Abstract
Immune interventions toward an HIV cure have focused on rejuvenating adaptive immune responses. Herein we provide a framework that features epigenetic programming of myeloid and CD4+ T cells as a major mechanism that promotes decay of the HIV reservoir. Coordinate regulation of gene expression and chromatin accessibility of pathways of innate antiviral immunity was associated with decay of cell-associated viral DNA (CA-vDNA) following analytical treatment interruption in simian immunodeficiency virus-infected rhesus macaques (RMs) treated with anti-IL-10 and anti-PD-1. TGF-β/SMAD signaling in a subset of combo-treated CA-vDNAhi RMs, suppressed this antiviral activity through histone deacetylases, reducing chromatin accessibility of interferon regulatory factors (IRFs) and STATs. Addition of HDAC inhibitors in vitro restored antiviral response in the presence of TGF-β. Induction of IL-6 in CA-vDNAlo RMs amplified the antiviral network through IRF9. We identified an overlapping molecular cascade in HIV elite controllers, who maintain small HIV reservoirs without antiviral treatment. These data provide insights into strategies for HIV cure interventions.
Similar content being viewed by others
Main
Curing HIV infection remains one of the most formidable challenges in modern medicine. Multiple approaches including ‘shock and kill’ strategies1, immune interventions2 and cell-based therapies3,4 have had little impact on the reservoir or led to only transient control of viral rebound following interruption of antiretroviral therapy (ART). Most of these interventions aimed at restoring adaptive immunity (T and B cell responses), with little focus on rejuvenating the innate arm of the immune system known to be dysfunctional in people with HIV (PWH)5. Yet innate antiviral immune responses, specifically the interferon (IFN) pathway, play a critical role in the control of viral infections6 as they trigger the expression of IFN-stimulated genes (ISGs) that target viruses at multiple stages of their life cycle and are associated with control of HIV acquisition7.
TGF-β is a pleiotropic cytokine produced by several cells of the immune system, including natural and induced regulatory T (Treg) cells8,9. It controls T cell homeostasis and is critical to reduce the inflammation observed in acute and chronic viral infections10. Circulating TGF-β levels are increased in PWH11. Our group and others have shown the critical role of TGF-β in the establishment and maintenance of HIV persistence12,13. The mechanisms downstream of TGF-β that modulate antiviral defenses allowing reservoir establishment are yet to be defined.
HIV elite controllers (ECs) are a rare group of PWH (0.1–2.5%) who maintain undetectable HIV viral load (VL; <50 copies per ml) for over 2 years in the absence of ART14. Host factors such as specific HLA subtypes, higher levels of expression of p21 mRNA in T cells, and lower levels of expression of CCR5 (refs. 15,16,17) partially explain the control of viremia in the majority but not all ECs. The identification of additional immune effector mechanisms associated with antiviral immunity can provide a better understanding of VL control in HIV ECs.
Herein, we used cellular, molecular, and systems immunology approaches to identify, in RMs, the pathways downstream of IL-10 and PD-1 signaling blockade associated with lower levels of cell-associated viral DNA (CA-vDNA).
Results
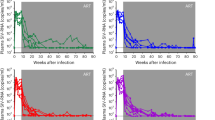
We previously demonstrated18 that infusion of a combination of monoclonal antibodies that block IL-10 and PD-1 signaling (combo treatment) in ART-treated rhesus macaques (RMs) (Extended Data Fig. 1a) led to (1) post-analytical treatment interruption (ATI) control of VL rebound in 90% of the RMs to <1,000 copies per ml over 6 months (Extended Data Fig. 1a); and (2) declining levels of Ca-vDNA (<100 cells per 106 CD4+ T cells) in 40% of RMs over the same period (so called CA-vDNAlo RMs; Extended Data Fig. 1b). A significant control of viremia was not observed when only IL-10 signaling was blocked (aIL-10). RMs in the combo-treated group could not be distinguished by the frequencies of cells with CA-vDNA pre-ATI18. Moreover, both CA-vDNAlo and CA-vDNAhi (cell-associated viral DNA high) RMs showed the same levels of plasma VL post-ATI and during the follow-up period18.
Antiviral gene expression signatures are induced by combo treatment and are associated with virologic control
We performed an unbiased analysis of the transcriptomic profile comparing the three groups (control (ART-only), aIL-10 and combo (aIL-10 + aPD-1))19 of RMs to define immunologically relevant pathways after combo treatment leading to viral rebound control. Module analysis (Hallmark, MSigDB; Extended Data Fig. 1c) and gene set enrichment analysis (GSEA; nominal P < 0.05) (Fig. 1a) revealed that IFN signaling (MSigDB Hallmark and Restriction Factors) was upregulated in lymph node mononuclear cells (LNMCs) and peripheral blood mononuclear cells (PBMCs) from combo-treated RMs pre-ATI when compared to RMs treated with aIL-10 alone and ART-only controls (Supplementary Tables 1 and 2 and Supplementary Fig. 1a–e). The network of differentially expressed genes (DEGs) from these pathways included known antiviral restriction factors (RFs) (APOBECs, TRIMs and ISG15; Fig. 1b), all of which are targets of the transcription factors (TFs) IRF7 and STAT1 (ref. 20). The inferred transcriptional activity of IRF7 and STAT1 in combo-treated RMs included heightened expression of genes that regulate cell proliferation, DNA damage and cell cycle arrest (PSMA5/6, PSMB8/10, RAD1 and BARD1) (Extended Data Fig. 1d,e). Sample-level enrichment analysis (SLEA) scores for all 28 RMs revealed elevated expression of the type-I and type-II IFN cascades in combo-treated CA-vDNAlo RMs (Fig. 1c). Expression of several ISGs (IRF7 and TRIMs) pre-ATI was inversely correlated with levels of CA-vDNA, cell-associated viral RNA (CA-vRNA) and two-long terminal repeats (2-LTR) circles 24 weeks post-ATI (Fig. 1d and Supplementary Table 3). The role of IFNs pre-ATI in defining virologic outcomes was further supported by high-dimensional flow cytometry showing that CA-vDNAlo RMs exhibited significantly higher levels of pIRF3/7 (particularly within LNMCs) compared to CA-vDNAhi animals (Extended Data Fig. 1f). pIRF3 expression levels in classical monocytes was inversely correlated with plasma VL at 24 weeks post-ATI, whereas pIRF7 expression in central memory CD4⁺ T cells (TCM) pre-ATI was inversely correlated with VL, CA-vDNA and CA-vRNA levels at the same time point (Extended Data Fig. 1g–h and Supplementary Fig. 2a–c). Integration of flow cytometry and bulk RNA sequencing (RNA-seq) data pre-ATI substantially improved prediction of lower CA-vRNA levels post-ATI in combo-treated RMs compared to either dataset alone (P = 0.0093 versus P = 0.021; Extended Data Fig. 1i). In LNMCs, increased levels of pIRF3 and pSTAT1 were observed in naive CD4+ and CD8+ TCM from combo-treated RMs 24 weeks post-ATI (Extended Data Fig. 2a). Of note, several IFN-related pathways, including Hallmark IFN α/γ signaling and Reactome IFN signaling, which were upregulated in CA-vDNAlo RMs pre-ATI, were maintained 24 weeks post-ATI (Fig. 1e and Supplementary Table 4) as compared to CA-vDNAhi RMs. IRF7, its target genes, host antiviral RFs ISG15/20, OAS1/L, MX1 and the genes encoding the chemokines CXCL10/11 were commonly upregulated in CA-vDNAlo RMs pre- and post-ATI (Fig. 1f). Taken together, these data reveal that combo treatment led to early induction (pre-ATI) of IFN-related pathways and antiviral genes whose expression levels in myeloid cells and T cells were inversely correlated with frequencies of cells that contained simian immunodeficiency virus (SIV) DNA. These signatures persisted for up to 24 weeks post-ATI in CA-vDNAlo RMs; conversely, they were significantly reduced in CA-vDNAhi RMs pre-ATI and 24 weeks post-ATI.
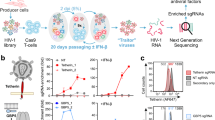
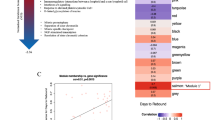
a, Pathways (MSigDB, Hallmark) significantly (P ≤ 0.05) differentiated in combo-treated RMs from control RMs or aIL-10 treated RMs pre-ATI in PBMCs and LNMCs. Heatmap illustrates GSEA NES ranging from high (red) to low (blue). Rows represent gene sets (IFN pathways) and columns represent contrasts in PBMCs and LNMCs. n = 28. b, Network inference (Cytoscape and Genemania) showing leading-edge genes (LEGs) (P < 0.05) of pathways in Fig. 1a upregulated in PBMCs and LNMCs of combo-treated RMs (red circular nodes). Connecting lines Edges: gene–gene interaction, color-coded by network type. c, Heatmap illustrating SLEA scores for significant (P < 0.05) IFN signaling pathways and RFs pre-ATI in LNMCs from control (black), aIL-10 only (red) and combo-treated RMs (blue). Rows represent pathways and columns represent individual RMs. Red-white-blue gradient for SLEA pathway scores from low and high relative levels of expression, respectively. Top annotation: levels of CA-vRNA, CA-vDNA and 2-LTR circles (white to blue, green, purple, respectively) 24 weeks post-ATI. The rectangle outline indicates the subset of combo-treated RMs (n = 10). n = 28. d, Heatmap illustrating gene expression in LNMCs pre-ATI correlated with viral outcomes (log10 SIV CA-vDNA, log10 CA-vRNA and log10 2-LTR circles) 24 weeks post-ATI (false discovery rate (FDR)-adjusted P < 0.05) (left). Two-sided Spearman correlation, P values adjusted for multiple comparisons with Benjamini–Hochberg (BH) procedure. Rows represent genes and columns represent individual RMs: controls (black), aIL-10 only (red), and combo-treated (blue). Standarized gene expression (z-score): blue-white-red gradient, for low to high relative expression, respectively. n = 25 RMs with virological readouts available. Network representing the top ISGs with antiviral capacity negatively correlated (NES = −1.44, P = 0.021) to viral outcomes at week 24 post-ATI (right). Blue nodes represent inverse correlation with viral outcomes. e, IFN and antiviral pathways (P ≤ 0.05) persistently upregulated in CA-vDNAlo RMs pre- and post-ATI. Heatmap illustrates NES for IFN-related pathways. Rows represent pathway sets and columns represent comparisons at both pre- and post-ATI time points in LNMCs. n = 10 combo-treated RMs. f, Network inference (Cytoscape and Genemania) showing common LEGs upregulated at weeks 0 and 24 in CA-vDNAlo RMs. Red nodes represent upregulated genes. Edges highlight gene interactions. g, Network inference (Cytoscape and Genemania) showing the IFN-associated genes upregulated in PBMCs and LNMCs (significant LEGs, P < 0.05). Red circular nodes represent upregulated genes in combo-treated RMs compared to aIL-10-treated RMs edges highlight gene–gene interactions. h, Experimental design. CD4⁺ T cells from healthy donors (n = 6) were pretreated with IFN-β and divided into two pools. Unlabeled cells were infected with HIV by spinoculation, while uninfected cells were labeled with CTV. On day 2, infected and CTV-labeled cells were mixed (1:1) and co-cultured for 4 days in the presence of saquinavir. Intracellular p24 expression was measured by flow cytometry. Figure created in BioRender; Pereira Ribeiro, S. https://biorender.com/qr7isln (2026). i, Frequencies of HIV p24⁺ cells were measured by flow cytometry 4 days post-infection in CTV⁻ and CTV⁺ cells. Gray bars, untreated; blue bars, IFN-β treated. Circles, HIV-infected CD4⁺ T cells; triangles, CTV-labeled bystander CD4⁺ T cells. Comparison of groups means was performed with a nonparametric Friedman test and P values were adjusted for multiple comparisons with a Benjamini, Krieger and Yekutieli (BKY) procedure. Error bars represent mean ± s.e.m. *P < 0.05, **P < 0.005, paired t-test.
Administration of aPD-1 is required for the induction of antiviral signatures
Previous reports showed that treatment with aPD-1 alone only had a transient impact on VL21,22,23,24,25,26. In our study, RMs treated only with aIL-10 also did not show VL control post-ATI18 (Extended Data Fig. 1a), suggesting that it was the combination of aPD-1 and aIL-10 that led to the control of VL and decay of CA-vDNA levels. Bulk RNA-seq data showed a total of 2,361 (626 upregulated) and 1,488 (938 upregulated) DEGs were significant modulated in LNMCs and PBMCs from combo-treated RMs, respectively, compared to RMs treated only with aIL-10 (Extended Data Fig. 2b and Supplementary Table 1). Functional analysis of these DEGs showed that RFs and IFN-induced pathways (IFN-β and IFN-γ) were enriched pre-ATI in LNMCs and PBMCs from combo-treated RMs (Fig. 1a), and included DEGs such as APOBECs, TRIMs and MX2 (Fig. 1g). Upregulation of these genes in LNMCs from combo-treated RMs distinguished between RMs with low and high levels of CA-vDNA, CA-vRNA and 2-LTR circles 24 weeks post-ATI (Extended Data Fig. 2c).
To experimentally confirm the role of IFNs in protecting bystander CD4+ T cells from infection, CD4+ T cells from healthy human volunteers were left unstimulated or were pretreated with IFN-β. Each culture was subdivided into two pools: cells in the first pool were infected with HIV, while those in the second pool were labeled with CellTrace Violet (CTV) and used as bystander targets (Fig. 1h and Supplementary Fig. 3a). Pretreatment with IFN-β before co-culture led to the upregulation of ISGs (e.g. IFIT1, APOBEC3G and MX2) compared to untreated conditions (Extended Data Fig. 3a). After 4 days of co-culture, untreated CD4+ T cells and untreated bystander (CTV-labeled) CD4+ T cells were both productively infected with HIV, whereas treated CD4+ T cells were protected from infection and presented with significantly lower frequencies of HIV p24+ cells than untreated CD4+ T cells. This led to significantly lower levels of infection of untreated CTV-labeled bystander CD4+ T cells. Treated CTV-labeled bystander CD4+ T cells were protected from infection in the presence of untreated, productively infected CD4+ T cells (Fig. 1i). Collectively, the induction of antiviral signatures pre-ATI in CA-vDNAlo RMs promoted refractoriness to infection, sparing bystander target cells from infection by rebounding virus post-ATI.
Plasma TGF-β levels and its downstream gene signatures are associated with a network of epigenetic enzymes that inhibit the IFN response
Among the 28 cytokines quantified in plasma, CA-vDNAhi RMs presented significantly higher levels of all TGF-β isoforms (TGF-β1, TGF-β2 and TGF-β3) pre-ATI compared to CA-vDNAlo RMs (Fig. 2a and Supplementary Table 5). TGF-β inhibits the production of type-I and type-II IFNs27. CA-vDNAlo RMs presented significantly higher plasma levels of MIP-1α, an inhibitor of HIV entry28 and IL-6, which triggers IFNs29,30. Supporting these findings, GSEA revealed that pathways downstream of TGF-β signaling, such as SMAD2/3 targets (AP-1: FOS/JUN) and/or inflammation (TNF signaling, NFKBIA, TNFSF9 and IL1A) were upregulated in LNMCs from CA-vDNAhi RMs (Fig. 2b,c and Supplementary Table 6). SMAD2/3 and AP-1 translocate to the nucleus to drive the expression of TGF-β19. This analysis revealed that pathways that regulate chromatin accessibility were upregulated in CA-vDNAhi RMs. The SWI/SNF complex (SMARCA4/BRG1 and SMARCB1) and HDAC11 are all inhibitors of both type-I and type-II IFN transcription31 and were associated with plasma levels of TGF-β and TGF-β signaling (SMAD3) (Fig. 2d). In contrast, the RF pathway19, including >50 genes with antiviral activity such as ISG15, APOBEC3A and TRIM7, and the C/EBP-β pathway, which controls the expression of IL-6 and inhibits the expression of AP-1, were specifically upregulated in CA-vDNAlo RMs. Of note, the normalized enrichment scores (NES) of TGF-β signaling (through SMAD2/SMAD3) and pathways that include enzymes involved in chromatin remodeling32 were significantly and negatively correlated with the IFN-induced antiviral pathways (P = 0.033, rho = −0.63; Fig. 2c). Conversely, the expression of RFs and C/EBP-β pathways were positively correlated with one another (P = 0.016, rho = 0.71; Fig. 2c). These data, point toward TGF-β and TGF-β signaling as upstream modulators of chromatin remodeling and suppression of antiviral responses.
a, Quantification of TGF-β isoforms in plasma from combo-treated RMs pre-ATI who became CA-vDNAlo or CA-vDNAhi 24 weeks post-ATI by Meso Scale Discovery (MSD) platform. PCA highlighting significantly different (two-sided Wilcoxon rank-sum test; P < 0.05) levels of plasma cytokines pre-ATI between CA-vDNAhi and CA-vDNAlo RMs (left). Jitter plots highlighting the different levels of all TGF-β isoforms (TGF-β1, TGF-β2 and TGF-β3), IL-6 and MIP-1a pre-ATI in CA-vDNAhi and CA-vDNAlo RMs (right). Combo-treated RMs with cell suspensions available for the single-cell multiome analysis are represented. In the boxplots, the central line indicates the median, with hinges at the first and third quartiles and whiskers extending 1.5× interquartile range. Groups (CA-vDNAhi n = 3 and CA-vDNAlo n = 4) were compared to a two-sided Wilcoxon rank-sum test. The BH method (*Padjusted < 0.05) was used to adjust for multiple testing. b, GSEA was used to test the enrichment of pathways (MSigDB Collections C2, C7 and Hallmark) that distinguished CA-vDNAlo from CA-vDNAhi RMs. The heatmap illustrates NES of pathways related to epigenetic modifications, SMAD2/3 targets, innate antiviral signatures, and inflammation (P < 0.05). The NES scale ranges from low (blue) to high (red). Columns represent relative enrichment in CA-vDNAhi (left) and CA-vDNAlo RMs (right), whereas rows represent pathways. n = 10 combo-treated RMs (6 CA-vDNAhi and 4 CA-vDNAlo). c, Network inference (Cytoscape and Genemania) was used to plot the significant LEGs of SMAD targets and epigenetic signatures (blue nodes), and RFs and C/EBP-β target genes (red nodes) pre-ATI in LNMCs thar were downregulated and upregulated in CA-vDNAlo versus from CA-vDNAhi combo-treated RMs, respectively. Edges highlight gene–gene interactions. The table highlights the two-sided Spearman correlation coefficients and P values between the RF pathways and the other pathways represented in the gene network. n = 10 combo-treated RMs. d, An integrated model was generated in the mixOmics R package59 using plasma cytokine and bulk RNA-Seq data. This integrated network highlights the two-sided Spearman correlation between gene expression (blue circular nodes) and plasma cytokines (blue square nodes). The thickness of connecting lines represents the correlation coefficient (rho) associated with marker pairs. e, Experimental design. CD4+ T cells isolated from healthy donors (n = 6) were pretreated with TGF-β, with or without an aTGF-β antibody for 24 h, followed by stimulation with IFN-β (24 h). CD4+ T cells were then infected with HIV by spinoculation. HIV p24 levels were evaluated by flow cytometry 4 days post-infection. Figure created in BioRender Pereira Ribeiro, S. https://biorender.com/qr7isln (2026). f, Bar graph depicting MFI of phosphorylated SMAD2/3 in live CD3+ cells from healthy human donors following stimulation with TGF-β and stimulation with TGF-β in presence of anti-TGF-β antibody. Statistical significance was assessed using a two-tailed paired t-test. Error bars represent mean ± s.e.m. g, Bar graph depicting MFI of phosphorylated STAT1 in CD3+CD4+CD45RA− cells from healthy human donors under unstimulated conditions, stimulation with IFN-β, and pre-stimulation with TGF-β followed by stimulation with IFN-β. h, Bar graph depicting frequencies of p24+ cells in cultures of CD3+CD4lo cells under different stimulation conditions (UNS versus TGF-β versus TGF-β + aTGF-β antibody). i, Bar graph depicting frequencies of p24+ cells in cultures of CD3+CD4lo cells under different stimulation conditions (IFN-β versus IFN-β + TGF-β versus TGF-β + aTGF-β antibody + IFN-β). Group comparison was performed with one-way analysis of variance (ANOVA) with Tukey’s multiple-comparisons test (g–i). Error bars represent mean ± s.e.m. *P < 0.05, **P < 0.005; ***P < 0.0005; ****P < 0.0001; NS, not significant; PCA, principal-component analysis; UNS, unstimulated.
To confirm experimentally that plasma TGF-β was upstream of the increased levels of HDAC11, shown to correlate with the frequency of cells with higher levels of cell-associated viral DNA (CA-vDNAhi) in a subset of the combo-treated RMs post-ATI, we treated freshly isolated memory CD4+ T cells from healthy human donors (n = 5) with plasma samples obtained from one CA-vDNAhi RM (RFl17, TGF-β concentration of 11,873 pg ml−1) and one CA-vDNAlo RM (RRh17, TGF-β concentration of 2,084 pg ml−1). Experimental controls included plasma from an uninfected healthy RM (TGF-β concentration of 1,260 pg ml−1), recombinant TGF-β (rTGF-β; 2,000 pg ml−1) and no plasma. Flow cytometry analysis revealed that higher concentrations of TGF-β in plasma from the CA-vDNAhi RM induced significantly higher levels of phosphorylation of SMAD2/3 (pSMAD2_3) in memory CD4+ T cells compared to the other treatments conditions (Extended Data Fig. 4a,b). Presence of anti-TGF-β (aTGF-β) significantly reduced the per-cell level of expression (median fluorescence intensity; MFI) of pSMAD2_3 in all conditions. After 16 h, plasma from the CA-vDNAhi RM led to significantly higher per-cell levels of expression of HDAC11 (Extended Data Fig. 4c,d) as compared to other treatment conditions. Of note, the capacity of aTGF-β to block the induction of HDAC11 expression was significantly higher when cells were cultured in the presence of plasma from the CA-vDNAlo RM (Extended Data Fig. 4e), reflecting the complex epigenetic network triggered by the plasma from the CA-vDNAhi RM previously observed in vivo (Figs. 2c and 5a). We extended our analysis to evaluate the infectivity of cells in these different conditions. Following in vitro infection with the dual-tropic HIV (p89.6), memory CD4+ T cells cultured in the presence of plasma from the CA-vDNAhi RM tended to exhibit higher HIV infection rates in terms of frequency of CD4− HIV p24+ T cells as compared to plasma from the CA-vDNAlo RM (P = 0.0568) (Extended Data Fig. 4f,g). Expression levels of HDAC11 before infection were significantly correlated with infection rates 4 days post-infection (Extended Data Fig. 4h). Taken together, these data confirm that plasma TGF-β induces the expression of HDAC11 resulting in higher HIV infection rates relative to controls, similar to what was observed in CA-vDNAhi RMs in vivo.
Validation of the interplay between TGF-β, HDACs and IFN antiviral activity
To demonstrate the capacity of TGF-β to dampen the antiviral activity of IFN signaling, memory CD4+ T cells from healthy human volunteers were cultured under various conditions (untreated; TGF-β only; IFN-β only; pre-stimulated with TGF-β followed by IFN-β, with or without aTGF-β) (Fig. 2e and Supplementary Fig. 3b). Treatment in vitro with aTGF-β significantly decreased TGF-β-induced phosphorylation of SMAD2/3 (pSMAD2/3; Fig. 2f). Pre-stimulation of CD4+ T cells with TGF-β for 24 h impeded the upregulation of pSTAT1 by IFN-β (Fig. 2g). The CD4+ T cells from different culture conditions were infected in vitro with HIV. After 4 days in culture, expression of HIV Gag p24 was evaluated by flow cytometry. Cultures treated only with TGF-β displayed significantly higher frequencies of p24+ cells after 4 days in culture compared to untreated cultures or cells cultured in the presence of aTGF-β (Fig. 2h). Cultures stimulated with TGF-β before IFN-β treatment exhibited significantly higher frequencies of p24+ cells compared to cells treated with IFN-β alone or cultures containing aTGF-β (Fig. 2i).
Based on the knowledge that histone deacetylases (HDACs) can block IFNs31, we next validated if the capacity of TGF-β to block antiviral responses was due to the induction of HDACs (Extended Data Fig. 5a,b). TGF-β treatment resulted in significantly increased expression of HDAC1 and HDAC11 in memory CD4+ T cells (Extended Data Fig. 5c–f), which was reduced by aTGF-β to similar levels as unstimulated controls. Conversely we monitored the capacity of HDAC inhibitors (HDACi) to restore antiviral pathways in cells treated with TGF-β ± aTGF-β. The HDACi panobinostat, SAHA (both pan-HDACi), romidepsin (potent HDAC1 and HDAC2 inhibitor) and SIS17 (selective HDAC11 inhibitor) were tested for their ability to reverse the impact of TGF-β on the suppression of innate IFN-induced antiviral pathways (Fig. 3a). TGF-β significantly induced the phosphorylation of pSMAD2/3, and IFN-β or IFN-γ significantly induced the phosphorylation of pSTAT1 in CD4+ T cells, compared to untreated conditions (Extended Data Fig. 6a). In addition, IFN-β induced significant upregulation of the per-cell level (MFI) expression of IRF1, 24 h post-stimulation, and this was suppressed in cells pretreated with TGF-β (Fig. 3b–f); however, when each HDACi was added along with TGF-β and cells were then stimulated with IFN-β, higher expression of IRF1 was detected, similar to what was observed with IFN-β alone (Fig. 3b–d and Extended Data Fig. 6b–d). Similar data were obtained following IFN-γ stimulation in combination with panobinostat (Extended Data Fig. 6e).
a, Experimental design. PBMCs from healthy donors (n = 5) were treated with SAHA, romidepsin (FK228) or SIS17 (HDAC11 inhibitor) for 2 h, and then treated with TGF-β for 24 h before stimulation with IFN-β (24 h). pSMAD2/3, pSTAT-1 and IRF1 were evaluated by flow cytometry 24 h later. Figure created in BioRender; Pereira Ribeiro, S. https://biorender.com/qr7isln (2026). b–d, Bar plots with individual data points showing the induction or suppression of IRF1 under various TGF-β, SAHA, romidepsin (FK228) or SIS17 (HDAC11 inhibitor), and IFN-β, stimulation conditions. Data are presented as mean ± s.e.m., with each dot representing an individual donor. Group comparisons were performed with two-way repeated-measures ANOVA, followed by Dunnett’s multiple-comparison test comparing each condition to the unstimulated control. **** P < 0.0001. e, Coexpression network (Cytoscape) illustrates a set of ISGs upregulated 2 h after the first dose of vorinostat administration compared to baseline (no administration) (n = 9). Red circular nodes indicate significantly upregulated genes (DEG analysis: LIMMA t.test, nominal P < 0.05). Colored edges represent gene–gene interactions. f, Heatmap illustrates gene expression measured 2 h post-vorinostat administration correlated to cell-associated HIV RNA (white to black gradient) measured on day 84 (70 days after discontinuation of vorinostat) (n = 9). Rows represent genes (FDR, P < 0.05) and columns represent individual samples. Standarized gene expression (z-score) is represented in blue-yellow-red gradient, where blue represents low relative expression and red high relative expression. Correlation was calculated with a two-sided Pearson correlation (R2) method and P values were adjusted for multiple comparisons with a BH procedure. Data from Elliott et al.33 (e,f). LIMMA, linear models for microarray and RNA-seq data.
We confirmed the ability of HDACi to induce the expression of antiviral pathways by using bulk transcriptomic data from a clinical trial of PWH on ART who received the vorinostat33, an inhibitor of class I, II and IV HDACs. Induction of IFN signaling, including receptors (IFNAR2 and IFNGR1), upstream modulators (STAT1 and STAT3) and downstream effector molecules (IRF7, MX1/2 and APOBEC3F; Fig. 3e) was observed as early as 2 h post-vorinostat treatment (before its impact on virus reactivation in the periphery) and persisted up to the seventh dose of vorinostat (day 7 + 2 h). Of note, regression analysis revealed that expression levels of ISGs 2 h post-vorinostat administration were inversely correlated with CA-vRNA levels in the study participants at the end of follow-up (84 days following initiation of vorinostat treatment; Fig. 3f).
Altogether, these data demonstrate the inhibitory capacity of TGF-β to modulate the expression of antiviral pathways through the upregulation of HDACs, which can be reversed by the blockade of the TGF-β signaling cascade or by using US Food and Drug Administration (FDA)-approved drugs such as HDACi.
Higher gene expression of IRF/STAT1 targets pre-ATI in CA-vDNAlo RMs
Herein we show that combo treatment in a subset of combo-treated RMs provided a poised cellular environment, which was associated with the control of VL and reservoir decay; this parallels mechanisms involved in trained immunity34,35, a poised state of innate immune antiviral responses. We used the multiome platform that combines single-cell (sc) RNA-seq and ATAC-seq (Extended Data Fig. 7a,b and Supplementary Fig. 3c) on LNMCs from combo-treated RMs to (1) identify gene signatures in myeloid and lymphoid cell subsets that triggered the IFN antiviral response leading to lower CA-vDNA levels, and (2) determine whether the transcriptional differences were associated with changes in chromatin accessibility of the loci that control innate antiviral immunity36. Cells were annotated as previously described37,38 (Extended Data Fig. 7c–e). Expression of antiviral genes was observed across all innate (monocytes, dendritic cells (DCs), plasmacytoid dendritic cells (pDCs) and follicular dendritic cells (FDCs)) and adaptive (naive, memory and effector CD4+ and CD8+ T cells and B cells) immune subsets (Extended Data Fig. 7f and Supplementary Table 7). While only a small fraction (<2%) of CD4+ and CD8+ effector T cells expressed messenger RNA for IFN-γ (Fig. 4a), significantly higher frequencies of cells (>47%) expressed target genes of IFN-γ/STAT1 signaling, including CD4+ T cells and myeloid cells (Fig. 4b), which are highly susceptible to HIV/SIV infection. The spreading of IFN-γ signaling to other cell subsets in LNMCs was confirmed by using cell–cell communication analysis39 (Extended Data Fig. 7g). We identified TH1, effector CD8+ T cells, cycling CD4+ and CD8+ T cells and TFH cells in LNMCs as senders of IFN-γ, whereas the receivers of IFN-γ signaling included all CD4+, CD8+ and B cell subsets and to a lesser extent macrophages, FDCs and pDCs. Reduced chromatin accessibility and concomitant poor expression of type-I IFN genes was shown in LNMCs, suggesting that IFN-γ produced by CD4+ and CD8+ T cells was the driver of the upregulation of antiviral ISG (Extended Data Fig. 7h).
a,b, Uniform Manifold Approximation and Projection (UMAP) showing the normalized per-cell expression of IFN-γ and antiviral ISGs module scores in lymphoid and myeloid cells sourced from LNMCs, respectively. The antiviral ISG module was calculated using the AddModuleScore function from Seurat and a list of 50 antivirals ISGs sourced from Schoggins and Rice60. c, UMAP plot with CD4+ and CD8+ T cell subsets. CD4+ and CD8+ T cells from LNMCs were re-clustered and manual annotation of the clusters was performed based on expression of marker genes (Extended Data Fig. 7e). d, Over-representation analysis (ORA) of DEGs in T cell subsets between CA-vDNAlo and CA-vDNAhi RMs. Red dots represent pathways upregulated and blue dots represent pathways that are downregulated in CA-vDNAlo RMs. The size of the dots reflects the number of DEGs in each pathway. The ‘EC_vs_nonHIC’ pathways include DEGs from CD4+ and CD8+ T cells and monocytes of ECs obtained from dos Santos et al.42. The ‘Tirosh_ET_AL’ pathway is composed of genes from Tirosh et al.61. e, Heatmap showing the expression fold change of DEGs from antiviral (left) and TNF signaling pathways (right) in CD4+ and CD8+ T cell subsets in CA-vDNAlo RMs compared to CA-vDNAhi RMs. White to red color gradient indicates upregulated genes and white to blue color gradient indicates downregulated genes. f, UMAP plot showing expression levels per cell of the antiviral ISGs upregulated in CA-vDNAlo RMs OAS2, DDX60 and EIF2AK2 and the overall antiviral ISG module score in T cell subsets as described in b. Cells from all RMs are represented. g, UMAP plot showing the clustering of myeloid cells and FDCs from LNMCs from combo-treated RMs pre-ATI. Manual annotation of the clusters was performed based on expression of marker genes. h, ORA of DEGs in myeloid cell subsets and FDCs between CA-vDNAlo and CA-vDNAhi RMs. Red dots represent pathways upregulated and blue dots represent pathways that are downregulated in combo-treated CA-vDNAlo RMs. The size of the dots reflects the number of DEGs in each pathway. i, Heatmap showing the expression fold change in expression of DEGs from antiviral pathways (left) and TNF signaling pathways (right) in subsets of myeloid cells and FDCs between combo-treated CA-vDNAlo and CA-vDNAhi RMs. White to red color gradient indicates upregulated genes and white to blue color gradient indicates downregulated genes. j, UMAP plot showing expression levels per cell of the antiviral ISGs upregulated in CA-vDNAlo RMs DDX60, IFI44L, IFI6 and the overall antiviral ISG module score in myeloid cells and FDCs subsets as described in B. Cells from all RMs are represented. Only pathways with an FDR-corrected P < 0.05 and containing at least three DEGs were represented in the dot plots from ORA analysis (d–h). DEGs were defined as genes with an FDR P < 0.1 in each cell subset in the comparison of CA-vDNAlo versus CA-vDNAhi RMs (two-sided Wilcoxon rank-sum test). In the heatmaps, genes that did not meet the significance threshold (FDR-adjusted P < 0.1) are shown as gray boxes. n = 7 combo-treated RMs for all the figures in the panel. FC, fold change; CA-vDNAhi, RBf17, RFl17, Rym17; CA-vDNAlo, RBv17, RNy16, RWs17, RRh17.
We subclustered T cells (Fig. 4c) and myeloid cells (Fig. 4g) to identify genes and pathways that discriminated CA-vDNAlo and CA-vDNAhi RMs. Pathways involving the IFN-γ and IFN-α response, HIV RFs and antiviral ISGs (Fig. 4d), were significantly upregulated in T cell subsets (naive CD4+ and CD8+ T cells, regulatory T (Treg) cells and follicular helper T (TFH) cells; Fig. 4d) and myeloid cells (macrophages and pDCs; Fig. 4h) from CA-vDNAlo RMs (Supplementary Table 8). Representative DEGs of the ISG antiviral pathways for both lymphoid and myeloid cell lineages are shown (OAS2, DDX60 and EIF2AK2; Fig. 4e,f,i,j).
Conversely, T cells and myeloid cells from CA-vDNAhi RMs were enriched in TNF-induced pathways via NF-κB signaling (Fig. 4d,e,h,i), as well as in genes downstream of AP-1 and associated with the inflammatory response (TNF, FOS, JUN, DUSP1 and REL) (Extended Data Fig. 8a,b). Of note, T cells from both CA-vDNAlo and CA-vDNAhi RMs expressed pathways and effector molecules (GZMB, PRF1 and TBX21) involved in cell-mediated immune responses (ZAP70 and FYN) as compared to ART-control RMs, when VL was undetectable. These effector molecules were associated with the expression of IFN-γ and were negatively correlated with CA-vRNA levels (Extended Data Fig. c–e).
Higher chromatin accessibility for IRF/STAT1 binding sites pre-ATI in CA-vDNAlo RMs
scATAC-seq data obtained from LNMCs showed that enhanced transcriptional activity of IFN loci was concomitant with increased chromatin accessibility of STAT1, IRF3 and IRF7 in T cells (TFH, cycling, effector CD4+ and CD8+ T cells) from CA-vDNAlo RMs (Fig. 5a,b). This was associated with increased expression of their target genes (Extended Data Fig. 9a–c) along with elevated expression of C/EBP-β targets (IRF9 and LTBP1), and open chromatin state of IL-6, another C/EBP-β target (Extended Data Fig. d–f). C/EBP-β is downstream of type-II IFN and triggers IL-6, which in turn induces IRF9, resulting in the amplification of IFN responses40, whereas LTBP1 is known to maintain TGF-β in its latent form41. FOS and HDAC9, which are downstream of TGF-β signaling, were upregulated in CA-vDNAhi RMs (Extended Data Fig. 9d–f).
a, Average difference of TFs motif chromatin accessibility (motif activity scores), in CD4+ and CD8+ T cell subsets from LNMCs isolated from CA-vDNAlo RMs versus CA-vDNAhi combo-treated RMs pre-ATI. TFs from the STAT, IRF and AP-1 families are indicated by arrows. Differentially active motifs were identified by performing a two-sided Wilcoxon rank-sum test on the average differences of scaled TFs motif activity by cell subset and group. Only motifs with an FDR-adjusted P < 0.05 in at least two cell subsets are shown. b, UMAP representation of chromatin accessibility of motif-binding sites for SMAD2/3 and FOS/JUN and IRF7 and STAT1 in T cells subsets from CA-vDNAlo RMs pre-ATI. Lower levels of chromatin accessibility (SMAD2 and FOS/JUN) are in blue and higher levels of accessibility (IRF7 and STAT1) are in red. Cells from all RMs are represented. c, Average differences in TFs activity scores in myeloid cells and FDCs from LNMCs from CA-vDNAlo RMs versus CA-vDNAhi combo-treated RMs pre-ATI. TFs motif activity was calculated and compared between groups as described for the T cell subsets (a). The top five TFs per subset based on highest absolute fold change are indicated by arrows. d, UMAP representation of chromatin accessibility of motif-binding sites for IRF4, IRF7, FIS and RFX2 in myeloid cells and FDCs from combo-treated CA-vDNAlo RMs pre-ATI. Lower levels of chromatin accessibility are in blue and higher levels of accessibility are in red. Cells from all RMs are represented. n = 7 combo-treated RMs for all the figures in the panel.
T cell subsets from CA-vDNAlo RMs presented decreased chromatin accessibility of genes of the AP-1 family (Fos/Jun and MAF) and SMAD2/3 TF binding sites pre-ATI (Extended Data Fig. 9g–i). The comparative accessibility of motif-binding sites for these TFs in T cells is shown in Fig. 5a,b. In the myeloid compartment, the binding sites for IFN-associated TFs (IRF4, IRF7, IRF8 and IRF9) were accessible in macrophages in LNMCs from CA-vDNAlo RMs (Fig. 5c,d). In contrast, chromatin accessibility for IRF7 and STAT1 was observed mostly in T cells that exhibited low chromatin accessibility for the SMAD2–SMAD3/AP-1 complex. This highlights the reciprocal interplay between IFNs and TGF-β in modulating TF activity and chromatin accessibility of their target genes in innate and adaptive immune cells, resulting in the differential modulation of antiviral versus TGF-β pathways in CA-vDNAlo and CA-vDNAhi RMs.
Protective signatures found in CA-vDNAlo RMs are expressed by ECs
To demonstrate the generalizability and clinical relevance of the signatures observed in CA-vDNAlo RMs, we extended our findings to HIV ECs who control VL in the absence of ART (ECs, <75 copies RNA per ml). We obtained the gene signatures upregulated in ECs at the single-cell level in peripheral blood CD4⁺ T cells, CD8⁺ T cells and monocytes, as compared to non-HIV controllers (non-HIC, n = 30)42. We assessed whether these signatures were observed in T cells (Fig. 4d) and myeloid cells (Fig. 4h) from LNMCs of CA-vDNAlo and CA-vDNAhi RMs. Notably, 40.6% of genes that were overexpressed in ECs were also upregulated in CA-vDNAlo RMs, including antiviral ISGs, HIV RFs, and genes associated with IFN-α and IFN-γ responses, (such as ISG15, MX1, IFIT3 and IFI6), all of which are involved in long-term control of plasma VL and low levels of CA-vDNA in LNMCs from combo-treated RMs (Fig. 6a,c and Supplementary Table 9). Conversely, downregulated gene signatures in ECs versus non-HIC overlapped (48.5%) with signatures found in T cells and myeloid cells from LNMCs of CA-vDNAhi RMs, and predominantly included target genes of the TNF signaling cascade via NF-κB and mTORC1, AP-1 targets and TGF-β/SMAD-related genes (ANXA1, BHLHE40 and JUN) (Fig. 6b,d, respectively). These findings highlight the interplay between TGF-β and C/EBP-β target genes and the relevance of the suppression mediated by TGF-β signaling for the proper generation of IFN-induced antiviral signatures (Extended Data Fig. 10).
a, Upregulated pathways in T cell subsets from CA-vDNAlo RMs and ECs. Larger nodes represent pathways, and the size of the nodes reflects the number of DEGs in each pathway. Smaller nodes represent shared LEGs and labeled nodes indicate genes from ECs that overlap with the DEGs from the CA-vDNAlo RMs. IRF9, a target of C/EBP-β is indicated in red. Colors inside the nodes indicate the cell subset in which the genes or pathways are statistically significant. b, Downregulated pathways in T cell subsets from CA-vDNAlo RMs and ECs. Targets of C/EBP-β are labeled in blue. c, Upregulated pathways in myeloid subsets and FDCs from CA-vDNAlo RMs and ECs. d, Downregulated pathways in myeloid subsets and FDCs from CA-vDNAlo RMs and ECs. Significant DEGs and pathways were obtained from analysis described in the legends of Fig. 4. Additional information about ECs’ gene signatures is provided in the legends of Fig. 4d and Methods section. n = 7 combo-treated RMs for all the figures in the panel.
Discussion
Herein, we identified and provided experimental validation for molecular pathways and effector mechanisms associated with the decay of the size of the SIV reservoir (CA-vDNA) observed in 40% of combo-treated RMs. Our findings suggest that the triggering and amplification of the antiviral pathways by IFN-γ and C/EBP-β/IL-6, respectively, conferred protection from infection of bystander CD4+ T cells upon ATI, resulting in the CA-vDNAlo phenotype post-ATI. In contrast, higher plasma levels of TGF-β and the upregulation of the SMAD signaling pathway led to the epigenetic reprogramming of cells and the blockade of antiviral signaling pathways, resulting in the CA-vDNAhi phenotype.
Coordinate transcriptional activity of IRFs and C/EBP-β is required for the control of VL and SIV DNA decay
Combo treatment was the only study arm that showed the transcriptional upregulation of IRF7 and STAT1, two TFs that regulate IFN production and signaling in CD4+ T cells and myeloid cells20. The fact that the IFN-γ response and production of this cytokine were induced only in combo-treated RMs suggests that addition of aPD-1 synergized with aIL-10 to promote transcription and chromatin accessibility of ISGs and increased frequencies of ISG+ monocytes and T cells, including IRF/STAT target genes, resulting in lower frequencies of cells with SIV RNA and DNA. Our results suggest a role for C/EBP-β in promoting the innate antiviral immune response observed in CA-vDNAlo RMs. C/EBP-β is triggered by type-II IFN40 which in turn activates IL-6, which was upregulated only in CA-vDNAlo RMs. IRF9, another C/EBP-β target gene, is an upstream regulator of the expression of type-I and -II IFN genes29. Moreover, heterodimerization of C/EBP-β with AP-1 leads to the inhibition of the proinflammatory response triggered by AP-1 (ref. 43). While we do not show evidence for the AP-1/CEBPβ heterodimerization, concomitant expression of C/EBP-β and AP-1 was shown to be associated with inhibition of the expression of AP-1 target genes43, similar to our findings in CA-vDNAlo RMs. The synergy between C/EBP-β and IL-6 provides a mechanism whereby a few LNMCs (2%) that produced type-II IFN could induce expression of ISGs in many (>47%) LNMCs. This synergy between type-II IFN and IL-6 likely represents a key mechanism that leads to the protection against viral dissemination post-ATI that was observed in CA-vDNAlo RMs.
TGF-β blunts the IFN response
CA-vDNAhi RMs presented elevated plasma levels of TGF-β and exhibited higher chromatin accessibility of SMAD2/3 target genes pre-ATI; this was associated with molecular pathways that modulate chromatin structure (SMARCA4), accessibility (HDACs and BRD4) and acetylation (HDACs) of histones. This TGF-β signaling suppresses the production of type-I IFNs31, resulting in the maintenance of latent HIV reservoirs. Addition of an HDACi augmented IFN signaling and transcription of antiviral ISGs. These data support a model whereby expression of TGF-β triggers HDACs thereby inhibiting expression of ISGs and preventing the development of innate antiviral effector functions that can confer refractoriness to infection of CD4+ T cells. Panobinostat and romidepsin, both HDACi, upregulated a minor set of ISGs with antiviral function albeit at much lower levels (at least tenfold) than IFN-β, the positive control44. Recently, it was showed45 that panobinostat synergize with IFN-α to induce the upregulation of ISGs with antiviral functions (APOBEC, OAS and IFIT), albeit at lower levels than IFN-α. PWH who were infused with panobinostat and IFN-α showed a decay in levels of intact proviruses45 and selective inhibition of viruses at sites of H3K27ac, the histone mark that promotes enhanced chromatin accessibility. These findings are consistent with our results, although the mechanisms underlying the synergy between panobinostat and IFN-α and the inhibition of viral integration in regions of open chromatin remain to be dissected.
Downregulation of AP-1 and NF-κB and upregulation of antiviral trained immunity in CA-vDNAlo RMs
Our findings underscore (1) the importance of IFNs and the expression of ISGs by cells that are the main targets of HIV/SIV infection; and (2) the importance of downregulation of AP-1 and NF-κB in immune cells from CA-vDNAlo RMs. IL-1 and TNF are well-known drivers of HIV replication, hence their downregulation in CA-vDNAlo RMs provides a mechanism that can lead to decreased levels of viral seeding post-ATI. In CA-vDNAhi RMs, heightened levels of TGF-β result from the expression of SMAD2/3 and AP-1, which inhibit the expression of antiviral ISGs. We observed a coordinate transcriptional regulation of genes commonly targeted by SMAD and AP-1 complexes and showed that CA-vDNAhi RMs exhibited heightened activity of proinflammatory pathways, including TNF signaling and hypoxia. AP-1 and NF-κB signaling are critical for the expression of IL-1β and NLRP3, two of the key effector molecules in proinflammatory pathways. TGF-β, which is produced by multiple lineages of white blood cells, but predominantly by CD4+ Treg cells, may provide the balance between these distinct pathways that are present in the two subgroups of combo-treated RMs.
Our findings are generalizable and clinically relevant, as the signatures observed in CA-vDNAlo RMs overlapped with those specific to HIV ECs46. This imprinting, induced in vivo by combo treatment, protected bystander CD4+ T cells from infection by rebounding SIV post-ATI. Wimmers et al.47 and Lee et al.48 showed the importance of epigenetic reprogramming and transcriptional regulation of TF loci that control type-I and type-II IFN production and their role in inducing a state of refractoriness to SARS-CoV-2 infection in vivo and to Zika and Dengue virus infection in vitro. Their results and ours highlight substantial differences between innate immune pathways that confer resistance to bacterial and fungal infections (AP-1-driven pathways) and those involved in resistance to viral infections (IRFs and STAT1-driven pathways)35.
Many immune interventions previously evaluated for their ability to reduce the HIV reservoir were focused on stimulating T cell-mediated immune responses49. Herein we showed the critical role of an innate, trained immunity antiviral program in shaping immune responses that can inhibit viral replication and dissemination and stimulate the development of long-lasting and effective adaptive immune responses. CA-vDNAhi and CA-vDNAlo RMs received the same intervention and were similarly capable of developing adaptive effector functions to control VL rebound post-ATI18. Of note, TGF-β played a key role in modulating signatures of trained immunity that are associated to SIV reservoir decay. The negative impact of TGF-β on radiation cancer immunotherapy in mouse models and on anti-PD-1 immunotherapy in cancer patients was reported previously50,51. The mechanisms underlying this TGF-β-mediated interference with immune interventions have not been fully defined, but TGF-β was shown to suppress type-I and type-II IFN signaling and adaptive immune responses52,53,54; of note, TGF-β inhibitors in combination with PD-1-blockade have shown partial efficacy in clinical trials in patients with cancer55. Moreover, a TGF-β receptor inhibitor was reported to promote SIV latency reactivation in a subset of SIV-infected, ART-treated RMs; mechanisms mediating this reactivation were not defined12. Our studies provide a strong rationale and a clear path toward clinical interventions that combine PD-1 and HDAC blockade and/or Interferon agonists to restore immune responses and induce a decay in HIV reservoir size.
Mouse models and clinical trials showed that HDACi and aPD-1 synergize to clear tumors56. Wen et al.57 showed that aPD-1 and chidamide (an HDACi) enhance chemokine expression by T cells and augment the IFN-γ response in immunocompetent preclinical models of natural killer T cell lymphoma. Wang et al.58 suggested that the combination of an aPD-1, an HDACi and aVEGF antibodies promoted progression-free survival for 18 weeks in 43% of patients with unresectable, advanced or metastatic colorectal cancer. Based on results in the field of cancer immunotherapy and our own results, we propose that interventions, including HDACi and or IFN agonists that (1) reverse immune dysfunction; (2) stimulate antigen specific immune responses18; (3) enhance the production of IFNs pre-ATI; and (4) inhibit the production and downstream functions of TGF-β, can intercept viral rebound and promote viral reservoir decay post-ATI.
Study limitations
In this study, we identified mechanisms downstream of the dual blockade of IL-10 and PD-1 that led to long-term VL control post-ATI in 90% of treated RMs18 and decay of the size of the SIV reservoir in 40% of animals. This intervention leads to virological, immunological and molecular end points similar to those observed in HIV ECs, raising the likelihood that it can be used as a therapeutic strategy for HIV cure. Unfortunately, combo treatment cannot be promptly translated to human clinical trials due to the side effects observed in treated RMs18; however, the data presented herein dissect the mechanisms of action of this intervention and identify target molecules and pathways that are FDA-approved and readily translatable to human clinical trials. Our findings set a benchmark and a path for clinical interventions that could help achieve a cure for HIV
Methods
Materials availability
This study used novel deimmunized aIL-10 (JES3.12G8) and aPD-1 mAbs (1B8 LC3/HC1), which are proprietary reagents developed by Merck & Co. Both aIL-10 and aPD-1 mAbs were the subject of material transfer agreements restrictions as these biologics are currently under investigation in human clinical trials.
Study approval
This study was approved by the Emory University Institutional Animal Care and Use Committee via permit no. 201800047. Experiments were conducted following guidelines set forth by the National Institutes of Health (NIH) and Animal Welfare Act in regard to the housing and welfare of laboratory RMs. All possible efforts were taken to minimize the pain and discomfort experienced by RMs.
Experimental model
A total of 28 Indian-origin, specific-pathogen-free RMs (Macaca mulatta) (25 males and 3 females) were sourced from the Emory National Primate Research Center (ENPRC) colony and single-housed in an animal BSL-2 facility at ENPRC, as previously described1. RMs were between 38 and 50 months old at the time of infection and Mamu-B*07− and Mamu-B*17−. Some RMs were Mamu-A*01+. RMs were infected, treated and followed up for virological and immunological outcomes, as previously described18. The analysis presented herein was focused on immune responses and virological readouts in plasma, PBMCs and LNMCs pre-ATI and at 24 weeks post-ATI. Sample collection and processing, formulation of the aIL-10 and aPD-1 mAbs, pharmacokinetics (PK) and pharmacodynamics (PD) of aPD-1 and aIL-10 mAbs, levels of antidrug antibodies (ADAs), plasma IL-10 levels, aPD-1 receptor occupancy, complete blood counts, blood chemistry, SIV VL, measurement of CA-vRNA and DNA levels6, and measurement of 2-LTR circles were all performed as previously described18.
Bulk RNA-seq
A total of 100,000 PBMCs or LNMCs from RMs were lysed directly into 700 μl of QIAzol reagent. RNA was isolated using the RNeasy Micro kit (QIAGEN) with on-column DNase digestion. RNA quality was assessed using an Agilent Bioanalyzer and total RNA was used as input for cDNA synthesis using the Clontech SMART-Seq v4 Ultra Low Input RNA kit (Takara Bio) according to the manufacturer’s instructions. Amplified cDNA was fragmented and appended with dual-indexed barcodes using the Nextera XT DNA Library Preparation kit (Illumina). Libraries were validated by capillary electrophoresis on an Agilent 4200 Tape Station, pooled at equimolar concentrations and sequenced on an Illumina NovaSeq6000 at 100SR, yielding 20–25 million reads per sample. The quality of reads was evaluated using Fast QC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Reads were aligned using STAR (v.2.7.3)62. The STAR index was built by combining genome sequences for M. mulatta (Mmul_10 Ensembl release 100). Transcript abundance estimates were calculated internal to the STAR aligner using the htseq-count algorithm63. The ReadsPerGene files were used to generate counts in the htseq format using a custom script that also converted the Ensembl ID to gene names using the gtf file. These files were imported in DESeq2 using the DESeqDataSet- FromHTSeqCount function. DESeq2 was used for normalization64, producing both a normalized read count table and a regularized log expression table. Regularized log expression values were obtained using the rlog function with the parameters blind = FALSE and filtType = parametric. Thresholds of Padj < 0.05, fold change > 1.5 and log-fold-change standard error < 1 were used to identify genes with statistically significant differences in expression levels. The input for GSEA was the regularized log expression values obtained from DESeq2, which was filtered to remove genes with mean expression levels of 0. Regularized log expression values were also used to generate heatmaps using the Complex Heatmap R library65. GSEA was performed using a compiled set of pathways from public databases, including MSigDB v.5.1 (http://software.broadinstitute.org/gsea/msigdb/) and blood cell marker signatures. The GSEA Java desktop program was downloaded from the Broad Institute (http://www.broadinstitute.org/gsea/index.jsp) and used with GSEA preranked module parameters (number of permutations = 1,000; enrichment statistic = weighted; seed for permutation = 111; and 10 ≤ gene set size ≤ 5,000). We used the Dynet Analyzer application implemented in Cytoscape v.3.6.0 to generate gene interacting networks that highlight overlapping genes between the different enriched modules. SLEA66 was used to investigate the enrichment of pathways in individual RMs upon the different interventions. In brief, the expression of all genes in a specific pathway was averaged across samples and compared to the average expression of 1,000 randomly generated gene sets of the same size. The resulting z-score was then used to reflect the overall perturbation of each pathway in each individual sample. Data were visualized using ggplot2 (v.3.3.2) in RStudio (v.1.4.1103) with custom code.
Construction of single-cell multiome (GEX and ATAC) libraries
Samples from CA-vDNAhi and CA-vDNAlo combo-treated RMs were used as available: CA-vDNAhi: RBf17, RFl17, Rym17; CA-vDNAlo: RBv17, RNy16, RWs17, RRh17. Single-cell multiome ATAC and gene expression (GEX) libraries were prepared using the Chromium Single Cell Multiome ATAC+ Gene Expression platform (10x Genomics) with a target nuclei recovery of 8,000 nuclei. Isolated nuclei were transposed and partitioned into Gel Beads-in-emulsion (GEMs) using the 10x Chromium Controller and Next GEM Chip J. ATAC and GEX libraries were generated from the same pool of preamplified transposed DNA/cDNA. Representative traces and quantitation of both libraries were determined using Bioanalyzer High Sensitivity DNA Analysis (Agilent). ATAC libraries were sequenced on an Illumina NovaSeq S2 with a sequencing depth of 25,000 read pairs per nucleus and GEX libraries were sequenced on an Illumina NovaSeq S4 with a sequencing depth of 20,000 read pairs per nucleus. The 10x Barcodes in each library type were used to associate individual reads back to the individual partitions and thereby to each single nucleus.
Analysis of single-cell multiome data (scRNA-seq and scATAC-seq)
Raw reads from 10x Chromium Single Cell Multiome ATAC and Gene Expression libraries were aligned to the reference M. mulatta genome (Mmul_10) using the Cell Ranger ARC Count v.2.0.2 pipeline. The aggregated counts file was then imported into Seurat v.5.01. Quality control parameters were applied to define high-quality cells. These parameters included: RNA counts between 500 and 25,000, detected genes > 300, mitochondrial reads < 25%, fragment counts between 1,000 and 70,000, a ratio of mononucleosomal to nucleosome-free fragments < 2, transcription start site enrichment score > 2 and fragments in peaks > 20%. Following the removal of low-quality cells, DoubletFinder was used to identify and exclude doublet cells. 56,193 high-quality singlets (RBf17: 7,805, RBv17: 9,561, RFl17: 7,146, RNy16: 6,191, RRh17: 8,820, RWs17: 9,618, RYm17: 7,052) were used in downstream analysis. Peak calling was performed using the CallPeaks function. Data normalization and dimensional reduction were performed independently on the gene expression (RNA) and chromatin accessibility (ATAC) datasets and then integrated using the weighted-nearest-neighbor (WNN) method. Clustering was performed on the integrated UMAP dimension reduction and manual annotation was performed using cell type-specific gene markers. CD4+ and CD8+ T cells were further re-clustered to improve the identification of specific subsets of T cells. DEGs between conditions were identified per cell type using the two-sided Wilcoxon rank-sum test, requiring a minimum cluster cell frequency of 1%, and an adjusted P value (FDR) < 0.1. Pathway ORA was performed using the clusterProfiler R package (v.4.18.2) with gene sets from MSigDB Hallmark collection. In addition, DEGs (log2 fold change > 0.25 and FDR P value < 0.05) identified in peripheral blood CD4⁺ T cells, CD8⁺ T cells and monocytes, from HIV ECs compared to non-HICs, as reported by dos Santos et al.42 were included in the analysis. Gene sets derived from these analysis were labeled ‘EC_vs_nonHIC’. Similarly, the Tirosh et al. gene set refers to genes described in Tirosh et al.61. For ATAC data, a per-cell motif-binding site activity score was calculated using chromVAR, utilizing a collection of 746 TFs from the JASPAR database67. Differences in motif activity per time point and group were identified using the FindMarkers function with an adjusted P value (FDR) < 0.05.
Flow cytometry
Cryopreserved PBMCs and LNMCs (106 per test) were thawed and stained with anti-rhesus or anti-human mAbs that are known to be cross-reactive with RMs cells and were validated in databases maintained by the Nonhuman Primate Reagent Resource. An antibody panel to evaluate antiviral responses was used to stain PBMCs and LNMCs pre-ATI and included CD95 BV605, BioLegend cat. no. 305628, Clone DX2; CD3 BUV395, Becton-Dickinson (BD) cat. no. 564117, Clone SP34-2; Live/Dead BV510, BD cat. no. 564406; CD8 BUV496, BD cat. no. 612942, Clone RPA-T8; CD28 BUV737, BD cat. no. 612815, Clone CD28.2; CD14 BV786, BD cat. no. 563698, Clone M5E2; CD16 BUV661, BD cat. no. custom 3G8; CD4 PerCP/Cy5.5, BD cat. no. 552838, Clone L200; pSTAT3 PE, BD cat. no. 558557, Clone 4/PSTAT3; pIRF7 (pS477/pS479) A647, BD cat. no. 558630, Clone K47-671; p38 (pT180/pY182) PE-Cy7, BD cat. no. 560241, Clone 36/p38; pStat5 PE-CF594, BD cat. no. 562501, Clone 47/Stat5(pY694); pIRF3, Abcam cat. no. ab138449, polyclonal; pStat1 AF488, BD cat. no. 612596, Clone K51-856; and donkey anti-rabbit BV421, BioLegend cat. no. 406410, polyclonal Ig. Cell surface staining was performed at 37 °C for 20 min. Staining was stopped by adding 50 μl of cold 1× TFP Fix/Perm Buffer to the cells (BD cat. no. 565575) and mixing gently. Cells were then fixed and permeabilized for 40 min at 2–8 °C. Cells were washed and rehydrated twice (500g for 5 min) in 200 μl of PBS + 2% fetal bovine serum (FBS) buffer. Then, 50 μl of ice-cold Perm Buffer III was added, cells were mixed gently and incubated on ice for 15 min. Cells were then washed twice with 200 μl of 1× TFP Perm/Wash Buffer and all residual buffer was removed. Intracellular staining was performed by adding mAbs to 25 μl of master mix diluted in 1× TFP Perm/Wash Buffer, followed by gentle mixing. Cells were incubated for 30 min at 2–8 °C and washed with 200 μl 1× TFP Perm/Wash Buffer by centrifugation at 500g for 5 min at 2–8 °C. Finally, cells were resuspended in 100 μl of PBS 2% FBS. Acquisition was performed on a minimum of 100,000 live cells on an A5 Symphony flow cytometer driven by BD FACSDiva software (BD Biosciences).
Assays for validation
IFN signaling protects bystander CD4+ T cells from HIV infection in vitro
Memory CD4+ T cells were isolated from healthy human donors (n = 6). Cryopreserved PBMCs were thawed and rested overnight in AIMV medium + 10% Serum Replacement (Corning Life Sciences) + 10 mM HEPES. Memory CD4+ T cells were isolated using the EasySep Human CD4+ T Cell Enrichment kit (STEMCELL Technologies). One half of memory CD4+ T cells was stimulated for up to 16 h with 5 ng ml−1 of IFN-β (Peprotech) while the other half remained unstimulated. These cells were further divided into two pools: unlabeled CD4+ T cells and CD4+ T cells labeled with CTV. Unlabeled CD4+ T cells were infected with HIV-1 (p89.6 dual-tropic strain). After infection, cells were cultured in the presence of saquinavir and mixed with autologous CTV-labeled CD4+ T cells in a 1:1 ratio. Staining for HIV-1 p24 was performed on day 4 post-infection using BD Phosflow Fix Buffer I (BD) and BD Phosflow Perm Buffer III (BD) according to the manufacturer’s instructions. The staining panel comprised Live/dead BV510, Life Technologies cat. no. L34957; CD3 BUV615, BD cat. no. 612992, Clone UCHT1; CD4 BV605, BioLegend cat. no. 317438, Clone OKT4; CD8 BUV737, BD cat. no. 564629, Clone SK1; CD45RA BV650, BioLegend cat. no. 304136, Clone HI100; CD27 APC-eFluor 780, eBioscience cat. no. 47027942, Clone O323; CCR7 BUV563, BD cat. no. 741317, Clone 3D12; MX2 AF488, Santa Cruz Biotechnology cat. no. 271527, Clone H-7; IFIT1 APC, Novus Biologicals cat. no. NBP2-71005APC, Clone OTI3G8; APOBEC3G AF700, Novus Biologicals cat. no. NBP1-77206AF700, polyclonal; pSTAT1 (p701) PE-CF594, BD cat. no. AB_2737715, Clone 4a; pIRF3 (pSer396) PerCP, Bioss cat. no. bs-3195R-PerCP, polyclonal; pIRF7 (pSer471+pSer472) AF350, Bioss cat. no. bs-3196R-A350, polyclonal; HIV-1 core antigen FITC, Beckman Coulter cat. no. 6604665, Clone KC57 and CTV BV421, Thermo Fisher Scientific cat. no. C34571. Cell surface staining was performed at 37 °C for 20 min in 25 μl of PBS + 2% FBS buffer. Cells were washed with 150 μl of PBS + 2% FBS and were fixed in 50 μl of Phosflow Fix Buffer I (BD) at 4 °C for 30 min. Cells were washed again with 150 μl of PBS + 2% FBS and then permeabilized in 50 μl of Phosflow Perm Buffer III (BD) on ice for 15 min. Cells were washed twice with 150 μl of PBS + 2% FBS at 500g for 5 min at room temperature. Intracellular staining was performed by adding the intracellular antibody mix in 25 μl of master mix diluted in 1× permeabilization buffer and mixing gently. Cells were incubated for 60 min at 4 °C and washed with 150 μl of 1× permeabilization buffer. Cells were then resuspended in 100 μl of PBS 2% FBS. Acquisition was performed on a minimum of 100,000 live cells on an A5 Symphony flow cytometer driven by FACSDiva. Acquired data were initially analyzed using FlowJo v.10.8.1.
Plasma from CA-vDNAhi RMs induces HDAC11 expression and leads to increased HIV infection in vitro
Plasma samples from healthy control (HC) RMs (TGF-β concentration of 1,260 pg ml−1), CA-vDNAhi RMs (RFl17, TGF-β concentration of 11,873 pg ml−1) and CA-vDNAlo RMs (RRh17, TGF-β concentration of 2,084 pg ml−1) were heat-inactivated at 56 °C for 35 min for complement deactivation and used in this experiment at 25% dilution. This plasma concentration was used as cell viability was maintained above 95%. The final concentrations per well were healthy control TGF-β (315 pg ml−1); CA-vDNAhi TGF-β (2,968 pg ml−1); and CA-vDNAlo TGF-β (521 pg ml−1). After complement deactivation, plasma samples were cleaned using a 75-μm filter to remove clots. Recombinant TGF-β (rTGF-β; 2,000 pg ml−1) was used as positive control. TGF-β signaling blockade was performed by using anti-TGF-β (10 μg ml−1). AIMV medium was used. Fresh PBMCs were isolated from healthy human donors (n = 5) as described. Memory CD4+ T cells were isolated by using the EasySep Human CD4+ T Cell Enrichment kit (Stemcell Technologies) and rested overnight in AIMV medium at a concentration of 2.5 × 106 CD4+ T cells per ml. After resting, cells were subjected to the following conditions: (1) no plasma; (2) HC plasma; (3) CA-vDNAhi plasma; (4) CA-vDNAlo plasma; and (5) rTGF-β. In all cases, cells were cultured in the presence or absence of anti-TGF-β. At 20 min following the addition of the respective conditions, 50,000 cells were removed for the evaluation of SMAD2/3 phosphorylation. Remaining cells were cultured for an additional 16 h, when another 50,000 cells were removed for measuring HDAC11 expression. The remaining cells were spinoculated for HIV infection in vitro as described above. These cells were cultured in AIMV medium supplemented with IL-2 and saquinavir to limit infection to one round, as previously described. HIV protein expression was measured by flow cytometry 4 days post-infection. The staining panel comprised Live/dead BV510, Life Technologies cat. no. L34957; CD3 BUV615, BD cat. no. 612992, Clone UCHT1; CD4 BV605, BioLegend cat. no. 317438, Clone OKT4; CD8 BUV737, BD cat. no. 564629, Clone SK1; CD45RA BV650, BioLegend cat. no. 304136, Clone HI100; CD27 APC-eFluor 780, eBioscience cat. no. 47027942, Clone O323; CCR7 BUV563, BD cat. no. 741317, Clone 3D12; pSMAD2(pS465/pS467)/pSMAD3 (pS423/pS425) PE-CF594, BD cat. no. 562697, Clone O72-670; HDAC11 PE, Santa Cruz cat. no. sc-390737PE, Clone C-5; and HIV-1 (p24) APC, Medimabs cat. no. MM-0289-APC, Clone 28B7. Cell surface staining was performed at 37 °C for 20 min in 25 ml of PBS + 2% FBS buffer. Cells were washed with 150 ml of PBS + 2% FBS and were fixed in 50 ml of Phosflow Fix Buffer I (BD) at 4 °C for 30 min. Cells were washed again with 150 ml of PBS + 2% FBS and then permeabilized in 50 ml of Phosflow Perm Buffer III (BD) on ice for 15 min. Cells were washed twice with 150 ml of PBS + 2% FBS at 500g for 5 min at room temperature. Intracellular staining was performed by adding the intracellular antibody mix in 25 ml of master mix diluted in 1× permeabilization buffer and mixing gently. Cells were incubated for 60 min at 4 °C and washed with 150 ml of 1× permeabilization buffer. Cells were then resuspended in 100 ml of PBS 2% FBS. Acquisition was performed on a minimum of 100,000 live cells on an A5 Symphony flow cytometer driven by FACSDiva.
TGF-β abrogates IFN-induced antiviral signatures and makes cells susceptible to HIV infection
Memory CD4+ T cells were isolated from healthy human donors (n = 6) as described above and were then subjected to the following conditions: (1) unstimulated; (2) stimulated with 2 ng ml−1 TGF-β (Peprotech, cat. no. 100-21) for 24 h; (3) stimulated with 0.2 or 2 ng ml−1 IFN-β; and (4) stimulated with 2 ng ml−1 TGF-β in presence or absence of anti-TGF-β antibody (Merck & Co.) for 24 h and then with 0.2 or 2 ng ml−1 IFN-β for 24 h. Stimulated CD4+ T cells were then infected with HIV (p89.6 dual-tropic strain) by spinoculation in the presence of saquinavir. Activation of TGF-β and IFN-β signaling pathways (pSMAD2/3 and pSTAT1, respectively) was assessed at 20 min post-stimulation, induction or suppression of ISGs was assessed at 16 h post IFN-β and TGF-β stimulation, and HIV p24 levels were assessed 4 days post-infection. Cell surface and intracellular staining were performed as above. The staining panel comprised Live/dead BV510, Life Technologies cat. no. L34957; CD4 BV605, BioLegend cat. no. 317438, Clone OKT4; CD8 BUV737, BD cat. no. 564629, Clone SK1; CD45RA BV650, BioLegend cat. no. 304136, Clone HI100; CD27 BUV496, BD cat. no. 751678, Clone O323; CCR7 BUV563, BD cat. no. 741317, Clone 3D12; IFIT1 APC, Novus Biologicals cat. no. NBP2-71005APC, Clone OTI3G8; APOBEC3G AF700, Novus Biologicals cat. no. NBP1-77206AF700, polyclonal; pSTAT1 (p701) AF488, BD cat. no. AB_2737715, Clone 4a; pSMAD2(pS465/pS467)/pSMAD3 (pS423/pS425) PE-CF594, BD cat. no. 562697, Clone O72-670; p24 RD1, Beckman Coulter cat. no. 6604667, Clone KC57; PD-1 BV711, BD cat. no. 564017, Clone EH12.1; and BCL-2 BUV395, cat. no. custom, Clone Bcl-2/100. Acquisition and analysis of flow cytometry data were performed as above.
TGF-β induces expression of HDACs
To validate our ex vivo findings showing that TGF-β upregulate HDAC1 and HDAC11 expression. PBMCs were isolated from fresh blood from healthy human donors (n = 5). Cells were cultured in AIMV + 10% Serum Replacement (Corning cat. no. 355500) + 10 mM HEPES overnight. Cells were then subjected to the following conditions: unstimulated, stimulated with: TGF-β (0.2 ng, 2 ng and 20 ng ml−1; Gibco, cat. no. 100-21) for 16 h in absence or presence of anti-TGF-β (Merck, cat. no. customized) or anti-IL-10 antibodies (Merck, cat. no. customized). Cells were first stained with a viability dye, washed and fixed according to standard protocols. Staining was performed as per the BD staining protocol with BD Phosflow Fix Buffer I (BD, cat. no. 557870) and BD Phosflow Perm Buffer III (BD, cat. no. 558050). The staining panel included: Live/dead APC-Cy7, BD cat. no. 565388; CD3 BUV805, BD cat. no. 612895, Clone UCHT1; CD4 BV605, BioLegend cat. no. 317438, Clone OKT4; CD8 BUV737, BD cat. no. 564629, Clone SK1; CD45RA BV650, BioLegend cat. no. 304136, Clone HI100; CD27 BUV615, BD cat. no. 751685, Clone O323; CCR7 PE-CF594, BD cat. no. 562381, Clone 2-L1-A; HDAC11 PE, Santa Cruz cat. no. sc-390737PE, Clone C-5; HDAC1 FITC, Santa Cruz cat. no. sc-81598FITC, Clone 10E-2; HLA-DR BV786, BD Biosciences cat. no. 564041, Clone G46-6; CD19 BUV395, BD Biosciences cat. no. 563549, Clone SJ25C1; CD14 BV570, BioLegend cat. no. 301832, Clone M5E2; CD16 BUV661, BD Biosciences cat. no. 741693, Clone B73.1; CD56 PE-Cy5, BD Biosciences cat. no. 555517, Clone B159. Surface staining was performed at 37 °C for 20 min in 25 μl of PBS + 2% FBS buffer. Cells were washed with 150 μl of PBS + 2% FBS buffer. Cells were fixed in 50 μl of Phosflow Fix Buffer I (BD, cat. no. 557870) at 4 °C for 30 min. Cells were washed again with 150 μl of PBS + 2% FBS buffer and then permeabilized in 50 μl of cold Phosflow Perm Buffer III (BD, cat. no. 558050) on ice for 15 min. Cells were washed twice with 150 μl of 1× permeabilization buffer (Invitrogen cat. no. 00-8333), and all residual buffer was removed. All centrifugation steps for staining were performed at 500g for 5 min at room temperature. Intracellular staining was performed by adding the intracellular antibody mix in 25 μl of master mix (diluted in 1× permeabilization buffer (Invitrogen cat. no. 00-8333)) and mixed gently. Cells were incubated for 60 min at 4oC and washed with 150 μl of 1× permeabilization buffer (Invitrogen cat. no. 00-8333). Finally, cells were resuspended in 100 μl of PBS for acquisition. Acquisition was performed on a minimum of 100,000 live cells on a A5 Symphony (BD Biosciences) driven by BD FACSDiva software. Acquired data were initially analyzed employing FlowJo software (v.10.8.1). Representative staining is shown for each panel in the respective figures. Data analysis was performed by using manual gating frequencies and MFIs are shown.
TGF-β suppresses IFN-dependent antiviral state through HDACs and promotes HIV infection
To validate our ex vivo findings, PBMCs were isolated from fresh blood from healthy human donors (n = 5). Cells were cultured in AIMV + 10% Serum Replacement (Corning cat. no. 355500) + 10 mM HEPES overnight. Cells were then subjected to the following conditions: (1) unstimulated; or stimulated with HDAC inhibitors (HDACi), such as (2) panobinostat (100 nM), (3) SAHA (10 nM), (4) romidepsin (10 nM), and (5) SIS17 (100 nM) individually for 24 h only; (6) TGF-β only (2 ng ml−1) for 24 h; (7) IFN- β (1 ng ml−1) only for 24 h; (8) IFN-γ (1 ng ml−1) only for 24 h; (9) TGF-β (2 ng ml−1) for 24 h and then IFN-β (1 ng ml−1) for 24 h; (10) TGF-β (2 ng ml−1) for 24 h and then IFN-γ (1 ng ml−1) for 24 h; (11) individual HDACi for 2 h, then TGF-β (2 ng ml−1) for 24 h, and then, IFN- β (1 ng ml−1) for 24 h; and individual, (12) HDACi for 2 h, then TGF-β (2 ng ml−1) for 24 h, and then, IFN-γ (1 ng ml−1) for 24 h. HDAC inhibitors and TGF-β were removed after 24 h of stimulation, just before the start of IFN-β or IFN-γ stimulation. Activation of the TGF-β and IFN-β signaling pathways (pSMAD2/3 and pSTAT1, respectively) was assessed at 20 min post-stimulation. Induction or suppression of ISGs was assessed at 16 h post IFNs and TGF-β stimulation. Staining was performed as per BD staining protocol with BD Phosflow Fix Buffer I (BD, cat. no. 557870) and BD Phosflow Perm Buffer III (BD, cat. no. 558050). The staining panel included Live/dead BV510, Life Technologies cat. no. L34957; CD3 BUV805, BD cat. no. 612895, Clone UCHT1; CD4 BV605, BioLegend cat. no. 317438, Clone OKT4; CD8 BUV737, BD cat. no. 564629, Clone SK1; CD45RA BV650, BioLegend cat. no. 304136, Clone HI100; CD27 BUV615, BD cat. no. 751685, Clone O323; CCR7 PE-CF594, BD cat. no. 562381, Clone 2-L1-A; IRF7 AF488, Novus biologicals cat. no. NBP306987AF488, Clone 3D9; IFIT1 APC, Novus biologicals cat. no. NBP2-71005APC, Clone OTI3G8; pSTAT1 (p701) RB780, BD cat. no. 569144, Clone 4a; pSmad2 (pS465/pS467)/Smad3 (pS423/pS425) R718, BD cat. no. 567080, Clone O72-670; H3K27ac Pacific Blue, Cell signaling cat. no. 23349, Clone D5E4; IRF1 PE, BD cat. no. 566322, Clone 20/IRF1; PD-1 BV711, BD Biosciences cat. no. 564017, Clone EH12.1; HLA-DR BV786, BD Biosciences cat. no. 564041, Clone G46-6; CD19 BUV395, BD Biosciences cat. no. 563549, Clone SJ25C1; CD14 BV570, BioLegend cat. no. 301832, Clone M5E2; CD16 BUV661, BD Biosciences cat. no. 741693, Clone B73.1; and CD56 PE-Cy5, BD Biosciences cat. no. 555517, Clone B159. Surface staining was performed at 37 °C for 20 min in 25 μl of PBS + 2% FBS buffer. Cells were washed with 150 μl of PBS + 2% FBS buffer. Cells were fixed in 50 μl of Phosflow Fix Buffer I (BD, cat. no. 557870) at 4 °C for 30 min. Cells were washed again with 150 μl of PBS + 2% FBS buffer and then permeabilized in 50 μl of cold Phosflow Perm Buffer III (BD, cat. no. 558050) on ice for 15 min. Cells were washed twice with 150 μl of 1× permeabilization buffer (Invitrogen cat. no. 00-8333), and all residual buffer was removed. All centrifugation steps for staining were performed at 500g for 5 min at room temperature. Intracellular Staining was performed by adding the intracellular antibody mix in 25 μl of master mix diluted in 1× permeabilization buffer (Invitrogen cat. no. 00-8333) and mixed gently. Cells were incubated for 60 min at 4 °C and washed with 150 μl of 1× permeabilization buffer. Finally, cells were resuspended in 100 μl of PBS for acquisition. Acquisition was performed on a minimum of 100,000 live cells on a A5 Symphony (BD Biosciences) driven by BD FACSDiva software.
All flow cytometry data were analyzed using FlowJo v.10.8.1. Data analysis was performed by using manual gating frequencies and MFIs are shown or by using UMAP followed by cluster analysis. Representative cytograms are shown for each panel in the respective figures.
Measurement of cytokines
Meso Scale MULTI-ARRAY Technology (Meso Scale Discovery) was used for evaluation of cytokine levels. A cytokine panel containing the following analytes was screened: CTAK, ITAC, IL-10, IL-16, IL-17A, IL-18, IL-4, IP-10, MCP-1, MCP-2, MIP-3β, GROA, IL-22, IL-7, IL-8, TGF-β1, TGF-β2, TGF-β3, Fractalkine, IFN-α, IFN-β, IL-15, IL-2, IL-6, IL-9, MIP-1α and TNF using 25 μl of serum from each donor in duplicates and following the manufacturer’s instructions. Samples were randomized to avoid batch effects. Results were extrapolated from standard curves for each analyte and plotted in pg ml−1 using the DISCOVERY WORKBENCH v.4.0 software (Meso Scale Discovery).
Statistical analysis
Statistical analyses and plots of data generated in the assays for experimental validation were performed with GraphPad Prism v.9.4.0 (GraphPad Software). The statistical test, symbol representation of statistical significance and sample sizes are listed within the figure legends for each analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw and processed data files from bulk RNA-seq and scMultiome (scRNA-seq and scATAC-seq) are available at the Gene Expression Omnibus NCBI database (bulk RNA-seq data are under accession code GSE279313 and scMultiome data are under accession code GSE279314). Source data are provided with this paper.
Code availability
The code used for analysis is available via GitHub at https://github.com/sekalylab/TGFB-IFN_SIV_reservoir_control.
References
Darcis, G., Van Driessche, B. & Van Lint, C. HIV latency: should we shock or lock? Trends Immunol. 38, 217–228 (2017).
Carcelain, G. & Autran, B. Immune interventions in HIV infection. Immunol. Rev. 254, 355–371 (2013).
Rutishauser, R. L. & Trautmann, L. CD8+ T-cell responses in HIV controllers: potential implications for novel HIV remission strategies. Curr. Opin. HIV AIDS 17, 315–324 (2022).
McBrien, J. B., Kumar, N. A. & Silvestri, G. Mechanisms of CD8(+) T cell-mediated suppression of HIV/SIV replication. Eur. J. Immunol. 48, 898–914 (2018).
van der Heijden, W. A. et al. Chronic HIV infection induces transcriptional and functional reprogramming of innate immune cells. JCI Insight 9, a022236 (2021).
Stetson, D. B. & Medzhitov, R. Type I interferons in host defense. Immunity 25, 373–381 (2006).
Fourati, S. et al. Integrated systems approach defines the antiviral pathways conferring protection by the RV144 HIV vaccine. Nat. Commun. 10, 863 (2019).
Wan, Y. Y. & Flavell, R. A. Yin-Yang’ functions of transforming growth factor-β and T regulatory cells in immune regulation. Immunol. Rev. 220, 199–213 (2007).
Tran, D. Q. TGF-β: the sword, the wand, and the shield of FOXP3(+) regulatory T cells. J. Mol. Cell. Biol. 4, 29–37 (2012).
Sanjabi, S., Oh, S. A. & Li, M. O. Regulation of the immune response by TGF-β: from conception to autoimmunity and infection. Cold Spring Harb. Perspect. Biol. 9, a022236 (2017).
Yim, L. Y. et al. Transforming growth factor β signaling promotes HIV-1 infection in activated and resting memory CD4(+) T cells. J. Virol. 97, e0027023 (2023).
Kim, J. et al. TGF-β blockade drives a transitional effector phenotype in T cells reversing SIV latency and decreasing SIV reservoirs in vivo. Nat. Commun. 15, 1348 (2024).
Ghneim, K., et al. Microbiome and metabolome driven differentiation of TGF-β producing Tregs leads to senescence and HIV latency. Preprint at bioRxiv https://doi.org/10.1101/2020.12.15.422949 (2020).
Pantaleo, G. et al. Studies in subjects with long-term nonprogressive human immunodeficiency virus infection. N. Engl. J. Med. 332, 209–216 (1995).
Leng, J. et al. A cell-intrinsic inhibitor of HIV-1 reverse transcription in CD4(+) T cells from elite controllers. Cell Host Microbe 15, 717–728 (2014).
Moosa, Y. et al. Case report: mechanisms of HIV elite control in two African women. BMC Infect. Dis. 18, 54 (2018).
Ostrowski, M. A. et al. Expression of chemokine receptors CXCR4 and CCR5 in HIV-1-infected and uninfected individuals. J. Immunol. 161, 3195–3201 (1998).
Pereira Ribeiro, S. et al. Dual blockade of IL-10 and PD-1 leads to control of SIV viral rebound following analytical treatment interruption. Nat. Immunol. 25, 1900–1912 (2024).
Abdel-Mohsen, M. et al. Expression profile of host restriction factors in HIV-1 elite controllers. Retrovirology 10, 106 (2013).
Lee, C. K. & Bluyssen, H. A. R. Editorial: STATs and IRFs in innate immunity: from transcriptional regulators to therapeutic targets. Front. Immunol. 10, 1829 (2019).
Velu, V. et al. Enhancing SIV-specific immunity in vivo by PD-1 blockade. Nature 458, 206–210 (2009).
Velu, V. et al. PD-1 blockade following ART interruption enhances control of pathogenic SIV in rhesus macaques. Proc. Natl Acad. Sci. USA 119, e2202148119 (2022).
Rao, M., Valentini, D., Dodoo, E., Zumla, A. & Maeurer, M. Anti-PD-1/PD-L1 therapy for infectious diseases: learning from the cancer paradigm. Int. J. Infect. Dis. 56, 221–228 (2017).
Harper, J. et al. CTLA-4 and PD-1 dual blockade induces SIV reactivation without control of rebound after antiretroviral therapy interruption. Nat. Med. 26, 519–528 (2020).
Bekerman, E. et al. PD-1 blockade and TLR7 activation lack therapeutic benefit in chronic simian immunodeficiency virus-infected macaques on antiretroviral therapy. Antimicrob. Agents Chemother. 63, e01163–19 (2019).
Fromentin, R. et al. PD-1 blockade potentiates HIV latency reversal ex vivo in CD4(+) T cells from ART-suppressed individuals. Nat. Commun. 10, 814 (2019).
Sundqvist, A. et al. Specific interactions between Smad proteins and AP-1 components determine TGFβ-induced breast cancer cell invasion. Oncogene 32, 3606–3615 (2013).
Cocchi, F. et al. Identification of RANTES, MIP-1α, and MIP-1β as the major HIV-suppressive factors produced by CD8+ T cells. Science 270, 1811–1815 (1995).
Saunders, B. M., Frank, A. A., Orme, I. M. & Cooper, A. M. Interleukin-6 induces early γ interferon production in the infected lung but is not required for generation of specific immunity to Mycobacterium tuberculosis infection. Infect. Immun. 68, 3322–3326 (2000).
Velazquez-Salinas, L., Verdugo-Rodriguez, A., Rodriguez, L. L. & Borca, M. V. The role of interleukin 6 during viral infections. Front. Microbiol. 10, 1057 (2019).
Cao, J. et al. HDAC11 regulates type I interferon signaling through defatty-acylation of SHMT2. Proc. Natl Acad. Sci. USA 116, 5487–5492 (2019).
Nunez-Alvarez, Y. & Suelves, M. HDAC11: a multifaceted histone deacetylase with proficient fatty deacylase activity and its roles in physiological processes. FEBS J. 289, 2771–2792 (2022).
Elliott, J. H. et al. Activation of HIV transcription with short-course vorinostat in HIV-infected patients on suppressive antiretroviral therapy. PLoS Pathog. 10, e1004473 (2014).
Kalafati, L. et al. Innate immune training of granulopoiesis promotes anti-tumor activity. Cell 183, 771–785 e712 (2020).
Ochando, J., Mulder, W. J. M., Madsen, J. C., Netea, M. G. & Duivenvoorden, R. Trained immunity - basic concepts and contributions to immunopathology. Nat. Rev. Nephrol. 19, 23–37 (2023).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Huang, Q., Liu, Y., Du, Y. & Garmire, L. X. Evaluation of cell type annotation r packages on single-cell RNA-seq data. Genom. Proteom. Bioinform. 19, 267–281 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 e3529 (2021).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Li, H., Gade, P., Xiao, W. & Kalvakolanu, D. V. The interferon signaling network and transcription factor C/EBP-β. Cell Mol. Immunol. 4, 407–418 (2007).
Robertson, I. B. et al. Latent TGF-β-binding proteins. Matrix Biol. 47, 44–53 (2015).
dos Santos, J. et al. Persistent viral control status is associated with enhanced innate immune responses in people with HIV-1. iScience 29, 3114807 (2026).
Bein, K., Leight, H. & Leikauf, G. D. JUN-CCAAT/enhancer-binding protein complexes inhibit surfactant-associated protein B promoter activity. Am. J. Respir. Cell Mol. Biol. 45, 436–444 (2011).
Jonsson, K. L. et al. Histone deacetylase inhibitor romidepsin inhibits de novo HIV-1 infections. Antimicrob. Agents Chemother. 59, 3984–3994 (2015).
Armani-Tourret, M. et al. Selection of epigenetically privileged HIV-1 proviruses during treatment with panobinostat and interferon-α2a. Cell 187, 1238–1254 e1214 (2024).
Woldemeskel, B. A., Kwaa, A. K. & Blankson, J. N. Viral reservoirs in elite controllers of HIV-1 infection: Implications for HIV cure strategies. eBioMedicine 62, 103118 (2020).
Wimmers, F. et al. The single-cell epigenomic and transcriptional landscape of immunity to influenza vaccination. Cell 184, 3915–3935 e3921 (2021).
Lee, A. et al. BCG vaccination stimulates integrated organ immunity by feedback of the adaptive immune response to imprint prolonged innate antiviral resistance. Nat. Immunol. 25, 41–53 (2024).
Borducchi, E. N. et al. Antibody and TLR7 agonist delay viral rebound in SHIV-infected monkeys. Nature 563, 360–364 (2018).
Vanpouille-Box, C. et al. TGFβ is a master regulator of radiation therapy-induced antitumor immunity. Cancer Res. 75, 2232–2242 (2015).
Yi, M. et al. TGF-β: A novel predictor and target for anti-PD-1/PD-L1 therapy. Front. Immuno.l 13, 1061394 (2022).
Gauthier, T. & Chen, W. IFN-γ and TGF-β, crucial players in immune responses: a tribute to Howard Young. J. Interferon Cytokine Res. 42, 643–654 (2022).
Takaki, H. et al. TGF-β1 suppresses IFN-γ-induced NO production in macrophages by suppressing STAT1 activation and accelerating iNOS protein degradation. Genes Cells 11, 871–882 (2006).
Qing, J. et al. Transforming growth factor β/Smad3 signaling regulates IRF-7 function and transcriptional activation of the β interferon promoter. Mol. Cell. Biol. 24, 1411–1425 (2004).
Gulley, J. L. et al. Dual inhibition of TGF-β and PD-L1: a novel approach to cancer treatment. Mol. Oncol. 16, 2117–2134 (2022).
Woods, D. M. et al. HDAC inhibition upregulates PD-1 ligands in melanoma and augments immunotherapy with PD-1 blockade. Cancer Immunol. Res. 3, 1375–1385 (2015).
Wen, T. et al. Histone deacetylases inhibitor chidamide synergizes with humanized PD1 antibody to enhance T-cell chemokine expression and augment IFN-γ response in NK-T cell lymphoma. eBioMedicine 87, 104420 (2023).
Wang, F. et al. Combined anti-PD-1, HDAC inhibitor and anti-VEGF for MSS/pMMR colorectal cancer: a randomized phase 2 trial. Nat. Med. 30, 1035–1043 (2024).
Rohart, F., Gautier, B., Singh, A. & Le Cao, K. A. mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752 (2017).
Schoggins, J. W. & Rice, C. M. Interferon-stimulated genes and their antiviral effector functions. Curr. Opin. Viro.l 1, 519–525 (2011).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gundem, G. & Lopez-Bigas, N. Sample-level enrichment analysis unravels shared stress phenotypes among multiple cancer types. Genome Med. 4, 28 (2012).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022).
Acknowledgements
This work was supported by NIH grants: R37AI141258 to R.P.S., UM1AI164561 to R.P.S., R01AI179476 to S.P.R. and P01AI178376 to R.P.S. Antibody reagents used in this study were provided by the Nonhuman Primate Reagent Resource (P40 OD028116, U24 AI126683). This work was supported in part with federal funds from the National Cancer Institute under contract 607 75N91019D00024/HHSN261201500003.
Author information
Authors and Affiliations
Contributions
K.G. and F.T.C. performed bioinformatic analyses and data integration. A.C.S., M.B.L., D.A.D.D., T.G.S., P.M.D.R.E. and P.S. designed and executed the experiments. Z.S., J.H., S.J. and C.W. contributed to the design and execution of animal studies, including drug administration, sample collection, processing and storage. R.B. and J.D.L. scientifically reviewed the article and provided technological expertise. G.R., E.R., C.P., G.W., L.M., D.J.H., D.M.G. and B.J.H. developed, tested and provided deimmunized antibodies and performed the PK/PD experiments. L.F.M.S. and P.C.L.G. performed mathematical data analysis. J.D.S., M.G.N. and A.V.D.V. provided single-cell data from HIV ECs and scientifically reviewed the manuscript. G.S., A.A.S. and M.P. contributed to the scientific discussion. H.S., S.P.R. and R.P.S. conceptualized, planned, led and interpreted the RMs work, and contributed to writing the manuscript. R.P.S. conceptualized, planned, interpreted, wrote and funded the development of the project.
Corresponding authors
Ethics declarations
Competing interests
G.R., E.R., C.V.P., G.W., L.M., D.M.G., B.J.H. and D.J.H. are employed by and/or have financial interests in Merck & Co. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: P. Jauregui, in collaboration with the Nature Immunology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of gene expression in combo-treated CA-vDNAlo and CA-vDNAhi RMs.
a, Top: Twenty-eight RMs infected intravenously with SIVmac239. ART (TDF/FTC/DTG) initiated at day 42 post-infection and maintained for 14 months prior to immunotherapeutic interventions. RMs received aIL-10 mAb (red), aIL-10 mAb + aPD-1 mAb (combo, blue) or vehicle (ART-only, black), with infusions every 3 weeks. ATI occurred 12 weeks after first dose of immunotherapy. mAb infusions continued for 14 weeks after ATI. Bottom: Continuous lines represent the median VL per group. A significant reduction in VL was observed by week 36 (combo versus control, P = 0.02; combo versus aIL-10, P = 0.015, two-sided Wilcoxon rank-sum test). Created in BioRender. Pereira Ribeiro, S. https://BioRender.com/g6uf0rn (2026). b, CA-vDNA levels in LNMCs from combo-treated CA-vDNAlo and CA-vDNAhi RMs 24 weeks post-ATI. Two-sided Wilcoxon rank-sum test. Error bars represent mean ± s.e.m. n = 10. c, Module of IFN-related pathways enriched in PBMCs and LNMCs from combo-treated RMs pre-ATI (P < 0.05) when compared to RMs treated with aIL-10 alone and ART-only controls. Only pairs of pathways with a Jaccard similarity index above 0.25 are represented. Red circles depict the upregulated signaling pathways. Circle size reflects the size of the gene set. d, Volcano plot of the top target genes of IRF1 and IRF7 upregulated in PBMCs/LNMCs of combo-treated RMs compared to controls pre-ATI. e. Volcano plot of the top target genes of STAT1 upregulated in PBMCs/LNMCs of combo-treated RMs compared to controls pre-ATI. D-E. TFs target genes were sourced from CHEA and TRANSFAC databases. Genes in the plots are the union of upregulated genes (FC > 1.3, P < 0.05) in PBMCs/LNMCs from combo-treated RMs compared to controls. A moderated t-test combined with an Empirical Bayes method (eBayes) was used to identify DEGs. f, Heatmap (top) depicting levels of phosphorylated IRF3, IRF7 and STAT1 in immune subsets from LNMCs and PBMCs pre-ATI in combo-treated RMs. Only features significantly different between CA-vDNAhi and CA-vDNAlo RMs (nominal P < 0.05) are shown. Dot plots (bottom) show the raw values of the feature per group. Groups comparison was performed with two-tailed unpaired t-test. Error bars represent mean ± s.e.m. n = 10. g. Heatmap (top) depicting levels of phosphorylated IRF3, STAT3, and IRF7 from PBMCs pre-ATI correlated to CA-vRNA in LNMCs at 24 weeks post-ATI. Smaller heatmap on the right indicate negative correlation (nominal P < 0.05) to virologic readouts. Correlations were calculated with two-sided Spearman correlation method. The boxplots (bottom) show the raw values of features in the heatmap. The central line indicates the median, with hinges at the first and third quartiles and whiskers extending 1.5× interquartile range. Group differences were assessed using regression t‑statistics (FDR P < 0.05), with Holm–Bonferroni correction for multiple comparisons. Control n = 8, aIL-10 n = 9, aIL-10 + aPD-1 n = 10. h. Correlation between levels of phosphorylated IRF7 in CD4+ TCM cells pre-ATI and virologic readouts in plasma (log pVL, copies per ml) and LNMCs (log10 CA-vRNA, log10 CA-vDNA, log10 IPDA and log10 2-LTR) 24 weeks post-ATI. Two-sided Spearman correlation. Nominal p-values and correlation coefficient (rho) values are shown. i, Left: integrated model from mixOmics R package59 using flow cytometry and bulk RNA-Seq data pre-ATI. The model negatively correlates with levels of CA-vRNA in LNMCs (P = 0.0093). A two-sided Spearman correlation (rho) coefficient was calculated per gene-flow cytometry marker interaction (significance P < 0.05). The BH method was used to adjust for multiple comparisons. The network highlighting the model between gene expression data (blue circles) and flow cytometry data (black squares). The thickness of edges represent the correlation coefficient between pairs of markers. Right panel: the bar chart illustrates the strength of the statistical significance of the association between CA-vRNA levels in LNMCs and gene expression data (NES), flow cytometry data and the combination of both. ART, antiretroviral therapy; ATI, analytical treatment interruption; CA-vDNA, cell-associated viral DNA; CA-vRNA, cell-associated viral RNA; DEGs, differentially expressed genes; DTG, dolutegravir; FC, fold change; FTC, emtricitabine; IPDA, intact proviral DNA assay; LNMCs, lymph node mononuclear cells; mAb, monoclonal antibody; NES, normalized enrichment scores; PBMCs, peripheral blood mononuclear cells; RMs, rhesus macaques; SIV, simian immunodeficiency virus; TCM, central memory T cells; TDF, tenofovir disoproxil fumarate; TFs, transcription factors; VL, viral load.
Extended Data Fig. 2 Induction of antivirals pre-ATI in combo-treated RMs.
A. STAT and IRF3 phosphorylation levels correlated to virological readouts 24 weeks post-ATI. Only features modulated by combo treatment and correlated to all virological readouts are shown. Differences across groups were assessed with t-statistics of linear regression coefficients. The p-value was corrected for multiple comparisons with the Benjamini–Hochberg method and only features with adjusted p-value < 0.05 were selected for downstream steps. Correlations with virological readouts were calculated using a two-sided Spearman method and only features with an FDR-corrected p-value < 0.1 are shown. Arrows at the bottom of the heatmap indicate CA-vDNAlo combo-treated RMs. B. A bar plot was used to illustrate differences in the number of DEGs (LIMMA P < 0.05, Y axis) in LNMCs and PBMCs from RMs treated with aIL-10 alone and combo-treated RMs pre-ATI. Red represents upregulated genes and blue represents downregulated genes in combo-treated RMs versus aIL-10 treated RMs. The Y axis represents the number of DEGs. C. Heatmap illustrating the differential levels of relative expression of LEGs belonging to the IFN and RFs gene sets in LNMCs from RMs treated with aIL-10 alone (red annotation) and combo-treated RMs (blue annotation) pre-ATI (light green annotation). LEGs from IFN pathways were summarized into a SLEA representation (z-score per pathway per sample). Rows represent genes and columns represent individual RMs. A gradient was used to depict relative levels of gene expression, where blue represents low relative expression and red represents high relative expression. The levels of CA-vRNA, CA-vDNA and 2-LTR circles (white to blue, green and purple, respectively) are annotated at the top of the heatmap. Note the higher levels of expression of antiviral genes in a subgroup of combo-treated RMs with low CA-vDNA (boxed). n = 20 RMs. 2-LTRs: 2-long terminal repeats circles; ATI: analytical treatment interruption; CA-vDNA: cell-associated viral DNA; CA-vRNA: cell-associated viral RNA; DEGs: differentially expressed genes; IFN: interferon; LEGs: leading-edge genes; LIMMA: linear models for microarray and RNA-Seq data; LNMCs: lymph node mononuclear cells; PBMCs: peripheral blood mononuclear cells; RMs: rhesus macaques.
Extended Data Fig. 3 Interplay between IFN and TGF-β in the modulation of antivirals.
A. Cytograms (top panels) and bar graphs (bottom panels) representing the expression of IFIT1, APOBEC3G and MX2 at baseline (that is, with or without IFN-β stimulation, before labeling, HIV infection and co-culture) in cells from healthy human donors. Shown are FMO (gray), UNS (dotted lines) and IFN-β-stimulated (blue). Statistical significance was assessed using a two-tailed paired t-test. Error bars represent mean ± SEM. n = 6 healthy donors. Statistical significance was assessed using the Friedman test followed by uncorrected Dunn’s multiple-comparisons test; *P < 0.05, **P < 0.01, ***P < 0.001. FMO: fluorescence minus one; HDAC: histone deacetylase; IFN: interferon; PBMCs: peripheral blood mononuclear cells; TCM, TTM and TEM: central, transitional and effector memory T cells, respectively; UNS: unstimulated.
Extended Data Fig. 4 Plasma from CA-vDNAhi RMs induces HDAC11 expression and leads to increased levels of HIV infection in vitro.
A. Representative histogram showing the expression of pSMAD2_3 in isolated memory CD4 T+ cells from healthy human donors (n = 5) that were cultured in the presence of rTGF-β + /- aTGF-β (gray and red, respectively). B. Bar plots showing the MFI of pSMAD2_3 in isolated memory CD4+ T cells from healthy human donors (n = 5) 20 min after treatment with plasma from the CA-vDNAhi RM (25% dilution), the CA-vDNAhi RM (25% dilution), rTGF-β and anti-TGF-β antibody. The color of the dots represents individual donors. RM one-way ANOVA with Tukey’s multiple-comparisons test. Error bars represent mean ± SEM. C. Representative histogram showing the expression of HDAC11 following treatment with rTGF-β + /- anti-TGF-β (gray and red, respectively) and HDAC11 FMO control in isolated memory CD4+ T cells from healthy human donors (n = 5). D. Bar plots showing the MFI of HDAC11 in isolated memory CD4+ T cells from healthy human donors (n = 5) after 16 h of treatment with plasma from the uninfected healthy RM (25% dilution), the CA-vDNAlo RM (25% dilution), the CA-vDNAhi RM (25% dilution), rTGF-β and anti-TGF-β antibody. The color of the dots represents individual donors. RM one-way ANOVA with Tukey’s multiple-comparisons test. Error bars represent mean ± SEM. E. Bar plots showing the decay of HDAC11 expression as calculated by the FC in MFI of HDAC11 between treatment in presence or absence of anti-TGF-β. The higher the value, the higher the decay of HDAC11 expression following treatment with anti-TGF-β. The color of the dots represents individual donors. RM one-way ANOVA with Tukey’s multiple-comparisons test. Error bars represent mean ± SEM. F. Gating strategy for the identification of HIV-infected cells (CD4− p24+) from healthy human donors. Representative dot plots for each condition are shown: plasma from healthy control RM (25% dilution; black), CA-vDNAlo RM (25% dilution; blue), CA-vDNAhi RM (25% dilution; red) and treated with rTGF-β (green). G. Bar plots showing the FC in the frequencies of HIV-infected cells (CD4− p24+) between the plasma condition being tested over treatment with plasma from the healthy control RM. Frequencies of CD4− p24+ cells were assessed 4 days post-infection following treatment with plasma from the uninfected healthy RM (25% dilution), the CA-vDNAhi RM (25% dilution), the CA-vDNAhi RM (25% dilution), and r-b. Color of the dots represent individual donors. RM one-way ANOVA with Tukey’s multiple-comparisons test. Error bars represent mean ± SEM. H. Spearman correlation between pre-infection HDAC11 expression levels and frequencies of CD4− p24+ T cells 4 days post-infection following treatment with plasma from the uninfected healthy RM (25% dilution; black), the CA-vDNAlo RM (25% dilution; blue), the CA-vDNAhi RM (25% dilution; red), and rTGF-β (green). Two-sided Spearman correlation coefficient (rho) is shown in the figure. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. FC: Fold change; FMO: fluorescence minus one; HC: healthy control; HDAC: histone deacetylase; MFI: median fluorescence intensity; RMs: rhesus macaques; rTGF-β: recombinant TGF-β.
Extended Data Fig. 5 TGF-β induces HDAC expression.
A. Schematic overview of the experimental workflow. PBMCs from healthy donors (n = 5) were either left unstimulated or pretreated with aTGF-β for 4 hours, followed by stimulation with TGF-β (0.2, 2, or 20 ng/ml) for 16 h. Flow cytometry was performed by gating on CD4⁺ T cells. Created in BioRender. Sekaly, R. (2026) https://BioRender.com/6bvy0gv. B. Gating strategy for the identification of memory CD4+ T cells (CD45RA−). C. Representative histogram overlays show HDAC1 and HDAC11 protein expression across conditions. D-F. Bar plot showing the expression of HDAC1 and HDAC11 at per-cell level (MFI) upon TGF-β +/- aTGF-β stimulation in TCM, TTM and TEM CD4+ T cells, respectively. Quantification of HDAC1 and HDAC11 MFI in central memory (TCM), transitional memory (TTM), and (G) effector memory (TEM) CD4⁺ T cell subsets. Each dot represents an individual donor. Data are presented as mean ± SEM. Statistical significance was assessed using the Friedman test followed by uncorrected Dunn’s multiple-comparisons test; *P < 0.05, **P < 0.01, ***P < 0.001. n = 5. FMO: fluorescence minus one; HDAC: histone deacetylase; IFN: interferon; MFI: median fluorescence intensity; PBMCs: peripheral blood mononuclear cells; TCM, TTM and TEM: central, transitional and effector memory T cells, respectively; SEM: standard error of the mean; UNS: unstimulated.
Extended Data Fig. 6 HDACi unleash antiviral responses.
A. Cytograms (top panels) and bar graphs (bottom panels) representing the expression of pSMAD2/3 post-TGF-β stimulation, and pSTAT1 post- IFN-γ or IFN-β stimulation, respectively. FMO (gray), UNS (dotted lines), TGF-β (purple), and IFN-γ and IFN-β-stimulated (blue). Statistical significance was assessed using a two-tailed paired t-test. Error bars represent mean ± SEM, with each dot representing an individual healthy donor (n = 5). B-D. Cytograms representing the expression of IRF1 post-stimulation in various conditions in combination with IFN-β, TGF-β, anti-TGF-β and under various HDAC inhibitors (SIS17, SAHA, and PK228 (Romidepsin)). Statistical significance was assessed using a two-sided paired Wilcoxon signed-rank test. *P < 0.05. Error bars represent mean ± SEM. with each dot representing an individual healthy donor (n = 5). E. Bar graphs (left panel) and cytograms (right panel) representing the expression of IRF1 post-stimulation in various conditions in combination with IFN-γ. Error bars represent mean ± SEM. with each dot representing an individual healthy donor (n = 5). Statistical significance was assessed using the Friedman test followed by uncorrected Dunn’s multiple-comparisons test. * P < 0.05, ** P < 0.005, *** P < 0.0005, **** P < 0.0001. IFN: interferon; ISGs: interferon-stimulated genes. Error bars represent mean ± SEM. n = 5.
Extended Data Fig. 7 Single-cell multiome gene expression and ATAC-Seq.
A. Experimental design. LNMC samples obtained pre-ATI from 7 combo-treated RMs were selected for single-cell GEX and quantification of chromatin accessibility (single-cell ATAC-Seq) using the 10X Genomics Multiome protocol. Created in BioRender. Sekaly, R. (2026) https://BioRender.com/1u26ix1. B. UMAP dimensional reduction based on gene expression (RNA-Seq, left panel), chromatin accessibility (ATAC-Seq, middle panel) and integrated RNA-Seq and ATAC-Seq data (right panel) using the WNN method. Colors represent cells derived from different RMs that were included in the analysis. There is no visible evidence of batch effects associated with sample origin. C. UMAP plot showing cluster-based manual cell type annotation based on key marker genes. D. Average relative expression levels of marker genes in all major clusters of myeloid cells, B lymphocytes and T lymphocytes. Average expression levels range from low (blue) to high (red). The frequency of cells expressing the gene in each cell type is represented by circle size. E. Average relative expression levels of marker genes in T lymphocyte subsets. Average expression levels range from low (blue) to high (red). The frequency of cells expressing the gene in each cell type is represented by circle size. F. Heatmap showing the average expression of IFN-related DEGs in major myeloid and lymphoid cell types. The top 5 genes with the highest FC per cell type are indicated. DEGs were identified based on the two-sided Wilcoxon rank-sum test, logfFC threshold = 0.1, minimum percentage of cells in the clusters = 1% and adjusted p value (FDR) < 0.05. IFN-related genes were obtained from the Interferon Alpha Beta Signaling, Interferon Gamma Signaling, and Interferon Signaling gene sets (ReactomeDB), Interferon Type I Signaling Pathways, Type III Interferon Signaling and Type II Interferon Signaling IFN gene sets (WikiPathways), and Interferon Alpha Response and Interferon Gamma Response gene sets (MSigDB Hallmark). G. Cell-cell communication analysis of type-II IFN inferred by CellChat. The heatmap depicts the probability of communication between pairs of cells based on ligand/receptor expression. The left side bar indicates source cells, while the bottom bar indicates cell types that express the IFN-γ-receptor. H. Coverage plots representation of chromatin accessibility of IFN genes in major myeloid and lymphoid cell subtypes isolated from LNMCs of combo-treated RMs. Normalized peak accessibility varies from 0 (minimum) to 50 (maximum). All IFN genes with detected peaks are represented. DEGs: differentially expressed genes; FC: fold change; FDR: false discovery rate; IFN: interferon; LNMCs: lymph node mononuclear cells; RMs: Rhesus macaques; UMAP: uniform manifold approximation and projection; WNN: weighted-nearest neighbor. CA-vDNAhi: RBf17, RFl17, Rym17; CA-vDNAlo: RBv17, RNy16, RWs17, RRh17.
Extended Data Fig. 8 Single-cell gene expression and chromatin accessibility of CEBEP/β target genes in combo-treated RMs.
A. UMAP plot showing the downregulated (blue) expression of genes from the TNF pathway in T cells from CA-vDNAlo RMs. B. UMAP plot showing the downregulated (blue) expression of genes from the TNF pathway in myeloid cells and FDCs from CA-vDNAlo RMs. C. Expression of effector genes IFNG, GZMB, PRF1, TBX21, CD69, CD40LG and TNFRSF9 in T cells. The lighter regions in the density plots indicate higher density of cells expressing the selected genes. The dot plot shows the frequencies of different subsets expressing these markers and its average expression (red to blue: high to low expression). D. GSEA was used to identify effector pathways (MSigDB, Hallmark) that significantly (P ≤ 0.05) differentiated CA-vDNAhi (left) and CA-vDNAlo (right) from control RMs in LNMCs pre-ATI. Heatmap illustrates NES for pathways of the effector modules, the NES scale ranges from low (blue) to high (red). Columns represent relative enrichment in CA-vDNAhi (left) and CA-vDNAlo RMs (right), whereas rows represent pathways. n = 10 combo-treated RMs (6 CA-vDNAhi and 4 CA-vDNAlo). GSEA is a statistical method to determine whether members of a particular gene set preferentially occur toward the top or bottom of a ranked-ordered gene list where genes are ranked by the strength of their association with the outcome of interest. More specifically, GSEA calculates an enrichment score (NES) that reflects the degree to which a set of genes is overrepresented among genes differentially expressed. E. Network inference (Cytoscape) was used to plot the significant effector LEGs (P < 0.05) of the pathways in E that were significantly inversely correlated with CA-vRNA (blue diamond) 24 weeks post-ATI in LNMCs of combo-treated RMs. Connecting lines highlight gene-gene interactions. Line thickness represents the strength of gene interactions. CA-vDNA: cell-associated viral DNA; LEG: leading-edge genes; LNMCs: lymph node mononuclear cells; NES: normalized enrichment scores; RMs: rhesus macaques; UMAP: uniform manifold approximation and projection.
Extended Data Fig. 9 Chromatin accessibility of TFs in T cell subsets associated with CA-vDNA levels.
A. The heatmap depicts the relative expression of IRF7 target genes that are upregulated (red) in selected T cell subsets from CA-vDNAlo RMs. B. Chromatin accessibility at ISG15 locus across cell types in CA-vDNAlo and CA-vDNAhi RMs. Top panel: Normalized ATAC-seq signal tracks representing chromatin accessibility across cell types. Each colored track denotes signal intensity mapped to the genomic coordinates. Blue boxes (exon) and arrows (intron) indicate the genomic coordinates of ISG15 on chromosome 1. Gray boxes indicate the peaks from ATAC-seq data. C. UMAP representation of levels of expression of representative IRF7 target genes (ISG15, DDX60, IFIT2, NT5C3A) that are upregulated (red) in T lymphocytes from CA-vDNAlo RMs. D. Relative expression of representative C/EBP-β target genes that are upregulated (red) and downregulated (blue) in selected T cell subsets from CA-vDNAlo RMs. E. Coverage plots representation of chromatin accessibility of IL-6 gene, a C/EBP-β target gene, in major myeloid and lymphoid cell subtypes isolated from LNMCs of combo-treated RMs. Normalized peak accessibility varies from 0 (minimum) to 50 (maximum). F. UMAP representation of levels of expression of C/EBP-β target genes that are upregulated (red - IRF9, LTBP1) or downregulated (blue - FOS, HDAC9) in selected T cell subsets from CA-vDNAlo RMs. G. Relative expression of representative SMAD target genes that are downregulated (blue) in selected T cell subsets from CA-vDNAlo RMs. H. Chromatin accessibility at ADORA2B locus across cell types in CA-vDNAlo and CA-vDNAhi RMs. Top panel: Normalized ATAC-seq signal tracks representing chromatin accessibility across cell types. Each colored track denotes signal intensity mapped to the genomic coordinates. Green boxes (exon) and arrows (intron) indicate the genomic coordinates of ADORA2B on chromosome 16. Gray boxes indicate the peaks from ATAC-seq data. Blue lines connecting peaks indicate co-accessibility and regulatory potential of these genomic regions. I. UMAP representation of levels of expression of SMAD target genes that are downregulated (blue - ADORA2B, RORA, RUNX2, LAG3) in selected T cell subsets from CA-vDNAlo RMs. CA-vDNA: cell-associated viral DNA; IFN: interferon; ISG: interferon stimulated genes; log2FC: log2 fold change; RMs: rhesus macaques; TF: transcription factors.
Extended Data Fig. 10 Overarching model of the interactions between TGF-β, IFNs and IL-6 and their downstream TFs.
The interaction of TGF-β, IFNs and IL-6 and their downstream TFs directed the expression of epigenetics modifiers and antiviral ISGs, resulting in the dichotomic phenotype of CA-vDNAhi and CA-vDNAlo RMs. Created in BioRender. Pereira ribeiro, S. (2026) https://BioRender.com/ult0x4p.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3.
Supplementary Tables 1–9 (download XLSX )
Spreadsheet-based tables combined into a workbook with multiple tabs.
Source data
Source Data Fig. 1 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Fig. 2 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Fig. 3 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Fig. 4 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Fig. 5 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Fig. 6 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 1 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 2 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 3 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 4 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 5 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 6 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 7 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 8 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Source Data Extended Data Fig. 9 (download XLSX )
Numerical source data for graphs combined into a workbook with multiple tabs.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghneim, K., ten-Caten, F., Santana, A.C. et al. TGF-β mediates epigenetic control of innate antiviral responses and SIV reservoir size. Nat Immunol 27, 686–699 (2026). https://doi.org/10.1038/s41590-026-02458-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-026-02458-x