Abstract
Food, especially plant-based diet, has complex chemical diversity. However, large-scale phytonutrient-metabolizing activities of gut bacteria are largely unknown. Here we integrated and systematically analysed multiple databases containing information on enzymatic reactions and food health benefits, and 3,068 global public human microbiomes. Transformation of 775 phytonutrients from edible plants was associated with enzymes encoded by diverse gut microbes. In vitro assays validated the biotransformation activity of gut species, for example, Eubacterium ramulus. The biotransformation of phytonutrients demonstrated high interpersonal and geographical variability. Machine learning models based on 2,486 public case–control microbiomes, using the abundances of enzymes associated with modification of phytonutrients present in health-associated foods, discriminated the health status of individuals in multiple disease contexts, suggesting altered biotransformation potential in disease. We validated the association of microbiome-encoded enzymes with the anti-inflammatory activity of common edible plants by combining metagenomics and metatranscriptomics analysis in specific-pathogen-free and germ-free mice. These findings have implications for designing precise, personalized diets to guide an individual towards a healthy state.
Similar content being viewed by others
Main
Definitions of nutritious diets continuously evolve along with understanding of how foods, essential nutrients and other dietary components influence health and disease1. Adopting dietary patterns emphasizing plant-based foods, including fruits, vegetables, whole grains, legumes, seeds and nuts, rather than a conventional Western diet2, can enhance health3,4 and environmental sustainability5. However, the appealing concept of universally applicable dietary strategies to prevent or treat disease is undercut by the heterogeneous responses to identical foods6,7,8. One primary source of variability is the gut microbiome7, as its composition contributes to responses to foods (for example, glycaemic index9) and its metabolic output altered by diets impacts physiological homeostasis and metabolic disease progression10,11,12, emphasizing the need to consider microbiome in personalized dietary recommendations.
Previous dietary research was predominantly based on 150 macronutrients, such as fat, protein and carbohydrate13, and on correlating overall dietary patterns, including red meat, fish and vegetable consumption, with health outcomes14 rather than from a biochemical standpoint. Yet human food has vast chemical diversity, comprising >26,000 mainly plant-derived distinct small molecules13, many bioactive and, additionally, subject to biotransformation by the gut microbiota’s enormous enzymatic repository15.
Compared with research on gut microbial metabolism of drugs15,16, research on the biotransformation of plant-derived food compounds (hereafter termed phytonutrients) by gut microbiota and its relationship to human health is only just emerging17. Phytonutrients associated with certain bacterial classes have been identified18, yet most studies have focused on specific compounds or sets of microbes, but not on a large scale. Understanding the immense ‘black box’ of biotransformation reactions and building a scientific basis for developing a new generation of foods and pre-, pro- and post-biotics targeted to individuals to thwart disease remain future goals.
We systematically investigated this black box, linking 775 phytonutrients to microbial enzymes associated with diverse microbes from a global dataset of 3,068 human gut microbiomes. Our findings suggest that ~70% of gut microbial enzymes are potentially involved in phytonutrient biotransformation. We showed in mice that the benefits of a healthy diet largely depend on the presence and transcriptional activity of certain microbial enzymes. Furthermore, the ability of gut bacteria to biotransform a healthy diet differs between health and disease, suggesting that once a dysbiotic microbiota is established, broad ‘one-size-fits-most’ dietary recommendations may have limited value.
Results
Gut microbial enzymes are genomically linked to hundreds of phytonutrients
To assess microbial enzymatic potential for metabolizing plant-derived phytonutrients, we systematically mapped dietary compounds to the gut microbiome. We retrieved 7,825 low-molecular-weight phytonutrients associated with edible plants (from NutriChem 2.0) and attempted to link them to enzymatic reactions, successfully linking >1,500 compounds to enzymes with unique Enzyme Commission (EC) numbers. Filtering out compounds incorrectly designated as natural plant compounds left 1,388 phytonutrients linked to 4,678 enzymes (Fig. 1a).
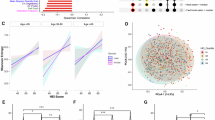
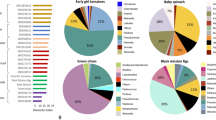
a, Graphical representation of the study design, methodology and databases used for linking the gut microbiota with phytonutrients. Metadata associated with the 3,068 shotgun metagenomic samples are shown (BMI, age, sex, sequencing depth (that is,reads number)). Created with BioRender.com. b, Barplot showing the chemical classes of phytonutrients. ‘Total’ refers to all phytonutrients present in NutriChem 2.0 linked with enzymes, and ‘microbiome-associated’ refers only to phytonutrients associated with gut bacterial enzymes. c, Barplot showing the enzyme classes. ‘Total’ indicates all phytonutrient-associated enzymes, and ‘microbiome-associated’ indicates phytonutrient-associated microbial enzymes. d, Left: phylogenetic tree of bacterial species with enzymatic potential to modify phytonutrients. Tracks correspond to (from inside to outside) species and associated phylum, signature species in each geographical region, relative abundance and prevalence of each species, and the number of phytonutrients potentially modifiable by each species. Right: scatterplots depicting the two-sided Spearman’s correlation of species abundance (left) and species prevalence (right) with the number of phytonutrients. White line and grey area show the linear regression line and 95% confidence interval. e, Barplot displaying the proportion of phytonutrients in 21 healthy edible plants biotransformed by gut bacterial enzymes. The proportion is calculated as the ratio of phytonutrients linked to gut bacterial enzymes to the total number of phytonutrients with a known enzymatic assignment. Size of bubbles corresponds to total number of phytonutrients for each edible plant.
Next, we identified phytonutrients associated with gut microbial enzymes, using a shotgun metagenomics cohort comprising 3,068 non-diseased human gut microbiomes from Europe (N = 1,379), Asia (N = 476), Africa (N = 326), Oceania (N = 103) and America (N = 784) (Supplementary Table 1). Taxonomic and functional profiling using MetaPhlAn3 and the stratified output of HUMAnN3 resulted in the annotation of 959 species and 2,855 enzymes. Accumulation curves indicated that plateaus were reached for every geographical region in both species and enzymes (Fig. 1a and Extended Data Fig. 1a). After linking the annotated gut microbiome enzymes with the 1,388 phytonutrients, ~67% of all enzymes annotated (N = 1,908) appeared to be involved in the potential biotransformation of 775 phytonutrients and 1,118 edible plants; ~64% (N = 1,226) were found only in gut microbial species but not in the human genome, according to the KEGG Enzyme database. To evaluate the robustness of the functional annotations, we compared the associated phytonutrients using HUMAnN3 and metagenome-assembled genomes (MAGs) in 200 randomly selected samples. The two methods showed high consistency, with 656 common phytonutrients (82.21% of MAGs and 91.49% of HUMAnN3 results; Supplementary Table 2). In addition, benchmarking against the ChocoPhlAn3 database showed that for most species, >90% of ECs in the reference pangenome of one species were also detected by our HUMAnN3 pipeline (Extended Data Fig. 1b).
Gut microbe-linked phytonutrients with known classification were preponderantly terpenoids, flavonoids and alkaloids, and the enzymes were mostly oxidoreductases, transferases and hydrolases (Fig. 1b,c). Taxonomic annotation of the enzymes suggests that all phyla are capable of phytonutrient biotransformation (Fig. 1d), with Proteobacteria (72.5%) and Bacteroidetes (71.2%) most commonly identified, followed by Firmicutes (58.4%) and Actinobacteria (48.3%), based on the species percentage per phylum (≥15 species and 200 phytonutrients). Abundance and prevalence of gut bacterial species correlated positively with their genomic capability to biotransform phytonutrients (Fig. 1d).
Among the 1,118 edible plants, we zeroed in on 21 common foods considered healthy dietary choices19, finding that more than half of their identified phytonutrients were linked to gut microbial enzymes (Fig. 1e). Some microbiome-associated biotransformations we identified were previously linked to health, including glycosylation of phloretin20 by EC 2.4.1.4 and conversion of phloretin to 3-(4-hydroxyphenyl)propanoic acid21 by chalcone isomerase (EC 5.5.1.6) (Supplementary Table 2). Overall, our analysis demonstrated vast phytonutrient-biotransformation potential among gut microbiota.
Sparse gut bacteria linked to high biotransformation potential
We next asked whether such biotransformation reactions are also performed by well-characterized, commercially available probiotics and whether they occur in vitro. Upon annotating the genomes of 59 probiotics (Supplementary Table 4), we found that enzymes in these strains can potentially biotransform 525 of the 775 phytonutrients linked to gut microbial enzymes (Extended Data Fig. 1c). Moreover, among the 186 phytonutrients in secondary-metabolism-related pathways (according to KEGG), 116 were modified by both probiotic and gut microbial enzymes, while 70 were uniquely associated with gut microbial enzymes (Fig. 2a; mean number of modifying species of 158 and 8, respectively, based on a 5% prevalence filter).
a, Pie chart showing the numbers of secondary-metabolism-related phytonutrients modifiable either exclusively by gut microbes (designated ‘gut-restricted’) or by gut microbes and probiotics (designated ‘shared’). The heat map at left depicts the enzymatic potential of gut microbial species to modify secondary-metabolism-related phytonutrients (green/light grey indicate presence/absence, respectively, of enzymatic potential). Selected species associated with ‘gut-restricted’ phytonutrients are shown. Examples of phytonutrients categorized into ‘shared’ and ‘gut-restricted’ space are shown, with the enriched functional groups highlighted in red. b, Left: stacked bar plots showing the minimum lists of gut bacteria and probiotic species needed to biotransform the vast majority of ‘shared’ phytonutrients, along with the cumulative number of biotransformed phytonutrients due to the inclusion of each species. Right: phytonutrient accumulation curve with combined biotransformation potential covering 43 ‘gut-restricted’ phytonutrients. c, Heat map showing the metabolism status of phytonutrient substrates across different conditions. Parentheses indicate the addition of reaction co-factors. The colour key depicts differences in the levels of phytonutrients between conditions with and without bacteria. The asterisks indicate both significant difference and bacterial metabolism (defined as the difference between 0 h and average levels at 6 and 24 h) >25%. Significant difference was determined by one-tailed t-test using pooled data at 6 and 24 h (FDR < 0.05). d, Chemical reactions involving the targeted phytonutrients. e, Biotransformation kinetics of the phytonutrient substrates butein and isoliquiritigenin and their products (for butein, butin and butin-FcrP; for isoliquiritigenin, liquiritigenin and davidigenin), as catalysed by E. ramulus. Two initial substrate concentrations (20 µM or 200 µM) were tested in the presence of E. ramulus, heat-killed (HK) E. ramulus, E. coli K-12, or bacteria-free medium control. Data shown are mean ± s.d of 3 independent replicates.
Given the increasing interest in the health benefits of fermented foods and postbiotics22, we attempted to select a minimal number of gut bacterial species with maximal potential to enzymatically modify those 186 phytonutrients. Looking first at the phytonutrients modified by both gut bacteria and probiotic species, a selection of 8 gut bacterial species or 4 probiotic strains sufficed to process most (>95%) phytonutrients (Fig. 2b, left). For the 70 phytonutrients enzymatically altered only by gut bacteria, we identified a selection of 11 bacteria (that is, minimum species) that could potentially modify 43 phytonutrients (the remainder were either modified by unannotated species or did not pass the filtration criteria) (Fig. 2b, right). We further compared phytonutrients potentially metabolized by these 11 species based on HUMAnN3 and RefSeq genomes. Over 90% overlapped, supporting the robustness of our annotation procedure (Supplementary Table 2). To complement our analysis of secondary-metabolism-related phytonutrients, we also grouped phytonutrients into non-primary-metabolism-related phytonutrients, defined as those that are not part of KEGG primary-metabolism-related pathways. A parallel analysis of non-primary-metabolism-related phytonutrients revealed substantial overlap (8 of the 11 minimum species), reinforcing their high biotransformation potential (Extended Data Fig. 1d).
Further comparison of the functional groups highlighted structural features enriched in the shared versus gut-restricted phytonutrients (Fig. 2a and Supplementary Table 5). Phytonutrients containing carboxylic acid and carbonyl groups, such as trans-cinnamic acid and naringenin, are particularly enriched in this shared chemical space (Fisher exact test, false discovery rate (FDR) < 0.05). In contrast, gut-restricted phytonutrients, including tricetin (metabolized by Bifidobacterium animalis, Streptococcus salivarius and so on) and fustin (metabolized by Firmicutes species), are enriched in those containing benzene rings and bicyclic groups.
We then investigated the capability of gut species to metabolize phytonutrients in vitro, testing representative strains from six gut species (Supplementary Table 6) with high biotransformation potential of secondary-metabolism-related phytonutrients compared with probiotics (Fig. 2b). We incubated these species with 36 phytonutrients (secondary-metabolism-related: 30) that they were predicted to metabolize, covering all enzyme classes mainly associated with the microbiome (EC1–EC5). E. ramulus, O. splanchnicus and B. uniformis incubation decreased phytonutrient levels, indicating potential biotransformation by the bacteria (Fig. 2c, one-tailed t-test FDR < 0.05; Supplementary Table 7 and Data 1 and 2). Notably, E. ramulus showed the strongest biotransformation activity, significantly metabolizing 11 of the 12 substrates. The marked predominance of flavonoids expands the known flavonoid-metabolizing repertoire of E. ramulus23 and suggests that its biotransformation activity may extend to other structurally related phytonutrients.
A more detailed investigation of two flavonoids, butein and isoliquiritigenin, previously reported to undergo spontaneous cyclization23, revealed a putative enzyme, EC 5.5.1.6, encoded in the E. ramulus genome that could catalyse their biotransformation (Fig. 2d). This inference is evidenced by accelerated biotransformation in a dose-dependent manner when live E. ramulus were present, compared with non-enzymatic conditions (Fig. 2e and Extended Data Fig. 2). Consistent with previous research23, the downstream products of these reactions were detected only in the presence of live E. ramulus. Using butin as substrate determined that the enzyme reaction directionality favoured chalcone-to-flavanone cyclization (Extended Data Fig. 2b). Overall, our analysis revealed shared capacity between gut bacteria and probiotics and highlighted specific gut bacteria with unique biotransformation potential.
Phytonutrient biotransformation shows inter-individual variability and geographical specificity
We next focused on whole microbiomes and their interactions with the phytonutrient space. On average, 70% of enzymes annotated in an individual microbiome and 90% of those in secondary-metabolism-related pathways were associated with phytonutrient biotransformation (Fig. 3a,b).
a, Left: ratio of phytonutrient-associated enzymes to the total number of microbial enzymes detected in the 3,068 metagenomes. Right: two-sided partial Spearman’s correlation (adjusted by sequencing depth) between species’ Shannon diversity and the enzyme ratio (left). The aqua points represent the enzyme ratio in each sample and the grey plot shows their distribution. b, As in a, but with only secondary-metabolism-related phytonutrients. c, Left: distributions of the number of phytonutrients associated with gut microbial enzymes from the 3,068 metagenomic samples; mean ± s.d. Right: upSet plot showing the total number of phytonutrients that can potentially be biotransformed by the gut microbiota of each continent (based on enzyme presence/absence and 1% prevalence cut-off per region) and their intersections. d, PCoA of Bray–Curtis dissimilarity between phytonutrient-associated gut bacterial enzymatic profiles. n = 1,936, Oceania was omitted because body mass index was unavailable. Arrows, phytonutrient classes significantly associated with enzymatic profile variation. e, Procrustes analysis between phytonutrient intake variation (Jaccard’s distance) and associated gut bacterial enzymatic variation (robust Aitchison’s distance) using 32 participants from ref. 7. One-tailed Procrustean randomization test was used with 999 Monte Carlo permutations. f,g, Microbial source-tracking-based similarity analysis of dietary records (f) and phytonutrient-associated enzymatic profiles (g), comparing American-born participants (n = 15) against Thai participants living in Thailand (Thailand, n = 15), who recently moved to the USA (NA, n = 11) or who have lived in the USA for >20 years (LTR, n = 14)12. For d, f and g, statistical significance was determined using PERMANOVA or two-sided Wilcoxon rank-sum test (**P < 0.01, ****P < 0.0001). Boxplots show the median (centre lines), first and third quartiles (box limits) and 1.5× interquartile range bounds (whiskers). OC, Oceania; AF, Africa; AM, America; EU, Europe; AS, Asia; LMWMs, low-molecular-weight molecules.
Microbiome alpha diversity was negatively correlated with the ratio of enzymes that were phytonutrient associated, suggesting that phytonutrient biotransformation is a common property of gut microbiota (Fig. 3a and Extended Data Fig. 3a). However, this correlation was positive for enzymes involved in secondary metabolism, suggesting that gut microbes have varying abilities to biotransform secondary-metabolism-related phytonutrients (Fig. 3b and Extended Data Fig. 3b).
We observed considerable inter-individual variability in phytonutrient biotransformation: the number of phytonutrients biotransformed ranged from 264 to 620 (IQR 32–52) among five geographical regions (Fig. 3c). Phytonutrient-associated microbiome uniqueness, adopted from a method indicating distinctness24, was significantly higher among samples from different individuals than those from the same individual (Wilcoxon rank-sum test, P < 0.05; Extended Data Fig. 3c).
Interestingly, when we aggregated individuals by continent, their total enzymatic machinery associated with the phytonutrient and food biotransformation space was highly similar across regions (Fig. 3c and Extended Data Fig. 3d). In total, 630 phytonutrients were commonly associated with gut microbial enzymes from all five regions, and only a few (for example, 10 for Asia, 2 for Oceania) showed high geographical specificity (Fig. 3c). For example, the flavonoid taxifolin—found in foods such as lychees (common in southern China) and Cudrania tricuspidata (a traditional medicine in East Asia)—and the associated enzyme EC 1.1.1.219 were annotated uniquely in individuals from Asia. The profiles of phytonutrient-associated enzymes (using presence/absence patterns or relative abundances) differed significantly (PERMANOVA, P < 0.05) across continents regardless of adjustment by age, sex and body mass index (BMI) (Fig. 3d and Extended Data Figs. 3e,f), reflecting considerable inter-region dietary variation. Specifically, Africa and Asia differed significantly from the other regions (Wilcoxon rank-sum test, P < 0.05). The enzymatic variation was significantly associated with specific chemical classes, including flavonoids and fatty-acid-related compounds (envfit, adjusted P < 0.05).
Distance-based redundancy analysis also showed associations of age, sex and BMI with phytonutrient-associated enzymatic profiles (Extended Data Fig. 3g). The phytonutrient-associated microbiome uniqueness and the ratio of phytonutrient-associated enzymes showed positive and negative correlations with age, respectively (Extended Data Fig. 3h,i), consistent with the increased compositional uniqueness during ageing24. Furthermore, 511 enzymes were significantly correlated with BMI (Spearman’s correlation, FDR < 0.05), particularly lyases, oxidoreductases and ligases (among classes with >10 enzymes) (Supplementary Table 8). We also observed a loss of phytonutrient-associated enzymes as BMI increased (partial Spearman’s correlation, R = −0.14, P < 0.05; Extended Data Fig. 3j).
The geographic differences in the beta-diversity comparisons (Fig. 3d) could have different causes, including genetic and dietary factors. To explore how diet shapes these patterns, we used existing data from a study with consecutive dietary information7 and paired shotgun metagenomic data. Procrustes analysis between the edible plant dietary records and the abundances of enzymes associated with the 1,665 phytonutrients of those edible plants showed significant agreement (Monte Carlo permutation test, P = 0.001; Fig. 3e), suggesting that edible plant intake is associated with the microbiome’s phytonutrient biotransformation potential. We then performed a similarity analysis, using a source-tracking algorithm25, of an existing dataset from US-born individuals living in the USA along with Thai individuals who lived in Thailand and had recently moved to the USA or had lived in the USA for ≥20 years12. Despite relatively modest adaptation in dietary habits, the edible-plant-associated enzyme composition of Thai participants became significantly more similar to that of US-born participants over time (Wilcoxon rank-sum test, P < 0.05; Fig. 3f,g and Extended Data Fig. 3k). Thus, even though certain core enzymes of phytonutrient metabolism are geographically conserved, both inter-individual and regional variability in phytonutrient biotransformation by gut bacterial enzymes exist, associated partly with dietary habits.
Potential to metabolize healthy foods is altered in disease
Because health status is a critical determinant of gut microbiome balance26, we hypothesized that healthy and diseased individuals may have different capacities to biotransform the phytonutrient space of edible plants, including those with demonstrated health benefits. To test this, we analysed public gut metagenomics datasets from case–control cohorts for inflammatory bowel disease (IBD), colorectal cancer (CRC) and non-alcoholic fatty liver disease (NAFLD). We linked gut bacterial enzymes to their respective beneficial phytonutrient spaces, identifying 608, 1,038 and 517 phytonutrient-associated enzymes for IBD, CRC and NAFLD, respectively.
Among these enzymes, 59.7% (n = 363), 49.9% (n = 518) and 22.2% (n = 115) showed significant differential abundances in IBD, CRC and NAFLD, respectively, compared with the corresponding controls (metagenomeSeq zero-inflated Gaussian model, FDR < 0.05, Supplementary Table 9). Then, we looked further into the altered biotransformation potential of gut microbiota for each beneficial food. On average, 63.0%, 33.6% and 25.3% of phytonutrient-associated ECs had significantly altered abundance in IBD, CRC and NAFLD, respectively (Extended Data Fig. 4a). Interestingly, great variation was observed among the three diseases and among different beneficial foods for the same disease, with some foods having over 50% significantly altered enzymes.
We then sought to pinpoint a combination of key phytonutrient-associated enzymes (species-stratified) that could be used to discriminate between healthy and diseased individuals using machine learning (ML). Our random-forest models included 18 phytonutrient-associated enzymes in IBD (area under the receiver operating characteristic curve (auROC) = 0.892), 28 for CRC (auROC = 0.763) and 22 for NAFLD (auROC = 0.95) (Extended Data Fig. 5a–c). We validated the models in external cohorts, achieving high accuracy across all three diseases (auROC = 0.72–0.79; Fig. 4a–c). In contrast, when we used the species-stratified output for non-dietary ECs (that is, not associated with phytonutrients), the model validation auROCs decreased (IBD, 0.779; CRC, 0.698; NAFLD, 0.584; Extended Data Fig. 4b).
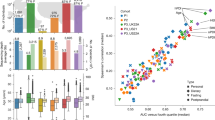
ROC curves and confusion matrices of machine learning models discriminating between health and disease states using the stratified abundances of enzymes associated with phytonutrients present in disease-associated edible plants (healthy foods). a–c, Diseases analysed were inflammatory bowel disease (IBD; a), colorectal cancer (CRC; b) and non-alcoholic fatty liver disease (NAFLD; c). Examples of SHAP dependence plots illustrate the effect of feature abundance on model prediction, with dark colours indicating disease cases and light colours representing controls. x axis, feature abundance; y axis, SHAP value for each feature indicating its contribution to control or disease prediction. Vertical dashed lines indicate optimal feature thresholds. Inset scatterplots (plotted with ranks) depict two-sided Spearman’s correlation between phytonutrient-associated enzyme abundances and metabolites involved in corresponding reactions, utilizing paired data from metagenomics and metabolomics analyses. The blue line represents the fitted linear regression (estimated mean response), and the grey area denotes 95% confidence interval.
To extract biological insights from the ML models, we used the SHapley Additive exPlanations (SHAP) method27 to compute the importance and prediction direction for each feature. In the IBD model, for example, we identified EC 4.1.99.1 of Alistipes finegoldii, which catalyses the conversion of tryptophan to indole and was predictive of healthy status (more abundant in controls) (Fig. 4a and Extended Data Fig. 5a). Tryptophan, abundant in edible plants such as garlic, reportedly benefit IBD patients only after being biotransformed by gut microbiota28. Notably, in the paired metabolomics–metagenomics datasets from the IBD cohort used for ML, we observed a significant negative correlation between the abundances of EC 4.1.99.1 and tryptophan (P < 0.05, r = −0.23, Spearman’s correlation; Fig. 4a), further supporting our hypothesis that the gut microbiome plays a crucial role in modifying phytonutrients to enable their beneficial properties for the host. Similarly, we identified associations of the levels of EC 1.4.3.5 from Porphyromonas asaccharolytica and tyrosine29 in CRC (Fig. 4b and Extended Data Fig. 5b), and the levels of EC 1.2.99.7 from Blautia wexlerae and trans-cinnamic acid30 in NAFLD (Fig. 4c and Extended Data Fig. 5c). Thus, the altered capacity of individuals with these conditions to biotransform healthy diets may compromise their diets’ beneficial effects, possibly in a food- and disease-dependent manner.
Effect of anti-inflammatory diet is associated with specific bacterial enzymes
To test in vivo whether gut microbial enzymes involved in phytonutrient biotransformation are essential for triggering the protective role of healthy diets, we investigated the effects of an anti-inflammatory food, strawberry31, in a mouse model of colitis under both specific-pathogen-free (SPF) and germ-free (GF) conditions (Fig. 5a). Indeed, 17-day supplementation of a normal diet with strawberries improved the overall health of dextran sodium sulfate (DSS) SPF mice, with significantly decreased body weight loss (day 14, two-tailed t-test, P = 0.019) and disease activity index (DAI) score (linear mixed model, P = 0.0042), along with attenuation of histological damage (two-tailed t-test, P = 0.037) (Fig. 5b,c and Extended Data Fig. 6a,b).
a, Experimental design for investigating the effects of strawberry supplementation on colitis in mice under SPF (n = 8) or GF (n = 6 or 7) conditions. Created with BioRender.com. b, Changes in body weight and DAI scores in SPF mice throughout the entire study. # indicates significant differences between ‘Vehicle’ and ‘DSS+vehicle’ groups (#P ≤ 0.05; ##P ≤ 0.01; ###P ≤ 0.001; ####P ≤ 0.0001); * indicates significant differences between ‘DSS+vehicle’ and ‘DSS+strawberry’ groups (*P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001, ****P ≤ 0.0001). c, Histological scores and representative images of colon tissue stained with H&E. Scale bars, 500 μm (top), 50 μm (bottom). Different-coloured arrows indicate signs of necrosis of the mucous layer (black), ulcerative lesion (dark green), neutrophil (light yellow), lymphocyte (blue), macrophage (red), fibrous tissue hyperplasia (orange) and submucosal oedema (light green). d,e, As in b,c, but in GF mice. In b–e, data are presented as mean ± s.e. and statistical differences were examined using two-sided unpaired t-tests. f,g, PCoA of robust Aitchison distance among strawberry-associated microbial enzymes between DSS+vehicle and DSS+strawberry groups (f), and between day 6 and day 18 of DSS+strawberry group (g) for DNA abundance and RNA expression levels. Significant differences were determined using PERMANOVA. h, Heat maps showing enzymes correlated with DAI scores and histological scores, and their median differences between the ‘DSS+strawberry’ and ‘DSS+vehicle’ groups, at DNA and RNA levels. *P < 0.05, two-sided Spearman’s correlations or Wilcoxon rank-sum test. Scatterplots show correlations, for example, of the strawberry-associated enzymes and grey area denotes 95% confidence interval.
Next, to assess whether this protective role depends on the gut microbiome, we studied the effects of a strawberry-supplemented diet in GF mice. Although the diet did not affect body weight, it delayed the occurrence of rectal bleeding and slightly improved stool consistency (Extended Data Fig. 6c), reflected in a reduced DAI score (linear mixed model, P = 1.5 × 10−4) (Fig. 5d). Interestingly, detailed histopathological analysis nonetheless found no differences between DSS GF mice on normal and strawberry-supplemented diets (Fig. 5e and Extended Data Fig. 6d), implying that strawberry supplementation only partially ameliorated pathological damage. Overall, our findings suggest that the protective role of strawberries in colitis progression is partly mediated by the gut microbiota.
To investigate the complex interactions between anti-inflammatory diets and gut microbiome, we performed stool shotgun metagenomics and metatranscriptomics analysis after 6 days of strawberry supplementation (but before DSS administration) and on the last day of DSS treatment in the different SPF mouse groups. Robust Aitchison distance analysis at day 6 revealed that strawberry supplementation changed the strawberry-associated enzymes profile between the DSS+strawberry and DSS+vehicle groups, at both DNA (PERMANOVA, P = 0.001) and RNA (PERMANOVA, P = 0.021) levels (Fig. 5f). Beta-diversity analysis comparing day 6 with day 18 in the DSS+strawberry group suggested that colitis significantly alters the profiles of strawberry-associated microbial enzymes (Fig. 5g, PERMANOVA; DNA level, P = 0.002, RNA level, P = 0.002) and strawberry metabolism at the phytonutrient level.
We then investigated specific gut microbiome-associated enzymatic activities potentially involved in alleviating colitis in the SPF group: 27 and 44 strawberry-associated microbial enzymes identified at the DNA and RNA levels, respectively, showed significantly higher median abundance in the DSS+strawberry group and also correlated with at least one of the two indicator scores for colitis (DAI and histological scores) (Fig. 5h, Spearman’s correlation, P < 0.05). At the DNA level, for instance, the enzyme 4.1.1.11 was negatively correlated with both scores (Fig. 5h). EC 4.1.1.11 is involved in the biotransformation of aspartate to beta-alanine, a metabolite reported to decrease inflammatory responses32, and the conversion of tryptophan to tryptamine, which can reduce colitis severity in mice33. At the RNA level, EC 6.3.1.1—involved in converting aspartate to asparagine, which helps maintain intestinal health34—was negatively correlated with histological score (Fig. 5h). These colitis-associated enzymes included 4.1.99.2 and 6.4.1.3 that discriminated between health and disease in the IBD ML model (Extended Data Fig. 5). Overall, our findings suggest that the full spectrum of anti-inflammatory benefits from strawberries depends on the abundance and expression of specific gut microbial enzymes.
Discussion
Previous studies have investigated how microbial metabolism of phytochemicals shapes gut microbiome composition35 or how particular transformations play beneficial or pathological roles in host health36,37. Extending these insights, we integrated multiple bioinformatics databases to systematically analyse the extensive interactions between plant diet and gut bacteria, focusing on microbial enzymes and main taxonomic drivers, revealing mechanistic links between human nutrition at the small-molecule level and microbiome. Integrating >3,000 global microbiomes and >1,300 phytonutrients, we made several observations: (1) 775 phytonutrients, including several with known bioactivities, are predicted to be metabolized by gut microbes, based on genomic content; (2) 67% of gut microbial enzymes annotated are potentially associated with phytonutrient biotransformation; and (3) phytonutrient biotransformation by gut bacteria is widespread across and varies among phyla. We also gained information about how these biotransformations vary by individual, health status and geographical region, and shape the health benefits of particular foods.
Our study could inform the development of next-generation probiotics. Probiotics have demonstrated efficacy in various disease contexts38,39,40, but expanding beyond Lactobacilli and Bifidobacteria is crucial to their broader application to human health41. Comparing the phytonutrient biotransformation of gut bacteria and 59 over-the-counter probiotics revealed a large dietary chemical space modifiable only by gut bacteria. Thus, identifying and isolating gut bacteria with specific beneficial properties based on their postbiotic capacity may provide more effective probiotic therapies well adapted to the human gastrointestinal environment for disease prevention and treatment42.
Our findings also suggest strategies for developing functional foods. The potential of microbial foods43, including fermented foods, to modulate human health through microbial metabolites is currently drawing interest22. However, research has been limited to analyses of the fermented food metabolome landscapes of just a few foods, including fermented dairy44,45 and kimchi46. Here we demonstrated the possibility to obtain small assemblies of gut bacterial species with maximal ability to biotransform target sets of phytonutrients: for example, a set of 11 gut bacterial species that can largely drive the biotransformation of 43 secondary-metabolism-related metabolites from common edible plants (which known probiotics lack the genetic potential to perform). This suggests a new direction for developing functional foods through fermentation with selected bacterial species, identified via mechanistic links between phytonutrients and gut bacteria.
Food choices can alter gut microbiome composition to improve host homeostasis7,19; conversely, the microbiome may influence diet’s impact on host homeostasis. We show here that phytonutrients from edible plants with known health benefits are biotransformed differently by gut bacterial enzymes in healthy and diseased individuals, highlighting the need for further studies in larger and more diverse cohorts to support or extend these observations. Furthermore, we observed that the full-spectrum effect of an anti-inflammatory plant food (strawberry) occurred only in mice with healthy microbiota, and identified key gut bacterial enzymes potentially biotransforming specific health-promoting phytonutrients. Our results, underscore the importance of revisiting the concept of a ‘healthy diet’, as the diet’s effectiveness may be significantly boosted by the presence of a healthy microbiome. Consuming edible plants with demonstrated health benefits may be insufficient for someone with an imbalanced microbiome capacity to biotransform health-associated phytonutrients. Thus, personalized nutrition may require a combination of specific foods and beneficial microbes, or ex vivo fermentation of food, to achieve the full nutritional potential of a plant diet. Consuming foods fermented by gut bacteria after rigorous safety testing could be extremely valuable for populations whose intestines may be unable to perform the modifications necessary to harness foods’ full nutritional value, including aged individuals and those with reduced microbiome diversity47,48. Interestingly, our analysis also showed a negative correlation between participant age and the ratio of phytonutrient-associated enzymes.
This study has some limitations. First, as a proof-of-concept study, it focused on single enzymatic reactions instead of chains of reactions that could allow us to characterize the metabolic fate of phytonutrients, leaving it unclear whether the metabolic products are absorbed by humans or further metabolized by other gut microbes. Second, due to the vast phytonutrient biotransformation reactions, we experimentally validated only a few, calling for future high-throughput assays. Finally, our enzyme-annotation-based framework is limited to known gene functions, neglecting unknown biotransformations. Better understanding of such functional ‘dark matter’ via additional computational and experimental approaches35,49,50 will facilitate the design of microbiome-based personalized nutrition at the microscale level.
Our large-scale systematic mapping of dietary phytonutrients and gut microbiota shows that microbial enzymes may biotransform hundreds of phytonutrients. Notably, this metabolic potential varies between healthy and diseased states, underscoring the central role of gut microbiota in mediating health effects of diet. This should open avenues for optimizing the nutritional value of plant-based diets through targeted microbial engineering and inform the development of next-generation probiotics, functional foods and personalized nutrition.
Methods
Global cohort collection
In this study, we collected publicly available Illumina-sequenced stool metagenomic data of non-diseased participants from 40 published studies, including microbiomes collected from five continents: Europe, Asia, Africa, Oceania and the Americas (here referred to as America) (Supplementary Table 1). Initially, age, sex and BMI data were obtained from the curatedMetagenomicData51 package in R, with further manual curation. If a cohort lacked recorded information, we supplemented it by extracting relevant metadata from corresponding publications.
Cohort collection for metagenomic sequencing data with paired dietary intake
We collected publicly available shotgun metagenomic sequencing data with paired dietary intake information from 2 microbiome projects. The studies included: (1) a cohort of 32 (2 individuals were excluded due to consuming only nutritional meal replacement beverage) healthy individuals with detailed dietary information and paired shotgun metagenomic data recorded for 17 consecutive days7 and (2) a cohort of healthy individuals consisting of US-born individuals living in the United States (N = 15), Thai individuals living in Thailand (N = 15), Thai individuals who had recently moved to the United States (N = 11) and Thai individuals who had lived in the United States for at least 20 years (N = 14)12.
Diseased cohort collection
We collected publicly available shotgun metagenomic sequencing data from three different diseases: inflammatory bowel disease (IBD), colorectal cancer (CRC) and non-alcoholic fatty liver disease (NAFLD). In each of these conditions, the gut microbiome has been extensively studied, with previous research demonstrating its key role in disease diagnosis and prevention11,52,53,54,55. The studies related to IBD included (1) an IBD cohort of IBD (N = 185) and non-IBD (N = 74) samples56 and (2) a cohort containing IBD (N = 728) and non-IBD (N = 271) samples57 for which paired metagenomics–metabolomics data were available.
The studies related to NAFLD included (1) an NAFLD cohort containing participants with NAFLD (N = 100)11, (2) a cohort containing NAFLD (N = 19) and non-NAFLD (N = 10) participants [BioProject ID: PRJNA732131] and (3) a non-NAFLD cohort (N = 207) containing paired metagenomics–metabolomics data58.
The studies related to CRC included (1) a cohort containing CRC (N = 46) and non-CRC (N = 61) participants59, (2) a cohort containing CRC (N = 27) and non-CRC (N = 28) participants60, (3) a cohort containing CRC (N = 52) and non-CRC (N = 52) participants61, (4) a cohort consisting of CRC (N = 61) and non-CRC (N = 53) participants62, (5) a cohort containing CRC (N = 60) and non-CRC (N = 57) participants63, (6) a cohort consisting of CRC (N = 74) and non-CRC (N = 53) individuals64, (7) a cohort containing CRC (N = 29) and non-CRC (N = 24) participants65, (8) a cohort containing CRC (N = 32) and non-CRC (N = 28) participants65, (9) a cohort containing CRC (N = 40) and non-CRC (N = 39) participants66 and (10) a CRC cohort (N = 76) for whom paired metagenomics–metabolomics data were available (in-house data).
Shotgun metagenomics data processing
Details on sample collection and sequencing of the original data can be found in the original studies. Quality control of the raw metagenomic reads was performed using the Sunbeam pipeline (v.2.1)67 as previously described68. Briefly, adapter sequences were removed and low-quality regions were trimmed. The filtered reads were then mapped to the human genome (GRCh38) with BWA, and mapped reads were removed as human DNA contamination. Taxonomic profiling of the high-quality reads was performed using MetaPhlAn369 with default settings, generating taxonomic relative abundances. Bacterial community profiles were then constructed at the species level for further analysis. Functional profiling was performed using the HUMAnN3 pipeline69. The quantified gene family abundances in units of reads per kilobase (RPKs) were then normalized to copies per million (CPM) units by the provided HUMAnN3 script, resulting in transcript-per-million-like (TPM) normalization. Normalized gene families were then regrouped to ECs for further analyses.
Linking phytonutrients with enzymes via bioreaction databases
To build the link between the 7,825 phytonutrients (that is, natural compounds present in plant-based foods) sourced in NutriChem 2.0 and the enzymes, we collected enzymatic reactions from 10 biochemical reaction databases: the Kyoto Encyclopedia of Genes and Genomes (KEGG)70, PubChem71, IntEnz72, BKMS-react73 (containing biochemical reactions collected from BRENDA, KEGG, MetaCyc and SABIO-RK), HMDB (4.0)74, EAWAG75, EzCatDB76, M-CSA77, SFLD78 and Transformer79. Data sources were accessed through either API-based query (KEGG), direct downloads of XML (HMDB), TXT (EzCatDB) or CSV files (PubChem ‘Biochemical Reactions’, M-CSA), upon query (BKMS-react) or web scraping (EAWAG, SFLD, Transformer) (for example, using the R packages KEGGREST (http://bioconductor.org/packages/KEGGREST/), XML (https://cran.r-project.org/web/packages/XML/) and rvest (https://rvest.tidyverse.org/)). To map the phytonutrients from NutriChem 2.0 to biochemical reactions, we used CID (PubChem Compound Identifier; PubChem, HMDB, EzCatDB, EAWAG, Transformer) or CHEBI (Chemical Entities of Biological Interest; IntEnz, SFLD) or both identifiers (KEGG, M-CSA) or InChIs (IUPAC International Chemical Identifiers; BKMS-react). The InChIs were obtained from SMILES using the cheminformatics toolkit RDKit (https://www.rdkit.org).
Fine curation of phytonutrient–enzyme links targeting natural products
For the 1,658 phytonutrients from edible plants linked with enzymes, we conducted additional filtering using a variety of natural product databases. First, we extracted the compound CID identifiers recorded in NutriChem 2.0 and used the PubChem71 database under content ‘Taxonomy’ as the source to examine whether they originated from the Viridiplantae domain in the natural product database LOTUS80, NPASS81 and KNApSAcK82. For the remaining compounds, we utilized their InChIs and SMILES to determine whether they were listed in the SuperNatural 3.0 database83. If there was a hit in the database that originated from the Viridiplantae domain based on NCBI taxonomy, we determined that the compound is a natural product. Finally, the remaining unmapped compounds underwent further manual curation of the compound–plant associations achieved by reviewing corresponding literature entries recorded in NutriChem 2.0. Through this process, we curated the final list of 1,388 phytonutrients associated with 4,678 enzymes with a unique EC identifier.
Acquisition of the microbial enzymes and human enzymes list
The microbial EC and affiliated species were obtained directly from HUMAnN3 stratified output. The EC information regarding human enzymes was retrieved from the KEGG database, specifically through their API: https://rest.kegg.jp/link/ec/hsa.
Metagenome-assembled genomes (MAG) construction and annotation
MAG construction was mainly based on the metaWRAP pipeline84. Quality control of 200 randomly selected (from the 3,068 cohort) samples’ raw paired-end reads was performed using the metaWRAP read_qc module with default parameters. Quality-controlled reads were then assembled individually with ‘–metaspades’ in metaWRAP (194 samples successfully assembled). Assembled contigs were further binned with MetaBAT2, MaxBin2 and CONCOCT using ‘–metabat2 –maxbin2 –concoct’ parameters. The resulting bin sets were refined using the ‘bin_refinement’ module in metaWRAP with parameters ‘-c 50 -x 10’. Dereplication was performed at 99% average nucleotide identity (ANI) using dRep85 with parameters ‘-comp 50 -con 5 -sa 0.99 -nc 0.3’ to cluster bins into strain-level non-redundant MAGs. Then MAGs were annotated using GTDB-Tk86. Gene prediction was performed with Prokka87, and functional annotation was conducted using eggNOG-mapper88, generating EC profiles.
Acquisition of healthy food list and anti-inflammatory compounds
The healthy food list was retrieved from a previous study19. For 23 edible plants designated as healthy foods, we retrieved phytonutrient contents from NutriChem 2.0. However, compounds linked with apples and bananas did not yield any matches with microbiome-associated enzymes, hence downstream analysis was performed for the phytonutrients derived from the remaining 21 healthy foods. To retrieve the list of phytonutrients with anti-inflammatory activity from those foods, we utilized the database InflamNat89, keeping only phytonutrients showing inhibition of nitric oxide (NO, an inflammatory factor) production (IC50 < 50 μM, as mentioned in the database). From this, we identified a list of 17 phytonutrients with anti-inflammatory activity: phloretin, quercetin, fisetin, daidzein, apigenin, genistein, luteolin, isorhamnetin, kaempferol, caffeic acid, protocatechuic acid, ellagic acid, curcumin, biochanin A, liquiritigenin, alpha-linolenic acid and isoliquiritigenin (Supplementary Table 3).
Probiotic collection and genome annotation
Probiotic strains were collected from five sources: three databases: Probio90 (https://bidd.group/probio/download.htm), Integrated Probiotic DataBase91 and AEProbio (https://usprobioticguide.com/)92; and two marketing sources: Chr. Hansen (https://www.chr-hansen.com/en/human-health-and-probiotics/our-probiotic-strains) and Optibac92 Probiotics (https://www.optibacprobiotics.com/professionals/probiotics-database). Fifty-nine genomes of these probiotic strains from the Actinobacteria, Firmicutes and Proteobacteria were successfully acquired from NCBI RefSeq by using ncbi-genome-download. Genomes were annotated using Prokka87 with default parameters and the ‘tab-separated file containing all annotated features (*.tsv)’ was used to extract the EC information.
Acquisition of the secondary-metabolism-related metabolites list and primary-metabolism-related metabolites
Metabolites were termed as secondary-metabolism-related if they belonged to KEGG PATHWAY classes ‘Metabolism of terpenoids and polyketides’ and ‘Biosynthesis of other secondary metabolites’ in the KEGG PATHWAY database. Metabolites were classified as primary-metabolism-related if they were from any of the following KEGG PATHWAY classes ‘Carbohydrate metabolism’, ‘Energy metabolism’, ‘Lipid metabolism’, ‘Nucleotide metabolism’,’Amino acid metabolism’, ‘Metabolism of other amino acids’, ‘Glycan biosynthesis and metabolism’, ‘Metabolism of co-factors and vitamins’. The metabolites and enzymes for each pathway were extracted using the ‘keggLink’ function in the R package KEGGREST.
Functional group analysis
For each phytonutrient, the functional group was identified by a module from rdkit.Chem.Fragments (https://www.rdkit.org). The enrichment analysis was conducted using Fisher’s exact test, with the false discovery rate adjusted using the Benjamini–Hochberg procedure.
Minimum species analysis
To identify the minimal set of bacterial species that encompass the potential for metabolizing phytonutrients related to secondary metabolites, we first linked enzymes from probiotics and gut microbes to these metabolites on the basis of the fine-curated phytonutrient–enzyme links above. To the gut microbial links only, we applied a three-step filtering process: we filtered out non-bacterial species, applied a 5% prevalence filter to retain commonly found species, and used a 10% contribution filter to keep species whose enzymes considerably contribute to the total enzyme abundance. This filtering ensures that we focus on the most relevant and prevalent bacteria. Next, we used a greedy algorithm implemented in the RcppGreedySetCover R package (https://github.com/matthiaskaeding/RcppGreedySetCover) to solve the set cover problem. This algorithm identified the smallest subset of species that collectively possess all the necessary enzymatic activities. This approach efficiently narrows down the species to those most critical for the metabolic coverage of the target secondary metabolites-related phytonutrients. We applied the same method to identify the minimum subset of gut microbial species to metabolize the non-primary-metabolism-related phytonutrients.
Comparison between genomic-based vs HUMAnN3-based results
-
(1)
HUMAnN3 vs MAGs: The EC profile produced from the MAGs (generated from 200 samples) was linked with phytonutrients using the phytonutrient–enzyme links from the previous step. The resulting MAGs-linked phytonutrients were then compared with the phytonutrients linked to the same set of samples by HUMAnN3.
-
(2)
HUMAnN3 vs pangenome database: We extracted UniRef90 identifiers from each ffn.gz file in the ChocoPhlAn3 databases and annotated them with ECs using the mapping file ‘map_level4ec_uniref90.txt.gz’ provided by HUMAnN3 (Uniref90 201901). Then for each species, a ratio was calculated as the number of species-specific ECs annotated by HUMAnN3 divided by the total number of ChocoPhlAn3-annotated ECs.
-
(3)
HUMAnN3 vs RefSeq microbial genomes: A total of 45,862 genomes of the 11 species were downloaded from NCBI RefSeq using NCBI Datasets command-line tools. The downloaded .fna files were annotated using Prokka to obtain the enzyme profile. The profile was then mapped to phytonutrients using our previous established phytonutrient–enzyme links. The total number of linked phytonutrients was then compared with that of the same 11 gut bacteria species based on HUMAnN3 species-stratified profiles from the 3,068 samples.
Species and strain selection for in vitro phytonutrient biotransformation assay
From the list of minimum species identified (for gut-restricted phytonutrients that are secondary-metabolism-related, see above), we looked into their biotransformed phytonutrients and rationally selected substrates for testing on the basis of commercial availability and technical feasibility. We preferred substrates involved in secondary-metabolism-related pathways, aiming to cover all enzyme classes mainly associated with microbiome (EC1–EC5). Then we selected their commercially available strains that encode predicted enzymes, based on the ‘map_level4ec_uniref90.txt.gz’ database from HUMAnN3 and the pangenome information for each species from PanPhlAn69. For Clostridium (Enterocloster) citroniae and Escherichia coli, we obtained the genome sequences of C. citroniae DSM 19261 strain from the DSMZ BacDive database, and E. coli K-12 strain (GCF_000005845.2) from the NCBI RefSeq database. We then used BLAST to verify the presence of specific enzymes of interest in the genomes. Overall, six gut strains including Bacteroides uniformis DSM 6597, C. citroniae DSM 19261, E. coli K-12, Eubacterium ramulus DSM 15684, Lactococcus lactis DSM 20481 and Odoribacter splanchnicus DSM 20712 were tested against the selected phytonutrients.
In vitro phytonutrient biotransformation assay
Anaerobic cultivation
Anaerobic cultivation was performed using an anaerobic chamber (Coy Laboratory Products) with 12% CO2, 2% H2 and 76% N2. Gut bacterial strains, including B. uniformis DSM 6597, E. ramulus DSM 15684, L. lactis DSM 20481, O. splanchnicus DSM 20712 and C. (Enterocloster) citroniae DSM 19261, were purchased from The Leibniz Institute DSMZ (Germany) in freeze-dried form, revived and validated by 16S ribosomal (r)RNA sequencing before further usage. E. coli K-12 (DSM 18039) was obtained from our in-house collection.
Phytonutrient metabolism assay
Frozen glycerol stocks (−70 °C) of bacterial strains were streaked onto brain–heart infusion (BHI; Becton Dickinson) agar plates supplemented with 10% horse blood and incubated anaerobically at 37 °C for 24–48 h until single colonies formed. For each strain, a pre-culture was prepared by inoculating single colonies into 4 ml of pre-reduced modified Gifu Anaerobic Broth medium (mGAM; HyServe) and incubating for 24 h (72 h for E. ramulus). The pre-culture for E. ramulus was further concentrated fourfold to enrich biomass. Heat-killed E. ramulus was prepared by incubating the pre-culture at 95 °C for 10 min. Biotransformation assays were initiated through the addition of 20 µl of pre-culture (or bacteria-free mGAM as control) into 176 µl of 5×-diluted mGAM and 4 µl of a phytonutrient substrate of interest in a 96-well plate (Nunc, 267544). For each phytonutrient–bacterium pair, two final concentrations of the phytonutrient (200 µM and 20 µM) were tested in two independent assay replicates. At 0, 6 and 24 h of incubation, 20 µl of the assay mixture were sampled and transferred to a V-bottom 96-well storage plate (Fisher Scientific, 10304513), snap-frozen and stored at −70 °C until further analysis. For the follow-up biotransformation kinetics assay with E. ramulus, samples were collected at 0, 15, 30, 45, 60, 120, 180, 360 and 540 min after incubation. The absence of enzyme 5.5.1.6 in E. coli K-12 was confirmed through BLAST analysis.
Sample preparation for mass spectrometry analysis
Liquid samples were processed through organic solvent extraction using the Agilent Bravo liquid-handling platform. In brief, 20 µl of sample were supplemented with 5 µl of internal standard mixture (containing caffeine-D9, diclofenac-D4, nafcillin-D5, oxfendazole-D3, phenylalanine-D5, tolfenamic acid-D4, tryptophan-D5 and warfarin-D5, each at 20 µM), extracted with 100 µl acetonitrile:methanol (1:1) and incubated at −20 °C for 1 h for protein precipitation. The extraction solutions were spun down at 4,347 × g at 4 °C for 15 min, and the supernatant was diluted 1:1 (v/v) with water for subsequent liquid chromatography–mass spectrometry (LC–MS) analysis.
Mass spectrometry instrument parameter
LC–MS analysis was performed on an Agilent 6550 iFunnel Q-TOF mass spectrometer coupled to an Agilent 1290 Infinity II UHPLC system. LC separation of most nutrients, except pipecolate, was performed on an InfinityLab Poroshell 120 HPH-C18 UHPLC column (2.1 × 100 mm, 1.9 µm) (695675-702) at 45 °C. The mobile phase consisted of solvent A (water with 0.1% formic acid) and solvent B (methanol with 0.1% formic acid). A linear gradient was applied, starting from 5% B to 95% B over 5.5 min at a flow rate at 0.6 ml min−1. For pipecolate separation, a Supel Carbon HPLC column (15 cm × 2.1 mm, 2.7 µm) (59987-U) was used at 60 °C. The gradient started with 5% B, held constant for the first minute, and then increased linearly to 100% B over the next 5 min. A dual AJS ESI source was applied with the following parameters: VCap at 3,500 V, nozzle voltage at 2,000 V, gas temperature at 225 °C, drying gas at 13 l min−1, nebulizer at 40 psi, sheath gas temperature at 275 °C, sheath gas flow at 12 l min−1. Data were acquired in positive-ion mode with a mass range of 100–1,700 m/z and acquisition rate of 1,000 ms per spectrum. Online mass calibration was performed with a reference solution containing purine ([M + H]+ = m/z 121.0509) and hexakis(1H,1H,3H-perfluoropropoxy)phosphazene ([M + H]+ = m/z 922.0098) via the secondary ESI source at a constant flow rate of 15 µl min−1. Agilent MassHunter Qualitative Analysis (v.10.0) was applied to examine the ion signals and retention times of nutrient substrates and their biotransformation products. Agilent MassHunter Quantitative Analysis (v.10.0) was applied to extract peak areas. RStudio (v.4.2.2) was applied for plotting and statistical analysis.
Microbiome diversity analysis
Alpha diversity (Shannon, Simpson and richness indices) and beta diversity (Bray–Curtis dissimilarity, Jaccard and robust Aitchison distances) were calculated using the vegan93 R package. Aitchison distance (beta diversity) was calculated using the robCompositions94 R package. Statistical comparisons of beta diversity were performed using the adonis function (999 permutations) of the vegan R package.
Microbiome uniqueness analysis
For the uniqueness analysis, we used the phytonutrient-associated enzymes profile. To calculate between-individual uniqueness, we randomly selected only one sample per individual. For within-individual uniqueness, we used multiple samples from the same individual. Bray–Curtis dissimilarity was calculated for the two parts separately. The minimum value for each sample (corresponding to the distance between each sample and its nearest neighbour) was then extracted as the uniqueness value as previously described24, as a measurement of distinctness of the gut microbiome.
Procrustes analysis and FEAST analysis
Procrustes analysis was performed using the procrustes function of the vegan93 R package. Monte Carlo P values for rotational agreement significance testing were determined from 999 permutations using the protest function of the vegan93 R package. Similarity analysis between US-born and Thai individuals was performed using fast expectation-maximization microbial source tracking (FEAST)25.
Acquisition of beneficial foods and phytonutrients for IBD, CRC and NALFD
We retrieved from NutriChem 2.0 all edible plants that, in previous studies, demonstrated beneficial impact on these diseases (potential ‘healthy foods’). On the basis of the 34, 83 and 29 healthy edible plants with benefits for IBD, CRC and NAFLD, we further retrieved from NutriChem 393, 711 and 444 phytonutrients, respectively. These phytonutrients were further linked to bacterial enzymes as above.
Random-forest classifiers
We built random-forest classifiers using the caret95 R package to discriminate diseased patients from healthy individuals on the basis of stratified enzyme abundances for enzymes that are involved in the biotransformation of phytonutrients present in foods that have been previously described to be beneficial for each disease. We randomly split the data into an 80% training set and a 20% test set. The training set was used to perform feature selection using the Boruta96 R package. Specifically, we conducted 100 independent Boruta runs, each yielding a ranked list of relevant features. From each run, we extracted the top 20 features and then formed the final feature set by taking the union of all selected features across iterations. Then a random-forest model was trained on the training set using this final feature set, and performance was evaluated on the test set. In addition, external validation was conducted using independent cohorts not involved in model training or feature selection. Model performance was assessed by computing receiver operating characteristic (ROC) and AUROC curve values using the pROC97 R package. Feature importance was calculated using the varImp function in the caret R package, which ranks features on the basis of the model-specific importance metric (mean decrease in accuracy). To interpret the contribution of individual features to the model’s predictions, we calculated Shapley values using the fastshap (https://cran.r-project.org/web/packages/fastshap) R package.
Animal model
To investigate whether the gut microbiota and its biotransformation of phytonutrients are essential for the protective effects of healthy diets, we selected IBD (including both ulcerative colitis and Crohn’s disease) among the three diseases as a relevant disease model because of its strong associations with both dietary patterns and the gut microbiome98,99. Among different in vivo models, we employed DSS-induced colitis due to its simplicity, reproducibility and similarity to human ulcerative colitis100. All mice were kept in the facility under a 12-h light/dark cycle, 8:00–20:00 light, 20:00–8:00 (the next day) dark, with unrestricted access to food (standard chow diet) and water. Colonies were maintained at 20–26 °C and 40–70% humidity. All animal experiments conducted in this study were approved by the Institutional Animal Care and Use Committee (IACUC) of GemPharmatech, ensuring compliance with ethics standards and guidelines for animal welfare (ethics approval numbers GPTAP20230904-2 and GPTAP20230904-3 for the SPF and GF mice, respectively). The institution is accredited by the Association for Assessment and Accreditation of Laboratory Animal Care International (AAALAC International) and possesses an animal use license issued by the Animal Management Committee of the Jiangsu Provincial Department of Science and Technology. Male C57BL/6J mice purchased from GemPharmatech were used in the experiments and were aged 5–6 weeks at the time of grouping.
SPF mice were weighed and randomly grouped on the basis of body weight when they were 5–6 weeks old. The date of grouping (Vehicle group, N = 8, DSS+vehicle group, N = 8, DSS+strawberry group, N = 8) was denoted as day 0. All mice in the ‘DSS+strawberry’ group were given test articles (5% strawberry powder in food) daily from day 0 until day 17. All mice in ‘DSS+vehicle’ and ‘DSS+strawberry’ groups were given 3% DSS via drinking water for 9 days (days 7–15) and then given normal water for 2 days (days 16–17). Body weight was measured twice a week before DSS induction (day 0, day 4) and daily after the first day of DSS induction. Disease activity index (DAI) scores, defined as (weight loss score + stool consistency score + rectal bleeding score)/3, were measured daily after DSS induction. Faecal samples from each mouse were collected on day 6 (before DSS induction) and at the study endpoint (day 18). All faecal samples were stored at −80 °C. Colon lengths were measured and photos were taken.
GF mice were similarly prepared as the SPF mice, with the following modifications: (1) All mice in ‘DSS+strawberry’ were given test articles (5% strawberry powder in food) daily from day 0 until day 15. (2) Given the higher susceptibility of GF mice101, a lower DSS concentration was used to induce colitis. All mice in the ‘DSS+vehicle’ and ‘DSS+strawberry’ groups were given 2% DSS via drinking water for 7 days (day 7–13) and then normal water for 2 days (days 14–15). The study endpoint was on day 15.
Haematoxylin and eosin (H&E) staining
Colon tissues were fixed with 4% paraformaldehyde (PFA) overnight. For paraffin embedding, the specimens were dehydrated in a series of ethanol dilutions, cleared in HISTO-CLEAR II solution, embedded in paraffin and sectioned into 3-μm slices. Paraffin sections were dewaxed, rehydrated in a series of ethanol dilutions and stained with haematoxylin. After being rinsed in tap water, the section was ‘blued’ by treatment with a weakly alkaline solution. Then, sections were stained with eosin, dehydrated, cleared and mounted with neutral balsam. Histological total scores were measured as the combination scores of the IBD severity level, hyperplasia, ulcers and lesion area.
Mouse stool samples library construction and sequencing
The stool samples were frozen at −80 °C and sent to Novogene UK for DNA extraction, RNA extraction and shotgun sequencing. DNA and RNA extraction were performed using TIANamp Stool DNA Kit DP328 (TIANGEN) and the Trizol-based method respectively, according to manufacturer instructions. DNA was randomly cut into short fragments to generate sequencing libraries. The fragments were end repaired, A tailed and ligated with Illumina adapter. The fragments with adapters were PCR amplified, size selected and purified. Libraries were assessed with Qubit and real-time PCR for quantification, and a bioanalyser for size distribution detection. Quantified libraries were pooled and sequenced on a NovaSeq 6000 platform S4 flow cell (Illumina) by Novogene UK. For RNA library preparation, rRNA was depleted with the standard method, followed by fragmentation into short fragments and reverse transcription into complementary (c)DNA. Sequencing ligands were ligated to the cDNA, and library fragments were purified, size selected and PCR amplified. Libraries were again checked with Qubit and real-time PCR for quantification, and a bioanalyser for size distribution detection. Quantified libraries were pooled and sequenced on a NovaSeq 6000 platform S4 flow cell (Illumina) by Novogene UK.
Functional profiling for mice metagenomics and metatranscriptomics data
Quality control of the raw reads was performed using the Sunbeam pipeline67. Functional profiling was performed using the HUMAnN3 pipeline69. To expand the list of strawberry-associated phytonutrients, we integrated strawberry-derived phytonutrients from additional sources102 into those from NutriChem. In addition, given the important role of amino acids in improving colitis, we added the strawberry-derived animo acids reported in ref. 103. The linked ECs were then retrieved using the phytonutrient–enzyme links above. We identified 396 enzymes in the murine gut microbiome linked with 50 phytonutrients in the metagenomics data and 416 enzymes linked with 53 phytonutrients in the metatranscriptomics data.
Statistical analysis
The indval function in the labdsv (https://cran.r-project.org/web/packages/labdsv) R package was used to get a list of signature species for each geographical region, according to the Dufrêne–Legendre model104. Spearman’s correlations were calculated using the cor_test function in the rstatix (https://cran.r-project.org/web/packages/rstatix/index.html) R package. Significant differences in source contributions to the sink were assessed using the Wilcoxon rank-sum test for the FEAST analysis25. Comparisons between the PC axes of the principal coordinates analysis (PCoA) plot were performed using Wilcoxon rank-sum test. Partial Spearman’s correlations between: the Shannon/Simpson diversity index and percentage of phytonutrients, age and percentage of phytonutrients, uniqueness and age, as well as between the richness diversity index and BMI were calculated using the ppcor105 R package. The relationships between phytonutrient classes and microbiome profile were calculated via envfit and considered significant if P < 0.05, with the false discovery rate adjusted using the Benjamini–Hochberg procedure. Distance-based redundancy analysis was employed to examine the relationship between host characteristics and the microbiome enzyme profile using the vegan R package. Significant differences in enzyme and species abundances between healthy and diseased groups were assessed with a zero-inflated Gaussian model using metagenomeSeq106 R package.
For mice experiments, two-sample unpaired t-tests were performed to compare the DAI scores, histological scores and body weights at each time point. Comparisons of body weights and DAI scores across multiple time points were analysed using a linear mixed model (value ~ Group × time + (1|AnimalID)) in the lmerTest107 R package. P values for fixed effect ‘Group’ were obtained by using the anova() function in the same package. Abundance was normalized to the centred-log-ratio (CLR) method with a pseudo count of half of the minimum abundance using the compositions108 R package before correlation analysis with DAI scores and histological scores. Significant differences in microbial enzymes between the DSS+strawberry and DSS+vehicle groups at different time points were assessed using the Wilcoxon rank-sum test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data associated with this study are documented in the paper and in the supplementary material (Supplementary Table 1), or are deposited in GitHub at https://github.com/lzhangxcode/Microbiome-Dietaryphytonutrients-interactions (ref. 109). The raw metagenomic and metatranscriptomic data have been deposited in ENA under Bioproject PRJEB82106. MAGs are available in figshare at https://doi.org/10.6084/m9.figshare.30347827 (ref. 110). Source data are provided with this paper.
Code availability
The analysis scripts were based on publicly available R packages as detailed in the Methods section and can be accessed in GitHub at https://github.com/lzhangxcode/Microbiome-Dietaryphytonutrients-interactions (ref. 109). Microbiome analyses were conducted using MetaPhlAn3 for taxonomic profiling and HUMAnN3 for functional potential profiling.
References
Gentile, C. L. & Weir, T. L. The gut microbiota at the intersection of diet and human health. Science 362, 776–780 (2018).
Christ, A., Lauterbach, M. & Latz, E. Western diet and the immune system: an inflammatory connection. Immunity 51, 794–811 (2019).
Nelson, M. E., Hamm, M. W., Hu, F. B., Abrams, S. A. & Griffin, T. S. Alignment of healthy dietary patterns and environmental sustainability: a systematic review. Adv. Nutr. 7, 1005–1025 (2016).
Cena, H. & Calder, P. C. Defining a healthy diet: evidence for the role of contemporary dietary patterns in health and disease. Nutrients 12, 334 (2020).
Bunge, A. C., Mazac, R., Clark, M., Wood, A. & Gordon, L. Sustainability benefits of transitioning from current diets to plant-based alternatives or whole-food diets in Sweden. Nat. Commun. 15, 951 (2024).
Berry, S. E. et al. Human postprandial responses to food and potential for precision nutrition. Nat. Med. 26, 964–973 (2020).
Johnson, A. J. et al. Daily sampling reveals personalized diet–microbiome associations in humans. Cell Host Microbe 25, 789–802.e5 (2019).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Rein, M. et al. Effects of personalized diets by prediction of glycemic responses on glycemic control and metabolic health in newly diagnosed T2DM: a randomized dietary intervention pilot trial. BMC Med. 20, 56 (2022).
Li, H. et al. Resistant starch intake facilitates weight loss in humans by reshaping the gut microbiota. Nat. Metab. 6, 578–597 (2024).
Ni, Y. et al. Resistant starch decreases intrahepatic triglycerides in patients with NAFLD via gut microbiome alterations. Cell Metab. 35, 1530–1547.e8 (2023).
Vangay, P. et al. U.S. immigration westernizes the human gut microbiome. Cell 175, 962–972.e10 (2018).
Barabási, A.-L., Menichetti, G. & Loscalzo, J. The unmapped chemical complexity of our diet. Nat. Food 1, 33–37 (2020).
Jew, S., AbuMweis, S. S. & Jones, P. J. H. Evolution of the human diet: linking our ancestral diet to modern functional foods as a means of chronic disease prevention. J. Med. Food 12, 925–934 (2009).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019).
Javdan, B. et al. Personalized mapping of drug metabolism by the human gut microbiome. Cell 181, 1661–1679.e22 (2020).
Kan, J. et al. Phytonutrients: sources, bioavailability, interaction with gut microbiota, and their impacts on human health. Front. Nutr. 9, 960309 (2022).
Beaver, L. M. et al. Promotion of healthy aging through the nexus of gut microbiota and dietary phytochemicals. Adv. Nutr. 16, 100376 (2025).
Asnicar, F. et al. Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nat. Med. 27, 321–332 (2021).
Pandey, R. P. et al. Enzymatic synthesis of novel phloretin glucosides. Appl. Environ. Microbiol. 79, 3516–3521 (2013).
Zhang, Y.-Y. et al. 3-(4-Hydroxyphenyl)propionic acid, a major microbial metabolite of procyanidin A2, shows similar suppression of macrophage foam cell formation as its parent molecule. RSC Adv. 8, 6242–6250 (2018).
Caffrey, E. B., Sonnenburg, J. L. & Devkota, S. Our extended microbiome: the human-relevant metabolites and biology of fermented foods. Cell Metab. 36, 684–701 (2024).
Braune, A., Gütschow, M. & Blaut, M. An NADH-dependent reductase from Eubacterium ramulus catalyzes the stereospecific heteroring cleavage of flavanones and flavanonols. Appl. Environ. Microbiol. 85, e01233-19 (2019).
Wilmanski, T. et al. Gut microbiome pattern reflects healthy ageing and predicts survival in humans. Nat. Metab. 3, 274–286 (2021).
Shenhav, L. et al. FEAST: fast expectation-maximization for microbial source tracking. Nat. Methods 16, 627–632 (2019).
Afzaal, M. et al. Human gut microbiota in health and disease: unveiling the relationship. Front. Microbiol. 13, 999001 (2022).
Lundberg, S. & Lee, S.-I. A unified approach to interpreting model predictions. In 31st Conference on Neural Information Processing Systems (NIPS 2017) https://papers.nips.cc/paper_files/paper/2017/file/8a20a8621978632d76c43dfd28b67767-Paper.pdf (2017).
Krishnan, S. et al. Gut microbiota-derived tryptophan metabolites modulate inflammatory response in hepatocytes and macrophages. Cell Rep. 23, 1099–1111 (2018).
Li, J. et al. Tyrosine and glutamine-leucine are metabolic markers of early-stage colorectal cancers. Gastroenterology 157, 257–259.e5 (2019).
Solanki, N. & Patel, R. Unraveling the mechanisms of trans-cinnamic acid in ameliorating non-alcoholic fatty liver disease. Am. J. Transl. Res. 15, 5747–5756 (2023).
Han, Y. et al. Dietary intake of whole strawberry inhibited colonic inflammation in dextran-sulfate-sodium-treated mice via restoring immune homeostasis and alleviating gut microbiota dysbiosis. J. Agric. Food Chem. 67, 9168–9177 (2019).
Chen, L. et al. Effects of β-alanine on intestinal development and immune performance of weaned piglets. Anim. Nutr. 12, 398–408 (2022).
Bhattarai, Y. et al. Bacterially derived tryptamine increases mucus release by activating a host receptor in a mouse model of inflammatory bowel disease. iScience 23, 101798 (2020).
Wang, X. et al. Asparagine attenuates intestinal injury, improves energy status and inhibits AMP-activated protein kinase signalling pathways in weaned piglets challenged with Escherichia coli lipopolysaccharide. Br. J. Nutr. 114, 553–565 (2015).
Culp, E. J., Nelson, N. T., Verdegaal, A. A. & Goodman, A. L. Microbial transformation of dietary xenobiotics shapes gut microbiome composition. Cell 187, 6327–6345.e20 (2024).
Kuziel, G. A. et al. Functional diversification of dietary plant small molecules by the gut microbiome. Cell 188, 1967–1983.e22 (2025).
Roichman, A. et al. Microbiome metabolism of dietary phytochemicals controls the anticancer activity of PI3K inhibitors. Cell 188, 3065–3080.e21 (2025).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008).
Aron-Wisnewsky, J. et al. Gut microbiota and human NAFLD: disentangling microbial signatures from metabolic disorders. Nat. Rev. Gastroenterol. Hepatol. 17, 279–297 (2020).
Miquel, S. et al. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 16, 255–261 (2013).
Khan, M. T. et al. Synergy and oxygen adaptation for development of next-generation probiotics. Nature 620, 381–385 (2023).
Ratiner, K., Ciocan, D., Abdeen, S. K. & Elinav, E. Utilization of the microbiome in personalized medicine. Nat. Rev. Microbiol. 22, 291–308 (2024).
Jahn, L. J., Rekdal, V. M. & Sommer, M. O. A. Microbial foods for improving human and planetary health. Cell 186, 469–478 (2023).
Drouin-Chartier, J.-P. et al. Systematic review of the association between dairy product consumption and risk of cardiovascular-related clinical outcomes. Adv. Nutr. 7, 1026–1040 (2016).
Gijsbers, L. et al. Consumption of dairy foods and diabetes incidence: a dose-response meta-analysis of observational studies. Am. J. Clin. Nutr. 103, 1111–1124 (2016).
Kim, E. K. et al. Fermented kimchi reduces body weight and improves metabolic parameters in overweight and obese patients. Nutr. Res. 31, 436–443 (2011).
Vich Vila, A. et al. Gut microbiota composition and functional changes in inflammatory bowel disease and irritable bowel syndrome. Sci. Transl. Med. 10, eaap8914 (2018).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Bae, M. et al. Metatranscriptomics-guided discovery and characterization of a polyphenol-metabolizing gut microbial enzyme. Cell Host Microbe 32, 1887–1896.e8 (2024).
Wu, Q. et al. Activity of GPCR-targeted drugs influenced by human gut microbiota metabolism. Nat. Chem. 17, 808–821 (2025).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Iliev, I. D., Ananthakrishnan, A. N. & Guo, C.-J. Microbiota in inflammatory bowel disease: mechanisms of disease and therapeutic opportunities. Nat. Rev. Microbiol. 23, 509–524 (2025).
Zheng, J. et al. Noninvasive, microbiome-based diagnosis of inflammatory bowel disease. Nat. Med. 30, 3555–3567 (2024).
Wong, C. C. & Yu, J. Gut microbiota in colorectal cancer development and therapy. Nat. Rev. Clin. Oncol. 20, 429–452 (2023).
Tito, R. Y. et al. Microbiome confounders and quantitative profiling challenge predicted microbial targets in colorectal cancer development. Nat. Med. 30, 1339–1348 (2024).
Hall, A. B. et al. A novel Ruminococcus gnavus clade enriched in inflammatory bowel disease patients. Genome Med. 9, 103 (2017).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Leung, H. et al. Risk assessment with gut microbiome and metabolite markers in NAFLD development. Sci. Transl. Med. 14, eabk0855 (2022).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Hannigan, G. D., Duhaime, M. B., Ruffin, M. T., Koumpouras, C. C. & Schloss, P. D. Diagnostic potential and interactive dynamics of the colorectal cancer virome. mBio 9, e02248-18 (2018).
Vogtmann, E. et al. Colorectal cancer and the human gut microbiome: reproducibility with whole-genome shotgun sequencing. PLoS ONE 11, e0155362 (2016).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Wirbel, J. et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat. Med. 25, 679–689 (2019).
Yu, J. et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66, 70–78 (2017).
Thomas, A. M. et al. Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat. Med. 25, 667–678 (2019).
Yachida, S. et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 25, 968–976 (2019).
Clarke, E. L. et al. Sunbeam: an extensible pipeline for analyzing metagenomic sequencing experiments. Microbiome 7, 46 (2019).
Seelbinder, B. et al. Candida expansion in the gut of lung cancer patients associates with an ecological signature that supports growth under dysbiotic conditions. Nat. Commun. 14, 2673 (2023).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife 10, e65088 (2021).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51, D1373–D1380 (2023).
Fleischmann, A. et al. IntEnz, the integrated relational enzyme database. Nucleic Acids Res. 32, D434–D437 (2004).
Chang, A. et al. BRENDA, the ELIXIR core data resource in 2021: new developments and updates. Nucleic Acids Res. 49, D498–D508 (2021).
Wishart, D. S. et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 46, D608–D617 (2018).
Gao, J., Ellis, L. B. M. & Wackett, L. P. The University of Minnesota Biocatalysis/Biodegradation Database: improving public access. Nucleic Acids Res. 38, D488–D491 (2010).
Nagano, N. et al. EzCatDB: the enzyme reaction database, 2015 update. Nucleic Acids Res. 43, D453–D458 (2015).
Ribeiro, A. J. M. et al. Mechanism and Catalytic Site Atlas (M-CSA): a database of enzyme reaction mechanisms and active sites. Nucleic Acids Res. 46, D618–D623 (2018).
Akiva, E. et al. The structure–function linkage database. Nucleic Acids Res. 42, D521–D530 (2014).
Hoffmann, M. F. et al. The Transformer database: biotransformation of xenobiotics. Nucleic Acids Res. 42, D1113–D1117 (2014).
Rutz, A. et al. The LOTUS initiative for open knowledge management in natural products research. eLife 11, e70780 (2022).
Zhao, H. et al. NPASS database update 2023: quantitative natural product activity and species source database for biomedical research. Nucleic Acids Res. 51, D621–D628 (2023).
Nakamura, K. et al. KNApSAcK-3D: a three-dimensional structure database of plant metabolites. Plant Cell Physiol. 54, e4 (2013).
Gallo, K. et al. SuperNatural 3.0—a database of natural products and natural product-based derivatives. Nucleic Acids Res. 51, D654–D659 (2023).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Zhang, R. et al. InflamNat: web-based database and predictor of anti-inflammatory natural products. J. Cheminformatics 14, 30 (2022).
Tao, L. et al. Database and bioinformatics studies of probiotics. J. Agric. Food Chem. 65, 7599–7606 (2017).
Tarracchini, C. et al. The Integrated Probiotic Database: a genomic compendium of bifidobacterial health-promoting strains. Microbiome Res. Rep. 1, 9 (2022).
Chan, P. L. et al. ProBioQuest: a database and semantic analysis engine for literature, clinical trials and patents related to probiotics. Database 2022, baac059 (2022).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Templ, M., Hron, K. & Filzmoser, P. in Compositional Data Analysis 341–355 (John Wiley & Sons, 2011).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Kursa, M. B. & Rudnicki, W. R. Feature selection with the Boruta package. J. Stat. Softw. 36, 1–13 (2010).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Adolph, T. E. & Zhang, J. Diet fuelling inflammatory bowel diseases: preclinical and clinical concepts. Gut 71, 2574–2586 (2022).
Pereira, G. V. et al. Opposing diet, microbiome and metabolite mechanisms regulate inflammatory bowel disease in a genetically susceptible host. Cell Host Microbe 32, 527–542.e9 (2024).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 104, Unit–15.25 (2014).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Huang, S., Ying Lim, S., Lau, H., Ni, W. & Fong Yau Li, S. Effect of glycinebetaine on metabolite profiles of cold-stored strawberry revealed by 1H NMR-based metabolomics. Food Chem. 393, 133452 (2022).
Zhang, J. et al. Metabolic profiling of strawberry (Fragaria×ananassa Duch.) during fruit development and maturation. J. Exp. Bot. 62, 1103–1118 (2011).
Dufrêne, M. & Legendre, P. Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol. Monogr. 67, 345–366 (1997).
Kim, S. ppcor: an R package for a fast calculation to semi-partial correlation coefficients. Commun. Stat. Appl. Methods 22, 665–674 (2015).
Paulson, J. N., Stine, O. C., Bravo, H. C. & Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 10, 1200–1202 (2013).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest package: tests in linear mixed effects models. J. Stat. Softw. 82, 1–26 (2017).
van den Boogaart, K. G. & Tolosana-Delgado, R. “compositions”: a unified R package to analyze compositional data. Comput. Geosci. 34, 320–338 (2008).
Zhang, L. Microbiome_Dietaryphytonutrients_interactions. GitHub https://github.com/lzhangxcode/Microbiome-Dietaryphytonutrients-interactions (2025).
Zhang, L. Assembled MAGs data. figshare https://doi.org/10.6084/m9.figshare.30347827 (2025).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (German Research Foundation; DFG) under Germany’s Excellence Strategy (EXC 2051) (project ID 390713860; G.P., Y.N., A.M.-S.), the BMBF-funded ‘PerMiCCion’ project (project ID 01KD2101A; G.P., A.D.-C.), EU Horizon-funded ‘NUTRIMMUNE’ (grant agreement no. 101162457; G.P.) and the Excellent Young Scientists Fund of the National Natural Science Foundation of China (Overseas), project ID 24HAA01325 (Y.N.). We acknowledge the generous support of L. Jeske from the Brenda team for extracting phytonutrient-related reactions in BKMS-react and S. Eichler for helping with the fine curation of the natural products.
Author information
Authors and Affiliations
Contributions
G.P. and Y.N. conceptualized and designed the study. L.Z., A.M.-S. and B.S. collected, processed and analysed the data. L.Z. and Y.N. analysed and interpreted data from the animal experiment. T.-H.K. and M.Z. performed in vitro work and interpreted the data. L.Z., A.M.-S. and T.-H.K. worked on the visualization of the chemical structures. G.P., L.Z., Y.N., A.M.-S. and A.D.-C. wrote the original manuscript draft. G.P., L.Z., Y.N., A.M.-S., B.S., A.D.-C., L.v.D., L.J.J., M.O.A.S., T.-H.K. and M.Z. edited the manuscript. G.P. and Y.N. led and supervised the research work. All authors made substantial contributions, reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Paolo Manghi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sample accumulation curve, and comparison of microbiome enzymatic potential between gut microbiota and probiotics.
a, Sample accumulation curve of microbial enzymes, phytonutrient-associated microbial enzymes and microbial species for different continents (Africa, Europe, America, Oceania and Asia). The x-axis represents the number of samples. b, Ratios of ECs detected by HUMAnN3 to the total ECs present in the corresponding species from the ChocoPhlAn database under different species prevalence thresholds (left) and different species abundance thresholds (right). The x-axis represents the applied threshold values; the y-axis represents the calculated ratio. The two horizontal dashed lines represent ratios of 0.90 and 0.95, respectively. c, Pie chart showing phytonutrients that can be modified either exclusively by gut bacteria (designated ‘gut-restricted’) or by gut bacteria and probiotics (designated ‘shared’). d, Phytonutrient accumulation curve with combined biotransformation potential covering 106 ‘gut-restricted’ non-primary metabolism related phytonutrients. The horizontal line represents 80% of the total phytonutrients and the vertical line marks the species rank at which the cumulative phytonutrient number just exceeds 80%. Boxplots show the median (centrelines), first and third quartiles (box limits) and 1.5x interquartile range bounds (whiskers).
Extended Data Fig. 2 Biotransformation kinetics of phytonutrient substrates butein, isoliquiritigenin and butin.
a, Biotransformation kinetics of the phytonutrient substrates isoliquiritigenin (top) and butein (bottom), along with their corresponding products (respectively, liquiritigenin and davidigenin; and butin and butin-FcrP, the product resulting from the Fcr enzyme reaction of butin), as catalysed by E. ramulus under assay conditions using M9 minimal medium. Two initial substrate concentrations (20 µM and 200 µM) were tested in the presence of E. ramulus, heat-killed (HK) E. ramulus, E. coli K-12 or a bacteria-free medium control. b, Biotransformation kinetics of the phytonutrient substrate butin, along with its corresponding products, under assay conditions using mGAM medium. The corresponding products include butein and butin-FcrP (the product resulting from the Fcr enzyme reaction of butin). Lines represent mean and error bars represent s.d. calculated from three independent assay replicates.
Extended Data Fig. 3 Geographical specificity of phytonutrient biotransformation.
a, Two-sided Partial Spearman’s correlation (adjusted by sequencing depth) between species’ Simpson diversity and the enzyme ratio. b, As in a, but including only phytonutrients that are part of the secondary metabolism. Red line and red area show the linear regression line and its 95% confidence interval in a – b. c, Boxplot comparing the uniqueness calculated when using samples from different individuals (n = 2531) than from same individuals (n = 719). Significance (P = 1.44e-40) is determined by two-sided Wilcoxon rank-sum test. d, Left: Distributions of the number of foods that can be biotransformed by the gut microbiota of each sample in each continent. Mean ± s.d. shown. Right: UpSet plot of the total number of foods that were biotransformed by the gut microbiota in each continent (1% prevalence cut-off of a region). The total number of foods is displayed as horizontal bars on the left part of the image. Intersection sizes are shown as black vertical bars. e, Principal coordinate analysis (PCoA) of Jaccard distance between phytonutrient-associated microbial ECs’ profiles (n = 1936). Significant differences among different continents were determined using PERMANOVA and were considered significant if P < 0.05. f, PCoA of Aitchison distance between phytonutrient-associated microbial ECs’ profiles. Significant differences were determined using PERMANOVA and were considered significant if P < 0.05. Vector matrices representing compound classes recorded in NutriChem are depicted by individual black lines with arrows. g, Host features explaining the variation of phytonutrient-associated enzyme profile by dbRDA analysis. Left: The total variation explained by the given features. Right: The dark bars represent the variation explained by each individual feature (all listed features are significant, P < 0.05, significance was determined using dbRDA analysis with FDR correction), while the light-coloured bars represent the cumulative variation explained by the given features. Features above the black line indicate the selected features by stepwise dbRDA analysis. h, Two-sided Partial Spearman’s correlation (adjusted by sequencing depth) between age and uniqueness. i, Two-sided Partial Spearman’s correlation (adjusted by sequencing depth) between age and the ratio of phytonutrient-associated enzymes to the total number of microbial enzymes detected in the 3068 metagenomic samples. j, two-sided Spearman’s correlation between BMI and the richness of phytonutrient-associated enzymes. The white line and grey area show the linear regression line and its 95% confidence interval in h–j. k, Boxplots showing the adaptation of dietary habits (grains, vegetables, fruits) of the Thailand’s cohort with paired metagenomic sequencing data and dietary intake. The y-axis indicates the amounts of the diet. The x-axis indicates different cohorts: Thailand (Thai individuals living in Thailand, n = 174), US-born (US-born subjects, n = 36), new arrivals (Thai individuals who have recently moved to the US, n = 300), and long-term residents (Thai individuals who have been living in the US for at least 20 years, n = 77) and 2nd generation (Thai individuals who were born in the US, n = 57). Boxplot shows the median (centerline), the lower and upper quartiles (box edges), and whiskers that extend to data points within 1.5× the interquartile range (IQR) from the quartiles.
Extended Data Fig. 4 Altered biotransformation potential of gut microbiota for each potentially beneficial food.
a, Bar plots illustrating the percentage of phytonutrient-modifying microbial enzymes that are significantly differentially abundant by each beneficial food for individuals with IBD (left), CRC (centre) and NAFLD (right). The y-axis represents beneficial foods for each disease (with at least 10 phytonutrient-associated microbial ECs). Significant differences were assessed by two-tailed Wilcoxon ranked sum test between healthy and diseased individuals. b, Receiver operating characteristic (ROC) curves of machine learning models discriminating between health and disease states using the stratified abundances of enzymes not associated with phytonutrients in IBD (left), CRC (centre) and NAFLD (left).
Extended Data Fig. 5 Selected features in the models between health and disease.
Selected features of the machine learning models. Left: Bar plots showing the importance of the selected features. Dark colours indicate disease-associated features; light colours represent healthy control–associated features. Middle: SHAP value plot illustrating the impact of each feature on the model output, with features ordered by their importance. Right: Heatmap depicting species and their associated ECs from the stratified features in the left panel, with colours indicating whether the species significantly differentially abundant species (S), significantly differentially abundant ECs (S), number of associated phytonutrients (C), number of associated foods (F) and type of enzyme classes (Class). (a–c) Diseases analysed were inflammatory bowel disease (IBD; a), colorectal cancer (CRC; b) and non-alcoholic fatty liver disease (NAFLD; c).
Extended Data Fig. 6 Changes in histological scores and DAI scores in the mouse model of DSS-induced colitis.
a, Changes in the four component scores used to calculate histological scores in SPF mice (n = 8, each group) were monitored at the study endpoint. Histological scores were determined by assessing four parameters: IBD severity level, hyperplasia, ulcers and lesion areas. b, Changes in the three scores used to calculate DAI in SPF mice were monitored throughout the entire study. The DAI score was calculated as the total score (body weight decrease + stool consistency + rectal bleeding) divided by 3. # indicates significant differences between the Vehicle and DSS+Vehicle group (#P ≤ .05; ##P ≤ .01; ###P ≤ .001; ####P ≤ .0001), and * indicates significant differences between the DSS+Vehicle and DSS+Strawberry group (*P ≤ .05; **P ≤ .01; ***P ≤ .001; ****P ≤ .0001), using two-sided two-sample unpaired t-tests. c,d, As in b,a, but in GF mice (n = 6 or 7, each group). All data are presented as Mean ± SE in a–d.
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Tables 1–9.
Supplementary Data (download XLSX )
Supplementary Data 1 and 2.
Source data
Source Data Figs. 1, 3–5 and Source Data Extended Data Figs. 3–5 (download XLSX )
Statistical source data for Figs. 1d, 3d–g, 4 and 5 and Extended Data Figs. 3k, 4 and 5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, L., Marfil-Sánchez, A., Kuo, TH. et al. Gut microbiome-mediated transformation of dietary phytonutrients is associated with health outcomes. Nat Microbiol 11, 94–110 (2026). https://doi.org/10.1038/s41564-025-02197-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02197-z