Abstract
Mucormycosis is an emerging, life-threatening human infection caused by Mucorales fungi1,2,3. Metabolic disorders uniquely predispose an ever-expanding group of patients to mucormycosis through poorly understood mechanisms1,2,4,5, suggesting that uncharacterized host metabolic effectors may confer protective immunity against this infection. Here we uncover a master regulatory role of albumin in host defence against Mucorales through the modulation of fungal pathogenicity. Our initial studies identified severe hypoalb uminaemia as a prominent metabolic abnormality and an independent biomarker of poor mucormycosis outcome across three distinct cohorts of patients with mucormycosis. Notably, purified albumin selectively inhibits Mucorales growth among a range of pathogens, and albumin-deficient mice display susceptibility specifically to mucormycosis. The antifungal activity of albumin is mediated by the release of bound free fatty acids (FFAs). Albumin prevents FFA oxidation, which otherwise abolishes their antifungal properties, and sera from patients with mucormycosis display high levels of oxidized FFAs. Physiologically, albumin-bound FFAs suppress the expression of key virulence factors by inhibiting protein synthesis, the reby rendering Mucorales avirulent in vivo. Overall, we identify a host defence mechanism that directs the pathogen to suppress its pathogenicity program in response to unfavourable metabolic cues regulated by albumin. These findings have major implications for the pathogenesis and management of mucormycosis.
Similar content being viewed by others
Main
Mucorales fungi cause mucormycosis—an emerging, life-threatening, opportunistic infection with limited therapeutic options and incompletely understood pathogenesis1,6,7. The overall mortality of mucormycosis exceeds 50% and approaches 100% in patients with disseminated disease1. The clinical hallmark of mucormycosis is massive tissue necrosis induced by the fungus, which impedes antifungal agents from reaching the sites of infection and often necessitates radical, disfiguring surgery to control the disease1,8. In contrast to other fungal infections, mucormycosis predominantly affects an ever-expanding group of patients with metabolic abnormalities through incompletely characterized mechanisms1,4,5. Specifically, poorly controlled diabetes mellitus (DM), acidosis, acquired iron overload syndromes, malnutrition and immunometabolic dysregulation induced by COVID-19 uniquely predispose individuals to the development of mucormycosis1,2,3,4,6,8. Thus, uncharacterized metabolic host defence mechanisms may confer protective immunity against mucormycosis.
From the pathogen perspective, Mucorales senses cues in the tissue environment, triggering the expression of virulence factors that transform these saprophytic organisms into rapidly invasive and potentially lethal pathogens1. Specifically, the production of the potent mycotoxin mucoricin during Mucorales hyphae growth9 induces extensive tissue necrosis, whereas the binding of CotH invasins to specific host receptors promotes angioinvasion and fungal dissemination10,11,12. Notably, germinating spores of Mucorales evade phagocytosis and induce acute lethality within 24 h of pulmonary infection in immunocompetent mice13. Thus, it is essential to identify metabolic host effectors that prevent extracellular growth and modulate Mucorales pathogenicity during the early stages of infection.
Albumin selectively inhibits Mucorales
Human serum has important inhibitory effects against Mucorales, which remain molecularly unexplored14,15. We found that, compared with sera from healthy individuals, the ability of sera from patients with mucormycosis to inhibit hyphal growth of a clinical isolate of Rhizopus arrhizus var. delemar (hereafter, R. delemar) is almost completely lost (Fig. 1a). Albumin, the most abundant serum protein, regulates important physiological functions intravascularly, in the interstitial space, and on mucosal surfaces16. Furthermore, severe hypoalbuminaemia is a common finding in patients with diverse immunometabolic abnormalities predisposing for mucormycosis2,5,17. We therefore decided to comprehensively evaluate the physiological function of albumin against Mucorales.
a, The length of germinating R. delemar spores cultured for 5 h in RPMI medium (n = 12), serum from healthy individuals (n = 5) or serum from patients with mucormycosis before initiation of antifungals (n = 4). Statistical analysis was performed using one-way analysis of variance (ANOVA) with Tukey’s post hoc test; *P = 0.0213, **P = 0.0079. Representative images are shown. Scale bar, 100 µm. b, Serum albumin concentrations at diagnosis in contemporaneously matched controls (n = 33), patients with pulmonary aspergillosis (n = 50) and patients with mucormycosis (n = 81). Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test; NS, not significant (P = 0.466). c, Kaplan–Meier survival curves for patients with mucormycosis presenting with severe hypoalbuminaemia (≤2.5 g dl−1) compared with all other mucormycosis cases from independent clinical cohorts in the USA (n = 81), India (n = 101) and France (n = 26). Survival differences were assessed using the log-rank (Mantel–Cox) test. MDACC, MD Anderson Cancer Center. d, The effect of albumin depletion from human serum of healthy donors on the growth of R. delemar (n = 15) and A. fumigatus (n = 6). Statistical analysis was performed using two-sided Wilcoxon matched-pairs signed-rank test; NS, P = 0.156. e, Coomassie blue staining of intact human serum, albumin-depleted serum and albumin-enriched eluates. Representative of three independent experiments. f, Germ-tube elongation of R. delemar spores cultured in medium with or without purified HSA, quantified by time-lapse microscopy. n = 50–200 spores per timepoint; two independent experiments, performed in triplicate. Data are mean ± s.d. Statistical analysis was performed using two-way ANOVA with Sidak’s multiple-comparisons test. g, Representative images of Calcofluor White (CFW)-stained R. delemar spores after 8 h of culture in medium with or without HSA. Scale bars, 50 µm. h, Inhibition of R. delemar growth by increasing concentrations of HSA or BSA. n = 4 independent experiments performed in triplicate. Data are mean ± s.d. i, Quantification of the growth of different Mucorales species and other pathogens cultured in medium with or without HSA. Representative of three independent experiments in triplicate. Full scientific names are given in the Methods. ****P < 0.0001 (b,d,f). The diagram in f was created using BioRender.
We initially explored the associations between serum albumin levels and susceptibility to the development of pulmonary mucormycosis and its outcome in contemporaneous high-risk patients with haematological malignancy at a major tertiary care cancer centre in the United States (Supplementary Table 1). Notably, most patients who developed pulmonary mucormycosis had significantly lower albumin levels at diagnosis compared with control patients matched for the underlying disease who developed bacterial pneumonia or pneumonia caused by the major airborne human fungal pathogen Aspergillus fumigatus (Fig. 1b). Furthermore, patients with mucormycosis and very low albumin levels (≤2.5 g dl−1) had significantly lower survival rates compared with other patients with mucormycosis (Fig. 1c). These findings were independently validated through an analysis of serum albumin levels in a cohort of patients with pulmonary mucormycosis in a tertiary care centre from the Postgraduate Institute of Medical Education and Research (PGIMER), Chandigarh, India (Fig. 1c and Supplementary Table 1), who had DM and COVID-19 as main underlying risk factors, and a published cohort of patients with mucormycosis from France (Ambizygo Study18) who had different risk factors (Fig. 1c). Notably, multivariate survival analysis identified severe hypoalbuminaemia (≤2.5 g dl−1) as an independent predictor of poor outcome across all three clinical cohorts of patients with mucormycosis (Extended Data Tables 1–3).
We next analysed the functional relationship between the serum albumin concentration and antifungal activity in patients at high risk for mucormycosis. Notably, we detected a significant association between the degree of hypoalbuminaemia and the loss of inhibitory activity of serum against R. delemar hyphal formation in prospectively collected sera from patients with liver cirrhosis or haematological malignancy (Extended Data Fig. 1a and Supplementary Table 1).
To account for potential confounders related to the underlying disease, we performed albumin depletion in sera from healthy individuals using affinity chromatography19. Equilibration of Cibacron Blue chromatography columns was achieved with concentrated serum filtrate previously passed through a 50 kDa centrifugal filter unit, to ensure that the albumin-depleted flow-through was not diluted for other serum proteins. We assessed the effect of albumin depletion on the activity of serum against R. delemar and another major human respiratory fungal pathogen A. fumigatus7. Notably, albumin depletion resulted in a significant loss of antifungal activity of the serum selectively against R. delemar but not against A. fumigatus (Fig. 1d).
We next purified human albumin from the sera of healthy volunteers to directly evaluate its antifungal activity against Mucorales using a protocol based on affinity column chromatography19 (Fig. 1e and Extended Data Fig. 1b). We found no evidence of transferrin, a serum iron-transferring protein with important role in nutritional immunity against Mucorales20, in the albumin-containing eluted fractions by western blot analysis (Extended Data Fig. 1c). Notably, purified human albumin dissolved at physiological concentrations (around 3.5 g dl−1) in liquid culture medium had potent activity against R. delemar (Fig. 1f), an effect that was not observed with comparable concentrations of human IgG (Extended Data Fig. 1d). Moreover, purified albumin from different sources, including commercially available bovine serum albumin (BSA) and human serum albumin (HSA) used for therapeutic applications, inhibited Mucorales growth when added at physiologically relevant concentrations in culture medium (Fig. 1g,h). Notably, we found that albumin specifically blocks filamentous (hyphal) growth of Mucorales after the initial stage of isotropic growth (swelling) of fungal spores (Extended Data Fig. 2a). The antifungal activity of albumin was fully reversible after culture of inhibited Mucorales spores in fresh medium without albumin (Extended Data Fig. 2b). Albumin selectively inhibited a wide range of clinical isolates of Mucorales species at physiological serum concentrations (4.5 g dl−1), whereas it showed no significant activity against other major human bacterial or fungal pathogens (Fig. 1i and Extended Data Fig. 2c). Collectively, these results reveal the specialized activity of albumin against Mucorales and identify severe hypoalbuminaemia as an independent biomarker of poor outcome of mucormycosis.
Albumin-bound FFAs inhibit Mucorales
To examine whether the inhibitory activity of albumin requires direct interaction of the protein with fungal cells, we filtered culture medium containing inhibitory concentrations of albumin (4.5 g dl−1) and assessed the antifungal activity of the culture filtrate (flow-through). Notably, we found that the flow-through of albumin retains full inhibitory activity against R. delemar (Extended Data Fig. 3a). Nutritional immunity is an essential host defence mechanism against Mucorales1,13. In view of the ability of albumin to bind a wide range of compounds, endogenous molecules, iron and other transition metals21, we investigated the possibility that albumin inhibits Mucorales through depletion of essential nutrients from the culture medium. We therefore analysed the components of regular RPMI culture medium and found that the presence of albumin resulted in a significant reduction in the concentration of certain amino acids (Extended Data Fig. 3b and Supplementary Table 2). However, supplementation experiments with amino acids and different combinations of nutrients contained in RPMI medium did not affect the inhibitory activity of albumin flow-through against R. delemar (Extended Data Fig. 3c).
We next considered the possibility that the release of an inhibitory molecule bound to albumin could account for the antifungal activity of the albumin-rich culture filtrate. Serum albumin acts as the main shuttle of non-esterified middle- and long-chain fatty acids (FFAs) in extracellular fluids22 and FFAs possess antimicrobial properties23. We therefore performed fractionation of lipid-containing elutions from a BSA-containing culture filtrate and functionally characterized the inhibitory activity against R. delemar. We identified a fraction with significant inhibitory activity against Mucorales; this fraction contained high amounts of caprylic acid (C8:0) as determined using gas chromatography–mass spectrometry (GC–MS) and electrospray ionization high-resolution MS (ESI-HRMS) analysis (Extended Data Fig. 4a–c). We also confirmed that purified caprylic acid dissolved in ethanol has potent inhibitory activity against Mucorales at concentrations lower than those contained in the BSA filtrate (Extended Data Fig. 4d).
To further explore the physiological relevance of our findings, we performed lipidomic profiling in purified human albumin isolated from healthy donors before and after filtration (Fig. 2a). The albumin filtrate retained full inhibitory activity against Mucorales spores (Fig. 2a) and contained physiological middle- and long-chain FFAs (Fig. 2b and Supplementary Table 3). Furthermore, we found that purified serum FFAs dissolved in ethanol exerted potent anti-Mucorales activity at physiologically relevant concentrations24 (Fig. 2c). We also found that a broad range of FFAs of various carbon chain lengths and degrees of saturation display potent antifungal activity against Mucorales (Fig. 2d). Importantly, purified human albumin after charcoal treatment for the removal of bound FFAs25, or commercially available BSA free of fatty acids (FFA-free BSA), had no activity against Mucorales (Fig. 2e). Furthermore, charcoal-treated BSA complexed with physiologically relevant concentrations of oleic acid (OA) dissolved the FFA and fully restored its activity against Mucorales (Fig. 2f). Experiments with fluorescent-labelled albumin further demonstrated that, although albumin avidly binds to the fungal cell wall, it is not internalized by Mucorales spores (Fig. 2g). Collectively, these findings demonstrate that albumin binds to, dissolves, shuttles and facilitates the release of physiological FFAs to optimize their antifungal activity.
a, Schematic of the experimental workflow for generating flow-through and isolating albumin from human serum using a 3 kDa molecular weight cut-off (MWCO) centrifugal filter (top). Bottom, representative bright-field images from three independent experiments showing R. delemar spores cultured for 5 h in minimal medium alone or in medium supplemented with isolated HSA or albumin flow-through. FT, flow-through; HD, healthy donor; ND, not detectable. Scale bar, 50 µm. b, Lipidomic profiling of FFAs in isolated HSA and corresponding flow-through obtained as in a. c, Antifungal activity of the major serum FFAs against R. delemar. The red-striped areas indicate physiological serum concentrations of each FFA. Data are mean ± s.d., representative of n = 3 independent experiments. IC50, half-maximum inhibitory concentration. d, The dose-dependent inhibitory effect of short-, medium- and long-chain FFAs on R. delemar growth. e, Representative images of R. delemar spores cultured for 5 h in medium supplemented with isolated HSA or charcoal-stripped albumin (left). Scale bar, 50 µm. Right, quantification of R. delemar growth inhibition by BSA, FFA-free BSA or charcoal-stripped BSA. n = 4 independent experiments, performed in triplicate. Data are mean ± s.d. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple-comparison test; NS, P = 0.574. f, The inhibitory effects of charcoal-stripped BSA, OA or charcoal-stripped BSA reconstituted with OA on R. delemar growth. Data are a representative example of n = 3 independent experiments, shown as the mean ± s.d., performed in triplicate. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple-comparison test. g, Representative fluorescence microscopy images from three independent experiments showing CFW-labelled R. delemar spores cultured for 6 h in the presence of FITC-labelled albumin, performed in duplicate. Scale bar, 10 µm. ****P < 0.0001 (e,f). The diagram in a was created using BioRender.
Albumin protects FFAs from oxidation
We next performed targeted lipidomic profiling of the sera of patients with mucormycosis and matched controls to analyse the FFA composition and identify abnormalities associated with the loss of antifungal activity in the serum. Notably, we found that the sera of patients with mucormycosis contained a significantly greater proportion of oxidized forms of FFAs than the sera of control patients, who were matched for the underlying disease without infection or those who have been diagnosed with invasive aspergillosis (Fig. 3a and Supplementary Table 3). Furthermore, the sera of patients susceptible to mucormycosis due to underlying cirrhosis or haematological malignancy contained high amounts of oxidized FFAs, which was directly proportional to the severity of hypoalbuminaemia, particularly in cirrhosis (Fig. 3b,c). Importantly, in patients with cirrhosis, the degree of serum FFA oxidation was strongly correlated with the loss of inhibitory activity against Mucorales (Fig. 3d), whereas this association was less pronounced in the sera of patients with haematological malignancy. Collectively, these findings suggested that oxidized FFAs display attenuated activity against Mucorales.
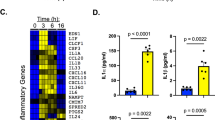
a, The relative concentrations of oxidized FFAs in sera from matched control patients (n = 6), patients with invasive aspergillosis (n = 6) and mucormycosis (n = 18). NS, P = 0.973; **P = 0.0061, **P = 0.0031. b, The relative concentrations of oxidized FFAs in sera from healthy controls (n = 21), patients with cirrhosis (n = 18) and patients with haematological malignancies (n = 20). NS, P = 0.576; *P = 0.0402, **P = 0.0066. c,d, Correlation between oxidized serum FFAs and albumin levels (c) or antifungal activity (d) of sera from patients with cirrhosis (n = 18) and haematological malignancies (n = 20) against R. delemar. Statistical analysis was performed using two-sided linear regression. e, Inhibitory effects of increasing concentrations of OA and oxidized OA on R. delemar growth. n = 2 independent experiments performed in triplicate. Data are mean ± s.d. f, Serum lipids from healthy individuals were oxidized and tested for antifungal activity. n = 13. Statistical analysis was performed using the two-sided Wilcoxon matched-pairs signed-rank test; ***P = 0.0002. g, CFW-labelled R. delemar spores cultured for 3 h with control or pre-oxidized C11-BODIPY (left). Scale bars, 8 µm. Right, quantification of C11-BODIPY mean fluorescence intensity (MFI) at 510 nm and 590 nm (n = 269–979 spores, three experiments). NS, P = 0.769. Em., emission. h, GC–MS analysis of non-oxidized OA in OA and BSA-conjugated OA before and after microwave oxidation. n = 3, triplicates. **P = 0.0041. i, R. delemar spores cultured for 5 h with oxidized OA or oxidized BSA–OA. Scale bar, 100 µm. Quantification of germling length is shown. n = 202–237 spores, three experiments. Statistical analysis was performed using two-sided Mann–Whitney U-tests. j, Area under the curve (AUC) quantification of caprylic acid in mock-treated and glycosylated BSA. n = 3, triplicates. Statistical analysis was performed using two-sided unpaired t-tests. k, The inhibitory effects of mock-treated and glycosylated BSA on R. delemar growth (n = 4, triplicates). Statistical analysis was performed using two-sided Mann–Whitney U-tests; *P = 0.0286. For i–k, data are mean ± s.d. For a, b, g and i, statistical analysis was performed using one-way ANOVA with Tukey’s test. ****P < 0.0001 (g–j).
To directly evaluate the antifungal activity of oxidized FFAs, we oxidized OA—a major physiological serum FFA—and assessed the effect on antifungal activity. We found that the oxidation of OA resulted in a greater than 100-fold decrease in its inhibitory activity against R. delemar (Fig. 3e). To establish a causal relationship between FFA oxidation and the loss of anti-Mucorales activity in serum, we isolated serum lipids from healthy individuals and assessed the effects of oxidation on their antifungal properties. Importantly, oxidation of serum lipids resulted in a significant decrease in their inhibitory properties against R. delemar spores (Fig. 3f).
As FFA oxidation diminishes their uptake by mammalian cells26, we evaluated the internalization of oxidized FFAs by fungal cells. We initially found that the uptake of a long-chain FFA fluorescent analogue C12-BODIPY by Mucorales spores is energy dependent, as it was abolished at 0 °C (Extended Data Fig. 5a). We next evaluated uptake of C11-BODIPY581/591 (hereafter, C11-BODIPY), a ratiometric reporter of lipid peroxidation that shifts its fluorescence after oxidation from red (around 590 nm; reduced) to green (around 510 nm; oxidized)27, in R. delemar spores with or without previous in vitro oxidation. Pre-oxidation almost completely abolished C11-BODIPY uptake by R. delemar, as indicated by a marked reduction in both reduced and oxidized fluorescence within fungal cells (Fig. 3g and Extended Data Fig. 5b). Furthermore, we measured OA uptake by R. delemar cells by fluorescence labelling with Nile Red lipid dye28. Notably, we detected a substantial degree of OA uptake by R. delemar spores within a few hours in culture, which was profoundly reduced after OA oxidation (Extended Data Fig. 5c,d).
Given the well-established antioxidant properties of albumin21, we reasoned that the binding of FFAs to albumin can protect them from oxidation. We therefore performed chemical oxidation of OA dissolved in ethanol or after complexation with albumin and assessed the degree of oxidation. We found that albumin significantly protected OA from oxidation (Fig. 3h) and retained its antifungal properties against Mucorales (Fig. 3i).
Albumin glycation induced by DM, a major predisposing factor for mucormycosis1, results in the dissociation of FFAs from their binding sites and increased oxidation29 We therefore performed in vitro glycation of albumin (BSA) and assessed its ability to inhibit Mucorales. BSA glycation led to near-complete dissociation of bound FFA (Fig. 3j and Extended Data Fig. 5e–g), resulting in significant loss of albumin antifungal activity (Fig. 3k). Collectively, these studies reveal that albumin protects FFAs from oxidation, which results in the loss of their antifungal activity against Mucorales. Moreover, we identified FFA oxidation as a prominent abnormality in mucormycosis sera.
FFAs target pathogenicity of Mucorales
We next investigated the physiological importance of albumin during in vivo infection with Mucorales after pulmonary infection of immunocompetent mice with swollen fungal spores13. Specifically, we allowed dormant spores of R. delemar to grow in culture medium with or without physiological concentrations (4.5 g dl−1) of albumin for around 4 h and performed intratracheal (i.t.) infection of the mice (Fig. 4a). We found that albumin pre-exposure rendered R. delemar spores almost completely avirulent in vivo (Fig. 4b). Notably, pre-exposure to albumin did not inhibit the in vivo germination of Mucorales during the early stages of infection in the lungs (Extended Data Fig. 6a). Instead, albumin pre-exposure completely abrogated massive tissue necrosis and tissue invasion induced by germinating fungal spores, as evidenced by staining for active caspase-3 in sections of the lung (Fig. 4c), and lung fungal burden determined by quantitative PCR (qPCR; Fig. 4d) and histopathology (Fig. 4e). These findings suggest a predominant effect of albumin in attenuating the virulence of Mucorales.
a, Schematic of i.t. instillation of dormant or swollen R. delemar spores. b, Survival of C57BL/6 mice infected i.t. with 2.5 × 106 dormant (n = 6), control-swollen (n = 18) or albumin-swollen (n = 12) spores as in a. Statistical significance was determined using the log-rank (Mantel–Cox) test. c, Representative lung histopathology on day 1 after infection with control- or albumin-swollen spores, stained for active caspase-3 (left), haematoxylin and eosin (H&E; middle) and Grocott’s methenamine silver (GMS; right). Scale bars, 100 µm. d, The fungal burden determined by qPCR, expressed as spores per g of lung tissue. n = 7 mice. Statistical analysis was performed using a two-sided unpaired t-test; *P = 0.0331. eq., equivalent. e, Quantification of GMS-stained R. delemar hyphae in lung tissue. n = 8 control-swollen and n = 7 albumin-swollen mice. Statistical analysis was performed using two-sided Mann–Whitney U-tests; ***P = 0.0006. HPF, high-power field. f, Workflow of RNA-seq analysis of dormant and swollen spores cultured as in a for 3 h and 6 h. g, Differentially expressed genes (DEGs) in albumin-swollen versus control-swollen spores. The red and blue dots indicate upregulated and downregulated genes, respectively. FC, fold change. h, Enriched GOs among DEGs identified by gene set enrichment analysis after culture of spores in albumin versus medium for 3 h. NES, normalized enrichment score. i, Representative fluorescence images of R. delemar spores cultured for 5 h in RPMI medium alone or supplemented with BSA (4.5 g dl−1) or caprylic acid (2 mM). Protein synthesis was assessed by OP-puro incorporation and 5-FAM-azide staining. Scale bars, 20 µm. j, Quantification of 5-FAM-azide MFI. n = 943–1,191 spores per group, three independent experiments. Statistical analysis was performed using one-way ANOVA with Tukey’s test. k, Differential mRNA expression of virulence-related genes after incubation in medium with or without albumin for 3 or 6 h. n = 3 biologically independent samples. Data show log2-transformed normalized counts per million (CPM) values. l, Confocal images of mucoricin expression in CFW-labelled control- and albumin-swollen R. delemar spores after 3 h (left). n = 107 control-swollen and n = 70 albumin-swollen spores, three experiments. Scale bars, 10 µm. Right, quantification of the mucoricin fluorescence intensity. Statistical analysis was performed using two-sided Mann–Whitney U-tests. Data are mean ± s.d. ****P < 0.0001 (j,l). The diagrams in a and f were created using BioRender.
We next performed RNA-sequencing (RNA-seq) analysis during the in vitro growth of R. delemar to examine the molecular mechanism of action of albumin on the pathogenicity program of the fungus (Fig. 4f). Volcano plot analysis revealed that albumin treatment differentially modulated the expression of a large number of genes within 3 h (Fig. 4g). Enrichment analysis (Gene Ontology (GO) (Fig. 4h) and KEGG (Extended Data Fig. 6b,c and Supplementary Table 3)) showed broad downregulation of protein-synthesis pathways, accompanied by induction of oxidative stress responses and lipid metabolism. To test whether albumin-bound FFAs directly impact fungal protein synthesis, we measured global translation in R. delemar using an assay based on incorporation of O-propargyl-puromycin (OP-puro) into nascent polypeptide chains, detected by click-chemistry with a fluorescent azide30. Both BSA and purified caprylic acid almost completely inhibited OP-puro incorporation in germinating R. delemar spores within 2 h of treatment (Fig. 4i,j). In view of the pronounced inhibitory effect of albumin-bound FFAs on protein synthesis, we analysed the transcriptional response of all characterized virulence factors of Mucorales, including CotH invasins1,10, the mycotoxin mucoricin9 and genes regulating the iron assimilation program of the fungus1. Notably, we identified mucoricin and CotH3 invasin10 as the genes most significantly downregulated by albumin at 6 h of R. delemar growth (Fig. 4k). We also found that pre-exposure of Mucorales to the albumin culture filtrate almost completely abolished the expression of mucoricin and CotH3 on the surface of swollen spores, as evidenced by immunostaining (Fig. 4l and Extended Data Fig. 6d); similarly, exposure of R. delemar spores to purified FFAs blocked mucoricin protein expression (Extended Data Fig. 6e). Finally, the silencing of mucoricin in R. delemar (mucoricin RNA interference (RNAi) strain9) resulted in a significant decrease in the pathogenicity of swollen spores after pulmonary infection of immunocompetent mice (Extended Data Fig. 6f). Overall, these results reveal that albumin-bound FFAs inhibit protein synthesis to modulate pathogenicity during in vivo fungal growth.
Albumin deficiency promotes mucormycosis
We next used a humanized model of albumin knockout (KO) transgenic mice to genetically validate the role of albumin in host defence against Mucorales31. These transgenic mice are double KO for albumin and the neonatal Fc receptor (FcRn), which regulates the recycling of albumin, and transgenic for human FcRn. The expression of human FcRn results in a prolonged half-life of human albumin following systemic administration31. We found that albumin KO (Alb−/−) mice were highly susceptible to disseminated and pulmonary R. delemar infection (Fig. 5a). However, Alb−/− mice displayed comparable susceptibility to control Alb+/+ mice after pulmonary infection with A. fumigatus in the neutropenic model of invasive aspergillosis (Fig. 5b and Extended Data Fig. 7a) or disseminated bloodstream infection with Candida albicans (Fig. 5c). Importantly, prophylactic or pre-emptive therapeutic administration of purified, FFA-free human albumin fully restored the resistance of Alb−/− mice to pulmonary mucormycosis (Fig. 5d). Histopathological sections of lungs from neutropenic Alb−/− versus Alb+/+ mice after pulmonary infection with R. delemar revealed invasive fungal growth (Fig. 5e). Alb−/− exhibited a trend toward higher pulmonary fungal burden (Extended Data Fig. 7b) and significantly higher expression of mucoricin on germinating hyphae, as evidenced by immunostaining of the tissue (Fig. 5f). Notably, albumin significantly attenuated mucoricin-induced cytotoxicity in epithelial cells ex vivo (Extended Data Fig. 7c). Sera (Fig. 5g) and bronchoalveolar fluid (Fig. 5h) obtained from Alb−/− mice exhibited significant loss of inhibitory activity against R. delemar growth, and serum lipids from these mice were less inhibitory towards Mucorales (Fig. 5i). Lipidomic analyses of serum FFAs revealed a significantly lower amount of non-oxidized FFAs in Alb−/− mice compared with in Alb+/+ mice, particularly at 72 h of infection (Extended Data Fig. 7d,e and Supplementary Table 3), consistent with previous reports32. Alb−/− sera also contained a significantly higher proportion of oxidized FFAs, although their absolute levels were comparable to those in Alb+/+ mice (Extended Data Fig. 7d,f and Supplementary Table 3). Consistent with these findings, albumin administration restored the inhibitory activity of Alb−/− serum (Fig. 5j) and led to a significant increase in a subset of non-oxidized serum FFAs (Fig. 5k). These results provide definitive evidence of the master regulatory role of albumin in host defence against Mucorales.
a, Survival of Alb+/+ and Alb−/− mice that were infected with R. delemar intravenously (i.v.) (left; n = 24–27; 1 × 105 spores) or i.t (right; n = 24; 1 × 106 spores). b, Survival of neutropenic Alb+/+ and Alb−/− mice infected i.t. with 1 × 106 A. fumigatus spores. n = 11. c, Survival of Alb+/+ (n = 13) and Alb−/− (n = 11) mice infected i.v. with 7.5 × 105 C. albicans blastoconidia. d, Outline of administration of FFA-free HSA (left). Right, survival of neutropenic Alb−/− mice with or without FFA-free HSA administered before infection (n = 10–13) or after 6 h of i.t. infection with 1 × 106 R. delemar spores (n = 10–11). CA, cortisone acetate; Ctx, cyclophosphamide. For a–d, statistical analysis was performed using the two-sided log-rank (Mantel–Cox) test. e, Representative lung histopathology on day 3 after infection with R. delemar in neutropenic Alb+/+ and Alb−/− mice. Scale bars, 100 µm. f, Representative fluorescence images on mucoricin expression in lung sections from the experiment in e (left). Scale bars, 10 µm. Right, quantification of mucoricin fluorescence. n = 220 spores per group, three experiments. ****P < 0.0001. g, Inhibitory effect of serum from Alb+/+ and Alb−/− mice on R. delemar growth (n = 8). ***P = 0.0006. h, The inhibitory effect of bronchoalveolar fluid (BALF) from Alb+/+ and Alb−/− mice on R. delemar growth (left). n = 8. ***P = 0.0007. Right, representative images of spores cultured 5 h in BALF. Scale bar, 100 μm. i, The inhibitory effect of serum lipids isolated from Alb+/+ and Alb−/− mice on R. delemar growth. n = 7. Statistical analysis was performed using one-way ANOVA with Tukey’s test; ***P = 0.0005. j, The inhibitory effect of serum from Alb−/− mice with or without FFA-free HSA supplemented as in d. n = 4. *P = 0.0286. k, Serum FFAs from Alb−/− mice with or without FFA-free HSA supplemented as in d. n = 8. Statistical analysis was performed using Wilcoxon rank-sum tests. Data are mean ± s.d. For f, g, h and j, statistical analysis was performed using two-sided Mann–Whitney U-tests. The diagram in d was created using BioRender.
Discussion
Our initial studies revealed an interesting association between low albumin levels and the development of mucormycosis in high-risk patients with haematological malignancies. Furthermore, severe hypoalbuminaemia independently predicted poor outcome of mucormycosis across patient cohorts with distinct risk factors, on three different continents. These compelling findings are consistent with previous reports on the increased susceptibility of patients with disease-related malnutrition to mucormycosis1,2,5,17,33,34. Given the lack of host biomarkers for prognostication of mucormycosis, albumin levels should be further investigated as a readily available, inexpensive and universal biomarker to stratify patients at risk and predict infection outcome.
Susceptibility studies after HSA depletion and purification provided unambiguous evidence of the specialized inhibitory activity of albumin against Mucorales. Functional analysis of lipid-containing fractions and lipidomic profiling of the albumin culture filtrate revealed that the antifungal activity of albumin is exclusively mediated by the release of bound physiological FFAs. The antimicrobial properties of naturally occurring FFAs have long been recognized23,35,36, particularly at epithelial surfaces where they reach concentrations that are orders of magnitude greater than those in serum and other extracellular fluids36,37,38,39. However, a host-protective role of albumin-bound FFAs has not been previously demonstrated. By contrast, bacterial pathogens exploit albumin as a source of FFAs to support growth40, and albumin can reprogram C. albicans metabolism independently of FFAs to promote pathogenicity ex vivo41. Consistent with these observations, we found that albumin lacks antimicrobial activity against human pathogens other than Mucorales.
Further experiments demonstrated that albumin has an essential role in dissolving, shuttling and protecting bound FFAs from oxidation. Notably, we also found that oxidized FFAs are not internalized by fungal cells and lose their antifungal properties. Whether the systemic antioxidant properties of albumin42 extend to other serum lipids not directly bound to it remains to be determined. Collectively, these unique properties of albumin promote FFA accumulation within Mucorales spores and result in the inhibition of fungal growth. Moreover, the extensive FFA oxidation in the sera of patients with mucormycosis is associated with the loss of antifungal activity, highlighting the physiological importance of albumin in host defence against Mucorales. Finally, the glycation of albumin, a modification associated with reduced FFA-binding capacity in patients with poorly controlled DM29, abolished its inhibitory activity against Mucorales due to dissociation of bound FFAs, providing mechanistic insights into the heightened mucormycosis risk in this patient population.
Notably, pre-exposure to albumin almost completely abrogated virulence of germinating R. delemar spores during in vivo infection of immunocompetent mice, as a result of inhibition of protein synthesis and the transcriptional downregulation of key virulence factors, especially CotH3 invasin10 and the potent mycotoxin mucoricin9. Future work will elucidate the precise molecular mechanism by which FFAs selectively inhibit protein synthesis in Mucorales.
In concordance with human mucormycosis cohorts, studies of disseminated and pulmonary infection by Mucorales and other major human fungal pathogens in albumin-KO humanized mice provided definitive evidence for a non-redundant, specialized role of albumin in immunity against Mucorales. The reversal of susceptibility in albumin-KO mice and the restoration of serum antifungal activity after albumin supplementation, provide strong rationale for correcting hypoalbuminaemia as (1) a low-cost preventive strategy in high-risk patients and (2) an adjunct therapeutic modality in established mucormycosis. The therapeutic value albumin in mucormycosis warrants evaluation in prospective clinical studies.
Our findings support a pathogenetic model of mucormycosis (Extended Data Fig. 7g) that explains the unique epidemiological features of this disease. Metabolic abnormalities that result in severe hypoalbuminaemia or chemical modifications that impair binding capacity of the protein for FFAs (such as glycation) lead to increased FFA oxidation and loss of their antifungal activity. Systemic oxidative stress induced by hyperinflammation, comorbidities or iron overload may further enhance FFA oxidation and susceptibility to mucormycosis1. Hypoalbuminaemia, a biomarker of poor outcome in multiple infectious43,44 and non-infectious45,46 diseases, probably reflects the broader immunometabolic functions of albumin and its bound FFAs. Defining the mechanistic basis and therapeutic potential of these interactions represents an important avenue for future research.
Finally, our work provides interesting cues on the mechanisms of selection pressure that account for the enigmatic evolution of albumin in vertebrates47, by linking its unique lipid-binding properties with the modulation of the pathogenicity of ubiquitous, toxin-producing phytopathogenic fungi. Collectively, these findings have implications for the immunopathogenesis and therapeutics of an emerging human fungal disease with limited treatment options.
Methods
Microorganisms and culture conditions
R. delemar (99-880)9, R. arrhizus (557969)48 and Cunninghamella bertholletiae (506313)48 have been previously described. Mucor circinelloides (JMRC:NRZ:0774), Rhizomucor pusillus (JMRC:NRZ:0496), Rhizopus microsporus (JMRC:NRZ:0680), A. fumigatus (ATCC, 46645), Aspergillus flavus (JMRC:NRZ:0756), Aspergillus terreus (JMRC:NRZ:0442), C. albicans (JMRC:NRZ:1000), C. glabrata (JMRC:NRZ:1006), C. albicans SC5314 and Fusarium proliferatum (JMRC:NRZ:0657) were obtained from the Leibniz Institute for Natural Product Research and Infection Biology, Hans Knöll Institute. All Mucorales isolates were cultured on potato dextrose agar (Becton Dickinson) plates for 7 days at 37 °C. R. delemar M16 is a previously described pyrF-null mutant derived from R. delemar 99-880 that is unable to synthetize uracil49. R. delemar transformed with RNAi targeting mucoricin expression and R. delemar transformed with empty plasmid (control) were derived from strain M16, as previously described9. For the experiments involving these RNAi strains, a synthetic medium containing yeast nitrogen base (YNB, BD) supplemented with a complete supplemental mixture without uracil (CSM−URA, MP Biochemicals) (that is, YNB + CSM−URA) was used.
All of the bacterial isolates used (Escherichia coli, Pseudomonas aeruginosa, Klebsiella pneumoniae and Staphylococcus aureus) were clinical isolates obtained from the University Hospital of Heraklion, Crete. All bacteria were streaked onto plates containing LB agar plates from a freshly prepared frozen glycerol stock. After growth on LB agar plates, single colonies were used to inoculate overnight cultures of LB (50 ml at 37 °C). The next day, a volume of 1 ml of each of these cultures was diluted 1:10 to a final volume of 10 ml in LB medium and incubated in conical flasks for 2 h, at 37 °C, with constant shaking, to reach mid-log phase of growth. A volume of 200 μl was taken from each culture and the optical density was measured at 600 nm (OD600). The desired volume from each culture was used and centrifuged at 2,000 rpm for 2–3 min. Bacteria pellets were washed three times with LB medium. Bacteria pellets were diluted in RPMI medium or RPMI plus 4.5 g dl−1 of albumin to the desired OD600 of 0.2. A volume of 200 μl from each test condition was transferred to a flat-bottom 96-well plate. The plates were incubated at 37 °C, with constant shaking. At regular time intervals (45 min), 10 μl from each culture condition was diluted in RPMI medium to a final volume of 100 μl and the OD600 was measured spectrophotometrically. The mean value of triplicate measurements of bacterial growth in regular medium at t = 340 min was defined as 100% growth.
For filamentous fungi, the effect of albumin on fungal growth was assessed by counting spore germination rates in RPMI medium compared with RPMI supplemented with 4.5 g dl−1 albumin at 6 h using light microscopy. At least 300 spores were counted per condition in biological triplicates. For the effect of human IgG (i.v. human IgG, Gaminex, 100 mg ml−1, Grifols) fungal spores were incubated in medium containing increasing concentrations of human IgG (from 1.5 g dl−1 to 4.5 g dl−1). At 6 h, spores were assessed by light microscopy and photos were taken.
For Candida isolates, the impact of albumin on growth was evaluated spectrophotometrically. C. albicans strain SC5314 and Candida glabrata (JMRC:NRZ:1006) were streaked from freshly prepared glycerol stocks onto Sabouraud dextrose agar plates and incubated overnight at 37 °C. Single colonies were used to inoculate Sabouraud dextrose broth (50 ml) and liquid cultures were grown overnight at 37 °C with agitation at 150 rpm. The OD600 of the overnight cultures was determined, and aliquots were diluted with phosphate-buffered saline (PBS) to an OD600 of 0.1. Cells were collected by centrifugation at 2,000 rpm for 10 min, washed three times with PBS, counted and resuspended in either RPMI medium alone or RPMI supplemented with 4.5 g dl−1 albumin to an OD600 of 0.1. For each condition, 200 μl of the cell suspension was transferred into wells of a flat-bottom 96-well plate and incubated at 37 °C with continuous shaking. After 8 h, 10 μl from each well was diluted in RPMI to a final volume of 100 μl, and the OD600 was measured spectrophotometrically. Growth in standard RPMI medium was assessed in triplicate, and the mean OD600 value at 8 h was defined as 100% growth for normalization.
Human albumin depletion and purification
Albumin was depleted from human serum using an Albumin Depletion Kit (Abcam) according to the manufacturer’s instructions with a modification in the rehydration step of Cibacron Blue 3G-A beads, which was performed using albumin-free serum filtrate (generated through serum filtration using Amicon 50 kDa molecular weight cut-off (MWCO) ultracentrifugal filters; Merck).
For HSA purification, Blue Sepharose 6 Fast Flow (GE Healthcare) was first rehydrated with an albumin-free serum filtrate, incubated with fresh human serum (3 ml) at 4 °C overnight with rotation and then packed back in a column. The first volume (3 ml) of the flow-through contained albumin-depleted human serum. The column was subsequently washed with 7 ml of wash buffer (20 mM Na2HPO4, 20 mM NaH2PO4, pH 8). Albumin was isolated from the column in six consecutive elutions using 7 ml of elution buffer (2 M NaCl, 20 mM Na2HPO4, pH 8) each time. Elution fractions 2–6 were pooled and further processed for in vitro experiments.
Owing to the increased amount of NaCl in the eluted fractions, dialysis was performed using a CelluSep T2 membrane (Orange Scientific, Cellusep T2 Tubings, 6,000–8,000 MWCO), which was embedded in the appropriate buffer (20 mM Na2HPO4, 20 mM NaH2PO4, pH 8) for 4 h at 4 °C to achieve a physiologically relevant NaCl concentration (150 mM). Elutions were then condensed using Amicon 3 kDa MWCO ultracentrifugal filters (Merck), to a final volume of 2 ml and filter-sterilized through 0.22 μm Spin-X centrifuge tube filters (Costar). The flow-through generated during the condensation procedure was pooled, measured for any remnants of albumin (no traces of albumin were found) and also filter-sterilized using the same 0.22-μm filters.
Chemical modifications of albumin
Highly oxidized albumin was prepared by the incorporation of cysteine into reduced albumin50. BSA was treated with a 50-fold molar excess of L-cysteine/cystine by mixing 80 ml of 0.06 mM BSA with 72 ml of 3 mM cysteine and 8 ml of 3 mM cystine. All of the solutions were diluted in 0.1 M sodium carbonate and hydrogen carbonate buffer (pH 10) and, after incubation at 37 °C for 48 h, the resulting mixture was lyophilized. Next, the residue was dissolved in PBS and purified from low-molecular-mass components (excess cysteine/cystine) by filtration through an Ultrafree-3000 Da membrane (Millipore) at 4 °C. Purified, highly oxidized albumin was lyophilized and stored at room temperature. Glycosylated albumin was prepared by diluting albumin in PBS containing 5 mM glucose51. The solution was incubated for 72 h under an atmosphere of 95% O2, and 5% CO2 to maintain the pH at 7.3–7.4. The mixture was then lyophilized, followed by solubilization in PBS and purification through ultrafiltration with an Ultrafree-3000 Da membrane. Purified, glycosylated albumin was lyophilized and stored at room temperature. The control solutions were prepared in PBS and subjected to the same lyophilization and filtration processes to increase the reliability of the results. The dried, equal-weight samples were derivatized with 20 μl of 20 mg ml−1 methoxy amine/pyridine and 50 µl of hexane. The mixture was vortexed and kept at 37 °C for 1.5 h, with vortexing every 30 min. Then, 90 μl (N-trimethylsilyl-N-methyl trifluoro-acetamide and trimethyl-chlorosilane (MSTFA + 1% TMS) were added and the samples were vortexed for 30 s and incubated at 37 °C for 1 h. The obtained solution was filtered through 0.45-μm syringe filters (nylon syringe filter, Membrane Solutions) and subjected to GC–MS/MS in multiple reaction monitoring (MRM) mode analysis according to previously published methods52,53. The obtained metabolites were imported into MetaboAnalyst (v.5.0; https://www.metaboanalyst.ca/home.xhtml). Hierarchical cluster analysis and partial least squares-discriminant analysis were then performed according to previous methods54.
The removal of FFAs from albumin (BSA and HSA isolated from donors) was performed with the use of activated charcoal, as previously described with some modifications25. Next, activated charcoal and albumin were mixed at a ratio of 1:2 for 1 h at 4 °C. The excess of activated charcoal was removed by ultracentrifugation at 20,200g for 30 min at 4 °C.
Loading of albumin with FFAs was performed as previously described55,56 with some modifications. In brief, 4.5 g dl−1 charcoal-treated albumin (diluted in RPMI medium) and 80 mM FFAs (diluted in ethanol) were heated at 55 °C for 30 min and 5 min, respectively. Then, FFAs and albumin were gently mixed at a ratio of 8:1. The FFA–albumin mixture was thoroughly mixed by vortexing and incubated at 37 °C for 1 h. Effective conjugation of FFAs to albumin resulted in a clear solution.
Extraction and quantification of caprylic acid in glycosylated and non-glycosylated BSA
Quantification of caprylic acid in glycosylated and non-glycosylated BSA was conducted by extracting caprylic acid using chloroform/methanol solvent (2:1, v/v)57. This was followed by centrifugation at 3,500 rpm to separate the organic layer, which was filtered through anhydrous sodium sulphate and then dried using a rotary evaporator (Büchi). The dried residue was subsequently derivatized and analysed using GC–MS/MS in MRM mode against standard caprylic acid (m/z 201.10, 117.10 and 75.00).
Fluorescence displacement assay of caprylic acid binding with glycosylated and non-glycosylated BSA
The fatty-acid binding affinity of glycosylated and non-glycosylated BSA was assessed using the 1-anilino-8-naphthalene sulfonic acid (ANS) fluorescence displacement assay according to a previously established protocol58. In brief, BSA solutions (2.5 μg in 100 μl PBS) were added into black 96-well flat-bottom microplates (Nunc Microwell Plates), followed by the addition of 50 μl ANS in PBS to a final concentration of (0, 1.25, 2.5, 5 and 10 μM) with incubation at room temperature in dark for 15 min. For the displacement assay, caprylic acid was added at increasing concentrations (0–10 μM) to the ANS–BSA complexes. A control without caprylic acid was included to assess the initial fluorescence. The mixtures were incubated at room temperature in the dark for an additional 15 min. Measurements were performed using a microplate reader (Synergy H1, Biotek) with excitation and emission wavelengths set at 360 nm and 460 nm, respectively. Fluorescence quenching, indicative of ANS displacement by caprylic acid, was recorded as the percentage change in fluorescence intensity (ΔF%) using the formula: %ΔF quenching = [(F0 − F1)/F0] × 100, where F0 is the fluorescence in the absence of caprylic acid and F1 is the fluorescence at each tested concentration of caprylic acid.
In vitro assessment of the inhibitory activity of albumin and human serum
Fungal conidia (spores) were collected by gentle shaking in the presence of sterile 0.1% Tween-20 in PBS, washed twice with PBS, filtered through a 40-μm-pore-size cell strainer (Falcon) to separate conidia from contaminating mycelium, counted using a haemocytometer and suspended at a concentration of 108 spores per ml. To achieve synchronized swelling of Rhizopus conidia, 1 × 106 per ml dormant conidia were incubated for 4–6 h at 28 °C in a six-well plate in RPMI-MOPS (pH 7.0) supplemented with 2% glucose with or without 4.5 g dl−1 BSA or other sources of albumin. To study the minimal growth requirements of R. delemar, a minimal-growth-requirements medium was used59 based on HBSS supplemented with 0.05% (w/v) MgSO4·7H2O, 0.1% (w/v) glucose and 0.1% (w/v) NH4Cl (minimal medium). For fluorescence labelling, the spores were stained with 20 μg ml−1 Fluorescent Brightener 28 (Calcofluor White; CFW, Sigma-Aldrich) in 0.1 M NaHCO3 pH 8.3 at room temperature in the dark for 1 h, with constant rotation.
In another set of experiments, R. delemar spores were incubated at 37 °C in RPMI-MOPS without phenol red adjusted to pH 7, for 2 h to start swelling. Then, 3.6 g dl−1 BSA and 900 μg ml−1 FITC-albumin (Sigma-Aldrich) were added to the medium and the spores were cultured in a 96-well μ-Plate (Ibidi) at 37 °C. After 5 h of culture, R. delemar live spores were collected, washed twice with PBS and then stained with 100 μg ml−1 CFW in 0.1 M NaHCO3 pH 8.3 at room temperature in the dark for 1 h, with constant rotation. Fungal cells were then imaged with a spinning-disk confocal microscope (Dragonfly 200, Andor).
To assess the inhibitory activity of human serum, 1 × 104 fungal spores were incubated in flat-bottom 96-well plates containing 100 μl of serum from healthy individuals or patients at 37 °C and 5% CO2 for 5–6 h. At least three different fields of each well (a total of >100 germinating spores) were imaged under an inverted microscope (Olympus) and the length of the germinating conidia was measured using ImageJ60. In certain experiments, time-lapse videos of fungal growth were acquired using an Operetta high screening content system (Perkin Elmer) at room temperature with 5% CO2. Photographs of each well were automatically taken approximately every 30 min for 18 timepoints and analysed using Harmony v.4.1 software (Perkin Elmer).
The inhibition of fungal growth was additionally quantitated via an XTT metabolic activity assay (Biotium). A solution of XTT tetrazolium salt (0.25 mg ml−1) and menadione (25 μM) was freshly prepared in PBS and added to the cell culture. Fungal spores were further incubated at 37 °C for another 1 h (ref. 61). Fungal-spore-free culture supernatants were collected, and their absorbances at 450 nm (OD450) and 620 nm (OD620) were measured using a microplate photometer (Multiscan, Thermo Fisher Scientific). Growth inhibition was calculated according to the following formula:
In vitro studies on the antimicrobial activity of FFAs
FFA stock solutions were initially dissolved in absolute ethanol to a final concentration of 80–1,000 mM and incubated at 95 °C for 5 min. 2× MOPS-RPMI medium was also heated at 95 °C for 30 min and serial dilutions of FFAs were made by slowly adding the desired volume of preheated FFAs into the medium. The final concentration of ethanol used under culture conditions was 5% to ensure optimal dilution of FFAs; we confirmed that 5% ethanol had no significant inhibitory effect on Mucorales growth. Fungal spores were incubated in FFA-containing MOPS-RPMI medium at 37 °C for 16–18 h and fungal germination was measured as previously described13.
Lipid extraction
Lipids were extracted using a CHCl3:CH3OH (2:1) solution according to a previously published method62, dried with anhydrous sodium sulfate, subjected to residual evaporation via a rotatory evaporator (Buchi) under an inert stream of nitrogen and then stored at −20 °C (ref. 63). The residue obtained was equally divided into two parts; one for the oxidation reaction and the other left under inert conditions to avoid auto-oxidation.
Microwave-assisted oxidation of FFAs
FFAs and BSA-bound FFAs were subjected to oxidation through an irradiation microwave reactor according to modified previously published methods64. The samples were dissolved in 0.2 mM H2O2 in methanol to a final volume of 10 ml. The oxidization reaction was performed in a sealed vial on a synergy microwave synthesizer (CEM) for 20 min at an operating temperature of 100 °C and a pressure of 200 psi. The reaction mixture was extracted with CHCl3, dried with anhydrous sodium sulfate, subjected to residual evaporation through a rotatory evaporator (Buchi) and stored at −20 °C.
Open-air-assisted oxidation of FFAs
FFAs (OA), human or mouse plasma lipid extracts were diluted in absolute ethanol and exposed to open air inside a laminar flow hood as previously described65. After 4 days, ethanol was evaporated, and the precipitates of the oxidized FFAs were rediluted to their initial volume in 0.01 M Tris-HCl pH 7.5 (human and mouse plasma lipids) or in absolute ethanol to a final concentration of 80 mM (OA) and stored at −20 °C.
Cell damage assay for the assessment of toxin neutralizing activity of albumin
Human alveolar epithelial cells (A549 cells) were obtained from a 58-year-old male Caucasian patient with carcinoma and procured from American Type Culture Collection (ATCC). The cells were propagated in F-12K Medium developed for lung A549 epithelial cells. The damage of A549 epithelial cells was quantified using a 51Cr-release assay9,66. In brief, confluent cells grown in 24-well tissue culture plates were incubated with 1 μCi per well Na251CrO4 (ICN) in F12K-medium for 16 h. On the day of the experiment, the medium was aspirated, and cells were washed twice with pre-warmed Hanks’ balanced salt solution (HBSS, ScienCell). Cells treated with mucoricin with or without the addition of albumin (free of FFAs BSA at a final concentration of 4.5 g dl−1) were suspended in 1 ml of F12K-medium supplemented with glutamine and incubated at 37 °C in a 5% CO2 incubator. Spontaneous 51Cr release was determined by incubating the untreated cells in the same volume of the culture medium supplemented with glutamine. At different timepoints, and after data were corrected for variations in the amount of tracer incorporated in each well, the percentage of specific cell release of 51Cr was calculated as follows: [(experimental release) − (spontaneous release)]/[1 − (spontaneous release)]66. Each experimental condition was tested at least in triplicate, and the experiment was repeated at least twice.
Ethics statement on human clinical studies
All research involving human participants was conducted in accordance with relevant ethical regulations and approved by the ethics committees of the respective participating institutions. All patients or their legal representative provided written informed consent for the collection of clinical samples analysed in this study, in accordance with the Declaration of Helsinki.
The medical records of contemporaneous patients who had been admitted at the Leukemia Department of the MD Anderson Cancer Center were retrospectively reviewed. Standardized EORTC/MSG criteria were applied for diagnosis of pulmonary mucormycosis and pulmonary aspergillosis67. Clinical information of matched control patients for the underlying disease who developed bacterial (Legionella) pneumonia were also reviewed. Albumin levels were retrieved from the medical records of all of the patients on the day of hospital admission. Information on demographics, underlying disease and risk factors for mucormycosis were collected. The study was approved by the Institutional Review Board (IRB) of The University of Texas MD Anderson Cancer Center (MDACC; protocol PA14-0802).
The medical records of consecutive patients admitted to the Department of Pulmonary Medicine, Department of Medical Microbiology, Institute of Medical Education and Research (PGIMER), Sector-12, Chandigarh, 160012, India over the past 8 years (2016–2023) with a diagnosis of pulmonary mucormycosis were retrospectively evaluated. The study was approved by the Institutional Ethics Committee (IEC; Intramural; INT) of the PGIMER (IEC/INT, 2023/study-1564) and registered with the Biological Research Regulatory Approval Portal (BIORRAP, POS36452025). All procedures were conducted in accordance with institutional and national guidelines and regulations.
Serum samples were prospectively collected from healthy individuals, patients with cirrhosis and patients with haematological malignancies at the University Hospital of Heraklion, Crete, and the Therapeutics Clinic of the General Hospital of Athens “Alexandra”. Demographics, clinical characteristics, serum albumin levels, total FFAs and oxidized FFA levels were measured in the sera of healthy individuals and patients. Approval for the collection of clinical information and blood samples from all individuals mentioned above was obtained from the Ethics Committee of the University of Heraklion, Crete, Greece (5159/2014, 10925/201 and 13-04-22/7970) and Therapeutics Clinic (122/18-2-2021).
For lipidomic and functional studies, serum samples from patients with mucormycosis or invasive pulmonary aspergillosis, and from matched control patients were obtained from patients with haematological malignancy admitted to the University Hospitals of Leuven, Belgium (IRB, S61797).
Vitamin quantification in culture medium using LC–MS
Samples were first filtered through 0.22-μm nylon syringe filters before undergoing LC–MS/MS analysis. The analysis was performed via an Elute UHPLC system linked to a Q-TOF mass spectrometer. Chromatography separation was performed on a Hamilton Intensity Solo C18 column, maintained at 35 °C. The sample mixture was 10 μl in a solvent system composed of 0.1% formic acid in deionized water (solvent A) and acetonitrile (solvent B). The elution gradient progressed from 1% to 99% solvent B over 20 min, with flow rates adjusted between 0.25 ml min−1 and 0.35 ml min−1. Vitamin detection followed established methods by employing standards of cyanocobalamin (B12), folic acid (B9), riboflavin (B2), thiamine (B1), biotin (B7), Ca-panthenoate (B5), pyridoxine (B6), nicotinamide (B3), choline chloride and PABA, all sourced from Sigma-Aldrich. Stock solutions of 5 mM for vitamins B1, B3, B5, B6, B7, B9 and B12, and choline chloride were prepared in water, with riboflavin (B2) in DMSO. Internal standard solutions were made at 10 mM, and working solutions comprised 200 mM of all vitamins in water and 100 mM of internal standards. Calibration curves were prepared using 0.1% formic acid in water, spanning six serial dilutions from 0 to 100 mM, each including 2.5 mM internal standards. Samples and calibration solutions underwent identical processing involving liquid-liquid extraction and drying before analysis, ensuring precise quantification of vitamins with detection limits in the low µg l−1 range. Each sample was injected three times.
Analysis of trace elements in culture medium using ICP-MS
Inductive coupled plasma MS (ICP-MS-9800 Series) was used to detect trace metals in the samples. The samples were collected and digested with hydrochloric acid, to ensure complete dissolution. Microwave-assisted digestion was used for rapid and efficient sample preparation. The digested samples were then diluted with deionized water such that the concentration was within the measurable range of the ICP-MS instrument. Calibration standards with known concentrations of Na, K, Ca, Mg, Cu and Fe were prepared, and isotope-labelled internal standards were used to correct for matrix effects and signal fluctuations. The prepared samples were injected into the ICP-MS instrument through a nebulizer, which converts the liquid sample into an aerosol. The ionized elements emit light at characteristic wavelengths, which are detected and quantified by a mass spectrometer. The concentrations of trace elements in the samples were calculated by comparing the sample signals to the calibration curve.
Analysis of amino acids in culture medium
The samples were centrifuged at 12,000 rpm for 10 min and the supernatants were analysed using the Waters Acquity ultra-performance liquid chromatography (UPLC) H-Class-Xevo TQD system equipped with ESI. HPLC was performed using the Acquity BEH C18 column (1.7 μm, 2.1 mm × 150 mm). The binary mobile phase consisted of solvent A (100% methanol) and solvent B (0.2% formic acid). The column gradient was as follows: 100% B for 10 min; 66.6% B for 0.5 min; returned to 100% B for 1 min; and maintained at 100% B for another 0.5 min. The flow rate was 0.2 ml min−1, the injection volume was 10 μl and the column effluent was monitored by MS. The MS analysis was performed in MRM mode in positive-ionization mode. The MS parameters were as follows: the dwell time was 0.02 s, and nitrogen was used as a desolvating gas at a flow rate of 600 l h−1. The ionization source conditions were as follows: desolvation temperature 350 °C; source temperature 150 °C; collision gas (argon) flow 0.1 ml min−1; and capillary voltage 3.0 kV. The parameters of mass analyser were set as follows: the LM1 and HM1 resolutions were 15 and 15, respectively, as ion energy 1; the LM2 and HM2 resolutions were 15 and 15, respectively, and the ion energy was 2. The amino acids in the samples were identified on the basis of the mass-to-charge ratio.
The quantification of amino acids was conducted using a standard mix calibration curve, with the UPLC–MS/MS system controlled by Lynx software (v.4.1, SCN 882). Data analysis was performed via the TargetLynx program. Each sample, which contained a mixture of amino acids at specific concentrations, was injected three times to ensure accuracy. The amino acid standard solution (AAS18, Sigma-Aldrich) included 2.5 μmol ml−1 of various amino acids such as L-alanine, L-arginine and L-valine, with L-cystine at 1.25 μmol ml−1. The samples were diluted in 0.1% formic acid in water for analysis. Using the MIDAS workflow, full scan linear ion trap MS/MS data confirmed the identity of the target analytes. MRM extracted ion chromatograms were used for all amino acids, especially at concentrations near their limits of detection, ensuring precise quantification and linearity. Limits of detection were accurately calculated for all amino acids, and their concentrations were measured.
Albumin ligand analysis and fractionation
BSA was added to minimal growth requirements medium to a final concentration of 4.5 g dl−1, left at room temperature for 0.5–1 h and then filtered through Amicon 3 kDa MWCO ultracentrifugal filters (Merck) to remove albumin. BSA after filtration (15 ml) was subjected to fractionation through C-18 solid-phase extraction. After conditioning with 6 ml of methanol and 6 ml of ultrapure water, 15 ml of the sample was applied, giving fraction 1 (F1). Subsequently, with gradual elution (6 ml of 5% methanol), fraction 2 (F2) was obtained. Finally, fractions 3 (F3) and 4 (F4) were obtained using 6 ml of 50% methanol and 6 ml of pure methanol, respectively. During the entire process, the flow rate was held constant at approximately 3 ml min−1. The solvents and reagents used were of LC-MS grade (Merck).
For fraction component identification through GC-MS analysis, the obtained fractions (1–4) were derivatized with methyl-chloroformate (Sigma-Aldrich). Specifically, 300 μl of every fraction was mixed with 80 μl of pyridine, 200 μl of methanol and 50 μl of methyl-chloroformate, vortexed for 30 s and incubated for 6 min at room temperature. After incubation, 1 ml of hexane was added to each reaction mixture and extracted via the liquid-liquid extraction technique. Finally, the hexane phase was subjected to GC-MS analysis.
Fractions F1–F4 were analysed through gas chromatography coupled with a single quadrupole MS (GC-MS) analyser. GC-MS analysis was performed with an Agilent gas chromatography instrument (model 8860) system coupled to a mass spectrometer (model 5970) using an electron ionization (EI) source.
The samples were qualitatively analysed and caprylic acid (or octanoic acid) was identified (NIST 14.0 EI spectral library) as the main component of F2. Furthermore, the caprylic acid structure was verified using a reference standard (Sigma-Aldrich), and its concentration was quantitatively determined in all four fractions, following the same derivatization protocol and a standard solution calibration curve. The entire analysis was conducted in duplicate. F2 was the richest in caprylic acid, with an average concentration of 194.1 μg ml−1, followed by F3, with a much lower concentration of 15.2 μg ml−1. In F1 and F4, only traces of caprylic acid were detected.
F2 was also analysed using HRMS without any derivatization steps to verify the structure of caprylic acid. Specifically, an UPLC system coupled with a HRMS analyser, especially a triple-TOF 5600+ (AB SCIEX) system, was used in negative ESI mode. LC analysis was performed on the ACQUITY H-Class UPLC system (Waters) equipped with a binary solvent manager and an FTN sample manager. A volume of 10 μl of F2 was injected into the system and separated by a linear gradient containing water and acetonitrile (mobile phases A and B, respectively, 5–100%), with the aqueous phase containing 0.1% formic acid. The solvents were of LC-MS grade and purchased from Merck. The analysis was performed with a Fortis Speedcore Biphenyl reversed-phase chromatography column (2.6 µm, 2.1 mm × 100 mm). The Triple-TOF platform was equipped with a DuoSpray ion source. The acquisition mode used was data dependent, covering a mass range of 80–500 m/z. Specifically, the tripleTOF parameters for acquisition were as follows: source temperature of 450 °C, source voltage of −4,500 V, exhaust gas pressure of 50 psi and curtain gas pressure of 35 psi. The caprylic acid structure was verified at m/z 143.1079 [M-H]- a, with a mass error of 1.1 ppm and degree of unsaturation (rings and double bond equivalents) of 1 (Extended Data Fig. 4).
FFA extraction from human plasma
FFAs were extracted from 100 μl of previously thawed plasma by the addition of 300 μl of cold methanol/ethanol (1:1) (Thermo Fisher Scientific). C17-Sphinganine was added as an internal standard. The samples were vortexed for 1 min, incubated on ice for 5 min and centrifuged at 16,100g for 20 min at 4 °C, after which 100 μl of the supernatant was transferred into LC–MS vials for analysis.
FFA extraction from the plasma of albumin-KO mice
FFAs were extracted from 100 μl of previously thawed plasma by the addition of 300 μl of cold methanol (Thermo Fisher Scientific). Next, 10 μl of 2 μM 9(S)-HODE-d4 solution (Cayman Chemical) was added as an internal standard. The samples were vortexed for 2 min and then centrifuged at 16,100g for 10 min at 4 °C, after which 100 μl of the supernatant was transferred into LC-MS vials for analysis.
Non-targeted analysis by LC–MS for FFAs
LC–MS analysis was performed on a UHPLC system 1290 Infinity II (Agilent Technologies) coupled with a 6546 QTOF MS detector in negative ESI mode. For separation, a volume of 1 μl was injected onto a Zorbax Rapid Resolution High Definition Extend-C18 column (Agilent Technologies, 2.1 × 50 mm, 1.8 μm) thermostated at 60 °C. The flow rate was 0.6 ml min−1, with a mobile phase composed of ultrapure Milli-Q water with 0.1% formic acid for A and acetonitrile with 0.1% formic acid for B. The chromatography gradient started from 5% B for the first min and increased to 80% B in 6.0 min, then to 100% by 11.0 min, and the starting conditions were returned to 1.0 min, allowing re-equilibration until 15.0 min. Data were collected in negative ESI ionization mode and operated in the range of m/z 100–600 and m/z 40–600 for MS/MS analysis using iterative Agilent mode. The nozzle voltage was set to 1,000 V, and the capillary voltage was −4,000 V with a scan rate of 1.2 scans per s. The drying gas was heated to 250 °C at a rate of 12 l min−1 and a pressure of 52.0 psi. Additional heating was applied using sheath-heated gas up to 370 °C with a flow rate of 11 l min−1 to improve ionization. For internal mass correction during data acquisition, one reference mass was infused continuously into the system throughout the whole analysis: m/z 112.9856 (proton-abstracted TFA anion). An external calibration with FA 18:0 was used for semi-quantification of fatty acid species. Calibration curve samples were prepared at different concentrations (1.0, 2.5, 5.0, 10.0, 13.0, 17.0 and 20.0 ppm), and 9(S)-HODE-d4 was included as an internal standard at the same concentration as the serum samples. The raw data collected by LC-MS were reprocessed with MassHunter Profinder software v.B.10.02. Calibration curves were built using normalized values of FA 18:0 by d4-9-HODE versus concentration in ppm. The results are expressed as the concentration in mM of the corresponding fatty acid in the serum. Detailed methods for identification of serum oxylipins and the standards used for the analysis of FFAs are provided in the Supplementary Methods.
RNA isolation from R. delemar cells
At the indicated timepoints of incubation in medium with or without albumin (0 h, 3 h and 6 h), the R. delemar cells were removed by scraping, centrifuged at 400g and lysed with 450 μl of RLT buffer + β-mercaptoethanol using the RNeasy Plant Mini Kit (Qiagen). Each sample was subsequently sonicated using a sonication probe on ice for 20 × 1 s (set 40). RNA was then isolated according to the manufacturer’s instructions.
RNA-seq data generation and analysis
RNA-seq libraries (strand-specific, paired-end) were generated from total RNA by using a TruSeq RNA sample prep kit (Illumina). In total, 150 nucleotides of sequence were determined from both ends of each cDNA fragment using the HiSeq 4000 platform (Illumina). Sequencing reads were aligned to the reference genome (R. delemar 99-880) using STAR aligner (v.2.7.10)68. The reads that mapped to genomic features were calculated using the featureCounts program (v.2.0.3)69. Statistical analysis of differential gene expression was performed using the DEseq2 R-based package70. A gene was considered differentially expressed if the absolute log fold change was greater than or equal to 1 and the false-discovery rate value for differential expression was below 0.05. The RNA-seq analysis was performed in biological triplicate. RNA-seq data were generated by Maryland Genomics at the Institute for Genome Sciences, University of Maryland School of Medicine.
RNA-seq enrichment analysis
The GO enrichment analysis performed using the gene set enrichment analysis (GSEA)71 method as previously implemented by the web-based application FungiFun3 (https://fungifun3.hki-jena.de/)72. The BlastKOALA automatic annotation server73 was used for the functional annotation and the assignment of the KEGG orthologies. The pathway enrichment analysis performed using the generally applicable gene-set enrichment method74 and functions of the gage R-based package. The genes of the selected KEGG pathways were identified using the KEGG PATHWAY database. R programming language was used for the visualization of the results. The heat maps were produced using the pheatmap package75. All of the P values were adjusted using the Benjamini–Hochberg method and adjusted P < 0.05 was considered to be statistically significant.
Animal studies
B6.Cg-Tg(FCGRT)32Dcr Albem12MvwFcgrttm1Dcr/MvwJ mice, referred to as Alb−/− mice throughout (obtained from The Jackson Laboratory), and C57BL/6 mice were maintained in grouped cages in a high-efficiency air-filtered, environmentally controlled, virus-free facility (24 °C, 50–60% relative humidity, 12 h–12 h light–dark cycle) and fed a standard chow diet and water ad libitum.
Ethics statement on animal studies
All experiments were approved by the local Ethics Committee of the University of Crete Medical School, the FORTH Ethics Committee, and the Directorate of Agricultural and Veterinary Policy of the Region of Crete, in accordance with national and European Union legislation (animal protocols 17/07/2017-147075 and 22/03/2023-90477). All efforts were made to minimize the number of animals used and their suffering.
In vivo studies of fungal infection
For virulence studies, 8–12-week-old, age- and sex-matched mice were infected either through i.v. inoculation or through i.t. instillation of 1–5 × 106 spores of R. delemar or A. fumigatus, and their survival was monitored for 14–25 days. After infection, two mice from each group were euthanized for inoculum verification. For histopathological evaluation, lungs were collected at different timepoints: 0 h, 6 h and 1 day after infection. In a separate experiment, age- and sex-matched C57BL/6 mice were infected as above and then euthanized on day +1 after infection. Lungs were collected and processed to determine tissue fungal burden by qPCR. Approximately half of the lung tissue was snap frozen in ethanol/dry ice and stored at −80 °C until analysed76. In brief, lungs were homogenized in Whirl Pak bags (Thermo Fisher Scientific). Approximately 1.5 ml of the homogenate was transferred to sterile screw-cap lysing matrix tubes with 1.4-mm-diameter glass beads (MP Biomedicals). The homogenate containing mouse tissues and fungal hyphae was mechanically disrupted using Fastprep FP120 (Bio Thermo Electro Corporation) with three bursts of 30 s at speed 4 (with incubation on ice between bursts). The supernatant was collected from the secondary homogenate by centrifugation at 800g for 5 min at 4 °C. DNA was extracted from secondary homogenate with the DNeasy tissue kit (Qiagen) according to the manufacturer’s instructions. DNA was recovered in 200 μl of elution buffer and stored at 20 °C until analysis by qPCR. Oligonucleotide amplification primers of the R. oryzae 18S rRNA gene (GenBank: AF113440) were designed with Primer Express software (v.1.5; Applied Biosystems) and synthesized commercially (Sigma-Aldrich). The sequences of these oligonucleotides are as follows: (1) sense amplification primer, 5′-GCGGATCGCATGGCC-3′; and (ii) antisense amplification primer, 5′-CCATGATAGGGCAGAAAATCG-3′. The qPCRs were performed as described previously76. Five-point standard curves were prepared by spiking uninfected lungs with known concentrations of R. oryzae sporangiospores, extracting total lung DNA, and then analysing R. oryzae-specific nucleic acid concentrations by qPCR. The standard curve created by spiking spores was used to calculate log10-transformed spore equivalents per gram of tissue.
For the establishment of immunosuppression/neutropenia, 8–12-week-old age- and sex-matched mice were administered three intraperitoneal injections of cyclophosphamide (Sigma-Aldrich, 150 mg per kg body weight on days −4 and −1, 100 mg per kg on day +3) and a subcutaneous injection of cortisone acetate (Sigma-Aldrich, 300 mg per kg on day −1), as previously described77. The mice were then infected i.t. with 1 × 106 spores of R. delemar. For the prophylactic model, 50 mg of FFA-free HSA was administered intraperitoneally on days −6, −4 and −2 prior to pulmonary infection with R. delemar. For the pre-emptive therapeutic model, 25 mg of FFA-free HSA was administered intraperitoneally daily for 6 consecutive days starting 6 h after pulmonary infection with R. delemar.
For the model of disseminated candidiasis, the C. albicans strain SC5314 was subcultured twice at 37 °C for 48 h on Sabouraud dextrose agar. After the second subculture, single C. albicans colonies were placed in liquid Sabouraud dextrose medium and grown for 16–18 h in a shaking incubator at 30 °C and 150 rpm. Candida blastoconidia were collected, centrifugated at 2,000 rpm for 10 min, washed three times in PBS, counted and reconstituted in PBS to a final concentration of 5 × 106 cells per ml. Then, 150 μl of C. albicans cells diluted in PBS (7.5 × 105) were used to i.v. infect 10–12-week-old, age- and sex-matched, wild-type C57BL/6 and Alb−/− mice, as previously described78. The viability of the inoculated C. albicans blastoconidia was assessed by performing serial dilutions in PBS and plating onto Sabouraud dextrose agar. Plated C. albicans cells were incubated at 37 °C overnight and the next day the number of colonies was counted and the inoculum validated.
Histopathology and immunohistochemistry/immunofluorescence studies
The mice were euthanized, and their lungs were excised and fixed with 10% formalin before being embedded in paraffin and cut into 5–7 μm sections. Lung tissue sections were deparaffinized in xylene and rehydrated through an ethanol gradient (100–70%). H&E or GMS (Sigma-Aldrich) staining was performed for histopathology and fungal burden assessment, according to the manufacturer’s instructions. For histological quantification of fungal burden79, FFPE lung sections were stained with GMS to visualize fungal elements. In certain experiments, fungal conidia were counterstained with periodic acid-Schiff (PAS). Images were acquired from at least ten non-overlapping fields per section at ×10 magnification. Fungal hyphae were enumerated per high-power field in a blinded manner by an experienced pathologist.
For active caspase-3 immunostaining in the lung tissue, heat-induced antigen retrieval was performed to deparaffinize and rehydrate the lung sections through incubation in sodium citrate buffer (10 mM, 0.05% Tween-20, pH 6) for 40 min at 90–95 °C using a steamer (Philips). The tissue sections were allowed to cool for 30 min, washed three times in PBS and blocked endogenous peroxidase with 3% H2O2 for 10 min. The slides were then incubated in blocking solution (serum-free protein block, Dako) for 20 min to block nonspecific binding. The primary antibody was added to the slides at a 1:200 dilution and incubated overnight in a humidified chamber at 40 °C Detection was accomplished using an Envision Horseradish Peroxidase Kit (Dako). Immunostaining was revealed using 3,3′-diaminobenzidine (Dako). The slides were lightly counterstained with haematoxylin, progressively dehydrated through graded alcohols and xylene, and finally covered with a coverslip after mounting in DPX mounting medium. Slides were examined under an Olympus light microscope that was equipped with a ×40 objective.
For mucoricin immunostaining in the lung tissue, heat-induced antigen retrieval was performed to deparaffinize and rehydrate the lung sections through incubation in sodium citrate buffer (10 mM, 0.05% Tween-20, pH 6) for 40 min at 90–95 °C using a steamer (Philips). The tissue sections were allowed to cool for 30 min, washed three times in PBS and permeabilized in 0.2% gelatin/Triton X-100 0.25% for 15 min at room temperature, followed by blocking in 5% BSA/5% normal goat serum (NGS)/PBS for 1 h at room temperature. The tissue slices were subsequently incubated at 4 °C overnight with an anti-mucoricin antibody (2 mg ml−1) in 1% BSA/1% NGS/PBS. The samples were washed three times with PBS then incubated with goat anti-rabbit CF488A or CF555 (1:500, Biotium) in the presence of 200 mg ml−1 CFW for 1 h at room temperature. The tissue sections were incubated for 10 min at room temperature with the nuclear dye TO-PRO-3 (1:2,000, Thermo Fisher Scientific), washed three times with PBS and quenched for autofluorescence by incubation in a solution of 10 mM CuSO4/50 mM NH4Cl.
For mucoricin and CotH3 immunostaining in R. delemar, spores were cultured at 30 °C in RPMI or RPMI supplemented with either 4.5 g dl−1 BSA, 4.5 g dl−1 BSA flow-through or 1 mM caprylic acid for 6-8 h until they swelled. The swollen spores were fixed in 4% paraformaldehyde for 10 min, followed by permeabilization in 0.1% Triton X-100 in PBS for 10 min at room temperature. The permeabilized spores were blocked with 5% NGS in PBS for 1 h at room temperature and subsequently incubated with the anti-mucoricin antibody (10 μg ml−1) or anti-CotH3 antibody (100 μg ml−1)10 for 2 h at room temperature. The spores were washed with Tris-buffered saline (0.01 M Tris HCl/0.15 M NaCl, pH 7.4) containing 0.05% Tween-20 and incubated with goat anti-rabbit CF488A (Biotium) diluted 1:500 in PBS for 1 h at room temperature.
Protein synthesis assessment in R. delemar
R. delemar spores were cultured in black 96-well cell culture plates with a #1.5 glass-like polymer cover slip bottom optimized for high resolution microscopy (P96-1.5P, IBL Baustoff+Labor) at a density of 1.5 × 105 per 200 μl per well. Fungal spores were cultured in RPMI/MOPS (pH 7) supplemented with 0.2% glucose with or without 4.5 g dl−1 BSA flow-through or 2 mM caprylic acid, at 37 °C and 5% CO2. After 2 h (RPMI only) or 3 h (RPMI + BSA flow-through or RPMI + caprylic acid) of culture, media were replenished with OP-puro working solution prepared in the respective media of each treatment group, according to the recommendations of the Protein Synthesis Assay Kit protocol (Cayman Chemical). After 2 h of culture at 37 °C under 5% CO2, R. delemar spores were fixed with the Cell-Based Assay Fixative, washed thrice with the cell-based assay wash buffer and incubated with 5 FAM-azide staining solution in the dark, at room temperature for 30 min. After three washes with cell-based assay wash buffer, R. delemar spores were preserved in 1× assay buffer and imaged directly with a spinning disk confocal microscope (Dragonfly 200, Andor), using the 488 nm/535 nm excitation/emission detection parameters.
Assessment of C12-BODIPY uptake by R. delemar
CFW-labelled R. delemar spores were cultured in RPMI/MOPS (pH 7) for 3 h at 37 °C. To assess whether R. delemar fungus uptakes lipids actively or passively, fungal spores were further cultured at 37 °C or 0 °C for 1 h, in RPMI/MOPS supplemented with 2 μM C12-BODIPY (Invitrogen). Active endocytosis events are inhibited at 0 °C, whereas passive diffusion events remain unaffected80,81. Subsequently, R. delemar spores were washed twice with ice-cold PBS and imaged in PBS using a spinning-disk confocal microscope (Dragonfly 200, Andor). The fluorescence intensity of C12-BODIPY inside R. delemar spores, which correlates with FFA accumulation, was quantified using IMARIS v.10.1 (Oxford Instruments Andor).
Assessment of C11-BODIPY uptake by R. delemar spores
C11-BODIPY (Invitrogen) was pre-oxidized by incubation in 20 mM H2O2/250 μM CuSO4 in UV-grade ethanol at 37 °C under mild shaking for 48 h (ref. 82). As a negative control, an equal amount of C11-BODIPY (40 μM) was incubated in UV-grade ethanol under identical conditions. CFW-labelled R. delemar spores were incubated in RPMI/MOPS (pH 7) without phenol red, supplemented with either pre-oxidized C11-BODIPY or 2 μM control C11-BODIPY, at 37 °C in 18-well μ-Plates (Ibidi). After 4 h, spores were washed twice and imaged in PBS using a spinning-disk confocal microscope (Dragonfly 200, Andor). To validate probe oxidation, spores were first incubated with control C11-BODIPY for 4 h, washed with PBS to remove unbound dye and subsequently treated with increasing concentrations of H2O2 to induce lipid peroxidation. Oxidation was confirmed by a fluorescence shift from 590 nm (red) to 510 nm (green), proportional to the degree of lipid oxidation.
Uptake of oxidized OA by R. delemar spores
For the analysis of OA uptake by R. delemar, fungal spores were cultured in 5% ethanol/RPMI-MOPS supplemented with the indicated concentrations of OA (Sigma-Aldrich) or oxidized OA at 37 °C for 5 h. The spores were collected by scraping, collected in low-affinity 1.5 ml tubes, washed twice with PBS and fixed with methanol-free 4% formaldehyde at room temperature for 20 min. The spores were subsequently washed twice with PBS, stained with 0.5 μg ml−1 Nile red (Thermo Fisher Scientific) at room temperature for 20 min, washed again twice with PBS and stained with 200 μg ml−1 CFW at room temperature for 20 min. Flow cytometry data were acquired using the FACSCanto II system and analysed using FlowJo v.10.6 software (BD Biosciences). All images were acquired using a spinning disk confocal system (Dragonfly 200, Andor) equipped with a motorized inverted Nikon Eclipse Ti2-E microscope, an Andor Sona sCMOS 4.2B-6 camera and four laser lines (405 nm, 488 nm, 561 mm and 633 nm). All images were obtained through z-stacks, deconvolved with Fusion software v.2.3.0.44 (Andor, Oxford Instruments) and further processed in Imaris v.10.1 (Andor, Oxford Instruments) for contrast adjustment, area selection, colour combining and scale bar addition. Certain images were acquired with a Leica TCS SP8 confocal microscope with a ×63 lens and analysed with the use of Fiji/ImageJ. Low-fluorescence immersion oil (Nikon) was used, and imaging was performed at room temperature. Quantitative analysis of fluorescence in confocal microscopy images was performed with IMARIS v.10.1 (Oxford Instruments Andor).
Western blot analysis
The six elutions containing isolated HSA were subjected to western blot analysis for assessment of the isolation protocol. Immunoblotting was performed according to the manufacturer’s instructions using simultaneous antibodies against human transferrin and albumin at a 1:1,000 dilution, followed by incubation at 4 °C overnight. The appropriate secondary antibodies (HRP-conjugated anti-mouse lgG and anti-goat lgG) were used at a 1:5,000 dilution for 1 h at room temperature, and immunoblots were developed by chemiluminescence (ECL; Thermo Scientific). BlueStar Plus Prestained Protein Marker (NIPPON Genetics) was used for the evaluation of protein molecular mass and images were acquired using the ChemiDoc XRSImager (BioRad) system.
Antibodies
Anti-mucoricin and anti-CotH3 antibodies were generated as previously described9. Antibodies against human albumin (sc-365871) and transferrin (sc-365871) were purchased from Santa Cruz. An antibody against mouse active caspase-3 was purchased from Cell Signaling (9661).
Statistical analysis
For univariate and multivariate analysis of the prognostic value of albumin in mucormycosis the primary outcome was all-cause mortality from the time of diagnosis of the infection in the two cohorts at the MDACC, USA and PGIMER, Chandigarh, India and from the time of inclusion to the end of participation of patients (S24 visit) in the Ambizygo study, France. Categorical variables were compared by χ2 or Fisher’s exact test, as appropriate. Continuous variables were compared using the Wilcoxon rank-sum test. Univariate time-dependent survival analysis was performed for the main variables of interest (for example, severe hyperglycaemia, persistent neutropenia, severe hypoalbuminaemia, status of underlying malignancy, steroid use) using Kaplan–Meier curves and the Cox model. The multivariate Cox model was used to identify independent prognostic factor of mortality. Therefore, variables with a P < 0.20 on univariate analysis were included in an initial multivariate model that was then reduced to the final model by backward elimination. Kaplan–Meier survival curves were generated and compared between patients with different albumin levels using the log-rank test. All tests were two-sided with a significance level of 0.05. Data analysis and visualization were performed using Microsoft Office Excel 365 (Microsoft), Prism v10 (GraphPad), SAS v.9.4 (SAS Institute), R (v.4.3.1, R Core Team) and SPSS (IBM, SPSS Statistics for Windows, v.22.0).
For serum lipidomics analysis in Alb−/− and Alb+/+ mice, significant differences between groups were assessed using the Wilcoxon rank-sum test. Data were row-centred and scaled, and visualized as heat maps using the pheatmap R package75. All other statistical analyses were performed using Prism v.10 (GraphPad) unless otherwise stated. P < 0.05 was considered statistically significant for all of the variables tested. A nonlinear regression three-parameter model—log(inhibitor) versus response (three parameters) was used to calculate IC50 values, according to the equation: Y = bottom + (top − bottom)/(\(1+{10}^{(X-\log [{\mathrm{IC}}_{50}])}\)). The FFA concentrations applied to generate the dose–response curves were based on physiological serum concentrations of each FFA in humans24. The statistical methods used to determine significance and the P values of each graph are provided in the figure legends.
Illustrations in Figs. 1f, 2a, 4a,f and 5d, and Extended Data Figs. 1b and 3a were created using BioRender.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All of the raw sequencing reads from this study are available at the NCBI Sequence Read Archive (SRA) under BioProject accession number PRJNA1141971. All data supporting the findings of this study are provided in the Article and its Supplementary Information. Source data are provided with this paper.
Code availability
No new code was generated for this study.
Change history
02 February 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41586-026-10188-1
References
Alqarihi, A., Kontoyiannis, D. P. & Ibrahim, A. S. Mucormycosis in 2023: an update on pathogenesis and management. Front. Cell Infect. Microbiol. 13, 1254919 (2023).
Ribes, J. A., Vanover-Sams, C. L. & Baker, D. J. Zygomycetes in human disease. Clin. Microbiol. Rev. 13, 236–301 (2000).
Hoenigl, M. et al. The emergence of COVID-19 associated mucormycosis: a review of cases from 18 countries. Lancet Microbe 3, e543–e552 (2022).
Hoenigl, M. et al. COVID-19-associated fungal infections. Nat. Microbiol. 7, 1127–1140 (2022).
Muthu, V. et al. Risk factors, mortality, and predictors of survival in COVID-19-associated pulmonary mucormycosis: a multicentre retrospective study from India. Clin. Microbiol. Infect. 30, 368–374 (2024).
Baldin, C. & Ibrahim, A. S. Molecular mechanisms of mucormycosis—the bitter and the sweet. PLoS Pathog. 13, e1006408 (2017).
Iliev, I. D. et al. Focus on fungi. Cell 187, 5121–5127 (2024).
Spellberg, B., Edwards, J. Jr & Ibrahim, A. Novel perspectives on mucormycosis: pathophysiology, presentation, and management. Clin. Microbiol. Rev. 18, 556–569 (2005).
Soliman, S. S. M. et al. Mucoricin is a ricin-like toxin that is critical for the pathogenesis of mucormycosis. Nat. Microbiol. 6, 313–326 (2021).
Gebremariam, T. et al. CotH3 mediates fungal invasion of host cells during mucormycosis. J. Clin. Invest. 124, 237–250 (2014).
Liu, M. et al. The endothelial cell receptor GRP78 is required for mucormycosis pathogenesis in diabetic mice. J. Clin. Invest. 120, 1914–1924 (2010).
Alqarihi, A. et al. GRP78 and integrins play different roles in host cell invasion during mucormycosis. mBio 11, e01087-20 (2020).
Andrianaki, A. M. et al. Iron restriction inside macrophages regulates pulmonary host defense against Rhizopus species. Nat. Commun. 9, 3333 (2018).
Gale, G. R. & Welch, A. M. Studies of opportunistic fungi. I. Inhibition of Rhizopus oryzae by human serum. Am. J. Med. Sci. 241, 604–612 (1961).
Eng, R. H., Corrado, M. & Chin, E. Susceptibility of Zygomycetes to human serum. Sabouraudia 19, 111–115 (1981).
Levitt, D. G. & Levitt, M. D. Human serum albumin homeostasis: a new look at the roles of synthesis, catabolism, renal and gastrointestinal excretion, and the clinical value of serum albumin measurements. Int. J. Gen. Med. 9, 229–255 (2016).
Kontoyiannis, D. P. et al. Zygomycosis in a tertiary-care cancer center in the era of Aspergillus-active antifungal therapy: a case-control observational study of 27 recent cases. J. Infect. Dis. 191, 1350–1360 (2005).
Lanternier, F. et al. Prospective pilot study of high-dose (10 mg/kg/day) liposomal amphotericin B (L-AMB) for the initial treatment of mucormycosis. J. Antimicrob. Chemother. 70, 3116–3123 (2015).
Jones, D. T. et al. Albumin activates the AKT signaling pathway and protects B-chronic lymphocytic leukemia cells from chlorambucil- and radiation-induced apoptosis. Blood 101, 3174–3180 (2003).
Artis, W. M., Fountain, J. A., Delcher, H. K. & Jones, H. E. A mechanism of susceptibility to mucormycosis in diabetic ketoacidosis: transferrin and iron availability. Diabetes 31, 1109–1114 (1982).
Ashraf, S. et al. Unraveling the versatility of human serum albumin—a comprehensive review of its biological significance and therapeutic potential. Curr. Res. Struct. Biol. 6, 100114 (2023).
van der Vusse, G. J. Albumin as fatty acid transporter. Drug Metab. Pharmacokinet. 24, 300–307 (2009).
Borreby, C., Lillebæk, E. M. S. & Kallipolitis, B. H. Anti-infective activities of long-chain fatty acids against foodborne pathogens. FEMS Microbiol. Rev. 47, fuad037 (2023).
Abdelmagid, S. A. et al. Comprehensive profiling of plasma fatty acid concentrations in young healthy Canadian adults. PLoS ONE 10, e0116195 (2015).
Chen, R. F. Removal of fatty acids from serum albumin by charcoal treatment. J. Biol. Chem. 242, 173–181 (1967).
Augé, N., Santanam, N., Mori, N., Keshava, C. & Parthasarathy, S. Uptake of 13-hydroperoxylinoleic acid by cultured cells. Arterioscler. Thromb. Vasc. Biol. 19, 925–931 (1999).
Pap, E. H. et al. Ratio-fluorescence microscopy of lipid oxidation in living cells using C11-BODIPY(581/591). FEBS Lett. 453, 278–282 (1999).
Kimura, K., Yamaoka, M. & Kamisaka, Y. Rapid estimation of lipids in oleaginous fungi and yeasts using Nile red fluorescence. J. Microbiol. Methods 56, 331–338 (2004).
Blache, D. et al. Glycated albumin with loss of fatty acid binding capacity contributes to enhanced arachidonate oxygenation and platelet hyperactivity: relevance in patients with type 2 diabetes. Diabetes 64, 960–972 (2015).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl Acad. Sci. USA 109, 413–418 (2012).
Low, B. E. & Wiles, M. V. A humanized mouse model to study human albumin and albumin conjugates pharmacokinetics. Methods Mol. Biol. 1438, 115–122 (2016).
Abdollahi, A., Dowden, B. N., Buhman, K. K., Zembroski, A. S. & Henderson, G. C. Albumin knockout mice exhibit reduced plasma free fatty acid concentration and enhanced insulin sensitivity. Physiol. Rep. 10, e15161 (2022).
Roden, M. M. et al. Epidemiology and outcome of zygomycosis: a review of 929 reported cases. Clin. Infect. Dis. 41, 634–653 (2005).
Zhang, J. et al. Risk factors for disease severity, unimprovement, and mortality in COVID-19 patients in Wuhan, China. Clin. Microbiol. Infect. 26, 767–772 (2020).
Dos Santos, P. T., Thomasen, R. S. S., Green, M. S., Færgeman, N. J. & Kallipolitis, B. H. Free fatty acids interfere with the DNA binding activity of the virulence regulator PrfA of Listeria monocytogenes. J. Bacteriol. 202, e00156-20 (2020).
Casillas-Vargas, G. et al. Antibacterial fatty acids: an update of possible mechanisms of action and implications in the development of the next-generation of antibacterial agents. Prog. Lipid Res. 82, 101093 (2021).
Coonrod, J. D., Lester, R. L. & Hsu, L. C. Characterization of the extracellular bactericidal factors of rat alveolar lining material. J. Clin. Invest. 74, 1269–1279 (1984).
Coonrod, J. D. & Yoneda, K. Detection and partial characterization of antibacterial factor(s) in alveolar lining material of rats. J. Clin. Invest. 71, 129–141 (1983).
Kenny, J. G. et al. The Staphylococcus aureus response to unsaturated long chain free fatty acids: survival mechanisms and virulence implications. PLoS ONE 4, e4344 (2009).
Egesten, A., Frick, I. M., Mörgelin, M., Olin, A. I. & Björck, L. Binding of albumin promotes bacterial survival at the epithelial surface. J. Biol. Chem. 286, 2469–2476 (2011).
Hitzler, S. U. J. et al. Host albumin redirects Candida albicans metabolism to engage an alternative pathogenicity pathway. Nat. Commun. 16, 6447 (2025).
Roche, M., Rondeau, P., Singh, N. R., Tarnus, E. & Bourdon, E. The antioxidant properties of serum albumin. FEBS Lett. 582, 1783–1787 (2008).
Zhang, L., Wu, T. & Jia, H. Invasive pulmonary aspergillosis in patients with lung cancer: risk factors for in-hospital mortality and predictors of clinical outcomes. J. Mycol. Med. 35, 101560 (2025).
Santos, A. et al. Hypoalbuminemia is a risk factor for invasive fungal infections and poor outcomes in infected kidney transplant recipients. Clin. Transplant. 37, e15052 (2023).
Gremese, E. et al. Serum albumin levels: a biomarker to be repurposed in different disease settings in clinical practice. J. Clin. Med. 12, 6017 (2023).
Leek, L. V. M. et al. Multi-omic analysis identifies hypoalbuminemia as independent biomarker of poor outcome upon PD-1 blockade in metastatic melanoma. Sci. Rep. 14, 11244 (2024).
Baker, M. E. Albumin, steroid hormones and the origin of vertebrates. J. Endocrinol. 175, 121–127 (2002).
Chamilos, G. et al. Drosophila melanogaster as a model host to dissect the immunopathogenesis of zygomycosis. Proc. Natl Acad. Sci. USA 105, 9367–9372 (2008).
Mertens, J. A., Skory, C. D. & Ibrahim, A. S. Plasmids for expression of heterologous proteins in Rhizopus oryzae. Arch. Microbiol. 186, 41–50 (2006).
Kawakami, A. et al. Identification and characterization of oxidized human serum albumin: a slight structural change impairs its ligand-binding and antioxidant functions. FEBS J. 273, 3346–3357 (2006).
Day, J. F., Thorpe, S. R. & Baynes, J. W. Nonenzymatically glucosylated albumin. In vitro preparation and isolation from normal human serum. J. Biol. Chem. 254, 595–597 (1979).
Soliman, S. S. M. et al. Effective targeting of breast cancer cells (MCF7) via novel biogenic synthesis of gold nanoparticles using cancer-derived metabolites. PLoS ONE 15, e0240156 (2020).
Semreen, M. H. et al. Metabolic profiling of candida auris, a newly-emerging multi-drug resistant Candida species, by GC-MS. Molecules 24, 399 (2019).
Xia, J. & Wishart, D. S. Metabolomic data processing, analysis, and interpretation using MetaboAnalyst. Curr. Protoc. Bioinformatics https://doi.org/10.1002/0471250953.bi1410s34 (2011).
Alsabeeh, N., Chausse, B., Kakimoto, P. A., Kowaltowski, A. J. & Shirihai, O. Cell culture models of fatty acid overload: problems and solutions. Biochim. Biophys. Acta 1863, 143–151 (2018).
Evans, M. et al. Conjugated linoleic acid suppresses triglyceride accumulation and induces apoptosis in 3T3-L1 preadipocytes. Lipids 35, 899–910 (2000).
Ferraz, T., Fiuza, M., Dos Santos, M., De Carvalho, L. P. & Soares, N. M. Comparison of six methods for the extraction of lipids from serum in terms of effectiveness and protein preservation. J. Biochem. Biophys. Methods 58, 187–193 (2004).
Yamazaki, E., Inagaki, M., Kurita, O. & Inoue, T. Kinetics of fatty acid binding ability of glycated human serum albumin. J. Biosci. 30, 475–481 (2005).
Ekundayo, J. A. & Carlile, M. J. The germination of sporangiospores of Rhizopus arrhizus; spore swelling and germ-tube emergence. J. Gen. Microbiol. 35, 261–269 (1964).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Ioannou, P. et al. Albumin enhances caspofungin activity against aspergillus species by facilitating drug delivery to germinating hyphae. Antimicrob. Agents Chemother. 60, 1226–1233 (2015).
Davison, A. & Wajda, M. Analysis of lipids from fresh and preserved adult human brains. Biochem. J. 82, 113 (1962).
Rose, H. G. & Oklander, M. Improved procedure for the extraction of lipids from human erythrocytes. J. Lipid Res. 6, 428–431 (1965).
Srinivasan, A. et al. Microwave-enhanced advanced oxidation treatment of lipids and food wastes. Water Air Soil Pollut. 229, 227 (2018).
Iuchi, K., Ema, M., Suzuki, M., Yokoyama, C. & Hisatomi, H. Oxidized unsaturated fatty acids induce apoptotic cell death in cultured cells. Mol. Med. Rep. 19, 2767–2773 (2019).
Ghannoum, M. A., Filler, S. G., Ibrahim, A. S., Fu, Y. & Edwards, J. E. Jr. Modulation of interactions of Candida albicans and endothelial cells by fluconazole and amphotericin B. Antimicrob. Agents Chemother. 36, 2239–2244 (1992).
Donnelly, J. P. et al. Revision and update of the consensus definitions of invasive fungal disease from the European Organization for Research and Treatment of Cancer and the Mycoses Study Group Education and Research Consortium. Clin. Infect. Dis. 71, 1367–1376 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Garcia Lopez, A., Albrecht-Eckardt, D., Panagiotou, G. & Schäuble, S. FungiFun3: systemic gene set enrichment analysis for fungal species. Bioinformatics 40, btae620 (2024).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 10, 161 (2009).
Kolde, R. & Kolde, M. R. pheatmap. R package version 1.0.12 (CRAN, 2015).
Luo, G. et al. Efficacy of liposomal amphotericin B and posaconazole in intratracheal models of murine mucormycosis. Antimicrob. Agents Chemother. 57, 3340–3347 (2013).
Wurster, S. et al. Blockade of the PD-1/PD-L1 immune checkpoint pathway improves infection outcomes and enhances fungicidal host defense in a murine model of invasive pulmonary mucormycosis. Front. Immunol. 13, 838344 (2022).
Chamilos, G. et al. Candida albicans Cas5, a regulator of cell wall integrity, is required for virulence in murine and toll mutant fly models. J. Infect. Dis. 200, 152–157 (2009).
Stolz, D. J., Sands, E. M., Amarsaikhan, N., Tsoggerel, A. & Templeton, S. P. Histological quantification to determine lung fungal burden in experimental aspergillosis. J. Vis. Exp. 9, 57155 (2018).
Varma, S. G., Mitra, A. & Sarkar, S. Self-diffusion is temperature independent on active membranes. Phys. Chem. Chem. Phys. 26, 23348–23362 (2024).
Tomoda, H., Kishimoto, Y. & Lee, Y. C. Temperature effect on endocytosis and exocytosis by rabbit alveolar macrophages. J. Biol. Chem. 264, 15445–15450 (1989).
Drummen, G. P., Gadella, B. M., Post, J. A. & Brouwers, J. F. Mass spectrometric characterization of the oxidation of the fluorescent lipid peroxidation reporter molecule C11-BODIPY(581/591). Free Radic. Biol. Med. 36, 1635–1644 (2004).
Acknowledgements
We thank E. Deligianni for assistance with time-lapse microscopy; K. Stylianou, C. Tsatsanis and M. Katrinaki for providing reagents and human samples, and performing measurements of albumin levels in human and mouse serum samples; A. Vatikioti for assistance with mouse sera collection; and I. Lagkouvardos for discussions and input on RNA-seq data analysis. A.P. was supported by the General Secretariat for Research and Technology (GSRT) and the Hellenic Foundation for Research and Innovation (H.F.R.I.) under the “1st call of H.F.R.I. for PhD Candidates” (project number: 1119); I.M. by the Hellenic Foundation for Research and Innovation (H.F.R.I.) under the “3rd Call for H.F.R.I. Research Projects to support Post-Doctoral Researchers” (project number: 7054); G.C. by an ERC Consolidator Grant (iMAC-FUN, 864957), an Horizon 2020–Research and Innovation Framework Programme from Europe, H2020-SC1-BHC-2018-2020 (HDM-FUN, 84750) and a ‘la Caixa’ Foundation Health Research Grant (TRANS-CPA, LCF/PR/HR17/52190003); G.C., E.K., A.E. and K.G. by the General Secretariat for Research and Innovation of Greece Grant PRO-sCAP (project code TAEDR-0541976) carried out within the framework of the National Recovery and Resilience Plan Greece 2.0 and funded by the European Union-Next Generation EU; A.C. by the Fundação para a Ciência e a Tecnologia (FCT) (UID/06304/2023, LA/P/0050/2020 and 2022.06674.PTDC) and by the ‘la Caixa’ Foundation under the agreement LCF/PR/HR22/52420003; V.G. and C.L. by the grant PID2024-160088NB-I00 funded by MICIU/AEI/10.13039/501100011033 and by ERDF/EU; A.S.I by the Public Health Service grant no. R01 AI063503; and V.M.B. by NIH grants U19 AI110820 and R01AI141360. D.P.K. acknowledges the Robert C. Hickey endowment.
Author information
Authors and Affiliations
Contributions
A.P., I.M. and V.N. performed and analysed most of the experiments in human samples and mice and participated in writing the manuscript. I.M. established protocols on the in vivo model of invasive fungal infections in albumin-KO mice, and in vitro assays on the assessment of antimicrobial activity of serum and BAL lipids and fungal protein synthesis. R.H. and S.S.M.S. analysed vitamins, trace elements and amino acids in culture medium, performed chemical modifications of albumin and serum lipids, and validated the efficiency of oxidation of FFAs. A.L.-L., M.M.-G. and S.M.C.-A. performed lipidomics analyses. V.M., R.A., S.M.R., F.L. and A.C. collected and analysed clinical data. M.H. and V.P. performed fractionation and identification of FFAs in albumin samples. M.P. and K.P. established the assays for human albumin depletion and purification from serum. I.K. performed in vivo studies and analysed data. Y.G. performed experiments on mucoricin expression and provided reagents. R.A., T.M., Y.V. and J.M. provided human samples and analysed data. S.-Y.C. and A.S. analysed clinical data. D.S. and E.K. collected human serum samples and analysed data. A.E. and K.G. analysed data. A.C. analysed data. O.K. provided clinical isolates and analysed data. E.D. and M.T. performed histopathological studies. V.M.B. performed RNA-seq analyses. E.I. performed gene enrichment analysis of RNA-seq data and analysed lipidomics data. V.G. and C.L. supervised the analysis on fungal transcriptomics. Y.J. and C.E. performed logistic regression analysis of clinical cohorts. C.B. supervised all of the lipidomics studies. D.P.K. provided clinical data and suggestions throughout the study. A.S.I. provided reagents, designed and supervised experiments, analysed data and participated in discussions and suggestions throughout the study. G.C. conceived and supervised the study, was involved in the design and evaluation of all of the experiments, and wrote the manuscript along with comments from all of the other authors.
Corresponding author
Ethics declarations
Competing interests
A.S.I. owns shares in Vitalex Biosciences, a start-up company that is developing immunotherapies and diagnostics for mucormycosis. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Andrew Hall, Tobias Hohl and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Correlation of serum albumin level with antifungal activity, albumin purification, and evaluation of other serum proteins.
a, Correlation between serum albumin levels and R. delemar growth in human sera obtained from patients with cirrhotic disease (n = 18, left) or haematologic malignancies (n = 20, right). Simple linear regression test was used to determine deviation from zero. b, Outline of human serum isolation using blue Sepharose columns. c, Immunoblot analysis of albumin and transferrin in human serum and eluted fragments following isolation. Representative image from 3 independent experiments. d, Representative images of R. delemar spores cultured for 8 h in medium alone or medium supplemented with increasing concentrations of human IgG. Representative images from 3 independent experiments performed in triplicate. Scale bar, 20 µm.
Extended Data Fig. 2 Selective fungistatic activity of albumin against Mucorales.
a, Representative images of R. delemar dormant spores (upper panel) or germinating spores (lower panel) cultured for 10 h in medium alone, medium + BSA, or medium + HSA. Scale bar, 50 µm. b, Assessment of R. delemar germination in medium alone, medium supplemented with human serum albumin (medium + albumin), or medium alone following initial incubation in albumin-containing medium (fresh medium) (n = 5 independent experiments). Data are mean ± s.d. ns = 0.437; ****p < 0.0001, one-way ANOVA with Tukey’s multiple comparisons post hoc test. c, Representative fluorescence images of different human fungal pathogens labelled with CFW-and cultured for 6 h in medium ± 4.5 g dl−1 albumin. Scale bar, 20 µm.
Extended Data Fig. 3 Analysis of antifungal activity and composition of albumin-rich medium following filtration.
a, Outline of albumin-rich medium filtration and generation of albumin-depleted flow through (left panel; created with BioRender.com). Assessment of R. delemar growth in medium, medium + albumin, or medium + albumin-depleted flow through (right panel) (n = 5 independent experiments). Data are mean ± s.d. *p = 0.022; ****p < 0.0001, one-way ANOVA with Tukey’s test. b, Heatmap of vitamins, amino acids (LC-MS), and trace elements (ICP-MS). Standard RPMI set to 1; red = low, green = high. c, Assessment of R. delemar growth in medium ± albumin (n = 3) and medium + flow through ± essential/non-essential amino acids (n = 3), vitamins, trace elements (n = 3), or all combined (n = 3), from independent experiments performed in triplicate. Data are mean ± s.d. ****p < 0.0001, one-way ANOVA with Tukey’s test. Data are included in Source Data file.
Extended Data Fig. 4 Analysis of activity and FFA composition in albumin filtrate.
a, Outline of solid-phase extraction (SPE) procedure. GC-MS TIC chromatographs of derivatized fractions (F2–F4) and assessment of R. delemar growth (16 h). Scale bar, 50 µm. Caprylic acid concentrations: F2 = 194 µg/mL (1.34 mM), F3 = 15 µg/mL (104 µM), F4 = 0.1 µg/mL (0.7 µM). b, EI-MS spectrum of F2 (caprylic acid methyl ester) compared to NIST14.0 reference. Molecular ion (M+·, m/z 158) and characteristic fragments annotated. c, ESI(-)-TOF-MS of underivatized F2: [M–H]− at m/z 143.1079 with isotopic distribution; EC and RDB eq. provided. Data are a representative example of n = 2 independent experiments. d, Inhibition of R. delemar by increasing purified caprylic acid in 5% EtOH (n = 6 independent experiments). Data are mean ± s.d.
Extended Data Fig. 5 Active uptake of non-oxidized FFAs by Mucorales.
a, Representative fluorescence images and cumulative C12-BODIPY MFI of CFW-labelled R. delemar spores at 37 °C or 0 °C (n = 316–334 spores, 3 independent experiments). Data are mean ± s.d. ****p < 0.0001, Mann–Whitney test. b, Fluorescence of R. delemar spores cultured with C11-BODIPY ± 0%, 1%, or 10% H2O2 (3 independent experiments). Scale bar, 10 µm. c–d, Nile Red staining of spores in medium ± oleic acid or oxidized OA. Representative images and cumulative MFI (n = 25–20 optical fields, 3 experiments). Data are mean ± s.d. ****p < 0.0001, one-way ANOVA with Tukey’s test. e–g, GC-MS, ANS binding, and fluorescence quenching of caprylic acid on glycosylated/non-glycosylated BSA (n = 3 independent experiments). Data are mean ± s.d. **p-values as indicated.
Extended Data Fig. 6 Albumin-bound FFAs suppress Mucorales virulence via protein synthesis arrest.
a, H&E histopathology of lungs at 0 h and 6 h post-infection with control (n = 8)- or albumin-swollen (n = 7) R. delemar. Scale bar, 100 µm. b, Enriched KEGG pathways for DEGs (GSEA). c, Differential expression of protein synthesis-related genes in dormant spores (T0) or after 3 h in medium ± albumin (T3/T3Alb) (n = 3 biologically independent samples). d–e, Representative confocal images and cumulative data for CotH3 and mucoricin expression (n = 85–128 spores or 18–29 optical fields, 3 independent experiments). Data are mean ± s.d. ****p < 0.0001, Mann–Whitney test. f, Survival of C57BL/6 mice infected i.t. with 2.5 × 106 swollen control-RNAi (n = 14) or mucoricin-RNAi (n = 15) spores. Log-rank (Mantel–Cox) test.
Extended Data Fig. 7 Albumin deficiency compromises serum FFA composition and increases susceptibility to mucormycosis.
a–b, Histopathology and cumulative hyphal counts of lungs from neutropenic Alb+/+ and Alb−/− mice infected with A. fumigatus or R. delemar (n = 4–6 mice). Data are mean ± s.d., two-sided Mann–Whitney test. Scale bar, 100 µm. c, Cell damage of A549 cells treated with mucoricin ± 4.5 g dl−1 FFA-free BSA (n = 14 mucoricin plus BSA, n = 15 mucoricin alone). Data are mean ± s.d. ***p = 0.0004, Mann–Whitney test. d, Serum non-oxidized/oxidized FFA concentrations in uninfected and infected Alb+/+ vs Alb−/− mice (n = 6). One-way ANOVA with Dunn’s test. e–f, Heatmaps of serum FFAs and free oxylipins (n = 10 mice, Wilcoxon rank sum test). g, Proposed model of albumin in host defence during Mucorales infection. Created with BioRender.com.
Supplementary information
Supplementary Methods (download PDF )
Detailed methods for the identification of serum oxylipins and the standards used for the analysis of serum FFAs.
Supplementary Table 1 (download XLSX )
The demographics and clinical characteristics of all clinical cohorts reported in the Article. Each sheet provides the data corresponding to the figure in which the cohort is presented.
Supplementary Table 2 (download XLSX )
The raw data supporting the analyses presented in Extended Data Fig. 3.
Supplementary Table 3 (download XLSX )
The raw lipidomics data corresponding to the relevant figures in the Article.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pikoulas, A., Morianos, I., Nidris, V. et al. Albumin orchestrates a natural host defence mechanism against mucormycosis. Nature 649, 693–702 (2026). https://doi.org/10.1038/s41586-025-09882-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09882-3
This article is cited by
-
Albumin protects against deadly fungal infection
Nature Reviews Immunology (2026)
-
Blood protein thwarts deadly fungal disease
Nature (2026)