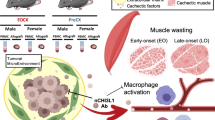
Abstract
Skeletal muscle wasting is a defining feature of cancer cachexia, a multifactorial syndrome that drastically compromises patient quality of life and treatment outcomes. Mitochondrial dysfunction is a major contributor to skeletal muscle wasting in cancer cachexia, yet the upstream molecular drivers remain elusive. Here we show that cancer impairs the activity of cAMP-dependent protein kinase A (PKA) and of its transcriptional effector CREB1 in skeletal muscle, ultimately contributing to the downregulation of a core transcriptional network that supports mitochondrial integrity and function. The restoration of cAMP–PKA–CREB1 signalling through pharmacological inhibition of the cAMP-hydrolysing phosphodiesterase 4 (PDE4) rescues the expression of mitochondrial-related genes, improves mitochondrial function and mitigates skeletal muscle wasting in male mice. Altogether, our data identify tumour-induced suppression of the cAMP–PKA–CREB1 axis as a central mechanism contributing to mitochondrial dysfunction in skeletal muscle during cancer cachexia. Furthermore, these findings highlight PDE4, particularly the PDE4D isoform, as a potential therapeutic target to preserve muscle mitochondrial function and counteract muscle wasting in cancer cachexia.
Similar content being viewed by others
Main
Cancer-associated cachexia affects the majority of patients with cancer and consists in a severe loss of skeletal muscle mass and strength. Although muscle wasting is a determinant feature of cachexia pathogenesis, its multifaceted nature is linked to a chronic inflammation status and to a disrupted metabolic interplay among various organs1,2. The prevalence of cachexia (defined as >5% weight loss within 6 months) varies considerably across different cancer types, with the highest rates—ranging from approximately 40% to 70%—observed in pancreatic, gastro-oesophageal, head and neck, lung and colorectal cancers3. Currently, effective pharmacological interventions to counteract cachexia progression are lacking in the clinical practice, and its occurrence in patients with cancer still heavily affects patients’ quality of life, reduces chemotherapy efficacy and is estimated to directly account for up to 30% of cancer-related deaths2,4. Cancer-associated cachexia remains an unmet clinical need that may be addressed by elucidating its underlying molecular mechanisms3,5.
A roster of cachectogenic factors released by the tumour microenvironment has been identified; however, clinical trials targeting these factors and their signalling pathways have failed to restore skeletal muscle function in cachectic patients. This suggests that cachexia is maintained by a redundant network of mediators, reflecting the heterogeneity and complexity of tumour–microenvironment interactions6,7. Nevertheless, a common feature of tumours is to deregulate host metabolism to divert nutrients from the skeletal muscle, the most abundant tissue in the human body and the main reservoir of protein-stored amino acids. Indeed, cachectic skeletal muscle undergoes transcriptional reprogramming driven by STAT3, NF-κB and SMAD2/3 transcription factors, which leads to the induction of muscle-specific E3 ubiquitin ligases responsible for myofibrillar protein breakdown3. However, although protein degradation is a key determinant of muscle wasting, mitochondrial dysfunction and energetic stress precede muscle atrophy and contribute to the overall wasting process8,9. For instance, AMP-activating protein kinase (AMPK), a key cellular energy sensor that orchestrates metabolic adaptation, becomes chronically activated under negative energy balance, thus leading to mTORC1 inhibition and contributing to muscle atrophy10.
Evidence from both genetically engineered mouse models and ectopic cancer models indicates that loss of mitochondrial function, unbalanced mitochondrial dynamics, impaired mitochondrial biogenesis and excessive mitophagy are common features of cachectic muscle, regardless of the primary cancer9,11,12,13,14. Yet, the molecular mechanisms linking tumour-derived cachectogenic factors and muscle mitochondrial dysfunction still remain to be identified.
Mitochondrial function, biogenesis and dynamics are regulated both transcriptionally and posttranslationally by the 3′,5′-cyclic adenosine monophosphate (cAMP) signalling pathway, which activates protein kinase A (PKA). PKA, in turn, phosphorylates a plethora of substrates, including proteins regulating mitochondrial dynamics, such as MIC60 complex, whose phosphorylation prevents mitophagy15, and DRP-1, whose PKA-mediated phosphorylation impairs its GTPase activity, thus hampering mitochondrial fission and increasing mitochondrial elongation16. In addition, PKA phosphorylates substrates in the electron transport chain, thus increasing mitochondrial ATP production17. Along with these posttranslational regulations, the cAMP–PKA axis extensively modulates gene transcription through the phosphorylation of the transcription factor cAMP-responsive element (CRE) binding protein 1 (CREB1)18. Of note, β2-adrenergic receptor (β2AR)-mediated activation of the cAMP–PKA signalling in skeletal muscle induces mitochondrial biogenesis by promoting nuclear localization and transcriptional activity of PGC-1α and PGC-1β, two master transcriptional co-activators, which interact with several transcription factors, including oestrogen-related receptors ERRα/β and myocyte enhancer factor 2C (MEF2C)19,20,21,22. Consistently, activation of the cAMP–PKA signalling pathway in several cell types, including myocytes, has been shown to induce mitochondrial biogenesis and enhances mitochondrial function both in vitro and in vivo23,24,25.
cAMP signalling is restricted by both cAMP hydrolysis carried out by phosphodiesterases (PDEs) and by its forced compartmentalization mediated by A-kinase-anchoring proteins26. Interestingly, PDE4 inhibition is effective in counteracting atrophy in burn-induced muscle wasting and unloading-induced atrophy27,28. Given the central role of cAMP signalling in controlling mitochondrial content and muscle oxidative capacity, which are strongly affected in cachectic muscles, we hypothesized that an impairment of cAMP signalling could be a determinant of the wasting process, and, in particular, of the oxidative dysfunction, during cachexia. To uncover the molecular mechanisms underlying mitochondrial dysfunction in the skeletal muscle of tumour-bearing mice, we integrated transcriptomic analysis, chromatin immunoprecipitation followed by sequencing (ChIP-seq), global serine/threonine kinase activity profiling and high-resolution mitochondrial respirometry. Our findings reveal that cancer suppresses the cAMP–PKA–CREB1 signalling axis in skeletal muscle, resulting in the transcriptional repression of key mitochondrial regulators. This disruption contributes to a collapse in mitochondrial integrity and oxidative capacity, hallmark features of cancer-induced muscle wasting. Importantly, we show that pharmacological inhibition of phosphodiesterase 4 (PDE4) in vivo restores cAMP–PKA–CREB1 signalling and the transcription of mitochondrial genes, improves mitochondrial respiration and attenuates muscle wasting. Moreover, among the PDE4 isoforms, PDE4D emerged as a key contributor to the mitochondrial dysfunction observed in cachectic muscle.
Altogether, our data uncover a mechanistic link between tumour-induced signalling defects and mitochondrial failure in cachexia and identify PDE4(D) as a potential therapeutic target for preserving muscle bioenergetics in cancer-associated muscle wasting.
Results
Cancer reduces PKA activity and impairs CREB1-mediated transcription of mitochondria-related genes in skeletal muscle
Ten days after C26 adenocarcinoma inoculation, mice featured a precachectic phenotype, with 6.7% of total weight loss and spleen enlargement (Extended Data Fig. 1a–d). Despite skeletal muscle atrophy being barely detectable (Extended Data Fig. 1e), mRNA expression of the key mediators of muscle atrophy, the E3-ubiquitin ligases Atrogin-1 (Fbxo32) and muscle-specific RING finger protein 1, MuRF1 (Trim63), was induced (Extended Data Fig. 1f), indicating an ongoing wasting process at the molecular level. As loss of mitochondrial function precedes cachexia8,9, we investigated the expression of key master regulators of mitochondrial biogenesis13,29. Interestingly, in precachectic muscles, mRNA expression of the master regulators of mitochondrial biogenesis PGC-1α (Ppargc1a) and PGC-1β (Ppargc1b), together with the expression of the members of the mitochondrial transcriptional regulatory network ERRα (Esrra) and ERRβ (Esrrb)20, were clearly reduced as compared with control mice (Extended Data Fig. 1g). Moreover, mRNA expression of the mitochondrial DNA (mtDNA) transcription factor Tfam (mitochondrial transcription factor A), of the PGC-1α regulator Perm1 (PPARGC1- and ESRR-induced regulator, muscle 1)30 and of Mef2c were also downregulated (Extended Data Fig. 1g). These transcriptional changes suggest that early molecular events could contribute to metabolic dysfunction in muscle during cachexia.
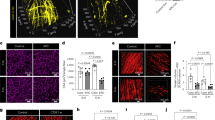
At a later stage (13 days after tumour cell inoculation), mice manifested a clear cachectic phenotype, showing severe total weight loss, epididymal adipose tissue loss, spleen enlargement and loss of skeletal muscle mass (Extended Data Fig. 1h–l). Transcriptomic analysis on gastrocnemius from late-stage cachexia showed that 64.2% of the whole transcripts were differentially expressed genes (DEGs, filtered for P adjusted value <0.05) between C26 and controls (Supplementary Table 1). Ingenuity pathway analysis (IPA) of these DEGs predicted ‘mitochondrial dysfunction’ as the top activated term (P adjusted value-based ranking) and ‘oxidative phosphorylation’ as the top inhibited term (Fig. 1a). Notably, EnrichR-based enrichment analysis revealed that loss of mitochondrial gene expression is also a cachexia hallmark induced by a KrasG12D/+;Lkb1f/f lung cancer (Extended Data Fig. 2a), a genetically engineered mouse (GEM) cachexia model, and not by fasting-induced muscle wasting (input data from Goncalves et al.31). Similar results were also recently reported in a pancreatic cancer GEM model14. Altogether, these data indicate that downregulation of mitochondrial gene expression is a common feature of cancer cachexia in different experimental models. In addition, enrichment analysis of the downregulated DEGs in cachectic muscles indicated a strong overlap with genes downregulated in CREB1 myocardial-specific loss-of-function mice, both in KrasG12D/+;Lkb1f/f lung cancer-induced cachexia (Extended Data Fig. 2b, gene set from Goncalves et al.31) and in C26-induced cachexia (Extended Data Fig. 2c), thus suggesting that a subset of CREB1 direct and indirect target genes are downregulated in cachectic muscles. Of note, analysis of the CREB1-dependent genes downregulated in C26-induced cachexia revealed a robust enrichment in mitochondrial-related metabolic processes, including ‘mitochondrial ATP synthesis coupled electron transport’, ‘aerobic respiration’ and ‘cellular respiration’ (Extended Data Fig. 2d). Consistent with a putative impairment in CREB1 transcriptional activity, several upstream components of the cAMP signalling pathway were differentially expressed in the muscles of cachectic mice compared with controls. In particular, the guanine nucleotide-binding protein G(αs) (Gnas), which activates the adenylyl cyclase-dependent cAMP production, was reduced at the transcriptional level in cachectic muscles, while Gαi3 (Gnai3), which antagonizes adenylyl cyclase activity, was instead induced (Fig. 1b). In addition, the two PKA catalytic subunits PKA Cα (Prkaca) and PKA Cβ (Prkacb) and the regulatory subunit PKA RIIα (Prkar2a) were downregulated in cachectic muscles at the transcriptional level (Fig. 1b). Finally, cachectic muscles featured higher expression of cAMP-hydrolysing PDE isoforms, including the most expressed Pde4d, Pde4b, Pde7a and Pde8a, while only Pde4a was downregulated (Fig. 1b and Extended Data Fig. 1m). Altogether these findings suggest that the tumour might impair cAMP–PKA signalling in the skeletal muscle, thus affecting CREB1-dependent transcription. We then used PamChip peptide array to measure global changes in serine/threonine kinase activity in cachectic muscles. Notably, three members of the PKA family—namely PRKACA, PRKX and PRKACB—ranked among the top 20 kinases with reduced activity in cachectic muscles as compared with controls, with PRKACA showing the third highest kinase score among inhibited kinases (that is, with negative median kinase statistics) (Fig. 1c and Supplementary Table 2). Among the top three highest-scored kinases, the cGMP-activated kinase PRKG2 was also predicted to be inhibited. Interestingly, despite its predicted impaired activity, PRKG2 is undetectable at the transcriptional level (Supplementary Table 1), yet it shares phosphorylation target sites with PKA32, suggesting that the predicted impairment of PRKG2 reflects the reduced PKA activity. Consistent with an impaired PKA activity, PKA phospho-substrate signal was reduced in muscle from cachectic mice versus controls, as shown in western blot (Fig. 1d,e). Therefore, we investigated whether CREB1 genomic distribution was altered in precachectic muscles. Chromatin immunoprecipitation (ChIP) of p-CREB1S133 followed by sequencing from muscles at early-stage cachexia (10 days from C26 cell injection) revealed that p-CREB1S133 occupancy was reduced in 1,748 peaks as compared with controls (signal ‘lower in C26’) (Fig. 1f,g and Supplementary Table 3). Conversely, p-CREB1S133 occupancy was increased in 70 peaks (signals ‘higher in C26’) (Fig. 1f,g and Supplementary Table 3). The majority of these binding sites occurred in candidate cis-regulatory elements (cCREs)33, either promoters (70.5%) or enhancers (24.3%) (Extended Data Fig. 3a,b), and showed enrichment for CRE binding motifs (5′-TGACGTCA-3′) (Extended Data Fig. 3c).
a, IPA of DEGs (filtered for P adjusted value <0.05) in gastrocnemius from C26 13 days (C26 13d) versus control (ctr). P adjusted value-based ranking top 10 terms are shown. n = 7 ctr, n = 7 C26 13d. b, Heatmap generated with bidirectional hierarchical clustering (gene- and sample-based) of normalized reads per kilobase per million mapped reads (RPKM) of cAMP signalling-related selected DEGs from RNA-seq in gastrocnemius from C26 13d versus ctr mice (for all genes listed: P adjusted value <0.01 and base mean >600). n = 7 ctr, n = 7 C26 13d. c, Serine/threonine kinase activity measured through PamGene peptide array in gastrocnemius muscle lysates from C26 tumour-bearing mice (day 13) versus non-tumour-bearing mice (ctr). Kinases are ordered by their kinase score, and the bubble colours represent the median kinase statistic score, calculated as the sum of the significance and specificity scores. Top 20 inhibited kinases (that is, with negative median kinase statistics) are shown. n = 6 mice for each experimental group. Red arrows highlight the PKA family members. d,e, Representative blot (d) and densitometric analysis of phospho(serine/threonine)-PKA substrates over GAPDH (e) of whole gastrocnemius lysate from non-tumour-bearing mice (ctr) or C26 13d. n = 5 for each experimental group. Two-tailed t-test. f, Volcano plot showing overall results of differential binding analysis of p-CREB1S133 ChIP-seq in C26 10d versus ctr. Each dot is a peak and is coloured according to differential binding status (based on P value ≤0.05 and log2FC lower or greater than 0): lower in C26, higher in C26, common (not changed). Differential binding analysis was performed using DiffBind (Methods). g, Heatmap of peak intensity from p-CREB1S133 ChIP-seq analysis in C26 10d versus ctr. Differential analysis defines two groups: lower or higher in C26. Signal is visualized within a ±2-kb window centred on the peak. Both quadriceps for each mouse were pooled together for the ChIP assay. n = 2 mice per experimental group. Two independent experiments with matched control and C26 samples were performed. h, Integrative Genomics Viewer (IGV) images showing representative p-CREB1S133 ChIP signals among the ‘lower in C26’ peaks (red bar) (Supplementary Table 3) aligned across the indicated gene. n = 2 mice per experimental group. i, Normalized RPKM of selected p-CREB1S133 target genes differentially expressed in RNA-seq of gastrocnemius from C26 13d versus ctr mice (Supplementary Table 1). n = 7 ctr, n = 7 C26 13d. Numbers indicate adjusted P values from DESeq2 analysis. Data are presented as mean values ± s.e.m. in e and i, and only significant P values (<0.05) are annotated in the graphs.
Among the genes featuring loss of p-CREB1S133 binding in cachectic muscle, we detected the key regulator of mitochondrial biogenesis PGC-1β (Ppargc1b), the regulator of mitochondrial ribosome assembly ‘ribosome binding factor A’ (Rbfa)34, the mitochondrial ribosomal component (Mrpl12), and subunits of the mitochondrial electron transport chain, including Ndufv1 and Ndufb2 (complex (C)I, NADH:ubiquinone oxidoreductase) and Cox5a (CIV, cytochrome c oxidase), and of the ATP synthase (CV) such as Atp5j2 (Fig. 1h). We also found reduced p-CREB1S133 binding to the transcription start site of mitofusin 2 (Mfn2), a mitochondrial membrane protein that promotes mitochondrial fusion, and of the mitochondrial superoxide scavenger enzyme ‘superoxide dismutase 2’ (Sod2)13 (Fig. 1h). Interestingly, the mRNA expression of the above-mentioned genes was reduced in cachectic muscles (Fig. 1i). More generally, 28% of genes associated with ‘lower in C26’ p-CREB1S133 binding overlapped with the list of the downregulated genes from RNA sequencing (RNA-seq) of C26 13 days versus controls (Extended Data Fig. 3d). Interestingly, analysis of this gene set revealed a strong enrichment in terms related to oxidative metabolism and mitochondrial function (Extended Data Fig. 3e). Collectively, the gene expression profile, together with p-CREB1S133 genome occupancy data in cachectic muscles, supports a coherent model of impaired CREB1-dependent transcriptional activity.
Notably, multiple cAMP–PKA–CREB-related terms (that is, ‘CREB signalling in neurons’, ‘Protein kinase A signalling’ and ‘cAMP-mediated signalling’) were predicted to be downregulated via IPA in a muscle transcriptomic analysis from 24 patients with pancreatic ductal adenocarcinoma-induced cachexia35 (Extended Data Fig. 4, IPA analysis and graph input data from Narasimhan et al.35), suggesting that the downregulation of cAMP–PKA–CREB1 signalling is a shared feature of muscle wasting in both preclinical models and patients with cachexia.
Altogether, these findings demonstrate that cachectic skeletal muscle features defective cAMP–PKA–CREB1 signalling coupled to an impaired expression of genes encoding mitochondrial proteins.
Pro-cachectic tumour-conditioned medium impairs cAMP–CREB1 signalling in myotubes
The observed impairment of the PKA–CREB1 signalling may depend on both cell autonomous and non-cell autonomous mechanisms. In the skeletal muscle, cAMP signalling is sustained by noradrenaline tonic release from the sympathetic innervation, which signals mostly through the β2AR22. However, muscle noradrenaline content in cachectic mice was not perturbed (Extended Data Fig. 5a) nor was the β2AR (Adrb2) mRNA expression modulated (Supplementary Table 1), suggesting that skeletal muscle sympathetic signalling was not impaired during cachexia. To uncover whether the defective PKA–CREB1 signalling relied on the tumour-induced cAMP dysregulation directly in the muscle, we used in vitro approaches. We analysed the cAMP signalling response in muscle cells treated with tumour-conditioned medium (TCM) from C26 colon carcinoma or Lewis lung carcinoma (LLC), two widely used pro-cachectic murine cancer cell lines5. Treatment of C2C12 myoblasts with C26 or LLC TCM for 24 h induced interleukin 6 (Il6) expression in a dose-dependent manner (Extended Data Fig. 5b,c) and STAT3- and NFκB-dependent transcriptional response (Extended Data Fig. 5d,e). Moreover, the two TCMs induced Il6 expression and atrophy in C2C12 myotubes, indicating that both were pro-cachectic in vitro (Extended Data Fig. 5f–h). We sought to investigate whether TCMs impair cAMP–PKA signalling in muscle cells in basal conditions and upon receptor(s) activation. First, we observed that the treatment with TCMs did not affect Adrb2 (β2AR) mRNA nor its surface expression in C2C12 myotubes (Extended Data Fig. 5i,j). Moreover, the two TCMs did not affect basal cAMP intracellular levels (Extended Data Fig. 5k) or basal PKA activity, as indicated by CREB1 and PKA substrate phosphorylation (Extended Data Fig. 5l–n). Interestingly, pretreatment with either C26 or LLC TCM inhibited the production of cAMP induced by isoproterenol (ISO)-dependent activation of β2AR, measured through the Förster resonance energy transfer (FRET)-based cytosolic cAMP sensor36 in C2C12 myoblasts (Fig. 2a–d) and through plate-based quantification in C2C12 differentiated myotubes (Fig. 2e). Consistently, signalling downstream cAMP was inhibited in C26 and LLC TCM pretreated myoblasts and myotubes. Indeed, ISO-induced CREB1 phosphorylation on serine 133, a PKA phosphorylation site37, was higher in control than in TCM-pretreated C2C12 myotubes (Fig. 2f,g) and myoblasts (Extended Data Fig. 6a–c). In addition, CRE-Luciferase reporter revealed that C26 and LLC TCM blunted the adrenergic-dependent CREB1 transcriptional activity (Fig. 2h,i).
a–d, cAMP dynamics following 10 µM ISO treatment measured through the FRET sensor Epac2-cAMP in C2C12 myoblasts treated for 24 h with control medium versus C26 TCM (a and b) or LLC TCM (c and d). Area under the curve (AUC) in b and d calculated from graphs in a and c from 50 s (ISO treatment) to 350 s. For a and b, n = 19 cells for each group from 3 independent experiments. For c and d, n = 40 control and n = 26 LLC TCM from 3 independent experiments. Two out of 26 data from LLC TCM exceed the y-axis range. For b, two-tailed t-test; for d, two-tailed Mann–Whitney test. e, Total cAMP measurement through plate-based quantification in C2C12 myotubes pretreated or not for 24 h with C26 TCM or LLC TCM, then treated with 10 µM ISO for 1, 5 or 10 min. n = 3 independent experiments. Two-way ANOVA. The statistical comparison between ISO and nt (P < 0.01 for all experimental conditions) is not shown in the graph. f,g, Representative blot (f) and densitometry analysis of phosphorylated over total CREB1 (g) after 5, 15 and 60 min from 10 µM ISO treatment in C2C12 myotubes pretreated for 24 h with C26 or LLC TCM. n = 3 independent experiments. Two-way ANOVA. h,i, CRE-Luciferase assay measuring CRE-dependent transcriptional response, quantified as relative light units (R.L.U.), following 6 h of 10 µM ISO treatment or not treated (nt) in C2C12 myoblasts pretreated for 24 h with C26 TCM (h) or LLC TCM (i). n = 5 independent experiments for h and n = 3 independent experiments for i. Two-way ANOVA. j,k, Ppargc1a (PGC-1α) (j) and Nr4a3 (NOR1) (k) mRNA expression following 10 µM ISO treatment at the indicated time in C2C12 myotubes pretreated with C26 TCM or LLC TCM for 24 h. n = 3 for each experimental group. Two-way ANOVA. l,m, Representative blot (l) and densitometry analysis of phosphorylated over total CREB1 (m) after 15 min from 10 µM ISO or 10 µM BAY60-6583 (BAY) treatment in C2C12 myotubes pretreated for 24 h with C26 TCM. n = 4 independent experiments. Two-way ANOVA. n, Nr4a3 (NOR1) mRNA expression following 1-h treatment of 10 µM ISO or 10 µM BAY treatment in C2C12 myotubes pretreated for 24 h with C26 TCM. n = 3 independent experiments. Two-way ANOVA. o,p, Representative blot (o) and densitometry analysis of phosphorylated over total CREB1 (p) after 15 min from 10 µM ISO treatment in human primary myotubes pretreated for 24 h with C26 or LLC TCM. n = 3 independent experiments. Two-way ANOVA. q, NR4A3 (NOR1) mRNA expression following 1 h of 10 µM ISO treatment in human primary myotubes pretreated for 24 h with C26 TCM or LLC TCM. n = 3 independent experiments. Two-way ANOVA. r,s, NR4A3 (NOR1) (r) and PPARGC1A (PGC-1α) (s) mRNA expression in muscle biopsies from precachectic (PC) and cachectic (C) patients with pancreatic cancer compared with controls. n = 51 controls, n = 34 PC, n = 48 C. One-way ANOVA. Data are presented as mean values ± s.e.m. in a–e, g–k, m, n and p–s, and only significant P values (<0.05) are annotated in the graphs.
Consistently, C26 and LLC TCM also impaired mRNA expression of PGC-1α (Ppargc1a) and NOR1 (Nr4a3), whose promoters carry one and three CRE sites, respectively38,39, in both C2C12 myotubes (Fig. 2j,k) and myoblasts (Extended Data Fig. 6d–f). Moreover, TCM modulates cAMP signalling not only in response to adrenergic stimulation, but also downstream of the adenosine A2B receptor (ADORA2B), a key regulator of muscle metabolism40. Indeed, treatment with C26 TCM impaired CREB1 phosphorylation induced by the ADORA2B agonist BAY-606583, as well as Nr4a3 expression (Fig. 2l–n), indicating that the TCM-induced cAMP dysfunction could be a general mechanism downstream of the Gαs protein-coupled receptors (GPCRs).
Notably, C26 or LLC TCM treatment affected ISO-induced CREB phosphorylation and NR4A3 expression also in human primary myotubes (Fig. 2o–q). Interestingly, the mRNA expression of NR4A3 (NOR1) was slightly decreased in the precachectic stage in muscle biopsies from patients with pancreatic cancer as compared with controls, while reaching significant reduction in patients showing >5% weight loss in the last 6 months (that is, cachectic) (Fig. 2r; demographic and clinical data are presented in Table 1). Moreover, the expression of the master regulator of mitochondrial biogenesis PPARGC1A (PGC-1α) was significantly lower at both the precachectic and cachectic stages, in line with mitochondrial impairment being an early feature of cachexia pathogenesis (Fig. 2s). Following sex-based stratification, male patients showed a weaker modulation of the analysed genes, while female patients showed a significant reduction of both NR4A3 and PPARGC1A at precachexia and cachexia stages (Extended Data Fig. 6g–j). Whether the sex-specific gene modulation merely reflects a heterogeneous and low cohort size or indicates a deeper consequence of hormonal regulation remains an open question.
Altogether, these data demonstrate that tumour-derived factors make murine and human muscle cells resistant to the activation of the cAMP–PKA pathway and CREB1-driven transcriptional reprogramming triggered upon Gαs-coupled GPCR stimulation.
TCM impairs cAMP signalling in myotubes through a PDE4-dependent mechanism
The recruitment and activation of PDEs, which hydrolyse cAMP to AMP, represent a major negative feedback mechanism involved in the modulation of the intracellular cAMP levels18. The PDE4 subfamily accounts for most of the cAMP hydrolysis in skeletal muscle41 and represents over 70% of the total PDE superfamily at mRNA level, of which Pde4b and Pde4d are the most abundant (Supplementary Table 1). Consistent with their induction in C26-induced cachexia (Extended Data Fig. 1m), Pde4b and Pde4d expression was also increased in muscles from KrasG12D/+;Lkb1f/f lung cancer-bearing mice31. Moreover, incubation with either C26 or LLC TCM induced the expression of PDE4B and PDE4D transcripts in C2C12 myoblasts, C2C12 myotubes and primary human myotubes (Fig. 3a–c). We thus hypothesized that PDE4 might account for the cAMP–PKA signalling dysfunction observed in our in vitro models. To test this hypothesis, we blocked PDE4 activity with rolipram, a well-characterized pan-PDE4 inhibitor25. Interestingly, ISO-induced cAMP responses were significantly lower in C26 TCM-treated and LLC TCM-treated C2C12, while treatment with 10 µM rolipram 5 min before ISO challenge completely restored cAMP levels to those of controls (Fig. 3d–i). Consistent with the FRET analysis, plate-based cAMP measurement performed on control versus TCM-pretreated C2C12 myotubes showed that targeting PDE4 fully restored the defective ISO-induced cAMP production (Fig. 3j). Also, the analysis of CREB1 phosphorylation and Nr4a3 expression supports a major role of PDE4 in suppressing cAMP signalling upon TCM treatment (Fig. 3k–m). Remarkably, rolipram per se enhanced basal CREB1 phosphorylation and Nr4a3 expression in TCM-pretreated cells and not in control cells, supporting the notion that pro-cachectic factors induce PDE4 activation in myotubes (Fig. 3k–m). Altogether, these data indicate that targeting PDE4 restores the tumour-induced cAMP signalling dysfunction in muscle cells.
a, Pde4a, Pde4b and Pde4d mRNA expression in C2C12 myoblasts following 24-h exposure to C26 TCM or LLC TCM. n = 3 independent experiments for control and C26 TCM, n = 4 independent experiments for LLC TCM. One-way ANOVA. b, Pde4a, Pde4b, Pde4c and Pde4d mRNA expression in C2C12 myotubes following 24-h exposure to C26 TCM or LLC TCM. n = 3 independent experiments. One-way ANOVA. c, PDE4B and PDE4D mRNA expression in human primary myotubes following 24-h exposure to C26 TCM or LLC TCM. n = 3 independent experiments. One-way ANOVA. d–f, cAMP responses measured through the FRET sensor Epac2-cAMP following 10 µM ISO treatment in C2C12 myoblasts pretreated for 24 h with C26 TCM. Two minutes before ISO challenge, 10 μM rolipram or vehicle (DMSO) was added. cAMP dynamics in control or C26 TCM in the presence of DMSO (d) or rolipram (e). The AUC (f) was calculated from graphs in d and e. n = 15 cells in control DMSO, n = 12 cells in C26 TCM DMSO, n = 19 cells in control + rolipram, and n = 20 cells in C26 TCM + rolipram from 3 independent experiments. Two-way ANOVA. g–i, cAMP responses measured through the FRET sensor Epac2-cAMP following 10 µM ISO treatment in C2C12 myoblasts pretreated for 24 h with LLC TCM. 10 μM rolipram or vehicle (DMSO) was added 2 min before ISO challenge. cAMP dynamics in control or C26 TCM in the presence of DMSO (g) or rolipram (h). The AUC (i) was calculated from graphs in g and h. n = 23 cells in control DMSO, n = 18 cells in LLC TCM DMSO, n = 46 cells in control + rolipram, and n = 51 cells in LLC TCM + rolipram from 3 independent experiments. Two-way ANOVA. j, Total cAMP measurement through plate-based quantification in C2C12 myotubes pretreated or not for 24 h with C26 TCM or LLC TCM and then treated for 1 min with ISO 10 µM. Two minutes before ISO challenge, 10 μM rolipram or vehicle (DMSO) was added. n = 3 independent experiments. Two-way ANOVA. k,l, Representative blot (k) and densitometry analysis of phosphorylated over total CREB1 (l) after 10 min from 10 µM ISO treatment in C2C12 myotubes pretreated for 24 h with C26 or LLC TCM. Two minutes before ISO challenge, 10 μM rolipram or vehicle (DMSO) was added. n = 4 independent experiments. Two-way ANOVA. m, Nr4a3 (NOR1) mRNA expression following 1 h of 10 µM ISO in C2C12 myotubes pretreated for 24 h with C26 or LLC TCM. Two minutes before ISO challenge, 10 μM rolipram or vehicle (DMSO) was added. n = 3 independent experiments. Two-way ANOVA. Data are presented as mean values ± s.e.m. in a–j, l and m, and only significant P values (<0.05) are annotated in the graphs.
PDE4 targeting mitigates muscle wasting in C26-bearing mice
To investigate the role of tumour-induced defective cAMP–PKA–CREB1 signalling in cancer-induced muscle wasting, we targeted PDE4 in vivo, by treating C26 tumour-bearing mice with rolipram daily from day 4 to day 13 upon cancer cell inoculation. Rolipram did not affect tumour weight (~200 mg) (Fig. 4a), nor mRNA expression of the pro-cachectic cytokines interleukin 1β (Il1b), IL6 (Il6), TNF (Tnf) and Activin A (Inhba) in tumour bulk (Extended Data Fig. 7a), suggesting that PDE4 targeting did not significantly affect tumour biology. In addition, rolipram treatment did not affect total weight loss (Fig. 4b), nor spleen enlargement or epididymal adipose tissue loss (Fig. 4c,d). In skeletal muscle, rolipram treatment raised cAMP levels together with phospho-PKA substrates (Fig. 4e–g), emphasizing a critical role for PDE4 in restraining cAMP signalling in this tissue. Initial analysis showed that PDE4 targeting in C26 tumour-bearing mice did not significantly protect against muscle weight loss of gastrocnemius and quadriceps compared with vehicle-treated animals (Fig. 4h,i), nor in the tibialis anterior, extensor digitorum longus (EDL) and soleus (Extended Data Fig. 7b–d). However, in cachectic gastrocnemius, rolipram treatment decreased tumour-induced expression of the major mediators of muscle atrophy, namely Trim63 (MuRF1), Fbxo32 (Atrogin1) and Fbxo30 (MUSA1)42,43 (Fig. 4j–l). This suggests that a putative anti-atrophic activity of rolipram may be masked by the variability in total body weight loss, which probably reflects differences in cachexia severity and may influence the kinetics of muscle wasting. To account for this potential confounding factor, we measured fat and lean body mass using EchoMRI and defined the experimental endpoint as the point at which mice had lost approximately 20% of their starting body weight. At this fixed timepoint, although rolipram treatment did not affect loss of fat and lean mass (Fig. 4m,n), a deeper analysis of myofibre cross-sectional area (CSA) in gastrocnemius muscle indicated that PDE4 targeting moderately counteracted myofibre shrinkage. In particular, the rolipram effect was more evident in type IIx intermediate and type IIb fast-glycolytic fibres, which are the fibre types most sensitive to atrophy during cancer cachexia (Fig. 4o,p and Extended Data Fig. 7e).
a, Tumour weight at 13 days from C26 cell subcutaneous injection in vehicle- versus rolipram-treated mice (C26 veh versus C26 Rol). n = 8 mice for each experimental group. Two-tailed t-test. b, Percentage of total weight loss at 13 days from cancer cell injection over the starting weight (day 0) of C26 veh versus C26 Rol. n = 8 mice for each experimental group. Two-tailed t-test. c,d, Spleen (c) and epididymal fat (d) weight at 13 days from tumour cell injection in controls (ctr), C26 veh and C26 Rol mice. n = 5 ctr mice, n = 8 C26 veh mice and n = 8 C26 Rol mice. One-way ANOVA. e, cAMP measurement through plate-based quantification from gastrocnemius muscle of non-tumour-bearing mice (ctr), C26 veh and C26 Rol mice. cAMP amount (pmol) was normalized over milligram of tissue. n = 6 ctr mice and n = 7 C26 veh and C26 Rol mice. One-way ANOVA. f,g, Representative blot (f) and densitometry analysis of phospho(serine/threonine)-PKA substrates over GAPDH (g) of whole gastrocnemius lysate from controls (ctr), C26 veh and C26 Rol mice. n = 5 mice for each experimental group. One-way ANOVA. h,i, Gastrocnemius (h) and quadriceps (i) weight normalized over mouse starting weight in controls (ctr), C26 veh and C26 Rol mice. n = 5 ctr mice, n = 8 C26 veh and C26 Rol mice. One-way ANOVA. j–l, mRNA expression fold change over control of Fbxo32 (Atrogin1) (j), Trim63 (MuRF1) (k) and Fbxo30 (MUSA1) (l) at day 13 after C26 tumour cell injection in gastrocnemius muscle of control (ctr), C26 veh and C26 Rol mice. n = 6 ctr mice, n = 8 C26 veh and C26 Rol mice for j and k. n = 8 mice for each experimental group for l. One-way ANOVA. m,n, EchoMRI analysis of fat mass (m) and lean mass (n) percentage change over starting point. n = 9 mice for each experimental group. One-way ANOVA. o,p, Representative immunofluorescence staining (o) and CSA quantification (p) of myofibre type I, IIa, IIx and IIb in gastrocnemius sections of control (ctr), C26 veh and C26 Rol mice. n = 5 mice for each experimental group. One-way ANOVA. Data are presented as mean values ± s.e.m. in a–e, g–n and p, and only significant P values (<0.05) are annotated in the graphs.
Moreover, we explored through western blot whether rolipram affects classical pathways involved in muscle wasting during cancer cachexia. These include the expression of autophagy markers (that is, phosphatidylethanolamine-conjugated LC3B-II and Beclin-1) and inflammation markers (that is, STAT3 and NF-κB), together with downstream targets of the mTOR (mechanistic target of rapamycin) signalling (that is, AKT, 4EBP1 and S6), which sustains protein synthesis and inhibits degradation3,43. While p-S6S235/236/S6 ratio and p-4EBP1T37/T46 were unaffected in cachectic muscles, western blot analysis revealed reduced p-AKTS473 and, as a possible compensatory response, increased total AKT in cachectic muscles (C26 vehicle versus controls) (Extended Data Fig. 7f–j). Rolipram did not affect AKT phosphorylation but prevented AKT accumulation seen in untreated C26 mice (Extended Data Fig. 7f,i,j). Moreover, both LC3B-II and Beclin-1 levels were elevated in cachectic muscles; however, only Beclin-1 accumulation was reduced by rolipram treatment, suggesting that PDE4 inhibition modulates autophagy (Extended Data Fig. 7f,k,l). Regarding the inflammation markers, rolipram decreases NF-κB phosphorylation, while it did not alter STAT3 expression nor phosphorylation (Extended Data Fig. 7f,m–o). As PDE4 inhibition affects glucose homeostasis in mice44, we performed a glucose tolerance test in controls and vehicle- or rolipram-treated C26 tumour-bearing mice. C26 mice exhibited a clear hypoglycaemic phenotype at time 0 (before glucose injection), while showing a glycaemic profile after glucose injection comparable to the control (Extended Data Fig. 7p,q). PDE4 targeting did not affect either the hypoglycaemic phenotype or the glycaemic dynamics (Extended Data Fig. 7p,q).
Overall, these observations suggest that rolipram marginally affects multiple pathways implicated in muscle cachexia and that restoring cAMP–PKA signalling by PDE4 targeting in vivo moderately mitigates muscle wasting in tumour-bearing mice.
PDE4 targeting restores mitochondrial- and oxidative metabolism-related gene expression in skeletal muscle from cachectic mice
To assess the contribution of defective cAMP–PKA–CREB1 signalling to tumour-induced reprogramming of the skeletal muscle transcriptome, we performed bulk RNA-seq analysis on gastrocnemius muscles from control mice and from vehicle- or rolipram-treated C26 tumour-bearing mice at 13 days after cancer cell injection (Supplementary Table 1). Principal component analysis (PCA) revealed a clear clustering of cachectic muscles versus controls through the PC1 axis (63% variance), while the PC2 axis (8% variance) discriminated between rolipram- versus vehicle-treated cachectic mice (C26 Rol versus C26 veh) (Fig. 5a). Out of 8,395 DEGs in C26 veh versus control, 23.4% (1,965) were also controlled by rolipram (shared DEGs with the C26 Rol versus C26 veh comparison) (Supplementary Table 1). Heatmap clustering analysis of this gene subset indicated that rolipram-regulated genes in cachectic mice strongly cluster towards controls (Fig. 5b). Indeed, out of 1,965 shared DEGs, 1,816 genes (92.4%) were ‘rolipram-rescued’ genes, that is, genes either downregulated in cachexia and upregulated by rolipram treatment or vice versa. We identified 854 downregulated genes in cachectic muscles whose expression was induced by rolipram, and 962 upregulated genes in cachectic muscles whose expression was reduced by rolipram treatment. IPA of these two groups of genes revealed that the top term activated by rolipram was ‘oxidative phosphorylation’, while the top inhibited was ‘mitochondrial dysfunction’ (Fig. 5c). Thus, we focused our attention on mitochondrial-related pathways intersecting our dataset with the web-based platform MitoXplorer45. Rolipram-rescued genes are 51.8% of the ‘oxidative phosphorylation‘ MitoXplorer list (Fig. 5d). In particular, rolipram treatment rescued the expression of genes encoding subunits of CI, such as Ndufa11, Ndufb10 and Ndufb5, the CII assembly protein Sdhaf4, subunits of the CII Sdhb, Sdhc and Sdhd, subunits of CIII (such as Uqcrfs1 and Uqcrq), subunits of CIV (such as Cox5a and Cox5b) and components of the mitochondrial ATP synthase CV (such as Atp5o and Atp5c1), which were all downregulated in cachectic muscles (Fig. 5d). Moreover, rolipram rescued the expression of mitochondrial dynamic-related genes (37% of ‘mitochondrial dynamics’ MitoXplorer list), including Mfn2, Mtfr1l, Oma1, Mtfp1, Mief2 and Rhot2 (Fig. 5e). Among the rolipram-rescued genes, we also found key mediators of muscle mitophagy (38.1% of ‘mitophagy’ MitoXplorer list), including Prkn and Ulk146 (Fig. 5f). Notably, rolipram treatment rescued the expression of 52 out of 184 genes of the MitoXplorer mitochondrial ‘translation’ list, including the key mediator of mitochondrial ribogenesis (Rbfa) and several mitochondrial ribosomal proteins (Mrpl and Mrps) (Extended Data Fig. 8a). Of note, several rolipram-rescued genes belonging to the MitoXplorer list showed reduced p-CREB1S133 binding in cachectic muscles, suggesting a causative link between the impaired cAMP–PKA–CREB1 signalling and the transcriptional downregulation of mitochondrial function-related genes in cachectic muscles. Indeed, out of 168 rolipram-rescued genes featuring lower p-CREB1S133 binding in precachectic muscle, 49 were mitochondrial-related genes. These genes included several subunits of the electron transport chain and ATP synthase such as Ndufb4, Ndufb5, Ndufb1, Ndufb2, Sdhc, Sdhd, Cox5a, Atp5j2, Atp5b and several key players of mitochondrial translation, dynamics and reactive oxygen species defence such as Rbfa, Mrpl12, Mrpl2, Mrpl34, Mff, Mfn2 and Sod2 (Supplementary Tables 1 and 3).
a, PCA of RNA-seq of gastrocnemius muscle from control (ctr), and C26-bearing mice treated with vehicle or rolipram (C26 veh and C26 Rol) at 13 days from cancer cell inoculation. The confidence ellipses have been obtained with the stat ellipse function, which draws a 95% confidence level for a multivariate t-distribution. n = 7 ctr and C26 veh mice and n = 5 C26 Rol mice. b, Heatmap generated following bidirectional hierarchical clustering (gene- and sample-based) of normalized RPKM of DEGs common between C26 veh versus ctr and C26 Rol versus C26 veh comparisons. n = 7 ctr and C26 veh mice and n = 5 C26 Rol mice. c, IPA canonical pathway of genes rescued by rolipram (down in C26 veh versus ctr and up in C26 rol versus C26 veh or vice versa). The P value ranking-based top 10 terms are shown. n = 7 ctr and C26 veh mice and n = 5 C26 Rol mice. d–f, Heatmap of normalized RPKM of rolipram-rescued DEGs selected from MitoXplorer gene lists: ‘oxidative phosphorylation’ (d), ‘mitochondrial dynamics’ (e) and ‘mitophagy’ (f). n = 7 ctr and C26 veh mice and n = 5 C26 Rol mice.
Altogether, our transcriptomic analysis revealed that boosting defective cAMP signalling rescues the expression of a core transcriptional network encoding mitochondrial proteins, which are downregulated in cachectic muscles. Furthermore, the most remarkable effect of PDE4 targeting was the regulation of several genes that coordinate transcription and translation of mitochondrial components, thus suggesting that defective cAMP signalling in cachectic muscle may contribute to the mitochondrial and oxidative dysfunction.
Inhibition of PDE4 counteracts mitochondrial and oxidative metabolic dysfunction in muscles of cachectic mice
As rolipram rescued the expression of key players of mitochondrial dynamics and translation, we measured mtDNA copy number through quantitative polymerase chain reaction (qPCR) relative quantification. C26-bearing mice displayed a ~50% decrease in mtDNA copy number compared with controls, while rolipram treatment counteracted this decline, sparing ~75% of mtDNA copy number (Fig. 6a). Accordingly, protein expression of voltage-dependent anion channel (VDAC) (a mitochondrial mass marker) and of mitochondrial electron transport chain subunits was reduced in muscle of C26-bearing mice and rescued by rolipram treatment (Fig. 6b,c).
a, mtDNA over nDNA qPCR analysis in gastrocnemius muscles from control (ctr) and vehicle- or rolipram-treated C26-bearing mice (C26 veh and C26 Rol). Primers are specific for mt-Nd6 (for mtDNA) and Hk2 (for nDNA) genes. n = 9 ctr mice, n = 8 C26 veh and C26 Rol mice. One-way ANOVA. b, OXPHOS blot of gastrocnemius lysate from ctr, C26 veh and C26 Rol mice showing the protein expression of subunits of the mitochondrial respiratory chain complexes CI, CII, CIII and CV, of VDAC and vinculin. Representative of 2 independent blots of 6 mice for each experimental group. c, Densitometry analysis of VDAC expression normalized over vinculin. n = 6 mice for each experimental group. One-way ANOVA. d, Representative trace of high-resolution respirometry, O2 concentration (black line) and O2 flux (light-blue line), during the SUIT protocol for mitochondrial respiratory capacity in permeabilized mouse gastrocnemius muscle fibres. PM, pyruvate + malate, leak respiration; ADP, OXPHOS capacity; Cyt C, cytochrome c, integrity of outer mitochondrial membrane; G, glutamate; S, succinate, uncoupler FCCP; Rot, rotenone; Ama, antimycin A; ROX, residual oxygen consumption; TMPD, N,N,N′,N′-tetramethyl-p-phenylenediamine; As, ascorbate; Az, azide. e,f, OCR normalized over tissue weight (pmol s−1 mg−1) (e) and OXPHOS CII specific flux calculated as the subtraction of OXPHOS CI from OXPHOS CI + CII (f) measured in explanted gastrocnemius muscles from ctr, C26 veh and C26 Rol mice. n = 3 ctr mice for e and f. n = 5 C26 veh and n = 6 C26 Rol mice for e and n = 4 C26 veh and n = 5 C26 Rol mice for f. One-way ANOVA. g, Representative traces of high-resolution respirometry during the SUIT protocol for mitochondrial respiratory capacity in intact C2C12 myotubes. Oxygen flux in the routine state (R), in the leakage state (L) after addition of oligomycin (an inhibitor of ATP synthase complex) and in maximum respiratory capacity (ETS) after the addition of carbonyl cyanide p-trifluoromethoxyphenylhydrazone (FCCP). P, pyruvate; Omy, oligomycin; Rot, rotenone; Ama, antimycin A; ROX, residual oxygen consumption. h,i, OCR normalized over protein content (pmol s−1 mg−1) of the routine state, leakage and maximum capacity (ETS) of myotubes pretreated for 48 h with C26 TCM in the presence of rolipram or vehicle (DMSO) (h). Oxygen consumption linked to ATP production, that is, oligomycin-sensitive respiration obtained by the subtraction of L from R (i). n = 4 ctr, n = 5 C26 TCM, n = 4 C26 TCM + Rol independent experiments. One C26 TCM Leak value exceeds the negative y-axis limit. Two-way ANOVA for h (differences of ETS versus Routine and ETS versus Leak are not indicated in the graph, for all P < 0.0001) and one-way ANOVA for i. j,k, Representative picture (j) and quantification of the postsynaptic NMJ volume (k) following rendering of Leica SP5 z-stack projection of α-bungarotoxin 488 staining in EDL muscle, in ctr, C26 veh and C26 Rol mice. n = 70 ctr, n = 104 C26 veh, n = 101 C26 Rol NMJ(s) quantified from 3 mice for each experimental group. One-way ANOVA. l, Peak force of five sequential grip test measurements at day 0, day 6 and day 13 for each mouse. Peak force is indicated as a percentage of the starting strength. n = 8 mice for each experimental group. Two-way ANOVA, *P value in C26 veh versus C26 Rol; #P value in C26 veh or Rol versus starting force. Data are presented as mean values ± s.e.m. in a, c, e, f, h, i, k and l, and only significant P values (<0.05) are annotated in the graphs.
To assess the overall muscle respiratory capacity, we evaluated the oxygen consumption rate (OCR) in saponin-permeabilized muscle fibres using a multiple substrate–uncoupler–inhibitor titration (SUIT) protocol47,48 (Fig. 6d). Following the addition of ADP in saturating concentrations, we measured the oxidative phosphorylation (OXPHOS) state, that is, maximum ATP-coupled respiration through CI (after the addition of glutamate) and through both CI and CII (after the addition of succinate). OXPHOS CI + CII was significantly lower in cachectic muscles and rescued in rolipram-treated mice (Fig. 6e). Coherently, this result was recapitulated in OXPHOS CII obtained by the subtraction of OXPHOS CI from OXPHOS CI + CII (Fig. 6f). Maximum electron transfer system capacity (ETS CI + CII, following the addition of the exogenous uncoupler FCCP) as well as submaximal ETS CII respiratory state (after CI inhibition by adding rotenone) were also reduced in cachectic gastrocnemius and recovered by the rolipram treatment (Fig. 6e). Finally, the hampered CIV activity was partially preserved by rolipram (Fig. 6e). Consistent with impaired mitochondrial oxidative function, cachectic muscles exhibited increased phosphorylation of the energy sensor AMPK, indicative of activation under energetic stress—a response that was fully normalized in skeletal muscle from rolipram-treated mice (Extended Data Fig. 8b,c). Consistently with the ex vivo data, rolipram treatment of C2C12 myotubes prevented the defective respiration capacity of the electron transport system (ETS) and ATP-linked respiration induced by C26 TCM (Fig. 6g–i), highlighting a muscle cell-autonomous effect. Altogether, the ex vivo and in vitro data strongly indicate that rescuing the defective cAMP signalling in cachectic muscles restores mitochondrial mass and function.
Emerging evidence indicates that oxidative metabolism and mitochondrial fitness in the skeletal muscle deeply modulate neuromuscular junction (NMJ) structure and turnover as well as muscle functionality49,50,51, two important aspects of muscle physiology that are affected during cancer cachexia52. Consistently, rolipram treatment counteracted postsynaptic NMJ disruption observed in cachectic mice (Fig. 6j,k) and improved muscle force production, tested through the hind-limb grip test (Fig. 6l). Moreover, vehicle-treated tumour-bearing mice showed a significant impairment in endurance compared with controls, while rolipram-treated mice exhibited an intermediate endurance phenotype, with a partial improvement compared with vehicle-treated C26 mice that approached, but did not reach, statistical significance (P = 0.06) (Extended Data Fig. 8d). Overall, these results indicate that PDE4 inhibition in mice undergoing cachexia restores muscle mitochondrial mass and the overall muscle oxidative capacity and muscle functionality.
PDE4D isoform is a critical mediator of cAMP and metabolic dysfunction in skeletal muscle in cancer cachexia
To dissect the specific contribution of different PDE4 isoforms in tumour-induced defective cAMP signalling, we generated CRISPR–Cas9-deleted C2C12 for either Pde4b or Pde4d (Extended Data Fig. 9a–c), the two PDE4 isoenzymes whose mRNA expression was induced in C2C12 and human primary myotubes upon incubation with pro-cachectic media and in cachectic muscles (Fig. 3a–c and Extended Data Fig. 1m). Thus, we compared the activation of the cAMP–CREB1 signalling upon ISO-induced β2AR stimulation in wild type (WT), Pde4b−/− and Pde4d−/− C2C12. Genetic ablation of Pde4d, but not of Pde4b, strongly increased ISO-induced cAMP levels in C2C12 myoblasts and completely rescued cAMP levels in TCM-pretreated cells (Fig. 7a–c). Notably, while rolipram induced a rapid increase in cAMP levels in WT and Pde4b−/− C2C12 (Fig. 7a), Pde4d−/− C2C12 cells were completely insensitive to rolipram treatment (Fig. 7b), suggesting that PDE4D plays a major role in restricting cAMP signalling downstream of β2AR in these cells. Consistently, deletion of Pde4d—and not of Pde4b—in C2C12 myoblasts enhanced ISO-induced CREB1S133 phosphorylation (Extended Data Fig. 10a–d) and the expression of NOR1 (Nr4a3) (Extended Data Fig. 10e). These findings are consistent with previous data from mouse embryonic fibroblasts, which showed that PDE4D—but not PDE4B—mediates cytosolic cAMP degradation, thereby restricting CREB1-dependent transcriptional activity53. Notably, treatment of primary human myotubes with the specific PDE4D inhibitor zatolmilast26 rescues the defective CREB1S133 phosphorylation in the presence of C26 TCM comparably to the pan-PDE4 inhibitor rolipram (Fig. 7d,e). Altogether, these data indicate that PDE4D, rather than PDE4B, is responsible for tumour-induced impairment of cAMP signalling.
a,b, cAMP dynamics measured through the FRET sensor Epac2-cAMP in WT, Pde4b−/− (a) or Pde4d−/− (b) cells pretreated with control or C26 TCM for 24 h. At the indicated timepoint, 10 μM ISO and 10 μM rolipram (Rol) were added. Experiments in a and b were done together, with the same WT as matched control, but divided into two graphs to simplify the visualization. n = 25 WT control cells, n = 27 WT C26 TCM cells, n = 30 Pde4b−/− control cells, n = 20 Pde4b−/− C26 TCM cells, n = 20 Pde4d−/− control cells, n = 32 Pde4d−/− C26 TCM cells from 3 independent experiments. c, AUC calculated from cAMP dynamics measured from 100 s (ISO treatment) to 350 s. n = 23 WT control, n = 29 WT C26 TCM, n = 28 Pde4b−/− control, n = 18 Pde4b−/− C26 TCM, n = 18 Pde4d−/− control and n = 30 Pde4d−/− C26 TCM cells. One-way ANOVA. d,e, Representative blot (d) and densitometry analysis of phosphorylated over total CREB1 (e) after 10 min from 10 µM ISO treatment in human primary myotubes pretreated for 24 h with C26 TCM. Two minutes before ISO challenge, 10 μM rolipram, 10 μM zatolmilast or vehicle (DMSO) was added. n = 3 independent experiments. Two-way ANOVA. f,g, Relative frequency percentage of Pde4b (f) and Pde4d (g) transcript variant over total Pde4b or Pde4d in gastrocnemius muscle from control and cachectic mice (C26 13d). Data are represented as means from n = 7 mice for each experimental group. h, Representative image of whole tibialis anterior after 3 weeks from AAV9-mediated transduction with shRNA against Pde4d expressing an EGFP reporter. Merge of immunostaining anti-laminin (grey signal) with EGFP endogenous signal. Scale bar, 1 mm. i, mRNA expression of Pde4b and Pde4d at the endpoint, on bulk tibialis anterior from non-tumour-bearing mice transduced with shCtr (Sham shCtr) and from C26 tumour-bearing mice transduced with shCtr (C26 shCtr) or shPde4d (C26 shPde4d). n = 8 sham shCtr mice, n = 10 C26 shCtr and n = 8 C26 shPde4d mice. Two-way ANOVA. j, cAMP measurement through plate-based quantification from tibialis anterior from Sham shCtr, C26 shCtr and C26 shPde4d mice. cAMP amount (pmol) was normalized over milligram of tissue. n = 5 Sham shCtr mice, n = 8 C26 shCtr mice and n = 6 C26 shPde4d mice. One-way ANOVA. k,l, Oxygen consumption of OXPHOS CI (k) and OXPHOS complexes I + II (l) rate normalized over tissue weight (pmol s−1 mg−1) measured in explanted tibialis anterior from sham shCtr, C26 shCtr and C26 shPde4d mice. n = 4 mice for each experimental group. One-way ANOVA. Data are presented as mean values ± s.e.m. in a–c, e and i–l, and only significant P values (<0.05) are annotated in the graphs.
Notably, different Pde4b and Pde4d transcript variants, originating from alternative splicing events or alternative promoter usage, generate long, short and super-short isoforms according to the presence of distinct N-terminal sequences and to the inclusion or not of the regulatory UCR1 and UCR2 domains26. These variants differ in subcellular compartmentalization, dimerization and intramolecular regulation of the enzymatic activity26. We thus analysed data from our RNA-seq for transcript variant differential expression (Supplementary Table 4), and we observed that, among the Pde4b variants, Pde4b-203, Pde4b-204 and Pde4b-207 were induced transcriptionally (Extended Data Fig. 10f), but their relative frequency distribution was not substantially affected (Fig. 7f). Intriguingly, the Pde4d super-short isoform Pde4d-204 and the long isoform Pde4d-213 were strongly upregulated (that is, 53- and 24-fold increase, respectively) in muscle from cachectic mice compared with controls, whereas the long isoform Pde4d-205 was downregulated (Extended Data Fig. 10g). This transcriptional reassortment resulted in major changes in the relative frequency of the different Pde4d transcripts (Fig. 7g). Of note, Pde4d-204, encoding the homologue of human PDE4D2, a super-short isoform that lacks the autoinhibitory domain26, shifted from the 1.68% to 36.11% of the total Pde4d transcripts in cachectic muscle (Fig. 7g). Pde4d-213, which encodes the homologue of human PDE4D8, a β2-adrenergic-associated long isoform, shifted from 0.35% to 3.42% in cachectic muscle. Pde4d-205, which encodes a highly regulated and compartmentalized long isoform, shifted its relative abundance from 41.22% to 5.39% in the cachectic muscles (Fig. 7g).
Altogether, these data support a possible central role of the PDE4D isoform in mediating cAMP dysfunction also in vivo. To test this hypothesis, we transduced tibialis anterior with AAV9 particles expressing Pde4d shRNA, together with an enhanced green fluorescent protein (EGFP) reporter, 2 weeks before C26 inoculation. At day 13 after cancer cell injection, most of the fibres were EGFP positive, although with different EGFP intensity levels (Fig. 7h), and the C26-induced upregulation of Pde4d, but not of Pde4b, was significantly reduced in Pde4d shRNA-transduced muscles (shPde4d) as compared with control shRNA (shCtr) (Fig. 7i), indicating a robust transduction efficiency. Moreover, muscle-specific knockdown of Pde4d led to a significant increase in total muscle cAMP levels compared with the control group, indicating an effective modulation of the signalling pathway (Fig. 7j). shCtr and shPde4d tumour-bearing mice showed comparable tumour weight, total weight change, spleen enlargement and tibialis anterior weight loss (Extended Data Fig. 10h–k). However, fibre CSA distribution highlighted a mild, yet significant, shift of shPde4d group towards non-tumour-bearing mice (Extended Data Fig. 10l). According to a moderate anti-atrophic effect, the expression of MuRF1 (Trim63) and Atrogin-1 (Fbxo32) was partially reduced upon Pde4d silencing, while MUSA1 (Fbxo30) expression was not affected (Extended Data Fig. 10m). Different to what we observed in gastrocnemius muscles from C26-bearing mice, mRNA expression of mitochondrial-related genes (Atp5j2, Mrpl12, Rbfa, Sdhc, Sod2, Cox5a, Mfn2 and Ndufb4) was highly variable and not reduced in cachectic tibialis anterior, nor was it changed upon Pde4d silencing (Extended Data Fig. 10n), possibly reflecting muscle-specific differences in the timing and dynamics of transcriptional remodelling during cachexia. Transduction of the tibialis instead of gastrocnemius was prompted by the technical limitation of achieving sufficient PDE4D silencing in the bulk muscle (Supplementary Fig. 1a–d). However, whole-muscle respirometry analysis of transduced tibialis anterior revealed that, as compared with the non-tumour-bearing group, the OXPHOS state through both CI and CII was significantly lower in C26 shCtr and was rescued upon Pde4d silencing (Fig. 7k,l). Similar results, although not reaching statistical significance, were obtained when measuring the OXPHOS state CI + CII, the maximum electron transfer system capacity (ETS) of CI + CII and the submaximal ETS CII respiratory state, while CIV activity was strongly impaired despite Pde4d silencing (Extended Data Fig. 10o–t). Collectively, these results underscore the muscle-cell-autonomous nature of our observations and suggest a potential, although not exclusive, role for the PDE4D isoform in cancer-induced muscle wasting and mitochondrial dysfunction. Altogether, PDE4D emerges as a promising target to alleviate muscle metabolic dysfunction in cancer cachexia.
Discussion
This study reports that impaired cAMP–PKA–CREB1 signalling drives metabolic dysfunction in skeletal muscle during cancer cachexia in mice. Physiologically, cAMP tonic signal within muscle fibres strongly depends on low-grade continuous activation of the β2AR, sustained by the noradrenaline release from the sympathetic nervous system22,54. Other Gαs-GPCR(s), such as the adenosine receptor A2B, contribute to the overall cAMP signalling modulation in the skeletal muscle, regulating metabolism and protein turnover40. Together, these upstream signals converge towards intracellular cAMP increase and consequent PKA-mediated activation of CREB1 transcription factor. Interestingly, pro-cachectic tumour-derived factors impair cAMP signalling downstream of both β2-adrenergic and adenosine 2B receptors in myotubes, suggesting a common mechanism acting on Gαs-GPCR(s) signalling pathway. In cachectic muscle, we observed reduced PKA activity coupled with a reduction of p-CREB1S133 occupancy on promoter of genes involved in mitochondrial function and encoding mitochondrial proteins, whose expression was also diminished. The muscle-cell-autonomous nature of our observations was supported by in vitro experiments on muscle cells exposed to pro-cachectic TCM, which linked cAMP–PKA–CREB1 signalling impairment to mitochondrial dysfunction.
Notably, the mRNA expression of the CREB1 target genes PGC-1α and NOR1, crucial for muscle oxidative function, was downregulated in muscle biopsies from cachectic patients with pancreatic cancer. Besides the analysis at the single gene level, a signature of cAMP signalling dysregulation was predicted through IPA in muscle from cachectic patients with pancreatic ductal adenocarcinoma in a previously published transcriptomic dataset35. Moreover, consistent with this predictive analysis, our data on human primary myotubes suggest that cAMP signalling dysfunction might also occur in human cachexia.
Besides demonstrating that pro-cachectic factors impair muscle cAMP–PKA–CREB1 signalling, our study identifies the functional consequences of this dysregulation. Indeed, stabilizing cAMP levels by targeting PDE4 effectively rescues mitochondrial respiratory capacity and muscle functionality. Given the multifaceted nature of cancer cachexia, current strategies are oriented towards a combinatorial therapeutic approach targeting muscle wasting at multiple levels2. Here, we identified cAMP signalling impairment as a molecular mechanism underlying primarily one aspect of muscle wasting, namely mitochondrial dysfunction, offering a promising framework for a multimodal therapeutical approach. Besides this main metabolic effect, PDE4 targeting mitigates type IIx and IIb glycolytic fibre shrinkage in cachectic muscles. This fibre-type-selective effect could arise from a broader dynamic range in terms of size decrease of glycolytic fibres during wasting conditions, thereby making it easier to detect even subtle or partial rescue effects. Moreover, the potential impact of mitochondrial dysfunction on the overall muscle wasting may be more pronounced in fibres with lower mitochondrial content, as their limited mitochondrial reserve heightens their vulnerability and makes them more likely to exceed the threshold at which mitochondrial dysfunction triggers muscle wasting.
Mechanistically, the upstream events leading to impairment of the cAMP pathway were herein only partially identified. However, alongside the transcriptional deregulation of several key players of the signalling, the cAMP-hydrolysing PDE enzymes—PDE4B and PDE4D—emerged as a feasible target to effectively reverse cAMP impairment. The pan-PDE4 inhibitor rolipram proved to be a valuable tool for enhancing muscle cAMP signalling and provided proof of concept for the molecular mechanism underlying mitochondrial dysfunction in cachexia. Besides cancer cachexia, mitochondrial dysfunction is a hallmark of other conditions related to chronic low-grade inflammation, including ageing-induced muscle wasting (that is, sarcopenia)13. Whether cAMP signalling dysfunction might represent an underlying mechanism of mitochondrial impairment also during ageing remains an open and intriguing question. Interestingly, rolipram has been shown to improve mitochondrial mass and function in ageing mice25, reinforcing the potential relevance of this pathway beyond cancer-related conditions. However, from a translational perspective, rolipram is unlikely to represent a viable therapeutic option owing to the considerable side effects observed in the gastrointestinal tract and central nervous system, which have limited its clinical applicability26. Interestingly, selective silencing of PDE4D in skeletal muscle effectively improved mitochondrial function, highlighting this PDE4 isoform as a promising therapeutic target to counteract the metabolic dysfunction associated with muscle wasting. Of note, the PDE4D-selective inhibitor zatolmilast is currently in phase III clinical trial for cognitive-related disorders55, indicating that selective inhibition of this isoform is achievable and potentially safe.
While our transcriptomic data strongly support the dominant role of the PKA–CREB1 transcriptional axis in mediating the metabolic rescue, we cannot exclude a potential contribution from other cAMP sensors. For instance, the cAMP sensor Epac1/RAPGEF3 has been linked to AMPK activation25 and muscle metabolic adaptation during exercise56. However, its lower mRNA expression compared with PKA catalytic and regulatory subunits (Supplementary Table 1) suggests a limited role in mediating the observed metabolic effects in skeletal muscle. Moreover, the profound restoration of a core transcriptional network related to mitochondrial homeostasis following PDE4 inhibition underscores the centrality of the transcriptional branch in this specific pathological context. Nevertheless, in contrast to the observations in gastrocnemius muscles, in tibialis anterior from C26 tumour-bearing mice the impaired mitochondrial respiration was not coupled with a reduction of mitochondrial-related gene expression, at least at the experimental endpoint (Fig. 7k,l and Extended Data Fig. 10n–t). This does not contradict our central hypothesis but rather highlights the well-documented heterogeneity of the cachectic response across different skeletal muscles. Several non-mutually exclusive factors could explain this discrepancy. First, muscles such as the gastrocnemius and tibialis anterior have distinct fibre-type compositions, metabolic profiles and patterns of functional recruitment, all of which influence their susceptibility and adaptive responses to systemic stressors such as cachexia57. Second, it is plausible that we are observing a temporal snapshot of a dynamic process. An additional layer of complexity in the molecular alterations underlying cAMP dysfunction during cachexia may arise from the dramatic tumour-induced reassortment of the PDE4D transcript variants. Indeed, different PDE4D transcript variants, depending on the presence of unique N-terminal sequences and to the inclusion or not of the UCR1 and UCR2 domains, display distinct localization, regulation and activity26. Of note, we observed a strong induction of Pde4d-204, which encodes the homologue of the human PDE4D2, a super-short soluble isoform lacking the UCR1 autoinhibitory domain26. The induction of this super-short variant points to a sophisticated regulatory mechanism. Understanding the upstream signals and the transcriptional machinery (for example, alternative promoter usage or splicing events) that orchestrate this switch in response to the tumour microenvironment represents a critical and exciting direction for future research.
In conclusion, our study identifies a mechanism underlying mitochondrial dysfunction in skeletal muscle during cancer cachexia. In particular, we show that the cAMP–PKA–CREB1 signalling within muscle is impaired during cachexia onset. We identify several CREB1 target genes encoding key regulators of mitochondrial biogenesis and components of the mitochondrial electron transport chain, and we demonstrate that the overall reduction of CREB1 function contributes to the transcriptional collapse and consequent mitochondrial dysfunction observed in cachectic muscles. Moreover, boosting cAMP signalling by targeting PDE4 restores the oxidative metabolism-related transcriptional profile and oxidative function within the muscle during cancer-induced cachexia. Finally, we identify PDE4D isoform as a key contributor of tumour-induced mitochondrial dysfunction in skeletal muscle. Our findings position PDE4(D) inhibition as a potent strategy to counteract the metabolic collapse that precedes and exacerbates muscle wasting, rather than a standalone cure for cachexia. This highlights its potential for use in multimodal therapeutic approaches targeting distinct facets of the syndrome, such as inflammation or hypercatabolism.
Methods
Ethical statement
All research was conducted in compliance with relevant ethical regulations. Animal studies were carried out under protocols approved by the Italian Ministry of Health (authorization numbers 668/2017-PR, PI A.G.; and 572/2022-PR, PI M.S.). The human research component was approved by the Ethical Committee for Clinical Experimentation of Padova (protocol numbers 3674/AO/15, 5609/AO/22 and 5677/AO/23). All patients were enrolled in accordance with the principles of the Declaration of Helsinki, and written informed consent was obtained from all participants.
Cell lines
C2C12 myoblasts (ATCC) were cultured in high-glucose Dulbecco’s modified Eagle medium (DMEM) (Euroclone; ECM0103L) supplemented with 10% of foetal bovine serum (FBS; Euroclone). For differentiation of myoblasts to myotubes, myoblasts were cultured in DMEM 10% FBS until they reached 90% confluency, then differentiation was induced by switching to DMEM supplemented with 2% horse serum (HS; Euroclone) for 96 h. The C2C12 cell line was used at passages ranging from 10 to 25 following purchase from ATCC. Human primary cells were purchased by William Cook Europe ApS (catalogue number P01062-17M from 17-year-old male subject) and were cultured in MyotonicTM Basal Medium (William Cook Europe ApS; MB-2222) supplemented with 10% MyoTonic Growth Supplement (William Cook Europe ApS; MS-3333). To differentiate primary myoblasts in myotubes, cells were seeded with a density of approximately 10,000–15,000 cells cm−2; then, after 2 days, differentiation was induced by switching culture medium to MyoTonicTM Differentiation Medium (William Cook Europe ApS; MD-5555) for 5 days. Human primary cells were used at passages ranging from 2 to 9 from purchase. C26 colorectal adenocarcinoma (C26) and LLC cell lines (kindly gifted by Paola Costelli, University of Turin) were cultured in DMEM 10% FBS. All cell lines were cultured at 37 °C and 5% CO2.
Animals
C57BI/6J and Balb/c mice were purchased from Charles River Laboratories International and housed in a pathogen-free environment. Four to six littermates were housed together and were fed with standard chow diet (Special Diets Services, DS861912G10R) with access to drinking water, while housed under a 12 h light–dark cycle. For TCM production, LLC cells were injected subcutaneously in 2-month-old C57BI/6J male mice, while C26 colon carcinoma cells were injected in 2-month-old Balb/c male mice. Two-month-old Balb/c male were used for all the in vivo experiments. Littermates were randomly assigned to the different experimental groups. A total of 106 C26 murine adenocarcinoma cells in 100 μl of PBS were subcutaneously injected in the right flank of the mice. Mice were weighed daily, and starting from day 4 after tumour cell inoculation, C26-bearing animals were injected intraperitoneally with rolipram (3 mg kg−1) or vehicle (1.86% ethanol in saline). For the AAV9-shRNA experiment, tibialis anterior was injected longitudinally with 30 µl of PBS with 1011 genome copies of AAV9 shRNA scramble or shRNA Pde4d, 2 weeks before cancer cell injection. Total body weight was measured daily after cancer cell inoculation. Tumour growth was monitored by palpation and by checking for the possible presence of ulcers. The tumour typically becomes detectable as a small, barely palpable mass around 8–9 days after injection. Our institutional ethical regulations did not define a maximal tumour size or burden but specified humane endpoint criteria: impaired mobility or inability to feed, signs of tumour ulceration and total body weight loss >20%. Mice were euthanized at the planned experimental endpoint or earlier if they reached one of the above-mentioned humane endpoint criteria. Tissues were weighted and properly collected for molecular and histological analysis.
Human skeletal muscle biopsies
The study enrolled consecutive patients (age >18 years) with pancreatic cancer surgically treated at General Surgery 1, Padova University Hospital, Padova, Italy, from 2016 to 2024. The samples originate, with some additions, from a previous study52. Patients with cancer were classified as cachectic in cases of >5% weight loss in the 6 months preceding surgery, >2% weight loss with either body mass index (BMI) <20 kg m−2 or low muscle mass defined by the skeletal muscle index cut-offs58. Skeletal muscle index values were quantified using the preoperative computed tomography scans as previously described52. The study also enrolled control healthy donors undergoing elective surgery for non-neoplastic and non-inflammatory diseases. Patients with presence of active inflammatory or infectious diseases, known myopathies or viral infections were excluded. The muscle biopsy was performed within 30 min after the start of the planned surgery by a cold section of a rectus abdominal fragment (1 × 0.5 cm). The fragment was immediately frozen and conserved in liquid nitrogen for gene expression analysis. The patient population’s characteristics reported in Table 1 include the sex of the participants. The cohort studied includes both sexes, and it is balanced between the three subgroups (control, precachectic and cachectic). Sex was determined based on self-reporting. Demographics and clinical data, including medications and comorbidities noted as having potential confounding effects on skeletal muscle homeostasis were collected from all patients (Table 1). Muscle biopsies and/or RNA/cDNA samples are stored at Veneto Institute of Molecular Medicine (Padova, Italy). Contact details for sharing request: roberta.sartori@unipd.it or marco.sandri@unipd.it.
C26 and LLC TCM preparation
C26 colorectal adenocarcinoma (C26) tumour was induced by subcutaneous injection of 106 C26 cells in the right flank of 2-month-old Balb/c male mice. C26 tumour was explanted after 13 days, when the cachectic phenotype (that is, ~15% of body weight loss and ~15% of skeletal muscle weight loss) was evident. LLC tumour was induced by subcutaneous injection of 5 × 106 LLC cells in the right flank of 2-month-old C57BI/6J male mice. LLC tumour was explanted after 24 days, when the cachectic phenotype (that is, ~15% of body weight loss and ~15% of skeletal muscle weight loss) was evident. Both C26 and LLC tumour bulk were minced in ~5-mm3 parts and plated in serum-free DMEM supplemented with penicillin–streptomycin solution (Gibco; 15140-122). Twenty-four hours later, the medium was collected and cells (or debris) were eliminated through two sequential centrifugations at 500g and at 5,000g, then filtered with a 0.22-μm pore size filter. The same procedure was repeated the second day, incubating tumour residues with fresh DMEM. The TCM concentrations for cell treatments (that is, 15% C26 TCM and 3% LLC TCM) were selected based on Il6 mRNA expression (Extended Data Fig. 5b,c). Pro-atrophic effects of the selected concentration for C26 and LLC TCM were assessed by quantifying myotubes diameters after their incubation in TCM for 24 h (Extended Data Fig. 5g,h). For myotube diameter quantification, pictures were taken with phase-contrast microscopy (Zeiss Primovert) at 10× magnification and mean diameter was measured manually through ImageJ software (https://imagej.nih.gov/ij/). Both C26 and LLC TCM were then stored in aliquots at −80 °C.
In vitro treatments
For in vitro treatments, myoblasts or myotubes were cultured with TCM in the appropriate growth medium for 24 h, then serum-deprived 1 h before drug treatment (with or without TCM). Cells were treated with 10 μM ISO (Sigma-Aldrich; I6504) or 10 μM BAY-606583 (Cayman Chemical; 17127) diluted in PBS for the indicated time. When indicated, before ISO treatment, cells were pretreated for 2 min with 10 μM rolipram (Sigma-Aldrich; R6520) or 10 μM zatolmilast (MedChemExpress; HY-117571) or vehicle (DMSO). For STAT3 and NF-κB luciferase reporter assay, myoblasts were cultured for 24 h in DMEM 10% FBS. As positive controls, 100 ng ml−1 IL6 (Prepotech; 216-16) or 10 ng ml−1 TNF (Peprotech; 315-01 A) were used.
Knockout clone generation
Pde4b−/− C2C12 and Pde4d−/− C2C12 cell lines were generated through CRISPR–Cas9 technology. Single guide RNA (sgRNA) sequences were designed to target Pde4b and Pde4d through UCSC CRISPR–Cas9 Sp. Pyog. target tracks (https://genome.ucsc.edu/) based on Doench et al.59. The guide sequences (listed in Supplementary Table 5) were synthesized by Metabion Company (Metabion International AG) and cloned in pSpCas9(BB)-2A-Puro (PX459) V2.0 plasmids. This plasmid was a gift from Feng Zhang (Addgene plasmid #62988; http://n2t.net/addgene:62988; RRID: Addgene_62988)60. Plasmids have been purified with NucleoSnap Plasmid Midi kit for plasmid DNA (Macherey-Nagel; 740410.50) according to the manufacturer’s protocol. 100,000 C2C12 cells were transfected with 2 μg of plasmid with Lipofectamine3000 reagent (Invitrogen; L3000001) used with a 3:2 Lipofectamine:DNA (µl:µg) ratio, following the manufacturer’s instructions. Transfected C2C12 cells were selected with a 48-h treatment with 3.5 µg ml−1 puromycin (Sigma-Aldrich; P8833). Selected C2C12 cells were recovered and clonally expanded in a puromycin-free DMEM 10% FBS medium. The clonal knockout of Pde4b and Pde4d was screened by western blot analysis for protein expression and by Sanger sequencing after PCR amplification using DreamTaq PCR master mix (Thermo Scientific; K1071). sgRNA sequences and primer sequences for screening are listed in Supplementary Table 5.
RT–qPCR analysis
Total RNA, from cells and snap-frozen tissues, was extracted using RNAzol reagent (Sigma-Aldrich; R4533). Tissues were homogenized using TissueRuptor (Qiagen). After quantification at NanoDrop2000, 1,000 ng of RNA was retro-transcribed with High-Capacity cDNA Reverse Transcription kit (Applied Biosystems; 4368813) and real-time qPCR was performed with 7900HT Fast Real-Time PCR system or with QuantStudio 6 Flex Real-Time PCR system (Applied Biosystems) using Luna Universal qPCR Master Mix (New England Biolabs; M3003L) or PowerUp SYBR Green Master Mix for qPCR (Applied Biosystems; A25742). TaqMan-based reverse transcription quantitative PCR (RT–qPCR) was performed using Universal Probe qPCR Master Mix (New England Biolabs; M3004E) with the indicated TaqMan probes. Relative mRNA levels were calculated by the ΔΔCt method and normalized to Gusb mRNA in the case of in vitro samples and to Actb (or ACTB for human samples) in the case of in vivo samples. For mtDNA content, total DNA was extracted from approximately 20 mg of gastrocnemius muscles. The tissues were incubated overnight at 56 °C on a rocking platform in 200 μl of mouse tail lysis buffer (100 mM Tris–HCl (pH 8.0), 5 mM EDTA, 200 mM NaCl, 0.2% (w/v) SDS and Proteinase K; Thermo Scientific, EO0491, 100 μg ml−1) and the following day the debris was removed by centrifugation at 15,000g for 10 min. To precipitate the DNA, 180 μl of the supernatant were transferred into a fresh tube containing 180 μl of 2-propanol and gently mixed; the precipitated DNA was collected, transferred to a fresh tube containing 100 μl of TE buffer (10 mM Tris (pH 8) and 1 mM EDTA) and incubated at 37 °C for 15–30 min. The samples were then incubated at 72 °C for 10 min to inactivate Proteinase K. mtDNA over nuclear DNA (nDNA) qPCR analysis was performed with 7900HT Fast Real-Time PCR system using Luna Universal qPCR Master Mix (New England Biolabs; M3003L) and primers specific for Hk2 nuclear gene (nDNA) and Nd1 mitochondrial gene (mtDNA) based on Quiros et al.61. The sequences of the SYBR primers, together with the codes of the TaqMan probes used for RT–qPCR, are listed in Supplementary Tables 6 and 7.
Western blotting
For western blot analysis, cells were lysed in ice-cold RIPA buffer (20 mM Tris–HCl (pH 7.4), 160 mM NaCl, 50 mM NaF, 1 mM EGTA, 1 mM EDTA, 0.1% SDS, 0.1% Na deoxycholate and 1% Triton X-100) supplemented with a protease inhibitor cocktail (Sigma-Aldrich; P8340) and 1 mM sodium orthovanadate (Sigma-Aldrich; S6508). For western blot analysis of tissues, 30 slices (20 μm each) of snap-frozen muscles were resuspended in the following lysis buffer: 50 mM Tris pH 7.5, 150 mM NaCl, 10 mM MgCl2, 0.5 mM dithiothreitol, 10% glycerol, 1% SDS and 1% Triton X-100 supplemented with protease inhibitor cocktail (Sigma-Aldrich; P8340) and 1 mM sodium orthovanadate (Sigma-Aldrich; S6508), then incubated 10 min at 70 °C. DNA were removed by centrifugation of lysates at 12,000g for 15 min at 4 °C. Protein concentration was determined by Pierce BCA Protein Assay Kit (Thermo Scientific; 23227). Samples were denatured in sample buffer (final concentration: 62.5 mM Tris–HCl (pH 6.8), 2% SDS, 10% glycerol and 0.01% bromophenol blue) supplemented with 50 mM dithiothreitol, then incubated for 5 min at 95 °C. For OXPHOS western blot analysis (Fig. 6b), samples were incubated for 10 min at 70 °C. Equal amounts of total proteins, ranging from 5 to 15 µg, were loaded and then separated by using Bolt Bis-Tris Plus Mini Protein Gels, 4–12% (Invitrogen) or 4–15% Mini-PROTEAN TGX Precast Protein Gels (Bio-Rad) and transferred to Hybond P Western blotting membranes (Amersham; GE10600023) through Trans-Blot Turbo Transfer System (Bio-Rad). Membranes were blocked for 1 h at room temperature with Tris-buffered saline (TBS)–bovine serum albumin (BSA) solution (recipe: 20 mM Tris, 150 mM NaCl, pH to 7.4 with 12 M HCl; supplemented with 3% m/v BSA; Sigma-Aldrich, A4503) and then probed by using the primary antibodies (Supplementary Table 8) diluted according to the manufacturer’s instructions in TBS–3% BSA and incubated overnight at +4 °C. The appropriate horseradish peroxidase-linked secondary antibody (Revvity; NEF812001EA, NEF822001EA) was used at 1:3,000 in TBS–0.25% Tween-20 (Sigma-Aldrich; P1379) and incubated for 1 h at room temperature. Immunoreactive bands were visualized by Western Lightning-ECL & Western Lightning Plus (Revvity; NEL104001EA) and acquired at Chemidoc touch (Bio-Rad). Image Lab Software (Bio-Rad) was used for the densitometry analysis.
FRET analysis
For FRET analysis, 30,000 C2C12 cells were plated in 35-mm optical dishes (Ibidi; 81156), then transfected with the ICUE3 Epac2-cAMP plasmid36. For transfection, ViaFect transfection reagent (Promega; E4982) was used at a 4:1 ratio (transfection reagent:DNA), following the manufacturer’s instructions. After 12 h, C2C12 cells were treated with LLC TCM or C26 TCM for 24 h, then serum-deprived 1 h before drug treatment. When indicated, before ISO treatment, cells were treated or pretreated for 2 min with rolipram (Sigma-Aldrich; R6520) or vehicle (DMSO). FRET signals were measured in live cells using a Leica SP5 confocal microscope, with CFP and YFP fluorescence emissions detected accordingly. Levels of cAMP were calculated with Las-X Software (Leica) as changes in CFP over YFP (FRET) ratio in a single cell area.
cAMP extraction and quantification
Snap-frozen gastrocnemius muscles were used for total cAMP measurements. Muscles were powdered in liquid nitrogen, and cAMP was extracted with cold 6% trichloroacetic acid. Samples were sonicated for 10 s, incubated at 4 °C under gentle agitation for 10 min and then centrifuged at 13,000g at 4 °C for 10 min. Supernatants were washed four times with five volumes of water-saturated diethyl ether and lyophilized. cAMP content was detected with the Cyclic AMP ELISA Kit (Cayman Chemical; item 581001) according to the manufacturer’s protocol. cAMP quantification was normalized over milligram of tissue power used. For cAMP quantification from cell cultures, cells were lysed with 0.1 M HCl, and cAMP content was subsequently measured using the Cyclic AMP ELISA Kit (Cayman Chemical; item 581001) according to the manufacturer’s protocol.
Cell surface β2AR quantification
C2C12 myoblasts were seeded in 96-well plates and differentiated for 4 days in DMEM supplemented with 2% HS. On day 4, cells were treated for 24 h with C26 or LLC tumour-derived TCM. Myotubes were fixed with 3% paraformaldehyde, supplemented with 4% sucrose in PBS 1×, pH 7.4, for 5 min at room temperature, then washed twice with PBS 1×. Blocking was performed with 1% BSA (Sigma-Aldrich; A1470) for 30 min at room temperature. Cells were incubated with 1.75 µg ml−1 of anti-β2AR (extracellular epitope) antibody (Supplementary Table 8) for 2 h at room temperature, followed by three washes with PBS 1×. Myotubes were incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibody (1:1,000; Revvity; NEF812001EA) for 30 min at room temperature followed by extensive washes with PBS 1×. Signal detection was performed by adding 100 µl of TMB substrate (Merck; T0440) per well. Absorbance was measured after 5 min at 600 nm using a plate reader (Promega; GloMax Microplate Reader). Only for permeabilized samples, after fixation, myotubes were permeabilized 10 min with cold HEPES–Triton buffer (20 mM HEPES, 300 mM sucrose, 50 mM NaCl, 3 mM MgCl2 and 0.5% Triton X.100, pH 7.4) before the incubation with primary antibody to measure total β2AR levels. The background signal was determined using control samples incubated with the secondary antibody only, under the same conditions as the experimental samples. This allowed the assessment and subtraction of non-specific binding during signal quantification.
Luciferase assays
For Luciferase assay analysis, 40,000 C2C12 cells were plated in 12-well plates and transfected with a pGL4.29[luc2P/CRE/Hygro] vector (CRE-Luciferase reporter) (Promega; E847A) or STAT3 reporter vector (pGL4.47[luc2P/SIE/Hygro]; Promega; E4041) or NF-κB (pGL4.32[luc2P/NF-kB-RE/Hygro]; Promega; E8491) and pRL-null Vector (Renilla Luciferase Control Reporter Vector) (Promega; E2231), with a 10:1 ratio (luc/renilla). For transfection, ViaFect transfection reagent (Promega; E4982) was used at a 4:1 ratio (transfection reagent:DNA). Twenty-four hours after transfection, C2C12 cells were treated with TCM, then serum-deprived 1 h before ISO treatment (10 μM). After 5 h of drug treatment, C2C12 cells were lysed using the Passive Lysis Buffer (Promega; E1941) following the manufacturer’s instructions. We used the Dual-Glo Luciferase Assay System Kit (Promega; E2980) to quantify the luminescence signal at the GloMax. Renilla luciferase was used as an internal control to normalize results.
Kinase activity profiling
PamChip peptide arrays (PamGene International BV)62 was used to measure kinase activity in muscle lysates from C26-bearing mice or controls. Around 15 mg of gastrocnemius muscle was lysed with TissueRuptor (Quiagen) in 600 µl of M-PER Mammalian Extraction Buffer (Thermo Fisher Scientific; 78501) supplemented with Halt Phosphatase Inhibitor Cocktail (Thermo Fisher Scientific; 78420); Halt Protease Inhibitor Cocktail, EDTA free (Thermo Fisher Scientific; 87785). Lysates were incubated for 30 min on ice, then centrifuged 15 min at 4 °C. Supernatant was collected, and protein concentration was determined through the Pierce BCA Protein Assay Kit (Thermo Scientific; 23227). Kinase activity profiling was conducted using the PamStation-12 platform 10.0 (PamGene International BV). Serine/threonine kinase arrays were performed strictly following the manufacturer’s instruction. Fluorescent signal intensities were recorded and analysed using BioNavigator software. Upstream kinase activity analysis was performed via the PamApp module, incorporating curated kinase–substrate databases (for example, PhosphoSitePlus, Reactome and phosphoNET) and computational prediction models, generating kinase-specific activity, significance and specificity scores. Three parameters were computed: (1) mean kinase statistic, representing the direction and magnitude of activity change between test and control samples, calculated as the median signal-to-noise ratio of its known substrate peptides; (2) mean significance score, reflecting the likelihood that the observed kinase activity change is not due to sample label permutation; (3) mean specificity score, indicating the probability that the observed effect is specific to the kinase’s substrate peptides and not due to random peptide selection. A combined kinase score (median final score), obtained by summing significance and specificity scores, was used to rank kinases. Kinases with negative median kinase statistic (that is, inhibited in C26 versus control) are shown (Fig. 1c and Supplementary Table 2).
Grasping test
To assess skeletal muscle force, mice were subjected to the grasping test on day 0, day 6 and day 13 using a grip strength meter (Ugo Basile, cat. no. 47200). Each test was performed by the same operator and consisted of five measurements of the forelimb strength. The analysis represents the maximal peak force recorded at each test, normalized over mouse weight.
Glucose tolerance test
For glucose tolerance evaluation, mice were fasted for 6 h and injected intraperitoneally with D-glucose (BDH Laboratory Supplies; 101174Y) dissolved in saline water (1.5 g kg−1 body weight). Blood glucose concentration was measured via tail vein blood in the fasted state (time 0) and at 15, 30, 60 and 120 min after glucose administration using the ONETOUCH Verio Reflect blood glucometer (Lifescan).
Treadmill test
Muscle performance was evaluated using a treadmill-based exhaustion protocol (Panlab Harvard Apparatus; LE8710M), with a maximum test duration of 60 min. Before the exercise performance test, mice were acclimated to the treadmill with a 10-min run at a constant speed of 14 cm s−1 once per day for two alternate days. The day of the experiment, each mouse was placed into an exercise test regimen of 14 cm s−1 for 10 min. The speed was then increased by 2 cm s−1 every 2 min, up to a maximum of 46 cm s−1, which was maintained until exhaustion or after 60 min of test. Exhaustion was defined as the inability to continue running despite repeated electrical stimuli. At that moment, total running time and distance were recorded.
AAV9 shRNA constructs
Plasmids encoding short hairpin RNAs (shRNAs) targeting Pde4d or a non-targeting control sequence were generated using VectorBuilder (VectorBuilder). The shRNA targeting Pde4d (Vector ID: VB240301-1258hts) was designed to target the sequence 5′-GCAAAGACAATCTTTAAGGAA-3′, while the control shRNA (Vector ID: VB240301-1259hqf) targeted 5′-GTTCAGATGTGCGGCGAGT-3′. Each plasmid contains a single shRNA expression cassette under appropriate regulatory control together with an EGFP-encoding sequence.
The plasmids were subsequently packaged into adeno-associated virus particles of serotype 9 (AAV9) by VectorBuilder using their medium-scale AAV packaging service with ultrapurification. Vector details, including sequence maps and full construct information, are available from the VectorBuilder website (www.vectorbuilder.com) using the respective vector IDs.
Body composition analysis
Body composition analysis was performed at day 0 and at the endpoint after cancer cell inoculation (20% of body weight loss) by quantitative magnetic resonance using an EchoMRI-100 (EchoMRI LLC) without the use of anaesthesia.
Noradrenaline measurement
For noradrenaline measurement, 40 mg of gastrocnemius muscle was homogenized using a TissueRuptor (Qiagen) in a buffer containing 0.01 M HCl, 1 mM EDTA and 4 mM K2S2O5 in water. The homogenate was then sonicated twice with 30% amplitude for 10 min each. To pellet the non-lysed tissue, the samples were centrifuged 10,000g for 5 min, and only the supernatant was kept for the noradrenaline measurement. For the quantification, noradrenaline high-sensitive ELISA kit (LDN; BA E-5200R) was used following the manufacturer’s instructions. As a technical control, sympathectomized gastrocnemius from 2-month-old Balb/c male mice was used. Chemical sympathectomy was induced by injecting the femoral biceps with 6-hydroxydopamine (Sigma-Aldrich; H4381) at a dose of 100 µg g−1 mouse with a concentration of 100 µg µl−1 or with vehicle (0.3% m/v ascorbic acid in H2O gassed for 20 min with N2) twice a week, for 13 days.
Muscle preparation for high-resolution respirometry
For high-resolution respirometry, excised gastrocnemius muscles were cryopreserved following the protocol described by Kuznetsov AV et al.63. Dissected muscles were immersed in 200 µl of BIOPS solution (50 mM K⁺-MES, 20 mM taurine, 0.5 mM dithiothreitol, 6.56 mM MgCl2, 5.77 mM ATP, 15 mM phosphocreatine, 20 mM imidazole (pH 7.1, adjusted with 5 N KOH), 10 mM Ca-EGTA buffer (2.77 mM CaK2EGTA + 7.23 mM K2EGTA; 0.1 µM free calcium)) containing 30% DMSO and 10 mg ml−1 fatty-acid-free BSA, and equilibrated with cryopreservation solution for 5 s. Gastrocnemius was then immediately frozen in liquid nitrogen and kept at −80 °C. Before OCR analysis, tubes were placed in a water bath at 37 °C, and when the cryopreservation medium was completely thawed, muscles were immediately transferred and washed in the respiration medium MiR05 (0.5 mM EGTA, 3 mM MgCl2, 60 mM lactobionic acid, 20 mM taurine, 10 mM KH2PO4, 20 mM HEPES, 110 mM sucrose and 1 g l−1 BSA, pH 7.1) containing 2 mg ml−1 BSA to minimize time of contact with DMSO. OCR analysis in skeletal muscle tissue requires plasma membrane permeabilization to ensure the washout of free cytosolic molecules including adenylates, substrates and cytosolic enzymes but also to make externally added compounds accessible to mitochondria64. For the permeabilization of plasma membranes, thawed gastrocnemius was placed in a Petri dish containing ice-cold BIOPS and mechanically minced with sharp forceps into fibre bundles (~2 mg each) under a dissecting microscope. Fibre bundles were then individually positioned into a 12-well plate filled with 2 ml BIOPS containing saponin (50 µg ml−1) and incubated with gentle shaking on ice for 20 min. Fibre bundles were then washed in 2 ml ice-cold BIOPS with gentle shaking for 10 min. Finally, samples were carefully dried on filter paper and weighed before being placed into the respirometer chambers containing mitochondrial respiration medium MiR05.
Histology and immunofluorescence
Muscles collected for histological analysis were mounted in Killik medium (BioOptica), frozen in liquid-nitrogen-cooled isopentane and stored at −80 °C. To determine myofibre CSA and fibre type composition, transverse sections (7 μm) were cut from the central part of gastrocnemius muscle with a cryostat at −20 °C. Fresh slices were then incubated for 1 h at room temperature in MOM blocking reagent (MKB-2213-1; Vector Laboratories), washed in PBS and incubated overnight at 4 °C with the following primary antibodies: mouse anti-MyHC Type I (Myh7) clone BA-D5, mouse anti-MyHC Type IIa (Myh2) clone SC-71, and mouse anti-MyHC Type IIb (Myh4) clone BF-F3, all at 1:100 dilution (Developmental Studies Hybridoma Bank), and rabbit anti-laminin (Z0097; DAKO) at 1:200 dilution. Tissue slices were then incubated for 1 h at room temperature with the appropriate antibodies herein listed: anti-mouse IgG2b 633 (ThermoFisher, A-21146), anti-mouse IgG1 488 (Invitrogen; A-21121), anti-mouse IgM 546 (Invitrogen; A-21045) and anti-rabbit H + L 405 (Invitrogen; A31556). Images were acquired using a Leica SP8 confocal microscope. CSA quantification and fibre type composition were automatically evaluated with ImageJ software (https://imagej.net/ij/download.html) using MorphoLibJ65. Triple-negative fibres were defined as type IIx.
The whole EDL muscle was used to visualize and quantify postsynaptic NMJ structures. EDL muscles were fixed in 4% paraformaldehyde (pH 7.4) immediately after collection and conserved in PBS at +4 °C. For staining, muscles were incubated in PBS containing 1:500 fluorescent α-bungarotoxin (Invitrogen; B13422) for 90 min at room temperature to label postsynaptic NMJs. After staining, 1-μm-thick Z-stacks were acquired using a Leica confocal SP5 microscope to reconstruct the entire postsynaptic NMJ. Volume quantification was performed using ImageJ software. The EGFP endogenous signal of the whole tibialis anterior was acquired using a Leica SP8 confocal microscope.
ChIP and library preparation
ChIP was performed on quadriceps from control and tumour-bearing mice, as previously described66,67. In brief, frozen tissues were pulverized using Covaris tissue TUBEs (Covaris; 520001) with a chilled hammer on a metallic block placed in dry ice. The resulting powdered tissue was transferred to a glass vial and resuspended in 10 ml 1× PBS, followed by cross-linking in 1% formaldehyde (Ted Pella) for 15 min at room temperature. After blocking the cross-link with 0.125 M glycine for 5 min and washing with PBS containing protease and phosphatase inhibitors, the pellets were resuspended in Farnham buffer (5 mM PIPES pH 8.0, 85 mM KCl and 0.5% NP-40) and briefly homogenized with a homogenizer. Subsequently, cells were lysed in RIPA buffer (1× PBS, 1% NP-40, 0.5% sodium deoxycholate and 0.1% SDS). Chromatin was sonicated to achieve fragment lengths of approximately 0.5 kb and then subjected to immunoprecipitation using 5 μg of phospho-CREB1S133 antibody and Protein G Dynabeads (Invitrogen; 10003D). Washes were performed as previously described67. The eluted DNA was reverse-crosslinked, digested with 20 μg of Proteinase K for 1 h at 55 °C and purified using the Qiagen MinElute PCR purification kit. The ChIP-eluted samples were used to prepare the library following the manufacturer’s instructions of NEBNext ChIP-Seq Library Prep Reagent Set for Illumina (New England Biolabs; E6240L). Libraries were sequenced on Illumina NextSeq 1000 System (paired-end 61 + 61 bp reads).
ChIP-seq data analysis
Following quality control with FastQC v0.11.2, paired-end sequencing reads were aligned to the mouse reference genome (mm10/GRCm38) using Bowtie v2.3.5.1 (ref. 68). Duplicated alignments (identified by Picard MarkDuplicates, https://broadinstitute.github.io/picard) and low-quality alignments/multi-mapping reads were excluded using SAMtools v1.6. Immunoprecipitation and corresponding control (input DNA) datasets were treated identically. Peak calling was performed for each immunoprecipitation against its matched Input DNA using MACS v2.1.4 (ref. 69) (command: callpeak -g mm --nomodel -f BAMPE -q 0.05). Input-normalized ChIP-seq fold-enrichment signal tracks were obtained using MACS v2.1.4 (ref. 69) (command: callpeak -g mm --nomodel -f BAMPE -q 0.05 --SPMR -B and bdgcmp -m FE). Differential binding analysis was performed using the DiffBind v3.10.0R/Bioconductor package70, using edgeR as a backend. In brief, starting from a merged set of peaks obtained from overlapped peaks between biological replicates in each condition, and the processed alignments for both immunoprecipitation and control samples, library size normalization was used for quantification of signal in peaks, and the generalized linear model framework was used for batch integration (formula: ~Condition+Batch). The following parameters were used for normalization and statistical analysis: normalize = DBA_NORM_LIB, library = DBA_LIBSIZE_DEFAULT, background = BKGR_FALSE, AnalysisMethod = DBA_EDGER. Peaks with P value ≤0.05 and absolute value of log2 fold change (log2FC) ≥0 were considered as significantly differentially bound in C26 samples as compared with control. Signal heatmaps in peak regions were generated using the deepTools71 computeMatrix and plotHeatmap utilities. Annotation of binding sites to genes and genomic regions (ENCODE ccREs33) was performed using rGREAT72 and BEDtools v2.30.0 (ref. 73). Motif enrichment was performed using the HOMER74 utility ‘findMotifsGenome.pl’ (parameters: -size given), considering a fixed window (±50 bp) centred on peak summits (retrieved using the BEDtools v2.30.0 bedtools slop utility).
RNA-seq analysis
RNA was isolated from snap-frozen gastrocnemius muscle using RNAzol agent (Sigma-Aldrich; R4533). The RNA-seq was performed by the Center for Omics Sciences (IRCCS Ospedale San Raffaele, Milan, Italy). RNA quality was assessed using a BioAnalyzer 2100. Base-calling was performed on NovaSeq 6000 Illumina machine, obtaining 30 million single end reads per sample on average. Reads were aligned to the reference mouse genome mm10, using STAR aligner (version 2.5.3a). Uniquely mapped reads were assigned to the corresponding genes using featureCounts (version1.6.4) and annotated according to Gencode basic annotations (Gencode version M22). Differential gene expression analysis was performed using DESeq2 R package with a false discovery rate correction of 0.05 for multiple test hypotheses.
Quantification and analysis of isoforms from RNA-seq data were performed using the RSEM computational pipeline75, with the STAR program76 used to align reads to the mm10 mouse reference genome. The Ensembl database v100 (ref. 77) was used as a source for isoform annotations.
Enrichment analysis and graphs
Enrichment analyses were performed using IPA (Qiagen) or, when indicated, through the online software EnrichR R package (https://maayanlab.cloud/Enrichr). Enrichment bubble plots and cluster heatmap plots were generated with SRplot78.
Statistical analysis
All statistical analyses were performed using GraphPad Prism version 10.4.1. For comparisons between two groups, unpaired two-tailed t-tests were used when data were normally distributed, as assessed by the Shapiro–Wilk test (P > 0.05). When at least one group did not meet the assumption of normality (P < 0.05), the non-parametric Mann–Whitney test was applied. For comparison between three or more groups, data distribution was assumed to be normal, but this was not formally tested. One-way analysis of variance (ANOVA) was used to compare more than two groups for a single variable, and two-way ANOVA was used when two independent variables were involved. Post-hoc comparisons were performed using Fisher’s least significant difference test following ANOVA analyses. P values <0.05 were considered statistically significant. Data are presented as mean ± standard error of the mean (s.e.m.). No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications52,79,80. For the in vivo experiments, mice were randomly assigned into the experimental group and data collection was performed in a blind manner. Data collection and analysis for in vitro assays were not performed blind to the conditions of the experiments, except for myotube diameter quantification. Data were excluded a priori based on the degree of cachexia (that is, less than 10% of weight loss at 13 days after C26 injection) or technical issues (insufficient Pde4d silencing or poor quality of the sample) as specified in the ‘Reporting summary’.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq and ChIP-seq data have been deposited in the NCBI Gene Expression Omnibus (GEO) portal and are available under the SuperSeries accession number GSE271831, with the following subseries accession numbers: GSE271492 (RNA-seq data) and GSE271830 (ChIP-seq data). The data were analysed using standard pipelines as previously described. Source data are provided with this paper.
References
Wyart, E., Carrà, G., Angelino, E., Penna, F. & Porporato, P. E. Systemic metabolic crosstalk as driver of cancer cachexia. Trends Endocrinol. Metab. https://doi.org/10.1016/j.tem.2024.12.005 (2025).
Berriel Diaz, M., Rohm, M. & Herzig, S. Cancer cachexia: multilevel metabolic dysfunction. Nat. Metab. 6, 2222–2245 (2024).
Baracos, V. E., Martin, L., Korc, M., Guttridge, D. C. & Fearon, K. C. H. Cancer-associated cachexia. Nat. Rev. Dis. Primers 4, 17105 (2018).
Fearon, K. C. H., Glass, D. J. & Guttridge, D. C. Cancer cachexia: mediators, signaling, and metabolic pathways. Cell Metab. 16, 153–166 (2012).
Ferrer, M. et al. Cachexia: a systemic consequence of progressive, unresolved disease. Cell 186, 1824–1845 (2023).
Marceca, G. P., Londhe, P. & Calore, F. Management of cancer cachexia: attempting to develop new pharmacological agents for new effective therapeutic options. Front. Oncol. 10, 298 (2020).
Lee, S.-J., Bhasin, S., Klickstein, L., Krishnan, V. & Rooks, D. Challenges and future prospects of targeting myostatin/activin a signaling to treat diseases of muscle loss and metabolic dysfunction. J. Gerontol. A 78, 32–37 (2023).
Delfinis, L. J. et al. Muscle weakness precedes atrophy during cancer cachexia and is linked to muscle-specific mitochondrial stress. JCI Insight 7, e155147 (2022).
Brown, J. L. et al. Mitochondrial degeneration precedes the development of muscle atrophy in progression of cancer cachexia in tumour-bearing mice. J. Cachexia Sarcopenia Muscle 8, 926–938 (2017).
Hardee, J. P., Montalvo, R. N. & Carson, J. A. Linking cancer cachexia-induced anabolic resistance to skeletal muscle oxidative metabolism. Oxid. Med. Cell. Longev. 2017, 8018197 (2017).
Huot, J. R. et al. Formation of colorectal liver metastases induces musculoskeletal and metabolic abnormalities consistent with exacerbated cachexia. JCI Insight 5, e136687 (2020).
Halle, J. L. et al. Tissue-specific dysregulation of mitochondrial respiratory capacity and coupling control in colon-26 tumor-induced cachexia. Am. J. Physiol. Regul. Integr. Comp. Physiol. 317, R68–R82 (2019).
van der Ende, M. et al. Mitochondrial dynamics in cancer-induced cachexia. Biochim. Biophys. Acta Rev. Cancer 1870, 137–150 (2018).
Gicquel, T. et al. Integrative study of skeletal muscle mitochondrial dysfunction in a murine pancreatic cancer-induced cachexia model. eLife 13, RP93312 (2024).
Akabane, S. et al. PKA regulates PINK1 stability and Parkin recruitment to damaged mitochondria through phosphorylation of MIC60. Mol. Cell 62, 371–384 (2016).
Chang, C.-R. & Blackstone, C. Cyclic AMP-dependent protein kinase phosphorylation of Drp1 regulates its GTPase activity and mitochondrial morphology. J. Biol. Chem. 282, 21583–21587 (2007).
García-Bermúdez, J. et al. PKA phosphorylates the ATPase Inhibitory Factor 1 and inactivates its capacity to bind and inhibit the mitochondrial H+-ATP synthase. Cell Rep. 12, 2143–2155 (2015).
Berdeaux, R. & Stewart, R. cAMP signaling in skeletal muscle adaptation: hypertrophy, metabolism, and regeneration. Am. J. Physiol. Endocrinol. Metab. 303, E1–E17 (2012).
Wu, Z. et al. Transducer of regulated CREB-binding proteins (TORCs) induce PGC-1alpha transcription and mitochondrial biogenesis in muscle cells. Proc. Natl Acad. Sci. USA 103, 14379–14384 (2006).
Hock, M. B. & Kralli, A. Transcriptional control of mitochondrial biogenesis and function. Annu. Rev. Physiol. 71, 177–203 (2009).
Xing, F. et al. The anti-Warburg effect elicited by the cAMP–PGC1α pathway drives differentiation of glioblastoma cells into astrocytes. Cell Rep. 23, 2832–2833 (2018).
Khan, M. M. et al. Sympathetic innervation controls homeostasis of neuromuscular junctions in health and disease. Proc. Natl Acad. Sci. USA 113, 746–750 (2016).
Zuo, L., Li, Q., Sun, B., Xu, Z. & Ge, Z. Cilostazol promotes mitochondrial biogenesis in human umbilical vein endothelial cells through activating the expression of PGC-1. Biochem. Biophys. Res. Commun. 433, 52–57 (2013).
Zhao, Z. & Pu, Y. Lixisenatide enhances mitochondrial biogenesis and function through regulating the CREB/PGC-1α pathway. Biochem. Biophys. Res. Commun. 508, 1120–1125 (2019).
Park, S.-J. et al. Resveratrol ameliorates aging-related metabolic phenotypes by inhibiting cAMP phosphodiesterases. Cell 148, 421–433 (2012).
Paes, D. et al. The molecular biology of phosphodiesterase 4 enzymes as pharmacological targets: an interplay of isoforms, conformational states, and inhibitors. Pharmacol. Rev. 73, 1016–1049 (2021).
Balasubramaniam, A., Sheriff, S., Friend, L. A. & James, J. H. Phosphodiesterase 4B knockout prevents skeletal muscle atrophy in rats with burn injury. Am. J. Physiol. Regul. Integr. Comp. Physiol. 315, R429–R433 (2018).
Lira, E. C. et al. Phosphodiesterase-4 inhibition reduces proteolysis and atrogenes expression in rat skeletal muscles. Muscle Nerve 44, 371–381 (2011).
Cho, Y. et al. Perm1 enhances mitochondrial biogenesis, oxidative capacity, and fatigue resistance in adult skeletal muscle. FASEB J. 30, 674–687 (2016).
Cho, Y., Hazen, B. C., Russell, A. P. & Kralli, A. Peroxisome proliferator-activated receptor γ coactivator 1 (PGC-1)- and estrogen-related receptor (ERR)-induced regulator in muscle 1 (Perm1) is a tissue-specific regulator of oxidative capacity in skeletal muscle cells. J. Biol. Chem. 288, 25207–25218 (2013).
Goncalves, M. D. et al. Fenofibrate prevents skeletal muscle loss in mice with lung cancer. Proc. Natl Acad. Sci. USA 115, E743–E752 (2018).
Chikuda, H. et al. Cyclic GMP-dependent protein kinase II is a molecular switch from proliferation to hypertrophic differentiation of chondrocytes. Genes Dev. 18, 2418–2429 (2004).
ENCODE Project Consortium et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Rozanska, A. et al. The human RNA-binding protein RBFA promotes the maturation of the mitochondrial ribosome. Biochem. J. 474, 2145–2158 (2017).
Narasimhan, A. et al. Profiling of adipose and skeletal muscle in human pancreatic cancer cachexia reveals distinct gene profiles with convergent pathways. Cancers 13, 1975 (2021).
Ghigo, A. et al. A PI3Kγ mimetic peptide triggers CFTR gating, bronchodilation, and reduced inflammation in obstructive airway diseases. Sci. Transl. Med. 14, eabl6328 (2022).
Gonzalez, G. A. & Montminy, M. R. Cyclic AMP stimulates somatostatin gene transcription by phosphorylation of CREB at serine 133. Cell 59, 675–680 (1989).
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011).
Nomiyama, T. et al. The NR4A orphan nuclear receptor NOR1 is induced by platelet-derived growth factor and mediates vascular smooth muscle cell proliferation. J. Biol. Chem. 281, 33467–33476 (2006).
Gnad, T. et al. Adenosine/A2B receptor signaling ameliorates the effects of aging and counteracts obesity. Cell Metab. 32, 56–70.e7 (2020).
Bloom, T. J. Cyclic nucleotide phosphodiesterase isozymes expressed in mouse skeletal muscle. Can. J. Physiol. Pharmacol. 80, 1132–1135 (2002).
Sartori, R. et al. BMP signaling controls muscle mass. Nat. Genet. 45, 1309–1318 (2013).
Bodine, S. C. & Baehr, L. M. Skeletal muscle atrophy and the E3 ubiquitin ligases MuRF1 and MAFbx/atrogin-1. Am. J. Physiol. Endocrinol. Metab. 307, E469–E484 (2014).
Irelan, D. et al. Acute PDE4 inhibition induces a transient increase in blood glucose in mice. Int. J. Mol. Sci. 24, 3260 (2023).
Marchiano, F., Haering, M. & Habermann, B. H. The mitoXplorer 2.0 update: integrating and interpreting mitochondrial expression dynamics within a cellular context. Nucleic Acids Res. 50, W490–W499 (2022).
Romanello, V. & Sandri, M. The connection between the dynamic remodeling of the mitochondrial network and the regulation of muscle mass. Cell. Mol. Life Sci. 78, 1305–1328 (2021).
Lemieux, H., Blier, P. U. & Gnaiger, E. Remodeling pathway control of mitochondrial respiratory capacity by temperature in mouse heart: electron flow through the Q-junction in permeabilized fibers. Sci. Rep. 7, 2840 (2017).
Holmström, M. H., Iglesias-Gutierrez, E., Zierath, J. R. & Garcia-Roves, P. M. Tissue-specific control of mitochondrial respiration in obesity-related insulin resistance and diabetes. Am. J. Physiol. Endocrinol. Metab. 302, E731–E739 (2012).
Anagnostou, M.-E. & Hepple, R. T. Mitochondrial mechanisms of neuromuscular junction degeneration with aging. Cells 9, 197 (2020).
Dupuis, L. et al. Muscle mitochondrial uncoupling dismantles neuromuscular junction and triggers distal degeneration of motor neurons. PLoS ONE 4, e5390 (2009).
Arnold, A.-S. et al. Morphological and functional remodelling of the neuromuscular junction by skeletal muscle PGC-1. Nat. Commun. 5, 3569 (2014).
Sartori, R. et al. Perturbed BMP signaling and denervation promote muscle wasting in cancer cachexia. Sci. Transl. Med. 13, eaay9592 (2021).
Blackman, B. E. et al. PDE4D and PDE4B function in distinct subcellular compartments in mouse embryonic fibroblasts. J. Biol. Chem. 286, 12590–12601 (2011).
Rodrigues, A. C. Z. et al. The sympathetic nervous system regulates skeletal muscle motor innervation and acetylcholine receptor stability. Acta Physiol. 225, e13195 (2019).
Lusardi, M., Rapetti, F., Spallarossa, A. & Brullo, C. PDE4D: a multipurpose pharmacological target. Int. J. Mol. Sci. 25, 8052 (2024).
So, W.-K. et al. Exchange protein directly activated by cAMP (Epac) 1 plays an essential role in stress-induced exercise capacity by regulating PGC-1α and fatty acid metabolism in skeletal muscle. Pflugers Arch. 472, 195–216 (2020).
Schiaffino, S., Dyar, K. A., Ciciliot, S., Blaauw, B. & Sandri, M. Mechanisms regulating skeletal muscle growth and atrophy. FEBS J. 280, 4294–4314 (2013).
Caan, B. J. et al. Explaining the obesity paradox: the association between body composition and colorectal cancer survival (C-SCANS Study). Cancer Epidemiol. Biomarkers Prev. 26, 1008–1015 (2017).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Quiros, P. M., Goyal, A., Jha, P. & Auwerx, J. Analysis of mtDNA/nDNA ratio in mice. Curr. Protoc. Mouse Biol. 7, 47–54 (2017).
Chirumamilla, C. S. et al. Profiling activity of cellular kinases in migrating T-cells. Methods Mol. Biol. 1930, 99–113 (2019).
Kuznetsov, A. V. et al. Cryopreservation of mitochondria and mitochondrial function in cardiac and skeletal muscle fibers. Anal. Biochem. 319, 296–303 (2003).
Doerrier, C. et al. High-resolution FluoRespirometry and OXPHOS protocols for human cells, permeabilized fibers from small biopsies of muscle, and isolated mitochondria. Methods Mol. Biol. 1782, 31–70 (2018).
Legland, D., Arganda-Carreras, I. & Andrey, P. MorphoLibJ: integrated library and plugins for mathematical morphology with ImageJ. Bioinformatics 32, 3532–3534 (2016).
Savic, D., Gertz, J., Jain, P., Cooper, G. M. & Myers, R. M. Mapping genome-wide transcription factor binding sites in frozen tissues. Epigenetics Chromatin 6, 30 (2013).
Segatto, M. et al. Epigenetic targeting of bromodomain protein BRD4 counteracts cancer cachexia and prolongs survival. Nat. Commun. 8, 1707 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Gu, Z. & Hübschmann, D. rGREAT: an R/bioconductor package for functional enrichment on genomic regions. Bioinformatics 39, btac745 (2023).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Harrison, P. W. et al. Ensembl 2024. Nucleic Acids Res. 52, D891–D899 (2024).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE 18, e0294236 (2023).
Okun, J. G. et al. Liver alanine catabolism promotes skeletal muscle atrophy and hyperglycaemia in type 2 diabetes. Nat. Metab. 3, 394–409 (2021).
Liu, X. et al. Activation of GPR81 by lactate drives tumour-induced cachexia. Nat. Metab. 6, 708–723 (2024).
Martin, L. et al. Diagnostic criteria for the classification of cancer-associated weight loss. J. Clin. Oncol. 33, 90–99 (2015).
Acknowledgements
We acknowledge the FIRC-AIRC 3-year fellowship to E.A. (22395); Project Age-It ‘Ageing Well in an Ageing Society’, NextGeneration EU, in the context of the National Recovery and Resilience Plan (DM 1557 11.10.2022), PRIN-2022 (9CJ27R), and AIRC (IG 17804 and 25702) to A. Graziani; AIRC (IG 23257 and 30366) to M.S.; AIRC (IG 29203) and PRIN-2022 (CLTAYH) to V.P.; AIRC (IG 21353) and Cariplo 2017-0604 to G.C.; AIRC (MFAG-25908) to A. Menga; PRIN-2022 (WFXCWM-A) to P.E.P.; PRIN-2022 (9CJ27R) and PRIN-2022-PNRR (P2022AFS8P) to D.C., who has received funding from NextGeneration EU – MUR –M4C2 1.1, CUP C53D23007570001; AGING Project (MUR) and Departments of Excellence 2023-2027 (Department of Translational Medicine, University of Piemonte Orientale) to N. Filigheddu. and D.C.; and NIH grants AR041164-18 and AR041126-26 to V.S. We acknowledge the Metabolism and Bioinformatics facilities of the University of Piemonte Orientale Interdepartmental Center Ipazia, Novara, Italy. Moreover, we thank A. Albano for ChIP-seq library preparation.
Author information
Authors and Affiliations
Contributions
E.A., L.B. and A. Graziani conceived the work and supervised the experiments. E.A. and L.B. designed the experiments and analysed the results. E.A., L.B., R.S., V.M., B.D’A., N. Formaggio, S.B., T.R., S.R., A. Murabito, M.N., F. Ferrero, C.P., G.R., D.T. and G.C. performed the experiments. A.L., F. Favero and D.C. performed bioinformatic analyses. R.S. and M.V. collected human patients’ biopsy samples. E.A., L.B. and A. Graziani wrote the manuscript. N. Filigheddu, A. Menga, E.H., V.P., A. Ghigo, M.S. and P.E.P. contributed to data interpretation. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A. Ghigo and E.H. are co-founders and shareholders of Kither Biotech, a pharmaceutical company focused on the development of PI3K inhibitors for airway diseases that are not in conflict with the content of this manuscript. The other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Maria Rohm and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of C26-induced cachexia at early and late stages.
a, Tumour weight at 10 days post C26 inoculation. n = 5. b, Total weight loss of control (ctr) vs. 10 days post C26 injection (C26 10d), percentage over starting weight. n = 7 ctr; n = 5 C2610 d. Two-tailed t-test. c, Epididymal fat weight in control (ctr) vs. 10 days post C26 injection (C26 10d). n = 7 ctr; n = 5 C26 10d. Two-tailed t-test. d, Spleen weight at 10 days post C26 injection. n = 7 ctr; n = 5 C26 10d. Two-tailed t-test. e, Muscle weight normalized over tibial length of tibialis anterior (TA), extensor digitorum longus (EDL), quadriceps (Quad), soleus (Sol), and gastrocnemius (Gas) muscles at 10 days post C26 injection (C26 10d) vs. control. n = 7 ctr; n = 5 C26 10d. Two-tailed t-test. f, mRNA expression of Fbxo32 (Atrogin1) and Trim63 (MuRF1) in gastrocnemius muscle of control mice (ctr) and C26-bearing mice 10 days after cancer cell injection (C26 10d). n = 5 ctr, n = 6 C26 10d. Two-tailed t-test for Fbxo32, Two-tailed Mann–Whitney test for Trim63. g, mRNA expression of Ppargc1a (PGC-1α), Ppargc1b (PGC-1β), Esrra (ERRα), Esrrb (ERRβ), Tfam, Perm1, and Mef2c in gastrocnemius muscle of control mice (ctr) and C26-bearing mice 10 days after tumour injection (C26 10d). n = 5 ctr, n = 6 C26 10d. Two-tailed t-test for Ppargc1a, Esrra, Tfam, Perm1, Mef2c; Two-tailed Mann–Whitney test for Ppargc1b, Esrrb. h, Tumour weight at 13 days post C26 injection. n = 8. i, Total weight loss of control (ctr) vs. 13 days post C26 injection (C26 13d), percentage over starting weight. n = 5 ctr; n = 8 C26 13d. Two-tailed t-test. j, Epididymal fat weight in control (ctr) vs. 13 days post C26 injection (C26 13d). n = 5 ctr; n = 8 C26 13d. Two-tailed Mann–Whitney test. k, Spleen weight in control (ctr) vs. 13 days post C26 injection (C26 13d). n = 5 ctr; n = 8 C26 13d. Two-tailed t-test. l, Muscle weight normalized over tibial length of tibialis anterior (TA), extensor digitorum longus (EDL), quadriceps (Quad), soleus (Sol), and gastrocnemius (Gas) muscles from C26 13 days (C26 13d) vs. control. n = 5 ctr; n = 8 C26 13d. Two-tailed t-test. m, RPKM normalized over control of Pde4a, Pde4b, Pde4c, Pde4d in gastrocnemius muscle of control mice (ctr) and C26-bearing mice 13 days after tumour cell injection (C26 13d). n = 7 for each experimental group. Numbers indicate adjusted P values from DESeq2 analysis. For a-m, data are presented as mean values ± SEM and significant P values are annotated in the graphs.
Extended Data Fig. 2 Enrichment analysis of the muscle transcriptome from KrasG12D/+; Lkb1f/f lung cancer and C26 colon carcinoma-induced cachexia mouse models.
a,b, Top 10 terms from enrichment analysis for GO Biological Process 2025 (EnrichR) (a) and for transcription factor loss-of-function (TF-LoF) (b) of genes downregulated in skeletal muscle of KrasG12D/+; Lkb1f/f (KL) mice and not in fasted mice. Data from published gene set from Goncalves et al.31. c, Top 10 terms from enrichment analysis for transcription factor loss-of-function (TF-LoF) (EnrichR) of genes downregulated in skeletal muscle of C26-bearing mice at 13 days from tumour injection (C26 13d) vs. control mice. d, Top 10 terms from enrichment analysis for GO Biological Process 2025 (EnrichR) of genes concomitantly downregulated in C26 13d vs. control and in Creb1 TF-LoF term.
Extended Data Fig. 3 Analysis of p-CREB1S133 binding and ChIP-seq integration with transcriptomic alterations in C26 cachexia.
a, Fraction of p-CREB1S133 ChIP-seq peaks occurring in candidate cis-regulatory elements (cCRE) for all the experimental groups. b, Number of p-CREB1S133 ChIP-seq peaks occurring in candidate cis-regulatory elements (cCRE) for each peak class resulting from differential binding analysis. c, Top enriched transcription factor motif (obtained using HOMER74) for each peak class resulting from differential binding analysis. d, Venn diagram showing overlap between genes associated with “lower in C26” p-CREB1S133 binding (Supplementary Table 3) and reduced expression in RNASeq of C26 13d vs. control (Supplementary Table 1). e, Top 10 terms from the GO Biological process 2025 enrichment analysis (EnrichR) of gene set intersection between downregulated DEGs in C26 13d vs. control (from RNAseq, Supplementary Table 1) and gene associated to “lower in C26” p-CREB1S133 binding signals from ChIP-seq analysis (Supplementary Table 3).
Extended Data Fig. 4 Ingenuity Pathways Analysis for differentially expressed genes in muscle biopsies from PDAC patients.
Top inhibited (that is z-score<0) 25 terms, ranked for number of genes, for Ingenuity Pathways Analysis (IPA) of differentially expressed genes in rectus abdominis biopsies from patients with pancreatic ductal carcinoma (PDAC) vs. controls. cAMP-PKA-CREB signalling-related terms are marked with red arrows. IPA analysis and input data taken from supplementary table S14 of Narasimhan et al.35.
Extended Data Fig. 5 Skeletal muscle noradrenaline content in C26-indued cachexia and characterization of tumour-conditioned media effects on C2C12 cells.
a, Noradrenaline content in gastrocnemius muscle at day 0 (control), day 10, and day 13 post C26 cell injection. As a technical control, noradrenaline in muscle after chemical sympathectomy (SE) is mostly undetectable. n = 6 C26 0, 10, 13 days, n = 3 SE. 1-way ANOVA. b,c, Il6 mRNA expression measured through qPCR following 24 h treatment of C2C12 myoblasts with C26 (b) or LLC (c) TCM at the indicated concentrations. Red squares indicate the concentration selected for further analysis, that is the concentration of TCM that induces ~15-fold increase in Il6 expression. n = 3 independent experiments; 1-way ANOVA. d,e, STAT3 (d) and NFκB (e) transcriptional activity measured through luciferase assay and quantified as relative light units (R.L.U.) in transfected C2C12 myoblasts, treated for 24 h with C26 TCM (15%) and LLC TCM (3%). Recombinant cytokines IL6 (100 ng/ml) and TNF (10 ng/ml) are used as controls of STAT3 and NFκB-dependent luciferase induction, respectively. n = 3 for each condition. 1-way ANOVA. f, Il6 mRNA expression fold change over control in C2C12 differentiated myotubes treated or not with the selected concentration of C26 TCM or LLC TCM for 24 h. n = 3 independent experiments; 1-way ANOVA. g,h, C2C12 myotube diameter in cells treated for 24 h with C26 (g) and LLC (h) TCM at the above-mentioned concentrations. n = 184 control and n = 178 C26 TCM myotube diameter from 3 independent experiments (g); n = 598 for control and n = 672 LLC TCM myotube diameter from 3 independent experiments (h). Two-tailed Mann–Whitney test. i, Adrb2 mRNA expression fold change over control in C2C12 differentiated myotubes treated or not with C26 TCM or LLC TCM for 24 h. n = 3 independent experiments; 1-way ANOVA. j, Surface expression of β2-adrenergic receptor in C2C12 myotubes treated or not with C26 or LLC TCM for 24 h. Permeabilized myotubes allowed to detect total β2-adrenergic receptor expression. Data are expressed as percentage over control. n = 3 independent experiments; 1-way ANOVA. k, Total cAMP measurement through plate-based quantification in C2C12 myotubes treated or not for 24 h with C26 TCM or LLC TCM. n = 3 independent experiments; 1-way ANOVA. l-n, Representative blot (l) and densitometry analysis of phosphorylated (S133) over total CREB1 (m) and phospho-serine/threonine PKA substrates over vinculin (n) in C2C12 myotubes treated or not for 24 h with C26 TCM or LLC TCM. n = 3 independent experiments; 1-way ANOVA. Data are presented as mean values ± SEM in a-k, m, and n; significant P values are annotated in the graphs.
Extended Data Fig. 6 Tumour-conditioned media-induced cAMP/CREB1 dysfunction in C2C12 myoblasts and gene expression data from human muscle biopsies of cachectic patients disaggregated by sex.
a-c, Representative blot (a) and densitometry analysis of phosphorylated (S133) over total CREB1 after 5 min, 15 min, and 60 min from 10 µM ISO treatment in C2C12 myoblasts pre-treated for 24 h with C26 (b) or LLC (c) TCM. n = 4 for each experimental group. 2-way ANOVA. One out of four replicates of the control group in graphs b and c is common for C26 and LLC TCM. d,e, Ppargc1a (PGC-1α) mRNA expression quantified by RT-qPCR following 10 µM ISO treatment at the indicated time in C2C12 myoblasts pre-treated with C26 (d) or LLC (e) TCM for 24 h. n = 3 for each experimental group. 2-way ANOVA. f, Nr4a3 (NOR1) mRNA expression quantified by RT-qPCR following 10 µM ISO treatment at the indicated time in C2C12 myoblasts treated with C26 TCM or LLC TCM for 24 h. n = 5 ctr, n = 4 C26 TCM, n = 3 LLC TCM for each experimental group. 2-way ANOVA. g-j NR4A3 (NOR1) and PPARGC1A (PGC-1α) mRNA expression in muscle biopsies from pre-cachectic (PC) and cachectic (C) pancreatic-cancer patients compared to controls in male (g,h) and female (i,j). For male cohort: n = 22 controls, n = 16 PC, n = 23 C. For female cohort: n = 29 controls, n = 18 PC, n = 25 C. 1-way ANOVA. Data are presented as mean values ± SEM in b-j and significant P values are annotated in the graphs.
Extended Data Fig. 7 Effects of rolipram on inflammation and muscle homeostasis-related pathways in C26-induced cachexia.
a, mRNA expression of interleukin 1 β (Il1b), interleukin 6 (Il6), tumour-necrosis factor (Tnf), and Activin A (Inhba) in tumour bulk from cachectic mice treated with vehicle (C26 veh) or rolipram (C26 Rol). n = 9 C26 veh and n = 8 C26 Rol for Il1b, Il6, Tnf. n = 9 C26 veh and n = 6 C26 Rol for Inhba. Two-tailed t-test. b-d, Tibialis anterior (b), extensor digitorum longus (EDL) (c), and soleus (d) muscle weight normalized over starting weight of C26-bearing mice at 13 days from cancer cell injection treated with vehicle (C26 veh) or rolipram (C26 Rol) as compared to controls (ctr). n = 5 ctr, n = 8 C26 veh and C26 Rol. 1-way ANOVA. e, Relative frequency of fibre type I, IIa, IIx, and IIb in gastrocnemius from ctr, C26 veh, and C26 Rol mice. n = 5 for each experimental group. 1-way ANOVA. f-o, Representative blot (f) and densitometry analysis of p-S6S235/236 over total S6 (g), p-4EBP1T37/T46 over GAPDH (h), p-AKTS473 over total AKT (i), AKT over GAPDH (j), LC3B-II over GAPDH (k), Beclin-1 over GAPDH (l), p-STAT3Y705 over total STAT3 (m), STAT3 over GAPDH (n), p-NFκBS563 over total NFκB (o) expressed as fold change over control, in control mice (ctr) and in C26 tumour-bearing mice treated with vehicle (C26 veh) or rolipram (C26 Rol). n = 5 for each experimental group. 1-way ANOVA. p,q, Glucose tolerance test (GTT) in ctr, C26 veh, and C26 Rol mice at 11 days post-cancer cell injection. In p are indicated absolute values. In q is indicated the percentage over time 0. n = 8 ctr, n = 6 C26 veh, n = 5 C26 Rol. Four values below the detection limit of the assay (20 mg/dL) were excluded. 2-way ANOVA, *P value in C26 veh vs. ctr. #P value in C26 Rol vs. ctr. Data are presented as mean values ± SEM in a-e and g-q, significant P values are annotated in the graphs.
Extended Data Fig. 8 Effects of rolipram on mitochondrial translation-related gene expression, on phospho-AMPK, and on endurance performance in C26-bearing mice.
a, Heatmap of normalized RPKM of rolipram-rescued DEGs selected from MitoXplorer gene lists “Translation” in vehicle- or rolipram-treated mice (C26 veh and C26 Rol) as compared to controls (ctr). n = 7 ctr, n = 7 C26 veh, and n = 5 C26 Rol mice. b,c, Representative image (b) and densitometry analysis (c) of p-AMPKT172, total AMPK, and vinculin of ctr, C26 veh, and C26 Rol mice. Densitometry of p-AMPKT172 was normalized over vinculin. n = 3 mice for each group. 1-way ANOVA. d, Endurance performance measured through Treadmill test; distance run in 60 min by ctr, C26 veh, and C26 Rol mice. n = 8 ctr, n = 7 C26 veh, n = 6 C26 Rol mice; 1-way ANOVA. Data are presented as mean values ± SEM in c and d, significant P values are annotated in the graphs.
Extended Data Fig. 9 Genotypic characterization and validation of Pde4b−/− and Pde4d−/− C2C12 clones.
a,b, Genotypic characterization of Pde4b−/− (a) and Pde4d−/− (b) through Sanger sequencing of C2C12 clones and WT C2C12. Single guide RNA (sgRNA) target sequence is highlighted in light blue while protospacer adjacent motif (PAM) sequence is highlighted in green. Sequencing of the “C4 clone” (a) revealed a homozygous single adenine insertion (in red) which generates a 1 bp frame-shift mutation. Sequencing of the “2A5 clone” (b) revealed heterozygous frame-shift mutations, in particular one 14 bp deletion and 1 bp single insertion (in red). c, Western blot analysis showing PDE4B and PDE4D protein expression in the above-mentioned knockout clones compared to WT C2C12. Vinculin was used as loading control. Genotyping characterization and validation by western blot analysis were performed in a single experiment.
Extended Data Fig. 10 Effects of tumour-conditioned medium on cAMP/CREB1 signalling in Pde4b−/− and Pde4d−/− C2C12, Pde4b and Pde4d transcript variants expression in muscle from C26 tumour-bearing mice, and analysis of in vivo Pde4d silencing.
a-d, Kinetic of CREB1 phosphorylation on serine 133 after 5 and 15 min from ISO treatment in WT, Pde4b−/−, or Pde4d−/− C2C12 treated with control or C26 TCM. Total CREB1 and vinculin (as housekeeping protein) are also shown. Representative blot and densitometry analysis (of phosphorylated over total CREB1) of WT and Pde4b−/− C2C12 (a,c) and of Pde4d−/− C2C12 (b,d). n = 3 independent experiments for c and n = 4 independent experiments for d. 2-way ANOVA. e, Nr4a3 (NOR1) mRNA expression at 1 h following 10 μM ISO treatment in WT, Pde4b−/−, or Pde4d−/− C2C12 treated with C26 TCM or control medium. n = 5 WT, n = 3 Pde4b−/−, n = 4 Pde4d−/−. 2-way ANOVA. f,g, Transcript per million (TPM) counts of Pde4b variants (Ensemble ID nomenclature) (f) Pde4d variants (Ensemble ID nomenclature) (g) in gastrocnemius from RNAseq analysis in cachectic (13 days post C26 injection) vs. control mice. TPM = 0 are not included in the graph. n = 7 mice for each experimental group. Two-tailed t-test for Pde4b-201, 203, 207 and Pde4d-204, 205, 207; two-tailed Mann-Whitney test for Pde4b-204 and Pde4d-201, 213. h, Tumour weight at 13 days from C26 cell s.c. injection in shCtr (C26 shCtr) and shPde4d (C26 shPde4d) mice. Mice were injected with 1011 AAV9 particles expressing a control shRNA sequence (C26 shCtr) or a Pde4d shRNA sequence (C26 shPde4d) 2 weeks prior tumour cell injection. n = 10 mice for C26 shCtr group and n = 9 mice for C26 shPde4d group; two-tailed t-test. i, Percentage of total weight loss at 13 days from tumour cell injection over the starting weight (day 0) of C26 shCtr vs. C26 shPde4d. n = 10 mice for C26 shCtr group and for C26 shPde4d group; two-tailed t-test. j, Spleen weight at 13 days from tumour cell injection in non-tumour bearing mice sham shCtr, and tumour bearing mice C26 shCtr and C26 shPde4d. n = 8 mice for sham shCtr group, n = 10 mice for C26 shCtr and for C26 shPde4d groups. 1-way ANOVA. k, Tibialis anterior weight normalized over mice starting weight in sham shCtr, C26 shCtr and in C26 shPde4d group. n = 8 mice for sham shCtr group, n = 10 mice for C26 shCtr and for C26 shPde4d groups. 1-way ANOVA. l, Cross-sectional area (CSA) frequency distribution measured in tibialis anterior from sham shCtr, C26 shCtr, and C26 shPde4d. n = 3 mice for sham shCtr group, n = 4 mice for C26 shCtr and C26 shPde4d. 2-way ANOVA; *p < 0.05 (800 μm2, 1800 μm2), ****p < 0,0001 (400 μm2, 600 μm2) in C26 shPde4d vs. C26 shCtr. p < 0.01 (400-1000 μm2 and 2000-3000 μm2) in C26 shCtr vs. sham shCtr (not shown in graph). m, mRNA expression of Fbxo32 (Atrogin1), Trim63 (MuRF1), and Fbxo30 (MUSA1) at day 13 in tibialis anterior of sham shCtr, C26 shCtr, and C26 shPde4d mice. n = 8 mice for sham shCtr and C26 shPde4d groups, n = 10 mice for C26 shCtr group. 1-way ANOVA. n, mRNA expression fold-change over control of Atp5j2, Mrpl12, Rbfa, Sdhc, Sod2, Cox5a, Mfn2, Ndufb4 expression in tibialis anterior from in sham shCtr, C26 shCtr and in C26 shPde4d groups. n = 8 mice for sham shCtr and C26 shPde4d groups, n = 10 mice for C26 shCtr group. 2-way ANOVA. o-t, Oxygen consumption rate (OCR) normalized over protein content (pmol/(s*mg)) of the leakage state (o), of OXPHOS complex II (p) and specific flux and maximum capacity (ETS) of CI (q), CI + CII (r), CII (s) and of CIV (t) measured in explanted tibialis anterior from sham shCtr, C26 shCtr, and C26 shPde4d. n = 4 mice for each experimental group. 1-way ANOVA. Data are presented as mean values ± SEM in c-t, significant P values are annotated in the graphs.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1
Supplementary Table 1 (download XLSX )
RNA-seq analysis of gastrocnemius muscles from control (ctr) and C26-bearing mice at 13 days after cancer cell inoculation treated with vehicle (C26 veh) or rolipram (C26 Rol). Sheet 1 reports the differential expression analysis comparing C26 veh versus ctr, while Sheet 2 shows the comparison between C26 Rol and C26 veh. The columns are organized as follows: (A) gene name, (B) base mean, (C) log2 fold change, (D) P value, (E) P adjusted value and (F) Mouse Genome Informatics (mgi) gene description. n = 7 ctr, n = 7 C26 veh and n = 5 C26 Rol.
Supplementary Table 2 (download XLSX )
Serine/threonine kinase activity profiling in gastrocnemius lysates from C26-bearing mice versus controls using PamChip peptide arrays; only inhibited kinases (that is, negative median kinase statistic) are shown. Columns: (A, B) kinase name and UniProt ID, (C, D) kinase group and family, (E) median kinase statistic, (F) mean significance score, (G) mean specificity score and (H) combined kinase score (sum of significance and specificity scores).
Supplementary Table 3 (download XLSX )
p-CREB1S133 differential binding sites detected by ChIP-seq in muscles from C26 at 10 days after cancer cell injection versus controls. A to D columns show peak coordinates; column E shows the peak name, column F the comparison score, column G the category (common, higher in C26, or lower in C26 signals), column H the associated gene, column I the distance from the transcription start site of the associated gene, and column J the cCREs associated to the peak.
Supplementary Table 4 (download XLSX )
Transcript per million (TPM) analysis from RNA-seq of gastrocnemius muscles from C26-bearing mice 13 days after cancer cell injection (C26) versus controls. Ensembl ID and gene name are shown in columns A and B.
Supplementary Table 5–8 (download XLSX )
Supplementary Table 5: sgRNA sequences and screening primers for Pde4b or Pde4d knockout generation. Supplementary Table 6: SYBR probe sequences. Supplementary Table 7: Taqman probes. Supplementary Table 8: Primary antibody list.
Source Data Supplementary Fig. 1 (download XLSX )
Statistical source data for Supplementary Fig. 1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download JPG )
Unprocessed western blots.
Source Data Fig. 2 (download JPG )
Unprocessed western blots.
Source Data Fig. 3 (download JPG )
Unprocessed western blots.
Source Data Fig. 4 (download JPG )
Unprocessed western blots.
Source Data Fig. 6 (download JPG )
Unprocessed western blots.
Source Data Fig. 7 (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 9 (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download JPG )
Unprocessed western blots.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Angelino, E., Bodo, L., Sartori, R. et al. Impaired cAMP–PKA–CREB1 signalling drives mitochondrial dysfunction in skeletal muscle during cancer cachexia. Nat Metab 7, 2548–2570 (2025). https://doi.org/10.1038/s42255-025-01397-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01397-5
This article is cited by
-
Reawakening cAMP signalling in cancer cachexia
Nature Metabolism (2025)