fig. 1f–m), highlighting the general role of daf-7 in cell non-autonomous UPRmt regulation. Notably, the daf-7 mutation did not affect the UPRmt induction in worms fed with cco-1 RNAi, suggesting that daf-7 is not involved in cell-autonomous UPRmt activation (Supplementary Fig. 1n, o)3. In contrast, the dbl-1 mutation, which abolishes TGF-β Sma/Mab pathway activity, did not suppress UPRmt induction in neuronal Q40::YFP animals (Supplementary Fig. 1p, q). Similarly, the mutation of sma-6, the gene encoding the type I receptor specific to the TGF-β Sma/Mab pathway27, did not affect the hsp-6p::gfp expression induced by neuronal cco-1 knockdown (Supplementary Fig. 1r, s). These results collectively indicate that loss-of-function mutation in daf-7, the ligand of the TGF-β Dauer pathway, suppresses systemic UPRmt activation in response to neuronal mitochondrial stress.
Cell non-autonomous stress communication extends beyond mitochondrial dysfunction to encompass endoplasmic reticulum (ER) or cytosolic (Cyt) protein homeostasis. Neuronal expression of the transcription factor xbp-1s, the spliced form of XBP-1, induces the unfolded protein response in the ER (UPRER) in the intestine28,29. Similarly, expression of the transcription factor hsf-1 in neurons activates the unfolded protein response in the cytosol (UPRcyt) in the intestine30,31. Interestingly, we observed that loss of daf-7 further induced UPRER in neuronal xbp-1s background, and also further induced UPRcyt in the neuronal hsf-1 expression background (Supplementary Fig. 1t–w). Therefore, daf-7 is required for the induction of the systemic UPRmt activation in response to neuronal mitochondrial perturbations.
DAF-7 acts in ASI chemosensory neurons to coordinate the systemic UPRmt activation
Previous studies have indicated that daf-7 is exclusively expressed in the ASI-ciliated chemosensory neurons18,32. However, data from the C. elegans Neuronal Gene Expression Map & Network (CeNGEN) show that daf-7 is expressed in several neurons, including ASI, OLQ, ASG, ADE, AWA, ASJ, and ASK neuron (Supplementary Fig. 1x)33,34,35. To identify the specific neuronal subtype where daf-7 is essential for UPRmt activation, we knocked down daf-7 by expressing double-stranded RNA under specific promoters for each neuron pair (Supplementary Fig. 1x)36,37,38,39,40,41.
When daf-7 was specifically knocked down in ASI neurons using the gpa-4 promoter36, we observed a strong suppression of intestinal UPRmt activation in animals with neuronal cco-1 knockdown in the sid-1 mutant background. The sid-1 mutation renders the animals defective in extracellular double-stranded RNA uptake, enabling tissue-specific RNAi treatment (Fig. 1d, e)33. This suppression was not observed with daf-7 knockdown in other neurons subtypes, indicating that daf-7 functions in ASI neurons to mediate the cell non-autonomous UPRmt.
To investigate whether daf-7 expression is affected under neuronal mitochondrial stress, we monitored its expression using the reporter daf-7p::gfp (FK181)20. First, we confirmed that daf-7 is primarily expressed in the ASI chemosensory neurons by colocalizing it with an ASI neuron marker (str-3p::mCherry) (Fig. 1f). Additionally, we observed that the fluorescence of daf-7p::gfp was induced by neuronal cco-1 knockdown (Fig. 1g, h), which is consistent with the up-regulated endogenous daf-7 mRNA level (Fig. 1i). These data suggest the TGF-β signaling is activated in ASI neurons in response to neuronal mitochondrial stress.
The TGF-β/Dauer signaling pathway is required for cell non-autonomous UPRmt activation
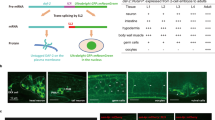
DAF-7 is a TGF-β ligand that governs the TGF-β/Dauer pathway in response to environmental cues. Secreted from sensory neurons, DAF-7 binds to the type I and type II receptors, DAF-1 and DAF-4, in target cells, leading to the phosphorylation of the R-Smads DAF-8 and DAF-14. This phosphorylation negatively regulates the Co-Smad DAF-3 and the Sno/Ski factor DAF-5 (Fig. 2a), promoting reproductive growth and blocking dauer entry22,42,43,44,45,46,47,48,49. The function of DAF-7 in preventing dauer entry is dependent on DAF-12, a nuclear hormone receptor50.
a Schematic diagram of TGF-β/Dauer pathway. Created in BioRender. Wang, Z. (2023) BioRender.com/i90a877. b Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, daf-1, daf-4, daf-8 and daf-14 animals in neuronal cco-1 knockdown background. c Fluorescence intensities of hsp-6p::gfp in animals shown in (b) were quantified using ImageJ. n = 30 worms. d Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, daf-7, daf-7;daf-3, daf-7;daf-5 and daf-7;daf-12 animals in neuronal cco-1 knockdown background. e Fluorescence intensities of hsp-6p::gfp in animals shown in (d) were quantified using ImageJ. n = 30 worms. p values were annotated via ordinary one-way ANOVA in (c) and (e). Error bars, SEM. Scale bar, 100 μm. Source data are provided as a Source Data file. See also Supplementary Fig. 2.
To examine the involvement of the entire TGF-β/Dauer signaling pathway in the cell non-autonomous UPRmt regulation, we introduced loss-of-function mutations in downstream signaling components in animals with neuronal cco-1 knockdown and measured their UPRmt activation. In line with the canonical TGF-β/Dauer signaling pathway, mutations in daf-1(m40), daf-4(m63), daf-8(e1393), and daf-14(m77) strongly suppressed intestinal UPRmt activation in animals with neuronal cco-1 knockdown or Q40::YFP expression (Fig. 2b, c and Supplementary Fig. 2a, b). Conversely, simultaneous depletion of the negative regulators DAF-3 or DAF-5 in the context of DAF-7 deficiency partially restored UPRmt induction under neuronal mitochondrial stresses (Fig. 2d, e and Supplementary Fig. 2c, d). Notably, daf-3(e1376) and daf-5(e1386) induced hsp-6p::gfp expression in the intestine (Supplementary Fig. 2e, f).
In contrast, the loss of daf-12 did not restore intestinal UPRmt induction with neuronal cco-1 knockdown (Fig. 2d, e) or Q40::YFP expression (Supplementary Fig. 2c, d). Thus, while both daf-3 and daf-12 are required for dauer entry when daf-7 is inactivated, suppression of cell non-autonomous UPRmt in these mutants only requires daf-3, suggesting a divergence in the molecular mechanisms regulating UPRmt and dauer entry. These findings indicate that the cell non-autonomous UPRmt activation requires the canonical TGF-β Dauer pathway mediated by DAF-7 but is not linked with its role in regulating dauer formation.
DAF-1 functions in RIM interneurons to mediate cell non-autonomous UPRmt
To identify the target cells that receive the DAF-7 signal from ASI neurons and transmit the signal to activate cell non-autonomous UPRmt in the intestine, we focused on DAF-1, the type I receptor of the Dauer pathway42,43. Both daf-1p::gfp and daf-1p::daf-1::bfp reporters showed predominant expression of daf-1 in the nervous system (Supplementary Fig. 3a). Restoring DAF-1 function using the daf-1 promoter and the pan-neuronal promoter rgef-1 in daf-1(m40) mutants significantly rescued the suppressed cell non-autonomous UPRmt (Fig. 3a, b and Supplementary Fig. 3b, c). Additionally, pan-neuronal daf-1 knockdown strongly suppressed intestinal UPRmt activation, confirming the requirement of neuronal DAF-1 for systemic UPRmt activation in response to neuronal mitochondrial stress (Supplementary Fig. 3d, e).
a Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, daf-1(m40), daf-1(m40);daf-1p::daf-1, daf-1(m40);rgef-1p::daf-1animals in neuronal cco-1 KD background. Scale bar, 100 μm. b Fluorescence intensities of hsp-6p::gfp in animals shown in (a) were quantified using ImageJ. n = 30 worms. c Schematic of subset neurons classification and their specific promoters. d Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, daf-1(m40) and DAF-1 subtype neurons rescue animals in neuronal cco-1 KD background. Scale bar, 100 μm. e Fluorescence intensities of hsp-6p::gfp in animals shown in (d) were quantified using ImageJ. n = 30 worms. f Representative fluorescence images of head region of animals expressing F23H12.7p::gfp (RIM, source from the CeNGEN project) and tdc-1p::mCherry (RIM and RIC). Scale bar, 10 μm. g Representative fluorescence photomicrographs of hsp-6p::gfp in neuronal cco-1 knockdown background with or without the expression of F23H12.7p::daf-1 hairpin(HP). Scale bar, 100 μm. h Fluorescence intensities of hsp-6p::gfp in animals shown in (g) were quantified using ImageJ. n = 30 worms. i Representative fluorescence photomicrographs of daf-8p::daf-8::gfp; F23H12.7p::mCherry reporter with or without neuronal cco-1 knockdown and daf-7 mutation. Scale bar, 5 μm. j Fluorescence intensities of daf-8p::daf-8::gfp expression in RIM neurons in animals shown in (i) were quantified using ImageJ. n = 10 neurons. p values were annotated via two-tailed unpaired t test in (h), via ordinary one-way ANOVA in (b), (e) and (j). Error bars, SEM. Source data are provided as a Source Data file. See also Supplementary Fig. 3.
To pinpoint the specific neuronal subset responsible for mediating cell non-autonomous UPRmt, we categorized all daf-1-expressing neurons into seven subsets based on previous studies (Fig. 3c)51,52,53,54,55. We then used specific promoters to drive daf-1 expression in these subsets. Our findings revealed that expressing daf-1 in set 1 (tdc-1p::daf-1) and set 2 (glr-1p::daf-1) neurons successfully restored UPRmt induction in daf-1 mutants with neuronal cco-1 knockdown (Fig. 3d, e) or neuronal Q40::YFP background (Supplementary Fig. 3f, g). Notably, overexpressing daf-1 in each set of neurons did not significantly induce hsp-6p::gfp expression in the daf-1(m40) background, suggesting that overexpression of the receptor alone is not sufficient to induce cell non-autonomous UPRmt (Supplementary Fig. 3h, i). These findings indicate that DAF-1 plays a critical role in RIM interneurons to mediate the cell non-autonomous UPRmt.
Next, we selected a RIM-specific expressing promoter, F23H12.7, from the C. elegans Neuronal Gene Expression Map & Network (CeNGEN) database33,34,35, and verified its expression through co-localization analyses with the tdc-1p::mCherry reporter, known to express in RIM, RIC, and UV-1 cells (Fig. 3f)51. Knocking down daf-1 in RIM interneurons (F23H12.7p::dsRNA-daf-1) strongly suppressed cell non-autonomous UPRmt (Fig. 3g, h). Furthermore, expressing DAF-1 in RIM neurons significantly rescued UPRmt activation in daf-1(m40) mutants with neuronal cco-1 knockdown (Supplementary Fig. 3j, k). Together, these results indicate that DAF-1 functions in RIM interneurons to coordinate the cell non-autonomous UPRmt.
DAF-8 accumulates in the nucleus in response to neuronal mitochondrial stress
To determine if TGF-β signaling is activated in response to pan-neuronal mitochondrial stress, we generated a transgenic strain expressing daf-8p::daf-8::gfp to monitor the protein level and subcellular localization of DAF-8, which relocates to the nucleus upon DAF-7 stimulation56,57. We specifically labeled RIM neurons using the marker F23H12.7p::mCherry, given our previous findings on the role of the receptor DAF-1 in these neurons in responding to neuronal mitochondrial stress. Neuronal cco-1 knockdown significantly increased the DAF-8 protein level and promoted its nuclear localization, as indicated by higher GFP fluorescence in the nuclei of RIM neurons (Supplementary Fig. 3l). Furthermore, the daf-7(e1372) mutation abolished the nuclear accumulation of DAF-8 in RIM neurons of animals with neuronal cco-1 knockdown (Fig. 3i, j). This finding suggests that the canonical TGF-β Dauer pathway component is activated in response to neuronal mitochondrial stress, leading to a notable nuclear accumulation of TGF-β/DAF-8 Smads in RIM neurons in a DAF-7-dependent manner.
ASI-specific mitochondrial perturbation is sufficient to activate systemic UPRmt in a daf-7-dependent manner
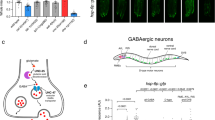
Given the essential role of the ASI-RIM neuron axis in TGF-β signal transduction for cell non-autonomous UPRmt activation, we investigated whether ASI-specific mitochondrial stress alone could drive this process in peripheral tissue. To explore this, we generated transgenic animals with ASI-specific cco-1 knocked down in the sid-1(qt9) mutant background (gpa-4p::cco-1 HP, referred to as ASI cco-1 KD). Remarkably, ASI cco-1 KD significantly induced UPRmt in the intestine, albeit to a lesser extent than pan-neuronal cco-1 KD, as evidenced by both the hsp-6p::gfp reporter and qPCR analyses of endogenous UPRmt target genes (Fig. 4a–c). However, UPRER and UPRcyt were not activated in ASI cco-1 KD animals, as accessed by fluorescence reporter (hsp-4p::gfp and hsp-16.2p::gfp) (Supplementary Fig. 4a–d) and qPCR analyses (Supplementary Fig. 4e). Furthermore, we generated strains with cco-1 KD in various neuron subtypes, including ADL, DVB, serotonergic, and GABAergic neurons. We observed that cco-1 KD in all these neuron types could drive cell non-autonomous UPRmt activation to varying extents, and this effect is also dependent on daf-7 (Supplementary Fig. 4f–m). These results suggest that mitochondrial stress in different neuron subtypes can induce the cell non-autonomous UPRmt, with a common requirement for daf-7.
a Representative fluorescence photomicrographs of hsp-6p::gfp with or without gpa-4p::cco-1 hairpin(HP). b Fluorescence intensities of hsp-6p::gfp in animals shown in (a) were quantified using ImageJ. n = 30 worms. c Relative mRNA levels of hsp-6 and hsp-60 in control and gpa-4p::cco-1HP animals. n = 3 biological replicates. d Representative fluorescence photomicrographs of daf-7p::gfp with or without gpa-4p::cco-1HP. Scale bar, 25 μm. e Fluorescence intensities of daf-7p::gfp in ASI neurons of animals shown in (d) were quantified using ImageJ. n = 12 neurons. f Relative mRNA level of daf-7 in control and gpa-4p::cco-1HP animals. n = 4 biological replicates. g Representative fluorescence photomicrographs of hsp-6p::gfp in gpa-4p::cco-1HP background with or without gpa-4p::daf-7HP or F23H12.7p::daf-7HP. h Fluorescence intensities of hsp-6p::gfp in animals shown in (g) were quantified using ImageJ. n = 30 worms. i Representative fluorescence photomicrographs of hsp-6p::gfp in gpa-4p::cco-1HP with or without unc-13, unc-31 or egl-21 mutations. j Fluorescence intensities of hsp-6p::gfp in animals shown in (i) were quantified using ImageJ. n = 30 worms. p values were annotated via two-tailed unpaired t test in (b), (e) and (f), via ordinary one-way ANOVA in (h) and (j), via two-way ANOVA in (c). Error bars, SEM. Scale bar, 100 μm. Source data are provided as a Source Data file. See also Supplementary Fig. 4.
Additionally, daf-7 expression was elevated in response to ASI cco-1 KD, as indicated by the transcriptional reporter daf-7p::gfp (Fig. 4d, e), and endogenous mRNA levels measured by qPCR (Fig. 4f). Moreover, the intestinal UPRmt activation requires daf-7 in ASI neurons and daf-1 in RIM neurons (Fig. 4g, h). These results suggest that systemic UPRmt activation in response to ASI cco-1 KD involves the ASI-RIM neuron axis of TGF-β/DAF-7 signaling.
Previous studies have identified several components involved in cell non-autonomous UPRmt activation, including neuropeptides flp-2, Wnt ligand egl-20, protein disulfide isomerase pdi-6, and G-protein-coupled receptor srz-754,11,12,58,59,60. To elucidate their roles in the ASI cco-1 KD system, we crossed these mutants into ASI cco-1 KD worms. Surprisingly, none of these mutants significantly suppressed the intestinal UPRmt induced by ASI cco-1 KD (Supplementary Fig. 4n, o).
To further understand the neuronal coordination network responding to ASI cco-1 KD, we investigated the involvement of neuronal transmission. The induction of cell non-autonomous UPRmt was significantly reduced in unc-13 mutants (Fig. 4i, j), a gene essential for the release of small clear vesicles (SCV) and neurotransmitters61. In contrast, mutants in unc-31, required for the release of dense core vesicles (DCV) containing mainly neuropeptides62, or egl-21, which encodes carboxypeptidase for neuropeptides maturation63, showed no discernable effect on UPRmt activation in response to ASI cco-1 KD (Fig. 4i, j). These findings suggest that neurotransmitters are essential for cell non-autonomous UPRmt activation in response to ASI cco-1 KD.
The UPRmt induced by ASI cco-1 KD requires dopamine and is negatively regulated by GABA
To further elucidate the mechanism behind ASI-specific mitochondrial stress and systemic UPRmt activation, we examined the roles of neurotransmitters by crossing ASI specific cco-1 KD animals with various neurotransmitter production or packaging mutants. We found that loss of cat-2, a gene involved in dopamine synthesis64, significantly suppressed the UPRmt induced by ASI cco-1 KD (Fig. 5a, b). Additionally, we investigated the effect of four dopamine receptors (dop-1, dop-2, dop-3, and dop-4) on UPRmt activation64,65. Loss of the D1-like receptor dop-1 strongly suppressed cell non-autonomous UPRmt, while loss of the invertebrate-specific receptor dop-4 partially suppressed it in ASI cco-1 KD animals (Fig. 5c, d). These suppressions were restored by overexpressing each receptor in the nervous system (Supplementary Fig. 5a–d), but not in the intestine (Supplementary Fig. 5a–d), suggesting that dopamine functions in neurons to activate systemic UPRmt.
a Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, cat-2, tbh-1, tph-1, unc-17, eat-4, tdc-1 and unc-25 animals in gpa-4p::cco-1HP background. b Fluorescence intensities of hsp-6p::gfp in animals shown in (a) were quantified using ImageJ. n = 30 worms. c Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, dop-1, dop-2, dop-3 and dop-4 in gpa-4p::cco-1HP background. d Fluorescence intensities of hsp-6p::gfp in animals shown in (c) were quantified using ImageJ. n = 30 worms. e Representative fluorescence photomicrographs of hsp-6p::gfp in wild-type, unc-47, unc-30, unc-49, lgc-35, exp-1, gbb-1 and gbb-2 animals in gpa-4p::cco-1HP background. f Fluorescence intensities of hsp-6p::gfp in animals shown in (e) were quantified using ImageJ. n = 30 worms. p values were annotated via ordinary one-way ANOVA in (b), (d) and (f). Error bars, SEM. Scale bar, 100 μm. Source data are provided as a Source Data file. See also Supplementary Fig. 5.
Surprisingly, loss of unc-25, a gene involved in GABA synthesis, further induced intestinal UPRmt activation in ASI cco-1 KD animals, suggesting that GABA negatively regulate systemic UPRmt activation (Fig. 5a, b). We confirmed that UPRmt activation in response to ASI cco-1 KD was also further induced in unc-47 (GABA vesicular transporter) and unc-30 (homeodomain transcription factor of UNC-25 and UNC-47) mutants (Fig. 5e, f)66,67. Moreover, we examined several ionotropic GABA-gated ion channels (unc-49, lgc-35, and exp-1) and metabotropic GABA-sensitive G protein-coupled receptors (gbb-1 and gbb-2)68,69,70,71, finding that the mutation of exp-1 further induced the UPRmt (Fig. 5e, f). Moreover, the UPRmt that is activated by unc-25 mutants is strongly suppressed by daf-7 mutation, and partially suppressed by cat-2 mutation (Supplementary Fig. 5e–h). Additionally, the cat-2 mutant did not suppress UPRmt induced by pan-neuronal cco-1 knockdown while the unc-25 mutant further induced it (Supplementary Fig. 5i, j). These results indicate that systemic UPRmt activation in response to ASI cco-1 KD requires dopamine but is suppressed by GABA.
Mitochondrial perturbation in ASI neurons alters various physiological characteristics of animals
To broadly investigate the effects of ASI-specific mitochondrial perturbations (ASI cco-1 KD), we analyzed several physiological characteristics of the animals. Both ASI cco-1 KD and pan-neuronal cco-1 knockdown resulted in a reduced brood size in WT background, but not further reduced in daf-7 mutant backgrounds (Supplementary Fig. 6a). Additionally, these animals exhibited slightly delayed development and a reduced oxygen consumption rate, which persisted even in daf-7 mutant backgrounds (Supplementary Fig. 6b, c). However, ASI neuronal cco-1 KD did not affect the bending rates or crawling ability of these animals (Supplementary Fig. 6d, e).
Next, we explored how ASI-specific mitochondrial perturbations influence systemic metabolic states and longevity. We first assessed lifespan and stress resistance in animals with ASI cco-1 KD. Mitochondrial perturbation in ASI neurons alone was sufficient to extend lifespan, dependent on daf-7 in ASI neurons and daf-1 in RIM neurons (Fig. 6a, b and Supplementary Fig. 6f). Loss of daf-7 also suppressed the extended lifespan in the pan-neuronal cco-1 knockdown background (Fig. 6c and Supplementary Fig. 6g). Furthermore, ASI cco-1 KD animals survived significantly longer than control animals when exposed to the pathogen Pseudomonas aeruginosa (PA14), a pathogen that kills C. elegans within days of exposure72,73, which is dependent on daf-7 in ASI neurons and daf-1 in RIM neurons (Fig. 6d, e). Loss of daf-7 similarly suppressed pathogen resistance in the pan-neuronal cco-1 knockdown background (Fig. 6f). These data confirmed the crucial role of ASI-RIM axis in coordinating cell non-autonomous UPRmt, lifespan extension, and pathogen resistance via TGF-β signaling.
a Lifespan analysis of gpa-4p::cco-1HP animals with or without gpa-4p::daf-7HP. b Lifespan analysis of gpa-4p::cco-1HP animals with or without F23H12.7p::daf-1HP. c Lifespan analysis of neuronal cco-1 knockdown animals with or without daf-7 mutation. d Survival analysis of gpa-4p::cco-1HP animals with or without gpa-4p::daf-7HP feeding with P. aeruginosa. e Survival analysis of gpa-4p::cco-1HP animals with or without F23H12.7p::daf-1HP feeding with P. aeruginosa. f Survival analysis of neuronal cco-1 knockdown animals with or without daf-7 mutation feeding with P. aeruginosa. g Representative photomicrographs of Oil red O staining of control and gpa-4p::cco-1HP animals with or without daf-7 mutation. h Intensities of Oil red O staining signals in the entire intestine of animals shown in (g) were quantified using ImageJ. n = 10 worms. i Relative mRNA level of cpt-4 in control and gpa-4p::cco-1HP animals with or without daf-7 mutation. n = 3 biological replicates. j Representative photomicrographs of Oil red O staining of neuronal cco-1 knockdown animals with or without daf-7 or daf-1 mutation and the expression of daf-7p::daf-7(cDNA) or F23H12.7p::daf-1(cDNA). k Intensities of Oil red O staining signals in the entire intestine of animals shown in (j) were quantified using ImageJ. n = 33 worms. p values were annotated via ordinary one-way ANOVA in (h), (i) and (k). Error bars, SEM. Lifespan and survival analysis have been repeated at least two times. Scale bar, 100 μm. Source data are provided as a Source Data file.See also Supplementary Fig. 6 and Supplementary Table 1.
ASI neurons in C. elegans are known to sense food and nutritional status, regulate lipid metabolism, and modulate endocrine signaling pathways such as IIS and TGF-β32,74,75,76,77. Considering the close relationship among lipid metabolism, longevity and immune responses78,79,80,81, we measured lipids levels using Oil Red O staining, which stains neutral triglycerides and lipids82,83. Both ASI cco-1 KD and pan-neuronal cco-1 KD animals had significantly lower lipid levels compared to control animals (Fig. 6g, h and Supplementary Fig. 6h, i). To investigate which fat metabolic pathways were affected by ASI-mitochondrial stress, we performed qPCR analyses to examine the expression of genes involved in lipid metabolism. cpt-4, a homolog of CPT1 (carnitine palmitoyl transferase) responsible for translocating fatty acids from the cytosol to the mitochondrial matrix during fatty acid β-oxidation84,85, was significantly increased in both ASI cco-1 KD and pan-neuronal cco-1 KD background, while other lipid metabolism genes were not significantly affected (Fig. 6i and Supplementary Fig. 6j). This phenotype was confirmed using the cpt-4p::gfp reporter (Supplementary Fig. 6k, l).
The decreased fat storage was not dependent on known mitokine regulators including egl-20, unc-31, unc-13, flp-2, or srz-75 (Supplementary Fig. 6h, i). daf-7 is required for the lipid depletion in ASI cco-1 KD animals (Fig. 6g, h). Additionally, the elevated transcription level of cpt-4 in ASI cco-1 KD animals was abolished by the daf-7 mutation (Fig. 6i). Notably, both daf-7 and daf-1 mutation completely suppressed the reduced fat storage in ASI cco-1 KD and pan-neuronal cco-1 knockdown animals, which is restored by daf-7p::daf-7 or RIMp::daf-1 rescue (Fig. 6j, k). These results suggest that neuronal mitochondrial perturbation reduces body fat storage via the ASI-RIM axis and TGF-β signaling.
In summary, our study revealed that targeted mitochondrial stress in ASI neurons leads to systemic UPRmt activation and subsequent alterations in various physiological characteristics of the animals, highlight the complex interplay between mitochondrial function, TGF-β signaling, and organismal physiology.
Discussion
In this study, we discovered that the TGF-β/DAF-7 signaling pathway in the ASI-RIM modulates the organism’s response to pan-neuronal mitochondrial stress, transmitting this stress to the intestine, which induces UPRmt responses, altering lipid metabolic status, and influences aging. Specifically, perturbing mitochondrial function solely in ASI neurons is sufficient to induce cell non-autonomous UPRmt, extend lifespan, and enhance pathogen resistance. The GABA suppresses systemic UPRmt activation in response to ASI-specific mitochondrial stress, while dopamine is necessary for this activation. Together, our findings highlight a central role for TGF-β/DAF-7 in regulating the systemic mitochondrial stress response and various physiological characteristics via the ASI-RIM axis (Fig. 7).
Neuronal mitochondrial stress initiated a TGF-β signaling cascade of ASI sensory neurons and RIM interneurons axis that ensure cell non-autonomous UPRmt induction in peripheral tissues. ASI-specific neuronal mitochondrial stress induced cell non-autonomous UPRmt which was mediated by dopamine and negatively regulated by GABA, confer increased lifespan, enhanced pathogen tolerance, fat storage depletion, and various physiological changes. Created in BioRender. Wang, Z. (2023) BioRender.com/h64v432.
Morphogens serve as key signaling molecules in inter-tissue and inter-cell communication by establishing concentration gradients across tissues, ensuring that the entire organism adapts to stress in a coordinated manner86,87. The identification of Wnt as the initial morphogen functioning as a “mitokine” signal, orchestrating systemic mitochondrial stress adaptation, and influencing the aging process invites speculation about the broader role of morphogens as crucial signaling molecules in inter-tissue and inter-cell communication11,26,88,89. In this study, we further identified a crucial morphogen, TGF-β, in the regulation of inter-tissue mitochondrial stress communication. Previous studies suggest that TGF-β exhibits dual roles in mitochondrial function: it can impair mitochondrial activity and elevate reactive oxygen species in certain conditions, such as idiopathic pulmonary fibrosis and tumor effusion90,91,92, while also enhancing OXPHOS and ATP production in others, like metastatic breast cancer cells93,94. Interestingly, TGF-β signaling can mediate UPRmt and mitochondrial fission induced by extracellular matrix remodeling in human cells95. These findings align with the evolving understanding of morphogens, which are traditionally seen as key contributors to developmental processes. It highlights their significant role as regulators in mitochondrial adaptive responses, particularly in the context of aging and pathological processes.
It is important to note that TGF-β plays distinct roles in various stress responses. In our study, the loss of TGF-β/daf-7 suppressed cell non-autonomous UPRmt activation. In contrast, daf-7 mutants further enhanced cell non-autonomous UPRER in a neuronal xbp-1s expression background. This aligns with previous findings showing that intestinal UPRER activation is further increased in daf-7 mutants in response to neuronal xbp-1s overexpression96. Additionally, daf-7 is required for transforming environmental 1-undecene odor signal to RIM/RIC neurons for cell non-autonomous UPRER regulation97. While TGF-β/daf-7 signaling appears to suppress UPRmt activation under mitochondrial stress, it paradoxically enhances UPRER under ER stress induced by neuronal xbp-1s overexpression. These findings suggest a complex regulatory mechanism where TGF-β signaling modulates different stress pathways in a context-dependent manner.
Neuronal coordination of systemic mitochondrial stress responses plays a fundamental role in organismal metabolism and aging98. Mitochondria are crucial for lipid metabolism, particularly fatty acid β-oxidation, which helps regulate fat storage and is associated with metabolic disorders and obesity99. Lipid metabolism is also interconnected with aging78,79 and is essential for immune activation and pathogen defense80,81. In this study, we observed that manipulating mitochondrial function in just one pair of ASI sensory neurons is sufficient to induce fat depletion via activation of β-oxidation in distal tissues. This indicates broader physiological alterations in response to ASI-mitochondrial perturbations. This finding suggests that targeting sensory inputs or corresponding receptors in specific sensory neurons could be a promising approach to maintaining mitochondrial homeostasis and balancing fat storage levels.
The nervous system’s complexity often involves multiple signals acting antagonistically on a particular phenotype. In C. elegans, dopamine and GABA exemplify this by regulating the egg-laying rate. Our study adds to this understanding by revealing that UPRmt activated by ASI-specific mitochondrial dysfunction is positively regulated by dopamine but negatively regulated by GABA. Moreover, the systemic activation of UPRmt induced by the loss of GABA also requires TGF-β/DAF-7 but is only partially dependent on dopamine. This suggests a complex interaction between GABA, dopamine, and TGF-β in regulating UPRmt. Ultimately, the output of neuronal coordination of mitochondrial stress response is the balanced result of both suppression and activation signals, highlighting the intricate interplay within the nervous system to control organismal-wide stress response.
Methods
Caenorhabditis elegans maintenance
Studies were undertaken with C. elegans hermaphrodites. If needed, nematode males are obtained by heat shock. Nematodes were maintained and experimentally examined at 20°C on standard nematode growth medium agar plates seeded with Escherichia coli OP50, unless otherwise indicated. See all strains used or generated in Supplementary Table 2.
Transgenic strain construction
KOD-Plus-Neo (TOYOBO Cat#KOD-401) and KOD-Plus-Mutagenesis Kit (TOYOBO Cat#SMK-101) was used for PCR, ClonExpress Ultra One Step Cloning Kit (Vazyme Cat#C115-02) and Trelief® Seamless Cloning Kit (TSINGKE Cat#TSV-S3) was used for recombination.
Transgenic strains were generated by microinjecting target constructs (5–100 ng/μL) mixed with a pRF4(rol-6) (60 ng/μL), myo-3p::DsRed (25 ng/μL), odr-1p::RFP (50 ng/μL), unc-122p::RFP (50 ng/μL) co-injection maker. Integrated lines were generated using UV irradiation and outcrossed six times with control animals. The original plasmid backbone is pNB23 with the 5’utr region of unc-54. More detail information of the plasmids were listed in Supplementary Table 2.
RNAi feeding
Synchronized worms were bleached and grown from hatch on Escherichia coli HT115 strains containing an empty vector control or double-stranded RNA targeting to different genes. RNAi strains were from the Vidal library if present, or the Ahringer library if absent in the Vidal library.
CRISPR/Cas9 mediated gene editing
To generate exp-1(yth226) mutants, two single-guide RNA (sgRNA) targeting sequence (5’-TGAACCACTTGGACGGGGAG-3’, 5’-ACTGAACACTGATTCCGTAT-3’) of exp-1 were used. The CRISPR/Cas9 technology in accordance with published protocols100. The sgRNA vectors (40 ng/µL), the Cas9 vector (40 ng/µL) and the co-injection marker Rol-6 (80 ng/µL) were injected. Roller F1 worms were singled into new plates and examined by PCR amplification and sequencing. 2 x Rapid Taq Master Mix (Vazyme Cat#P222-03) was used for PCR, MinElute PCR Purification Kit (Qiagen Cat#28004) was used for DNA purification and QlAprep Spin Miniprep Kit (Qiagen Cat#27104) was used for plasmid extraction.
Imaging and analysis of the fluorescence intensity
For fluorescence image, worms were anesthetized with 50 mM sodium azide as soon as possible, and photographs were taken in no more than 5 min using a Leica M165 FC dissecting microscope and LAS X software 3.3 (Leica). Micrographs of specific neurons colocalization images were taken using a Zeiss Imager M2 microscope and ZEN (Zeiss) software. Fluorescent daf-7 and daf-1 expression images were taken using a Leica TCS SP8 microscope and LAS X software. Exposure times were the same within each experiment. To quantify the fluorescent intensity of each UPR reporters, the entire intestine regions were outlined and quantified using ImageJ 1.53k (Wayne Rasband (NIH)) software. To quantify daf-7p::gfp fluorescent intensity, each ASI neuron body were outlined and quantified using ImageJ. To quantify cpt-4p::gfp fluorescent intensity, the whole worm were outlined and quantified using ImageJ.
RNA isolation and quantitative PCR analyses
Total RNA was isolated using TRIzol (Invitrogen Cat#15596026). Worms were synchronized and washed off the plates using M9 buffer, and 1 mL TRIzol were added to the samples and homogenized by repeated freezing and thawing using liquid nitrogen for 6 times. RNA was isolated according to manufacturer’s instructions. DNA was wiped off and cDNA was synthesized using the ReverTra Ace® qPCR RT Master Mix with gDNA remover (Toyobo Cat#FSQ-301). Gene expression levels were determined by real-time PCR using SYBR Green Realtime PCR Master Mix (Toyobo Cat#QPK-201) and Bio-rad CFX96/384 Real-Time PCR Detection Systems. Relative gene expression was normalized to act-3 (T04C12.4) or act-1 (T04C12.6) mRNA levels. Fold changes in gene expression were calculated using the comparative ΔΔCt method, and then normalized to the control for every single biological repeat. The primer sequences used in the quantitative PCR are shown in Supplementary Table 3.
Lifespan analysis
Lifespan experiments were performed on NGM plates at 20 °C as previously described101. While temperature shift experiments, synchronized eggs were seeded at 20 °C and then transferred to 25 °C at the L4 stage. To prevent progeny production, 150 µL 10 mg/mL 5-fluoro-20-deoxyuridine (FUdR (Aladdin Cat#F110732)) was added to seeded plates. Worms were synchronized by bleach and were grown on OP50 from hatch, and transited to FUdR plates from day 1 of adulthood, then transferred a second time at day 5 of adulthood. Animals were counted every second day and were scored as dead if they failed to respond when touched. Prism 10.2.3 software (GraphPad Software) was used for statistical analysis. Log-rank (Mantel-Cox) method was used to determine the significance difference. All survival data are available in Supplementary Table 1.
P. aeruginosa slow-killing assay
Slow-killing experiments were performed as previously described102. P. aeruginosa was cultured overnight in LB containing 50 µg/ml kanamycin at 37 C and then seeded onto slow-killing nematode growth medium (NGM) agar plates (with 0.35% peptone). Plates were allowed to dry at room temperature, incubated at 37 °C for 24 h and then balanced at room temperature for another 24 h. 100 μl 10 mg/ml FUdR was added in the slow-killing plates. Synchronized day 1 worms were transferred from E. coli NGM plates to slow-killing plates and maintained at 25 °C. Animals were counted at the described times and were scored as dead if they failed to respond when touched. Prism 10.2.3 software was used for statistical analysis. Log-rank (Mantel-Cox) method was used to determine the significance difference. Each experiment was repeated at least two times. All original data are available in Supplementary Table 1.
Brood size assay
At least 10 synchronized L4 worms were placed onto NGM plates with seeded OP50 at 20 °C and transferred to fresh NGM plates every day until they stopped laying eggs. The total number of progenies was counted for each strain. Each experiment was repeated for three times.
Developmental rate assay
Worms were synchronized and about 300 eggs of each strain were placed onto NGM plates with seeded OP50. These worms grow at 17.5 °C for 108 h and the number of worms at each developmental stage was calculated.
Motility assay
For bending ability test, day 1 worms were transferred to a drop of M9 buffer. After 1 min of adaptation, the number of body bends during 20 s was counted. A body bend was defined as change in the direction of bending in the middle of the body103.
For crawling ability test, day 1 worms were transferred to an empty NGM plate without OP50. After 1 min of adaptation, 20 s of crawling movies were captured via Wormaction1858 (Zebra Scientific Development) software, and the center point speed was analyzed via WormLab (MBF Bioscience) software.
Measurement of oxygen consumption rate
Oxygen consumption rate (OCR) was measured using a Seahorse XFe96 analyzer (Seahorse Bioscience) at 20 °C as described previously104. Fifteen synchronized D1 worms of each strain were transferred into each well of a 96-well microplate containing 200 μL M9, 5 wells per strain. Basal respiration was measured for a total of 90 min, in 9 min intervals that included a 3 min mix, a 3 min time delay and a 3 min measurement. Experiments were repeated at least three times for each strain.
Oil-red-O staining and quantification
The Oil-Red-O (ORO) staining was performed as previously described105. Briefly, synchronized day 1 worms were collected and washed three times with M9 + 0.1% Triton X-100 (Sigma Cat#T8787). Worms were suspended in 100 µL M9, and then fix worms by adding 100 µL 4% paraformaldehyde (PFA). After 1 h incubating, wash worms with M9 + 0.1% Triton X-100 3 times to remove PFA and resuspension in 1 mL 60% isopropanol to dehydrate. Worms were then incubated with rocking in 500 μL 60% ORO solution (Abcam Cat#ab146295, 0.5 g/100 mL isopropanol stock solution, freshly dilute with ddH2O, rock for several hours and filter with a 0.45-µm filter) for 2.5 h at 20 °C. Stained worms were washed with M9 + 0.1% Triton X-100 3 times, and then imaged using a Leica M165 FC dissecting microscope. To quantify ORO signals, the channels of images were split and the mean intensity in the red color channel of the post pharyngeal intestine was measured using Image J software.
Statistical analysis
The statistical analyses were performed using GraphPad Prism 10.2.3. Statistical significance was calculated by unpaired t-test, ordinary one-way ANOVA and two-way ANOVA as annotated. The data were presented as the means ± SEM with indicated p values.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Additional data related to this paper may be requested from authors. All strains synthesized in this manuscript are derivatives of N2 or other strains from CGC and are either made available on CGC or available upon request. Source data are provided with this paper.
References
Singh, J. & Aballay, A. Neural control of behavioral and molecular defenses in C. elegans. Curr. Opin. Neurobiol. 62, 34–40 (2020).
Srinivasan, S. Regulation of body fat in Caenorhabditis elegans. Annu Rev. Physiol. 77, 161–178 (2015).
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011).
Shao, L. W., Niu, R. & Liu, Y. Neuropeptide signals cell non-autonomous mitochondrial unfolded protein response. Cell Res 26, 1182–1196 (2016).
Berendzen, K. M. et al. Neuroendocrine coordination of mitochondrial stress signaling and proteostasis. Cell 166, 1553–1563.e1510 (2016).
Shen, K. et al. The germline coordinates mitokine signaling. Cell 187, 4605–4620.e4617 (2024).
Chen, L. T. et al. Neuronal mitochondrial dynamics coordinate systemic mitochondrial morphology and stress response to confer pathogen resistance in C. elegans. Dev. Cell 56, 1770–1785.e1712 (2021).
Dishart, J. G. et al. Olfaction regulates peripheral mitophagy and mitochondrial function. Sci. Adv. 10, eadn0014 (2024).
Kang, G. M. et al. Mitohormesis in hypothalamic POMC neurons mediates regular exercise-induced high-turnover metabolism. Cell Metab. 33, 334–349.e336 (2021).
Gomez-Valades, A. G. et al. Mitochondrial cristae-remodeling protein OPA1 in POMC neurons couples Ca(2+) homeostasis with adipose tissue lipolysis. Cell Metab. 33, 1820–1835.e1829 (2021).
Zhang, Q. et al. The mitochondrial unfolded protein response is mediated cell-non-autonomously by retromer-dependent Wnt signaling. Cell 174, 870–883 (2018).
Liu, Y. et al. Two sensory neurons coordinate the systemic mitochondrial stress response via GPCR signaling in C. elegans. Dev. Cell 57, 2469–2482.e2465 (2022).
Cheruku, H. R. et al. Transforming growth factor-β, MAPK and Wnt signaling interactions in colorectal cancer. EuPA Open Proteom. 8, 104–115 (2015).
Zhu, T. et al. The S100 calcium binding protein A11 promotes liver fibrogenesis by targeting TGF-beta signaling. J. Genet Genomics 49, 338–349 (2022).
Yousefi, F. et al. TGF-beta and WNT signaling pathways in cardiac fibrosis: Non-coding RNAs come into focus. Cell Commun. Signal 18, 87 (2020).
Dzialo, E., Tkacz, K. & Blyszczuk, P. Crosstalk between the TGF-beta and WNT signalling pathways during cardiac fibrogenesis. Acta Biochim Pol. 65, 341–349 (2018).
Zhang, K., Zhang, M., Luo, Z., Wen, Z. & Yan, X. The dichotomous role of TGF-beta in controlling liver cancer cell survival and proliferation. J. Genet Genomics 47, 497–512 (2020).
Ren, P. et al. Control of C. elegans larval development by neuronal expression of a TGF-beta homolog. Science 274, 1389–1391 (1996).
Gallagher, T., Kim, J., Oldenbroek, M., Kerr, R. & You, Y. J. ASI regulates satiety quiescence in C. elegans. J. Neurosci. 33, 9716–9724 (2013).
Meisel, J. D., Panda, O., Mahanti, P., Schroeder, F. C. & Kim, D. H. Chemosensation of bacterial secondary metabolites modulates neuroendocrine signaling and behavior of C. elegans. Cell 159, 267–280 (2014).
Moore, R. S., Kaletsky, R. & Murphy, C. T. Piwi/PRG-1 argonaute and TGF-beta mediate transgenerational learned pathogenic avoidance. Cell 177, 1827–1841 (2019).
Greer, E. R., Perez, C. L., Van Gilst, M. R., Lee, B. H. & Ashrafi, K. Neural and molecular dissection of a C. elegans sensory circuit that regulates fat and feeding. Cell Metab. 8, 118–131 (2008).
You, Y. J., Kim, J., Raizen, D. M. & Avery, L. Insulin, cGMP, and TGF-beta signals regulate food intake and quiescence in C. elegans: a model for satiety. Cell Metab. 7, 249–257 (2008).
Wexler, L. R., Miller, R. M. & Portman, D. S. C. elegans males integrate food signals and biological sex to modulate state-dependent chemosensation and behavioral prioritization. Curr. Biol. 30, 2695–2706.e2694 (2020).
Morita, K., Chow, K. L. & Ueno, N. Regulation of body length and male tail ray pattern formation of Caenorhabditis elegans by a member of TGF-beta family. Development 126, 1337–1347 (1999).
Zhang, Q. et al. The memory of neuronal mitochondrial stress is inherited transgenerationally via elevated mitochondrial DNA levels. Nat. Cell Biol. 23, 870–880 (2021).
Krishna, S., Maduzia, L. L. & Padgett, R. W. Specificity of TGFbeta signaling is conferred by distinct type I receptors and their associated SMAD proteins in Caenorhabditis elegans. Development 126, 251–260 (1999).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 8, 519–529 (2007).
Taylor, R. C. & Dillin, A. XBP-1 is a cell-nonautonomous regulator of stress resistance and longevity. Cell 153, 1435–1447 (2013).
Douglas, P. M. et al. Heterotypic signals from neural HSF-1 separate thermotolerance from longevity. Cell Rep. 12, 1196–1204 (2015).
Link, C. D., Cypser, J. R., Johnson, C. J. & Johnson, T. E. Direct observation of stress response in Caenorhabditis elegans using a reporter transgene. Cell Stress Chaperones 4, 235–242 (1999).
Schackwitz, W. S., Inoue, T. & Thomas, J. H. Chemosensory neurons function in parallel to mediate a pheromone response in C. elegans. Neuron 17, 719–728 (1996).
Hammarlund, M., Hobert, O., Miller, D. M. 3rd & Sestan, N. The CeNGEN project: The complete gene expression map of an entire nervous system. Neuron 99, 430–433 (2018).
Taylor, S. R. et al. Molecular topography of an entire nervous system. Cell 184, 4329–4347 (2021).
Barrett, A. et al. A head-to-head comparison of ribodepletion and polyA selection approaches for Caenorhabditis elegans low input RNA-sequencing libraries. G3 (Bethesda) 11, jkab121 (2021).
Jansen, G. et al. The complete family of genes encoding G proteins of Caenorhabditis elegans. Nat. Genet 21, 414–419 (1999).
Tobin, D. M. et al. Combinatorial expression of TRPV channel proteins defines their sensory functions and subcellular localization in C. elegans neurons. Neuron 35, 307–318 (2002).
Flames, N. & Hobert, O. Gene regulatory logic of dopamine neuron differentiation. Nature 458, 885–889 (2009).
Ortiz, C. O. et al. Searching for neuronal left/right asymmetry: genomewide analysis of nematode receptor-type guanylyl cyclases. Genetics 173, 131–149 (2006).
Sengupta, P., Chou, J. H. & Bargmann, C. I. odr-10 encodes a seven transmembrane domain olfactory receptor required for responses to the odorant diacetyl. Cell 84, 899–909 (1996).
Miranda-Vizuete, A. et al. Lifespan decrease in a Caenorhabditis elegans mutant lacking TRX-1, a thioredoxin expressed in ASJ sensory neurons. FEBS Lett. 580, 484–490 (2006).
Georgi, L. L., Albert, P. S. & Riddle, D. L. daf-1, a C. elegans gene controlling Dauer larva development, encodes a novel receptor protein kinase. Cell 61, 635–645 (1990).
Estevez, M. et al. The daf-4 gene encodes a bone morphogenetic protein receptor controlling C. elegans Dauer larva development. Nature 365, 644–649 (1993).
Shaw, W. M., Luo, S., Landis, J., Ashraf, J. & Murphy, C. T. The C. elegans TGF-beta Dauer pathway regulates longevity via insulin signaling. Curr. Biol. 17, 1635–1645 (2007).
Inoue, T. & Thomas, J. H. Targets of TGF-beta signaling in Caenorhabditis elegans Dauer formation. Dev. Biol. 217, 192–204 (2000).
Thomas, J. H., Birnby, D. A. & Vowels, J. J. Evidence for parallel processing of sensory information controlling dauer formation in Caenorhabditis elegans. Genetics 134, 1105–1117 (1993).
da Graca, L. S. et al. DAF-5 is a Ski oncoprotein homolog that functions in a neuronal TGF beta pathway to regulate C. elegans Dauer development. Development 131, 435–446 (2004).
Patterson, G. I., Koweek, A., Wong, A., Liu, Y. & Ruvkun, G. The DAF-3 Smad protein antagonizes TGF-beta-related receptor signaling in the Caenorhabditis elegans Dauer pathway. Genes Dev. 11, 2679–2690 (1997).
Tewari, M. et al. Systematic interactome mapping and genetic perturbation analysis of a C. elegans TGF-beta signaling network. Mol. Cell 13, 469–482 (2004).
Riddle D. L., Albert P. S. Genetic and environmental regulation of Dauer larva development. In: C. elegans II (eds Riddle D. L., Blumenthal T., Meyer B. J., Priess J. R.). 2nd edn (1997).
Alkema, M. J., Hunter-Ensor, M., Ringstad, N. & Horvitz, H. R. Tyramine functions independently of octopamine in the Caenorhabditis elegans nervous system. Neuron 46, 247–260 (2005).
Brockie, P. J., Madsen, D. M., Zheng, Y., Mellem, J. & Maricq, A. V. Differential expression of glutamate receptor subunits in the nervous system of Caenorhabditis elegans and their regulation by the homeodomain protein UNC-42. J. Neurosci. 21, 1510–1522 (2001).
Collet, J., Spike, C. A., Lundquist, E. A., Shaw, J. E. & Herman, R. K. Analysis of osm-6, a gene that affects sensory cilium structure and sensory neuron function in Caenorhabditis elegans. Genetics 148, 187–200 (1998).
Much, J. W., Slade, D. J., Klampert, K., Garriga, G. & Wightman, B. The fax-1 nuclear hormone receptor regulates axon pathfinding and neurotransmitter expression. Development 127, 703–712 (2000).
Schiffer, J. A. et al. Caenorhabditis elegans processes sensory information to choose between freeloading and self-defense strategies. eLife 9, e56186 (2020).
Park, D., Estevez, A. & Riddle, D. L. Antagonistic Smad transcription factors control the Dauer/non-Dauer switch in C. elegans. Development 137, 477–485 (2010).
Park, D. et al. Repression of a potassium channel by nuclear hormone receptor and TGF-beta signaling modulates insulin signaling in Caenorhabditis elegans. PLoS Genet 8, e1002519 (2012).
Li, X. et al. Protein disulfide isomerase PDI-6 regulates Wnt secretion to coordinate inter-tissue UPR(mt) activation and lifespan extension in C. elegans. Cell Rep. 39, 110931 (2022).
Tian, Y. et al. Mitochondrial stress induces chromatin reorganization to promote longevity and UPR(mt). Cell 165, 1197–1208 (2016).
Zhu, D. et al. NuRD mediates mitochondrial stress-induced longevity via chromatin remodeling in response to acetyl-CoA level. Sci. Adv. 6, eabb2529 (2020).
Madison, J. M., Nurrish, S. & Kaplan, J. M. UNC-13 interaction with syntaxin is required for synaptic transmission. Curr. Biol. 15, 2236–2242 (2005).
Speese, S. et al. UNC-31 (CAPS) is required for dense-core vesicle but not synaptic vesicle exocytosis in Caenorhabditis elegans. J. Neurosci. 27, 6150–6162 (2007).
Jacob, T. C. & Kaplan, J. M. The EGL-21 carboxypeptidase E facilitates acetylcholine release at Caenorhabditis elegans neuromuscular junctions. J. Neurosci. 23, 2122–2130 (2003).
Chase D. L., Koelle M. R. Biogenic amine neurotransmitters in C. elegans. WormBook, 1-15 (2007).
Suo, S., Sasagawa, N. & Ishiura, S. Cloning and characterization of a Caenorhabditis elegans D2-like dopamine receptor. J. Neurochem 86, 869–878 (2003).
Eastman, C., Horvitz, H. R. & Jin, Y. Coordinated transcriptional regulation of the unc-25 glutamic acid decarboxylase and the unc-47 GABA vesicular transporter by the Caenorhabditis elegans UNC-30 homeodomain protein. J. Neurosci. 19, 6225–6234 (1999).
McIntire, S. L., Reimer, R. J., Schuske, K., Edwards, R. H. & Jorgensen, E. M. Identification and characterization of the vesicular GABA transporter. Nature 389, 870–876 (1997).
Jobson, M. A. et al. Spillover transmission is mediated by the excitatory GABA receptor LGC-35 in C. elegans. J. Neurosci. 35, 2803–2816 (2015).
Beg, A. A. & Jorgensen, E. M. EXP-1 is an excitatory GABA-gated cation channel. Nat. Neurosci. 6, 1145–1152 (2003).
Dittman, J. S. & Kaplan, J. M. Behavioral impact of neurotransmitter-activated G-protein-coupled receptors: muscarinic and GABAB receptors regulate Caenorhabditis elegans locomotion. J. Neurosci. 28, 7104–7112 (2008).
Bamber, B. A., Beg, A. A., Twyman, R. E. & Jorgensen, E. M. The Caenorhabditis elegans unc-49 locus encodes multiple subunits of a heteromultimeric GABA receptor. J. Neurosci. 19, 5348–5359 (1999).
Tan, M. W., Mahajan-Miklos, S. & Ausubel, F. M. Killing of Caenorhabditis elegans by Pseudomonas aeruginosa used to model mammalian bacterial pathogenesis. Proc. Natl Acad. Sci. USA 96, 715–720 (1999).
Pellegrino, M. W. et al. Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection. Nature 516, 414–417 (2014).
Tataridas-Pallas, N. et al. Neuronal SKN-1B modulates nutritional signalling pathways and mitochondrial networks to control satiety. PLoS Genet 17, e1009358 (2021).
Dixit, A. et al. Neuronal control of lipid metabolism by STR-2 G protein-coupled receptor promotes longevity in Caenorhabditis elegans. Aging Cell 19, e13160 (2020).
An, L., Fu, X., Chen, J. & Ma, J. Application of Caenorhabditis elegans in lipid metabolism research. Int J. Mol. Sci. 24, 1173 (2023).
Srinivasan, S. Neuroendocrine control of lipid metabolism: lessons from C. elegans. J. Neurogenet. 34, 482–488 (2020).
Wang, M. C., O’Rourke, E. J. & Ruvkun, G. Fat metabolism links germline stem cells and longevity in C. elegans. Science 322, 957–960 (2008).
Han, S. et al. Mono-unsaturated fatty acids link H3K4me3 modifiers to C. elegans lifespan. Nature 544, 185–190 (2017).
Nandakumar, M. & Tan, M. W. Gamma-linolenic and stearidonic acids are required for basal immunity in Caenorhabditis elegans through their effects on p38 MAP kinase activity. PLoS Genet 4, e1000273 (2008).
Anderson, S. M. et al. The fatty acid oleate is required for innate immune activation and pathogen defense in Caenorhabditis elegans. PLoS Pathog. 15, e1007893 (2019).
Soukas, A. A., Kane, E. A., Carr, C. E., Melo, J. A. & Ruvkun, G. Rictor/TORC2 regulates fat metabolism, feeding, growth, and life span in Caenorhabditis elegans. Genes Dev. 23, 496–511 (2009).
O’Rourke, E. J., Soukas, A. A., Carr, C. E. & Ruvkun, G. C. elegans major fats are stored in vesicles distinct from lysosome-related organelles. Cell Metab. 10, 430–435 (2009).
Dasgupta, M. et al. NHR-49 transcription factor regulates immunometabolic response and survival of Caenorhabditis elegans during Enterococcus faecalis infection. Infect. Immun. 88, e00130–00120 (2020).
Lee, K., Kerner, J. & Hoppel, C. L. Mitochondrial carnitine palmitoyltransferase 1a (CPT1a) is part of an outer membrane fatty acid transfer complex. J. Biol. Chem. 286, 25655–25662 (2011).
Zhang, Q., Wang, Z. & Tian, Y. Inter-tissue communication of mitochondrial stress in aging. Yi Chuan 45, 187–197 (2023).
Li, J., Cui, J. & Tian, Y. Neuron-periphery mitochondrial stress communication in aging and diseases. Life Med. 1, 168–178 (2022).
Zhang, H. et al. Inter-tissue communication of mitochondrial stress and metabolic health. Life Metab. 2, 1–11 (2023).
Zhang, Q. & Tian, Y. Molecular insights into the transgenerational inheritance of stress memory. J. Genet Genomics 49, 89–95 (2022).
Yoon, Y. S., Lee, J. H., Hwang, S. C., Choi, K. S. & Yoon, G. T. G. F. beta1 induces prolonged mitochondrial ROS generation through decreased complex IV activity with senescent arrest in Mv1Lu cells. Oncogene 24, 1895–1903 (2005).
Dimeloe, S. et al. Tumor-derived TGF-beta inhibits mitochondrial respiration to suppress IFN-gamma production by human CD4(+) T cells. Sci. Signal 12, eaav3334 (2019).
Zhang, N. et al. TGF-beta changes cyto/mito-ribosome balance to target respiratory chain complex V biogenesis in pulmonary fibrosis therapy. Signal Transduct. Target. Ther. 8, 136 (2023).
Gabriel, S. S. et al. Transforming growth factor-beta-regulated mTOR activity preserves cellular metabolism to maintain long-term T cell responses in chronic infection. Immunity 54, 1698–1714 (2021).
Liu, Q. Q. et al. TGF‑beta1‑induced epithelial‑mesenchymal transition increases fatty acid oxidation and OXPHOS activity via the p‑AMPK pathway in breast cancer cells. Oncol. Rep. 44, 1206–1215 (2020).
Zhang, H. et al. The extracellular matrix integrates mitochondrial homeostasis. Cell 187, 4289–4304 (2024).
Ozbey, N. P. et al. Tyramine acts downstream of neuronal XBP-1s to coordinate inter-tissue UPR(ER) activation and behavior in C. elegans. Dev. Cell 55, 754–770.e756 (2020).
De-Souza, E. A., Thompson, M. A. & Taylor, R. C. Olfactory chemosensation extends lifespan through TGF-beta signaling and UPR activation. Nat. Aging 3, 938–947 (2023).
Chen, P. X., Zhang, L., Chen, D. & Tian, Y. Mitochondrial stress and aging: Lessons from C. elegans. Semin Cell Dev. Biol. 154, 69–76 (2024).
Littlejohn, N. K., Seban, N., Liu, C. C. & Srinivasan, S. A feedback loop governs the relationship between lipid metabolism and longevity. eLife 9, e58815 (2020).
Waaijers, S. et al. CRISPR/Cas9-targeted mutagenesis in Caenorhabditis elegans. Genetics 195, 1187–1191 (2013).
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002).
Kirienko, N. V., Cezairliyan, B. O., Ausubel, F. M. & Powell, J. R. Pseudomonas aeruginosa PA14 pathogenesis in Caenorhabditis elegans. Methods Mol. Biol. 1149, 653–669 (2014).
Calculli, G. et al. Systemic regulation of mitochondria by germline proteostasis prevents protein aggregation in the soma of C. elegans. Sci. Adv. 7, eabg3012 (2021).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Ching, WangF. Y. TT. Oil Red O Staining for Lipid Content in Caenorhabditis elegans. Bio Protoc. 11, e4124 (2021).
Acknowledgements
We thank Tian lab members for the insightful discussions and Y. Liu for helping with strain maintenance. In this study, Y.T. is supported by the National Key R&D Program of China (2022YFA1303000), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB39000000), the National Natural Science Foundation of China (31930023, 32225025, 92254305, 32321004), the CAS Project for Young Scientists in Basic Research (YSBR-076). Q.Z. was supported by the China National Postdoctoral Program for Innovative Talents (BX2021356), the National Natural Science Foundation of China (32471212, 32200624) and the China Postdoctoral Science Foundation (2021M703474). Several C. elegans strains used in this work were provided by the Caenorhabditis Genetics Center (CGC), funded by the NIH P40OD010440. Several deletion worm strains were also provided by the Japanese National BioResource Project.
Author information
Authors and Affiliations
Contributions
Y.T., Z.W., and Q.Z. conceived the study and wrote the manuscript. Z.W. performed the C. elegans crosses and strain generations. Z.W. performed fluorescence microscopy, RNAi, qPCR, P. aeruginosa slow-killing, motility, and oxygen consumption rate experiments. Z.W. and Q.Z. performed the Oil-Red-O staining, and life-span experiments. Z.W. and Y.J. performed plasmids construction and microinjection. Z.W., Q.Z. and J.Z. performed the brood size and developmental rate experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Z., Zhang, Q., Jiang, Y. et al. ASI-RIM neuronal axis regulates systemic mitochondrial stress response via TGF-β signaling cascade. Nat Commun 15, 8997 (2024). https://doi.org/10.1038/s41467-024-53093-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53093-9
This article is cited by
-
Mitochondrial stress orchestrates chromatin remodeling and longevity via phosphoregulation of the NuRD component LIN-40
Science China Life Sciences (2025)