Abstract
Spatiotemporal changes in the nuclear lamina and cell metabolism shape cell fate, yet their interplay is poorly understood. Here we identify lamin A/C as a key regulator of cysteine catabolic flux essential for proper cell fate and longevity. Its loss in naive mouse pluripotent stem cells leads to upregulation of the cysteine-generating and catabolizing enzymes, cystathionine γ-lyase (CTH) and cystathionine β-synthase (CBS), thereby promoting de novo cysteine synthesis. Increased cysteine flux into acetyl-CoA fosters histone H3K9 and H3K27 acetylation, triggering a transition from naive to primed pluripotency and abnormal cell fate and function. Conversely, the toxic gain-of-function mutation of Lmna, encoding lamin A/C and associated with premature ageing, reduces CTH and CBS levels. This reroutes cysteine catabolic flux and alters the balance between H3K9 acetylation and methylation, crucially impacting germ layer formation and genome stability. Notably, modulation of Cth and Cbs rescues the abnormal cell fate and function, restores the DNA damage repair capacity and alleviates the senescent phenotype caused by lamin A/C mutations, highlighting the potential of modulating cell metabolism to mitigate epigenetic diseases.
Similar content being viewed by others
Main
Pluripotent stem cells possess the capacity for self-renewal and differentiation into diverse cell types. Upon exposure to developmental cues, these cells undergo coordinated changes in chromatin architecture, gene expression, metabolism and cellular behaviour to execute precise lineage specification and tissue morphogenesis1,2,3,4. While the importance of cell metabolism and chromatin architecture in stem cell pluripotency, cell fate decisions and differentiation is widely recognized, their potential interplay in directing cell fate remains largely unexplored.
Various states of embryonic pluripotency, such as naive and primed states can be effectively maintained in culture. Naive and primed pluripotency states correspond to the pre- and post-implantation epiblast, respectively, and differ in their developmental potential5,6. Naive pluripotency, characterized by the unbiased ability to generate all somatic cell lineages and germ cells, is recapitulated in in vitro settings in mouse embryonic stem (mES) cells. In contrast, primed cells are poised for differentiation and cannot contribute to blastocyst chimeras. These two states are characterized by distinct transcriptional profiles, epigenetic landscapes and metabolic activity4,7. A close link between cell metabolism and transcriptional-epigenetic changes is important for both stem cell pluripotency and differentiation, as several metabolites act as cofactors for transcriptional and epigenetic modifiers8,9,10,11,12,13,14,15,16. For example, methyl groups required for the synthesis and methylation of DNA, RNA and associated proteins, as well as the biogenesis of amino acids, creatine, phospholipids, NADH and NADPH are dependent on one-carbon (1C) metabolism17. This pathway links the methionine and folate cycles with cysteine biosynthesis and catabolism, and is central to cellular function. Indeed, methionine metabolism is elevated in pluripotent stem cells and is required to maintain the high level of DNA and histone methylation required to maintain pluripotency. This is also reflected by the fact that methionine depletion reduces histone and DNA methylation and promotes pluripotent stem cells differentiation8. Oxidative metabolism is also high in ‘naive’ versus ‘primed’ stem cells9, which can cope with the increased generation of reactive oxygen species (ROS) by virtue of augmented levels of nonenzymatic antioxidants.
The dynamics of genome–nuclear lamina interactions play a pivotal role in determining cell fate decisions by either ‘locking’ or ‘unlocking’ genes that confer cell identity at the nuclear periphery2,18. Nuclear lamins build filamentous networks located between the inner nuclear membrane and the peripheral heterochromatin. These proteins are strategically positioned to perform vital roles in maintaining nuclear structure19,20,21 as well as orchestrating chromatin organization by anchoring heterochromatin regions termed lamina-associated domains (LADs) to the nuclear periphery22,23,24. Additionally, they serve as a scaffold for tethering protein complexes, facilitating DNA replication and repair25,26,27. Consistent with the important biological functions of nuclear lamins, numerous studies have shown that their loss of function (LOF) or mutation results in genomic instability, chromatin alterations, abnormal gene expression patterns28,29,30,31,32,33,34,35 and influence stem cell pluripotency, proliferation, longevity, cell fate choices and differentiation, as well as stress responses2,18,33,36,37. For instance, during the differentiation of mES cells, astrocyte-specific genes18 and cardiomyocyte-specific genes33,38 detach from mES cell LADs. This detachment leads to their activation or epigenetic priming for activation at a later stage33,38. Notably, lamin A/C is expressed in naive pluripotent stem cells but dramatically decreases with the acquisition of a primed state33. Abnormal cell fate upon lamin A/C loss or mutation has been reported in other tissues32,39,40,41, suggesting that aberrant activation of cell fate genes could be a common feature of laminopathic cells, carrying mutations in genes encoding nuclear lamins42,43,44.
Here, we present evidence of lamin A/C-mediated epigenetic control beyond LADs, which, exerted by lamin A/C-regulated cysteine flux into acetyl-CoA and histone acetylation, is crucial for proper germ layer formation, cell fate determination and ageing.
Results
Lamin A/C represses cysteine biosynthesis and catabolism in naive pluripotent stem cells
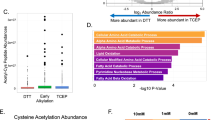
In a recent study, we found that lamin A/C plays a crucial role in three-dimensional (3D) chromatin organization in naive pluripotent stem cells, where it safeguards naive pluripotency and ensures proper cell fate choices during cardiogenesis33. A comparison of genes found in chromatin regions showing increased accessibility in Lmna−/− mES cells with genes located within lamin A/ lamin B1 LADs revealed that more than 85% of the former reside in non-LAD regions (Fig. 1a). To investigate how lamin A/C mutations could affect chromatin structure outside LADs we performed Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) pathway analysis of genes upregulated in Lmna−/− mES cells. This highlighted the impact of Lmna depletion on cell metabolism (Fig. 1b and Supplementary Fig. 1a), and prompted the hypothesis that metabolic adaption resulting from the lack of lamin A/C could alter the bioavailability of histone epimodifiers (such as acetyl and methyl groups), leading to maladaptive epigenetic and transcriptional remodelling. Indeed, a metabolomics approach to compare metabolism in Lmna-expressing and -deficient mES cells revealed significant alterations in key metabolic pathways, particularly in cysteine and methionine metabolism (Fig. 1b,c and Supplementary Fig. 1a–c and Supplementary Table 1). ʟ-Cystathionine, ʟ-cysteine and ʟ-cystine, which are all intermediates of the cysteine metabolism, were most altered by Lmna deficiency, whereas other pathways such as glycolysis, the tricarboxylic acid (TCA) cycle and glutaminolysis were not affected (Fig. 1d and Supplementary Fig. 1d–f).
a, Overlap of genes showing increased ATAC-seq signal in Lmna−/− mES cells with genes located within lamin A/B1-shared LADs as determined by lamin A and lamin B1 DamID. b, KEGG pathway analyses of genes upregulated in Lmna−/− mES cells. n = 3 biological replicates. c, Pathway impact analysis showing metabolic alterations between Lmna−/− and control groups. n = 6 biological replicates. d, Intracellular levels of metabolites in control and Lmna−/− mES cells determined by LC–MS or by fluorometric assay (acetyl-CoA). n = 6 biological replicates. e, Representative DNA FISH images of Cth (red), Cbs (red) and 4,6-diamidino-2-phenylindole (DAPI) (blue). f, ChIP–qPCR for lamin A/C at the Cth and Cbs promoter in control and Lmna−/− mES cells presented as % of input. n = 3 biological replicates. g, Representative immunoblots analysis of CTH and CBS levels in Lmna+/+ and Lmna−/− mES cells (n = 6 independent experiments). α-tubulin served as a loading control. h, Motif enrichment analysis of accessible chromatin regions within ±5 kb of the Cbs and Cth transcription start sites in Lmna−/− versus control cells. n = 3 biological replicates. Significance was calculated in HOMER using a two-sided hypergeometric test. i, ChIP–qPCR analysis of SP1 enrichment at the Cth and Cbs promoter in control and Lmna−/− mES cells; GD, gene desert region, used as a negative control. n = 4 biological replicates. j, qPCR analysis of Cth and Cbs mRNA level in control and Lmna−/− mES cells after Sp1 knockdown by siRNA. n = 4 biological replicates. k, Illustration presenting hallmarks of progeria. l, KEGG pathway analyses of genes downregulated in LmnaG609G/+ and LmnaG609G/G609G mES cells. m, Immunoblots analysis of CTH and CBS levels in control, LmnaG609G/+ and LmnaG609G/G609G mES cells. n, Relative levels of intracellular cysteine, pyruvate and acetyl-CoA in control, LmnaG609G/+ and LmnaG609G/G609G mES cells. n = 6 biological replicates. Data in d,f,i,j,n are mean ± s.d. Differences were assessed using an unpaired two-tailed Student’s t-test (d,f,i); a two-way ANOVA with Tukey’s correction (j); and a one-way ANOVA with Tukey’s correction (n). Significance was calculated in DAVID using a one-sided Fisher’s exact test (b) and a one-sided hypergeometric test in MetaboAnalyst v.5.0 (c). NS, not significant.
Two key cysteine metabolizing enzymes exist, which are cystathionine β-synthase (CBS) and cystathionine γ-lyase (CTH, also known as CSE), and the Cth gene localizes to the nuclear periphery (Fig. 1e) and resides within LADs (Fig. 1a). Lmna deletion in mES cells led to the relocalization of Cth from the transcriptionally inactive nuclear periphery to the nuclear interior, whereas the positioning of Cbs remained unchanged (Fig. 1e and Extended Data Fig. 1a). However, chromatin immunoprecipitation followed by quantitative PCR (ChIP–qPCR) analysis revealed significant enrichment of lamin A/C at the promoter regions of both Cbs and Cth (Fig. 1f), suggesting a direct role in regulating their expression. Further, we observed increased chromatin accessibility and transcriptional upregulation of both Cth and Cbs genes (Extended Data Fig. 1b–d) as well as elevated protein expression (Fig. 1g and Extended Data Fig. 1f,g) and enzyme activity upon lamin A/C loss (Extended Data Fig. 1e). In contrast, other enzymes involved in 1C metabolism were not significantly altered (Extended Data Fig. 1f,g).
To investigate the mechanisms underlying the upregulation of Cth and Cbs, we performed motif analysis at their promoter regions. In both cases, we identified SP1 as a key regulatory transcription factor (Fig. 1h). SP1 has previously been implicated in the transcriptional regulation of both Cth and Cbs45,46. We therefore examined whether lamin A/C influences SP1 binding and function. Notably, we observed increased chromatin binding by SP1 (Extended Data Fig. 1h), with significantly higher enrichment at the Cth and Cbs promoters in Lmna-deficient cells (Fig. 1i). Silencing Sp1 in Lmna−/− mES cells reduced Cth and Cbs expression to levels observed in control cells, whereas Sp1 knockdown in wild-type mES cells led to further reduction (Fig. 1j). In contrast, ATF4, a key transcription factor activated during cellular stress that has been shown to regulate the levels of CTH and CBS, did not show changes in expression levels or chromatin binding upon lamin A/C loss (Extended Data Fig. 1i). Together, these data support a central role for SP1 in the regulation of Cth and Cbs expression in mES cells and downstream of lamin A/C.
Lamin A/C restricts transcription factor binding through recruitment of repressive complexes, including the EZH-based PRC2 complex, SUV39H1 and HDACs41. We therefore tested whether these complexes regulate Cbs and Cth expression. Inhibition of Ezh1/2 with UNC1999 led to a significant increase in Cbs and Cth expression (Extended Data Fig. 1j and Supplementary Fig. 2a), whereas inhibition of Suv39h1 or HDACs had little effect (Supplementary Fig. 2b,c). Notably, H3K27me3, catalysed by the PRC2 complex, was reduced at both Cth and Cbs promoters in Lmna−/− mES cells (Extended Data Fig. 1k). Treatment with UNC1999 decreased H3K27me3 levels and increased Sp1 binding (Extended Data Fig. 1l), supporting a direct role for PRC2 in lamin A/C-mediated repression of Cth and Cbs and in limiting SP1 occupancy.
To further investigate the link between cysteine metabolism and the nuclear lamina, we studied the Lmna p.G609G mutation47, which leads to the production of a dominant gain-of-function isoform of lamin A, referred to as progerin. Progerin is constitutively farnesylated on its C terminus and accumulates in the nuclear lamina, thereby disturbing nuclear structure as well as nuclear lamina dynamics and function48 (Extended Data Fig. 2a,b) and giving rise to Hutchinson–Gilford progeria syndrome (HGPS), a severe form of accelerated ageing47 (Fig. 1k). A comparison of genes found in chromatin regions showing decreased accessibility in Lmna p.G609G mutant mES cells with genes located within lamin A/lamin B1 LADs revealed that more than 70% of the former reside in non-LAD regions (Extended Data Fig. 2c). KEGG pathway analysis of genes downregulated in Lmna p.G609G mutant cells highlighted the importance of 1C and cysteine and methionine metabolism, similarly to Lmna−/− mES cells (Fig. 1l). Moreover, the expression of CTH and CBS as well as intracellular cysteine, pyruvate and acetyl-CoA levels were all decreased in mES cells that were either heterozygous or homozygous for the mutation (Fig. 1m,n and Extended Data Fig. 2d,e). In contrast to our findings in the Lmna LOF, the Lmna p.G609G mutation led to decreased SP1 chromatin binding (Extended Data Fig. 2f), accompanied by significantly reduced SP1 occupancy and increased H3K27me3 and lamin A/C enrichment at the Cth and Cbs promoters (Extended Data Fig. 2g,h). Taken together, these data suggest that altering lamin A/C function affects cysteine biosynthesis and catabolism in naive mES cells.
CTH and CBS are upregulated during naive-to-primed state transition in a lamin A/C-dependent manner
Lamin A/C loss results in naive-to-primed pluripotency transition and abnormal cell fate choices during development18,33. To further assess the impact of nuclear lamina–cysteine catabolism axis on stem cell differentiation, we next used a metabolomics approach. Of note, methionine metabolism and homocysteine degradation were one of the most significant pathways enriched during the specification of stem cells in the three germ layers (Extended Data Fig. 3a). Notably, homocysteine, cystathionine and cysteine levels were markedly increased in day 3 embryoid bodies (EBs) compared with naive mES cells, whereas cystathionine and cysteine levels remained high at later stages, suggesting an important role of cysteine metabolism during the early stages of stem cell differentiation (Extended Data Fig. 3b).
To assess the impact of cysteine catabolism and the interplay between the nuclear lamina and stem cell pluripotency, naive embryonic stem (ES) cells were switched to conditions favouring a primed state. ES cell priming resulted in the loss of lamin A/C, increased the expression of CTH and CBS (Extended Data Fig. 3c) along with increased cellular cysteine levels (Extended Data Fig. 3d). Similar results were observed with human ES cells cultured in naive versus primed conditions7 (Extended Data Fig. 3e). Because these observations suggested that increased CTH and CBS levels were linked to induction of primed pluripotency, we determined whether depletion of Cth or Cbs could re-establish a naive pluripotent state in Lmna−/− mES cells. Indeed, knockdown of both genes in Lmna−/− mES cells restored colony morphology to the compact and round shape reminiscent of naive cells (Extended Data Fig. 3f and Supplementary Fig. 3a) and increased the number of cells expressing high levels of the pluripotency marker OCT4 (OCT4high) and NANOG (NANOGhigh) (Extended Data Fig. 3f,g and Supplementary Fig. 3b,c). In contrast, CTH- and CBS-overexpressing mES cells, formed flatter, irregular colonies that contained fewer cells expressing high levels of OCT4 and NANOG compared with control mES cells (Extended Data Fig. 3f,g and Supplementary Fig. 3b,c). In line with the observed shift in pluripotency state, markers of primed pluripotency were significantly upregulated in Lmna−/− mES cells. Silencing either Cth or Cbs reduced their expression to levels comparable with controls, with Cth knockdown having a more pronounced effect (Extended Data Fig. 3h).
Together, these data suggest that increased de novo cysteine synthesis and flux may drive the transition from naive to primed pluripotency. Supporting this hypothesis, cysteine depletion, either alone or combined with the CTH and CBS inhibitor AOAA (aminooxyacetic acid), increased the number of OCT4high and NANOGhigh mES cells (Extended Data Fig. 3i and Supplementary Fig. 3d).
Next, we tested the impact of lamin A–green fluorescent protein (GFP) overexpression49 using a TET-ON inducible system in wild-type and Lmna−/− mES cells, focusing on its effects on CTH and CBS expression levels and the acquisition of naive pluripotency. The GFP signal was detectable as early as 2 h (Supplementary Fig. 3e), and by 6 h, we observed a marked increase in lamin A protein levels, accompanied by a significant reduction in Cth and Cbs as well as cysteine levels (Extended Data Fig. 3j–l). Of note, cells expressing high levels of lamin A consistently showed lower levels of both CTH and CBS (Extended Data Fig. 3j). Despite these changes, the proportion of NANOGhigh mES cells remained unchanged at 6 h (Extended Data Fig. 3m), suggesting that the downregulation of CTH and CBS is a direct effect of lamin A overexpression, rather than a secondary consequence of altered pluripotency state. At the 24-h and 48-h time point, we observed a notable decrease in CTH and CBS levels (Extended Data Fig. 3j,l and Supplementary Fig. 3f,g), accompanied by an increase in NANOGhigh mES cells (Extended Data Fig. 3m and Supplementary Fig. 3h). Overexpression of lamin A in cells cultured under primed conditions also led to an increased number of NANOGhigh cells, indicating that lamin A has the capacity to promote naive features even if cultured in primed conditions (Extended Data Fig. 3n,o). Together, these data indicate that lamin A-mediated metabolic rewiring precedes and support the re-establishment of the naive pluripotency.
To further validate these findings and account for potential line-to-line variability in responsiveness, we ablated Lmna in R1 mES cells, an F1 hybrid of the 129X1 × 129S1 strains, using the same gene-targeting strategy employed to generate the Lmna knockout in E14 cells33. R1 mES cells expressed slightly, yet significantly, higher levels of lamin A/C compared with the E14 line (Extended Data Fig. 4a,b). Further, lamin A/C protein levels correlated with NANOG expression, and cells with low lamin A/C showed no detectable NANOG signal (Extended Data Fig. 4c). Moreover, culture in primed conditions led to reduced lamin A/C levels, increased expression of CTH and CBS and higher cysteine levels (Extended Data Fig. 4d,e). Similarly, lamin A/C loss in R1 cells led to increased CTH and CBS levels, along with elevated intracellular cysteine (Extended Data Fig. 4f–h). Moreover, Lmna-deficient R1 mES cells adopted a flatter morphology, a reduced number of NANOGhigh cells, and exhibited increased expression of primed pluripotency markers, consistent with a shift toward a more developmentally advanced state (Extended Data Fig. 4i–l).
Together, these data demonstrate that the upregulation of CTH and CBS is a conserved feature of the naive-to-primed transition and occurs in a lamin A/C-dependent manner.
The transition from pre- to post-implantation is marked by decreased lamin A/C expression coupled with increased CTH and CBS levels
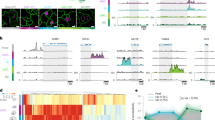
Next, we examined the relationship between lamin A/C and Cth and Cbs expression in preimplantation (E3.5 and E4.5) and post-implantation (E5.5–E6.75) mouse embryos, corresponding to the naive and primed pluripotency states, respectively (Fig. 2a). Single-cell RNA sequencing (scRNA-seq) data50 along with immunostainings revealed that while Lmna was highly expressed in early epiblast cells of E3.5 and E4.5 embryos, Cth and Cbs levels were very low. In contrast, Lmna expression decreased sharply from E5.5, coinciding with a dramatic increase in Cth and Cbs levels (Fig. 2b–e).
a, Schematic representation of the transition from naive to primed pluripotency. b, Feature plots of the expression of Lmna, Cth and Cbs across E3.5-E6.75 mouse embryonic stages. Trophectoderm cells expressing Eomes or Cdx2 were excluded from the analyses of E3.5 and E4.5 embryos. c, Violin plots visualizing expression levels of Lmna, Cth and Cbs in preimplantation (E3.5 and E4.5) and post-implantation (E5.5) mouse embryos. d,e, Representative immunostaining for lamin A/C (red) and CTH (green) (d) or lamin A/C (red) and CBS (green) (e) in E3.5 and E6.5 mouse embryos. n = 6 embryos. Scale bars, 20 µm. f, TOBIAS footprinting analysis showing significant enrichment of RA-responsive motifs in accessible chromatin regions within ±5 kb of the Lmna transcription start site in mES cells cultured under primed conditions compared with naive conditions (ArrayExpress, E-MTAB-7207). Significance was calculated using a two-sided hypergeometric test within the TOBIAS footprinting framework. g, Violin plots visualizing expression levels of RA-pathway-related genes in preimplantation (E3.5 and E4.5) and post-implantation (E5.5) mouse embryos. h, qPCR analysis of Lmna and Nanog expression in wild-type (WT) mES cells after 24 h treatment with all-trans-retinoic acid in the absence of LIF. n = 3 biological replicates. i, KEGG analysis of the genes upregulated in E5.5 compared with E3.5. Significance was calculated in DAVID using a one-sided Fisher’s exact test. j, Motif analysis of the promotor region (TSS ± 2 kb) of genes upregulated in E5.5 compared with E3.5 embryos. Significance was calculated in HOMER using a two-sided hypergeometric test. k, Dot-plot showing the expression level and frequency of Sp1 from E3.5 to E6.75. l,m, Representative immunostaining for lamin A/C (red) and CTH (green) (l) or lamin A/C (red) and CBS (green) (m) in Lmna+/+ and Lmna−/− mouse embryos. Scale bars, 20 µm. n, Quantification of fluorescence intensity of CTH (top) and CBS (bottom) in the inner cell mass and trophectoderm (TE) of Lmna+/+ and Lmna−/− mouse embryos. For CTH: Lmna+/+ (n = 7) and Lmna−/− (n = 5); for CBS, Lmna+/+ (n = 8) and Lmna−/− (n = 3). Data in h and n represent mean ± s.d. Differences in h and n were assessed using an unpaired two-tailed Student’s t-test.
To identify factors involved in the rapid downregulation of lamin A/C in primed compared with naive stem cells, we first performed TOBIAS footprinting analysis on the Lmna promoter. This revealed RARA:RXRG and RARB, retinoic acid (RA) receptors known to mediate RA signalling, as top candidate regulators (Fig. 2f). Previous studies have linked RA signalling to Lmna expression, which is also influenced by matrix stiffness51. Notably, the mechanical properties of the embryonic environment change markedly between the pre- and post-implantation stages, transitioning from a soft, extracellular matrix (ECM)-poor setting to a stiffer, ECM-rich environment52. Consistent with a developmental role for RA signalling, scRNA-seq data showed strong upregulation of Rara and Rxrg at E5.5 (ref. 50) (Fig. 2g). Furthermore, treatment of mES cells with RA led to a marked reduction in both Lmna and Nanog expression (Fig. 2h), supporting a model in which RA signalling contributes to the downregulation of lamin A/C during the transition from naive to primed pluripotency. Further KEGG pathway analysis of genes significantly upregulated in E5.5 compared with E3.5 embryos revealed enrichment in DNA replication, oxidative phosphorylation, amino acid biosynthesis, and cysteine and methionine metabolism (Fig. 2i). Consistent with our in vitro findings, we observed significant enrichment of the SP1 binding motif at genes upregulated in E5.5 versus E3.5 embryos (Fig. 2j), which further increased during development (Fig. 2k). Additionally, CTH and CBS levels were significantly elevated in Lmna−/− embryos (Fig. 2l–n), hereby validating the physiological relevance of our in vitro findings.
Cysteine serves as a major carbon source for histone acetylation at lysine 9 and lysine 27 in Lmna mutant mES cells
Cysteine is a precursor of important metabolites including taurine and glutathione, one of the most important mammalian nonenzymatic antioxidants, thereby serving an essential function in regulating the redox balance53 (Supplementary Fig. 1c). In addition, cysteine catabolism under specific conditions can also generate the bioactive signalling molecule hydrogen sulfide (H2S)54, which regulates persulfidation, a post-translational modification, of proteins. Persulfidation is essential for cell fate, function and longevity55,56,57. However, although CBS and CTH were metabolically active in Lmna−/− mES cells, they failed to generate H2S (Extended Data Fig. 5a), indicating that in Lmna−/− mES cells, the non-canonical catabolic pathway of cysteine is preferred54.
To assess the metabolic fate of cysteine in mES cells in greater detail we performed metabolic flux analyses and determined the contribution m + 3 and m + 2 mass isotopologues to the pyruvate and acetyl-CoA pools following supplementation with 13C3-ʟ-cysteine (Fig. 3a). 13C3-ʟ-cysteine levels were increased in Lmna−/− mES cells compared with control mES cells, indicating an increased cysteine uptake (Fig. 3b), a change that could be attributed to increased expression of cysteine transporters (Extended Data Fig. 5b). Further, the flux of cysteine-derived carbons was directed towards pyruvate and acetyl-CoA rather than towards glutathione (Fig. 3b), pointing to a previously unexplored role for cysteine catabolism in the regulation of acetyl bioavailability within mES cells. The bioavailability of nuclear acetyl-CoA used by histone acetyltransferases (HATs) to transfer an acetyl group to lysine residues in histone proteins, is an important regulator of chromatin accessibility and higher order chromatin organization. In line with increased total acetyl-CoA levels, we detected a major increase in nuclear acetyl-CoA (Extended Data Fig. 5c). Thus, we determined whether cysteine-derived carbons contribute to histone acetylation by tracing 13C3-ʟ-cysteine. Strikingly, only a limited number of lysine residues showed incorporation of 13C3-acetyl groups derived from 13C-cysteine in control mES cells. Notably, the highest incorporation was observed at H2A/BK15ac, a site with markedly low global acetylation levels, suggesting site-specific incorporation rather than a general, widespread labelling across histone lysines in mES cells (Fig. 3c). A number of acetylated sites showed significantly higher levels in Lmna−/− mES cells compared with control mES cells. Notably, H3K9, H3K27 and H4K12 showed remarkable incorporation of 13C3-acetyl derived from 13C-cysteine in Lmna−/− mES cells, whereas no incorporation was detected in control mES cells, indicating that cysteine-derived carbons promote acetylation at these sites specifically upon lamin A/C loss (Fig. 3c). Immunoblot analysis confirmed the general increase in histone 3 (H3) acetylation as well as the specific acetylation of H3K9 and H3K27 (Extended Data Fig. 5d). Such an effect was directly linked to the expression of CTH and CBS, as silencing of either Cth or Cbs by a small hairpin RNAs (shRNA) normalized the levels of H3K9ac, H3K27ac and pan-H3ac in Lmna−/− mES cells back to the levels in control cells (Fig. 3d,e and Extended Data Fig. 5e). The overexpression of CTH or CBS in wild-type mES cells, on the other hand, increased acetylation of the same sites (Fig. 3d,e). Similarly, treatment with the CTH and CBS inhibitor AOAA decreased H3K9ac, H3K27ac and pan-H3ac levels (Extended Data Fig. 5f). As methionine serves as the precursor for cysteine synthesis, we next examined the impact of methionine and cysteine depletion on histone acetylation. Deprivation of methionine alone or in combination with cysteine depletion successfully rescued the increased histone acetylation (Fig. 3f). Interestingly, histone residues that were not found decorated with cysteine carbons for example H3K56ac, were also not different between wild-type and Lmna deleted cells (Extended Data Fig. 5d). Notably, loss of Lmna did not lead to a global change of total protein acetylation (Extended Data Fig. 5g), suggesting that acetyl-CoA generated from cysteine is preferentially transferred to histones. Similar results were observed in control and Lmna−/− R1 mES cells (Extended Data Fig. 5h).
a, Schematic representation of the experimental setup to trace 13C-labelled cysteine. b, Mass isotopologue distribution and intracellular abundance of 13C-labelled cysteine, pyruvate, acetyl-CoA and glutathione in a 13C3-ʟ-cysteine flux assay using Lmna+/+ and Lmna−/− mES cells. n = 3 biological replicates. c, Enrichment of 13C3-ʟ-cysteine-derived carbon at acetylated histone sites in Lmna+/+ and Lmna−/− mES cells (left), as well as the distribution of global histone acetylation in WT mES cells (right), presented as area under the curve per milligram of total protein. n = 3 biological replicates. d,e, Immunoblots analysis of H3K9ac, H3K27ac and pan-H3ac levels in Lmna+/+ and Lmna−/− mES cells after shRNA mediated silencing with control shRNA or shRNA against Cth or Cbs or mES cells overexpressing CTH (CTH OE) or CBS (CBS OE) constructs (d), and quantification of the relative fold changes of protein levels from three independent experiments (e). f, Immunoblot analysis of acetylated histones in control and Lmna−/− mES cells cultured for 24 h in complete medium, or in medium lacking methionine (Met), cysteine (Cys) or both. g, Immunoblot analysis for H3K9ac, H3K27ac and pan-H3ac of histone extracts from control, LmnaG609G/G609G and LmnaG609G/G609G mES cells overexpressing both CBS and CTH. h, Immunoblot analysis for H3K9ac, pan-H3ac and total H3 of histone extracts from control and LmnaG609G/G609G mES cells cultured in the indicated concentration of cysteine for 48 h. i,j, Immunoblot analysis of H3K9ac, H3K27ac and H3K56ac levels in mES cells cultured in naive or primed conditions for 48 h (i) and relative fold changes from three biological replicates (j). H3 served as a loading control. k, Representative immunostaining for lamin A/C (red) and H3K9ac (green) in E3.5 and E6.5 mouse embryos. n = 6 embryos. Scale bars, 20 µm. Data in b,c,e,j are presented as mean ± s.d. Differences in b and c were assessed using an unpaired two-tailed Student’s t-test; Differences in e and j were assessed using one-way ANOVA with Tukey correction. NS, not significant.
In contrast with Lmna−/−, Lmna p.G609G mES cells showed a significant decrease in H3K9ac and H3K27ac, as well as pan-H3ac levels (Fig. 3g and Extended Data Fig. 5i), in line with the decrease of CTH and CBS and the major role in cysteine-mediated histone acetylation in ES cells. Notably, overexpression of CTH together with CBS was sufficient to increase histone acetylation levels in LmnaG609G/G609G mES cells to control levels (Fig. 3g and Extended Data Fig. 5i). Moreover, increasing the cysteine concentration in the medium (to 400μmol l−1) partially restored the decreased histone acetylation in p.G609G mES cells (Fig. 3h). In line with this, overexpression of lamin A in mES cells reduced H3K9ac levels after 24 h, occurring after the downregulation of CTH and CBS, suggesting that changes in histone acetylation occur downstream of changes in cysteine metabolism (Extended Data Fig. 5j).
To evaluate whether altered histone deacetylase (HDAC) activity contributes to the changes in histone acetylation levels, we measured the activity of class I HDACs, the main histone deacetylating enzymes. HDACs class I activity remained unchanged in Lmna−/− cells but was reduced in progeroid cells that had lower H3K9ac and H3K27ac levels, suggesting that changes in class I HDAC activity is unlikely to account directly for the increased acetylation (Supplementary Fig. 4a,b).
As we observed a major increase in CTH and CBS expression during the transition to primed pluripotency, we next analysed histone acetylation in naive versus primed cells both in vitro and in vivo. Consistently, the levels of H3K9ac and H3K27ac were significantly higher in primed cells (Fig. 3i,j). In contrast, H3K56ac, which is not labelled with cysteine-derived carbons, showed no significant change. Notably, H3K9ac exhibited the greatest changes in cell culture, and we observed higher H3K9ac levels in post-implantation compared with pre-implantation embryos (Fig. 3k).
As we detected increased cysteine uptake (Fig. 3b), a change likely mediated by cysteine transporters (Extended Data Fig. 5b), we next analysed the expression of Slc1a1, Slc1a5 and Slc7a11 transporters in pre- and post-implantation embryos. Slc1a1 and Slc1a5 were significantly upregulated in Lmna-deficient cells. While Slc1a1 expression was specific to E3.5, Slc1a5 levels gradually increased from E3.5 to E5.5. of note, the major cysteine transporter Slc7a11 showed only minimal expression (Supplementary Fig. 5a), similar to mES cells. Silencing of Slc1a1 and Slc1a5 in Lmna-deficient cells only partially reduced the upregulation of key transcription factors associated with disease phenotypes following lamin A/C loss (Supplementary Fig. 5b), but this effect was considerably weaker than that observed upon silencing Cth. Unlike the major cysteine transporter Slc7a11, Slc1a1 and Slc1a5 primarily transport glutamate, which may also influence pluripotent stem cell fate15.
Taken together, these data support the notion that nuclear lamina-regulated cysteine catabolism modulates histone acetylation during early embryonic development.
CTH activation results in epigenome remodelling and transcriptional activation
Next, we determined whether changes in chromatin accessibility in Lmna−/− mES cells relied on cysteine-dependent histone acetylation. ChIP-seq analysis revealed higher levels of H3K9ac and H3K27ac in Lmna−/− and CTH or CBS-overexpressing (CTH OE and CBS OE) mES cells at ATAC-seq peaks that were increased in Lmna−/− mES cells (Fig. 4a–d). Genes with significantly increased H3K9ac were associated with GO terms including cell adhesion, regulation of Pol II transcription, cell differentiation, pattern specification and calcium ion transport, whereas genes with increased H3K27ac were linked to regulation of Pol II-mediated transcription, cell differentiation, chromatin remodelling and protein phosphorylation (Supplementary Fig. 6a). In contrast, ATAC-seq peaks that were decreased upon lamin A/C loss exhibited decreased H3K27ac in Lmna−/− mES cells as well as decreased H3K9ac and H3K27ac in CTH and CBS OE mES cells (Supplementary Fig. 6b).
a, Integrative Genomics Viewer (IGV) tracks of H3K9ac ChIP-seq, H3K27ac ChIP-seq, ATAC-seq and RNA-seq of control (grey), and Lmna−/− mES cells (red) on genes showing increased chromatin accessibility and expression upon lamin A/C loss. b, Normalized H3K9ac and H3K27ac ChIP-seq signal intensity in control (grey) and Lmna−/− ES cells (red) at ATAC-seq peaks exhibiting increased (gain) chromatin accessibility in Lmna−/− mES cells versus control mES cells. n = 3 biological replicates. c, IGV tracks of H3K9ac ChIP-seq, H3K27ac ChIP-seq and RNA-seq of control (grey), CBS OE (green) and CTH OE mES cells (purple). d, Normalized H3K9ac ChIP-seq and H3K27ac ChIP-seq signal intensity in control (grey), CBS OE (green) and CTH OE mES cells (purple) at ATAC-seq peaks exhibiting increased (gain) chromatin accessibility in Lmna−/− mES cells versus control mES cells. n = 2 biological replicates. e,f Normalized H3K9ac and H3K27ac ChIP-seq signal intensity in control (grey) and Lmna−/− ES cells (red) (e) or control (grey), CBS OE (green) and CTH OE mES cells (purple) (f) at genes exhibiting increased chromatin accessibility and upregulation in Lmna−/− ES cells (‘activated genes’), or genes upregulated during cardiomyocyte differentiation (‘primed genes’)33. n = 2 biological replicates. g,h, Normalized H3K9ac ChIP-seq signal intensity in Lmna−/− ES cells overexpressing control shRNA or shRNA against Cth or Cbs at gained upon Lmna LOF peaks (g) or at genes either activated or primed following Lmna loss (h). n = 2 biological replicates. i, IGV tracks showing H3K9ac ChIP-seq, H3K27ac ChIP-seq, ATAC-seq, lamin A DamID, lamin B1 DamID and significant genomic interactions at H3K9ac and H3K27ac peaks determined by Hi-C in control (grey) and Lmna−/− (red) mES cells at non-LAD and LA/LB1 LAD genomic regions. j, Quantification of interaction frequency in LA/LB1 LADs and outside LADs (non-LADs) at H3K9ac ChIP-seq, H3K27ac ChIP-seq and ATAC-seq peaks with significantly gained or lost signal in Lmna−/− mES cells compared with control. n = 3 biological replicates. Boxplots show median (centre line), 25th–75th percentiles (box) and 1.5 × IQR (whiskers). NS, not significant. For NGS metaplots in b,d–h, lines indicate mean signal and shaded regions represent ±s.e.m. across biological replicates. RPM, reads per million mapped reads.
Genes that became transcriptionally activated upon lamin A/C loss exhibited increased H3K9ac and H3K27ac at their promoters and across gene bodies. In contrast, primed genes, those displaying increased chromatin accessibility without corresponding transcriptional upregulation, showed a more selective enrichment, with elevated H3K9ac confined to promoter regions (Fig. 4e). Notably, overexpression of CTH and CBS led to increased H3K9ac and H3K27ac specifically at transcriptionally activated genes, but not at primed genes (Fig. 4f). Moreover, silencing of Cth or Cbs reduced the elevated H3K9ac levels at gained ATAC-seq peaks in Lmna−/− mES cells (Fig. 4g). However, only Cth silencing led to a marked reduction in H3K9ac enrichment at primed and transcriptionally activated genes following lamin A/C loss (Fig. 4h). These findings suggest that elevated cysteine-derived acetyl-CoA levels promote histone acetylation in a transcription-coupled manner, but that lamin A/C loss is required to create a permissive chromatin environment.
We next examined whether the increased histone acetylation induced by lamin A/C leads to alterations in 3D chromatin architecture. We found a significant increase in Hi-C interactions at H3K27ac, H3K9ac or ATAC peaks with significantly elevated signal at non-LAD regions (Fig. 4i,j and Supplementary Fig. 6c). Conversely, decreased interaction frequencies were observed only at H3K27ac or H3K9ac peaks with reduced signal, but not at ATAC peaks, also restricted to non-LAD regions. These findings suggest that cysteine-dependent histone acetylation upon lamin A/C loss results in substantial modifications to the 3D genome architecture, beyond LADs.
Next, we assessed whether the inhibition of CTH or CBS could rescue the abnormal gene expression pattern in Lmna−/− mES cells. RNA-seq analyses revealed a major overlap between genes upregulated in Lmna−/− mES cells and those downregulated following Cth or Cbs depletion (52,8% and 61,6% overlap, respectively) (Extended Data Fig. 6a and Supplementary Table 2). This pattern was apparent in genes located in LADs as well as those in non-LAD regions (Extended Data Fig. 6b and Supplementary Fig. 6d). Conversely, the overexpression of CTH or CBS in wild-type mES cells led to a significant increase in the expression of both LAD and non-LAD genes upregulated in Lmna−/− mES cells (Extended Data Fig. 6c,d and Supplementary Fig. 6e). Overexpression of CTH led to upregulation of CBS, and conversely, CBS overexpression increased CTH expression (Supplementary Fig. 6f). There was a striking overlap in the genes upregulated in Lmna−/− mES cells and those genes upregulated following CTH overexpression (59.6%) or those upregulated upon CBS overexpression (41.4%) (Extended Data Fig. 6c and Supplementary Table 2). Fitting with the link to laminopathies and the clinical phenotype of patients carrying LMNA mutations, KEGG pathway analysis of genes upregulated upon CTH or CBS overexpression revealed enrichment in pathways related to hypertrophic and dilated cardiomyopathy, calcium signalling pathway and cardiac muscle contraction (Supplementary Fig. 6g). The associated biological processes included DNA damage response and repair, calcium ion transport, cell differentiation and platelet-derived growth factor signalling (Extended Data Fig. 6e). These terms were similar to those associated with Lmna deletion.
In line with an importance of cysteine metabolism also in progeroid cells, RNA-seq analyses revealed a major overlap between genes downregulated in p.G609G mES cells and those upregulated upon CTH or CBS overexpression (Extended Data Fig. 6f and Supplementary Table 3), while KEGG pathway analysis of the overlapping genes identified cysteine and methionine metabolism, base excision repair, cell cycle, Huntington disease and diabetic cardiomyopathy to be overrepresented (Supplementary Fig. 6h). The associated biological processes included calcium ion transport, cell differentiation and signalling pathways (Extended Data Fig. 6g). Notably, overexpression of CTH and CBS rescued the abnormal gene expression of both genes located in LADs (Dchs2 and Mycbp2) as well as those in non-LAD regions (Bmp4, Klf4, Klf5 and Smarcd3) (Extended Data Fig. 6h), further supporting the important role of CTH and CBS in the transcriptional alterations observed in progeroid cells.
Non-physiological levels of CTH and CBS result in abnormal cell fate
Lamin A/C loss results in abnormal cell fate choices during development and promote cardiomyocyte cell fate specification and differentiation18,33 (Fig. 5a). In line with this, overexpression of CTH or CBS promoted cardiac mesoderm commitment (Fig. 5b) as well as cardiomyocyte differentiation (Fig. 5c). In contrast, cysteine depletion impaired cardiac mesoderm formation, cardiac progenitor marker expression and cardiomyocyte differentiation (Fig. 5d and Extended Data Fig. 7a,b), leading to a reduced number of cardiomyocytes (Fig. 5e and Extended Data Fig. 7c). Furthermore, Cth silencing was sufficient to suppress the enhanced cardiomyocyte specification and differentiation potential observed in Lmna−/− mES cells, whereas Cbs silencing resulted in a less pronounced rescue (Fig. 5f and Extended Data Fig. 7d). Similarly, treatment with the CTH and CBS inhibitor AOAA mitigated the elevated cardiomyocyte differentiation of Lmna-deficient cells, suggesting that defects in cell fate depend on the catalytic activity of CTH and CBS (Fig. 5g and Extended Data Fig. 7e).
a, Cell fate alterations upon lamin A/C loss. b,c, qPCR analysis of cardiac mesoderm marker genes in D4 EBs (b) or CM marker genes in D10 EBs (c) differentiated from mES cells transiently transfected with control, CBS or CTH construct before differentiation. n = 3 biological replicates. d,e, qPCR analysis of cardiac mesoderm marker genes in D4 EBs (d) and percentage of troponin T+ CMs determined by FACS analysis of D10 EBs (e) differentiated from mES cells cultured in medium containing the indicated amounts of cysteine. n = 3 biological replicates. f,g, RT–qPCR analyses for CM genes in D10 EBs from control and Lmna−/− following knockdown with control, Cth, or Cbs shRNA (f) or of control and Lmna−/− mES cells non-treated or treated with 100 µM AOAA for 48 h before differentiation (g). n = 3 biological replicates. h, t-Stochastic neighbour embedding (t-SNE) plot of scRNA-seq in D10 EBs from control, Lmna−/− and Lmna−/− mES cells after Cth knockdown. CM, cardiomyocyte; EC, endothelial cell; Ep, epithelial cell; Ery, erythrocyte; Hep, hepatocyte; NPC, neuron progenitor cell; SMC, smooth muscle cell; LBM, limb bud mesenchyme; Mono/MΦ, monocytes/macrophages. i, Percentage of cells in populations shown in h. j, Schematic representation of the Lmna p.G609G mES cells phenotype. k,l, Relative germ layer marker expression in D4 EBs (k) and CM genes in D10 EBs (l) from mES cells after silencing with control shRNA or shRNA for Cbs and Cth. n = 3 biological replicates. m,n, Relative germ layer marker expression in D4 EBs (m) or CM markers in D10 EBs (n) from control, LmnaG609G/G609G mES cells or LmnaG609G/G609G mES cells overexpressing CTH or CBS. n = 3 biological replicates. o, Immunostaining for α-actinin (red) and DAPI (blue) in D10 EBs. p, Immunoblot analysis of CTH in whole-protein extracts of control and LmnaG609G/G609G mES cells cultured with the indicated cysteine concentrations for 48 h. α-tub, α-tubulin. q,r, Relative CM marker expression (q) and α-actinin (red)/DAPI (blue) immunostaining (r) in D10 EBs differentiated from control and LmnaG609G/G609G mES cells cultured in the indicated cysteine concentrations. n = 3 biological replicates. Differences in k and l were assessed using an two-tailed unpaired Student’s t-test; in b–g,m,n,q using one-way ANOVA with Tukey correction; in i were assessed using two-sided pairwise proportion tests. Data are presented as mean ± s.d. NS, not significant.
Next, we examined whether upregulation of CTH and subsequent cysteine-dependent histone acetylation impact cell fate choices upon lamin A/C LOF at a global level. To this end, single-cell RNA sequencing (scRNA-seq) of day 10 EBs differentiated from control, Lmna-deficient and Cth-Lmna-double deficient mES cells was performed, as Cth silencing resulted in a more effective rescue than Cbs. A total of 10,393 cells were obtained for analysis and clustering on the basis of known cell type-specific markers identified 15 major cell types (Fig. 5h and Extended Data Fig. 7f). Lamin A/C deficiency resulted in changes in the number of smooth muscle cells, epithelial cells and hepatocytes, erythrocytes, cardiomyocyte and neuronal progenitor cells (Fig. 5i). These changes were either partially (epithelial cells, erythrocytes) or completely rescued (cardiomyocytes, smooth muscle cells, hepatocytes and neuronal progenitor cells) by Cth depletion (Fig. 5i). Of note, not only cell number but also transcriptional alterations upon lamin A/C loss were rescued by Cth depletion (Extended Data Fig. 7g).
Unlike the effects of Lmna deletion or LOF, the Lmna gain-of-function leads to defects in germ layer specification33 (Fig. 5j). To investigate the potential contribution of decreased CTH and CBS expression, and decreased cysteine metabolism to germ layer formation and differentiation defects of Lmna p.G609G mES cells, we first employed an shRNA approach to knock down Cth and Cbs in mES cells. This resulted in a major decrease, particularly of mesodermal and endodermal marker genes, mirroring the pattern observed in Lmna p.G609G mutant cells (Fig. 5k and Extended Data Fig. 7h). Furthermore, cardiomyocyte differentiation was compromised, resembling the phenotype observed in progeroid cells (Fig. 5l and Extended Data Fig. 7i). As such, downregulating cysteine metabolism reproduced the phenotype of the Lmna p.G609G mutation in mES cells. Notably, the overexpression of CTH or CBS increased the expression of mesoderm, endoderm and later cardiomyocyte marker genes (Fig. 5m,n), and rescued the cardiomyocyte differentiation defects of Lmna G609G mES cells (Fig. 5o). Next, we tested whether addition of cysteine might be beneficial in progeria. Increasing cysteine concentration in the medium to 400 μmol l−1 partially rescued the decreased histone acetylation (Fig. 3h) and CTH levels (Fig. 5p) as well as the cardiomyocyte differentiation defects, as evidenced by the elevated expression of cardiomyocyte marker genes (Fig. 5q,r).
Silencing of Cth and Cbs reverses the Lmna-related cardiomyopathy phenotype, restoring normal cardiomyocyte differentiation and function
Lamin A/C loss results in premature cardiomyocyte differentiation, cell cycle withdrawal, abnormal contractility33 and increase the sensitivity to mechanical and oxidative stress58,59. Therefore, we next examined the impact of Cth and Cbs silencing on the Lmna-associated cardiomyopathy phenotype. Knockdown of either gene prevented premature binucleation and improved cardiomyocyte rhythmicity in Lmna-deficient cells (Fig. 6a–d). Moreover, it significantly reduced cardiomyocyte apoptosis (Fig. 6e,f). Given that increased apoptosis has been linked to the mechanoprotective actions of lamin A/C, we subjected mES cell to mechanical stretch. Lmna-deficient cells showed marked accumulation of γ-H2AX, a sensitive marker of DNA double-strand breaks, upon stretch, which was significantly reduced following silencing of either Cth or Cbs and further exacerbated by their overexpression (Fig. 6g,h).
a,b Immunostaining with α-actinin and DAPI (a) and quantification of the percentage of binucleated CM (b) in FACS-sorted Nkx2.5+ cells from D14 EBs. n = 3 independent experiments. Arrows in a indicate binucleated cardiomyocytes. c,d Representative contraction traces in spontaneously beating CMs at D10 extracted from image sequences using MUSCLEMOTION V1.0 (c) and quantification of proarrhythmic beating (d). e,f Representative FACS analysis of D10 CMs stained with Annexin V-APC and 7-AAD (e) and quantification of apoptotic and necrotic cells (f). n = 3 independent experiments. g,h Immunostaining for γH2AX (red) and DAPI (blue) in mES cells subjected to 15% mechanical stretch for 30 min (g), and quantification of the percentage of γH2AX-positive mES cells (h). n = 6 biological replicates. i, Dot plots showing expression of genes related to DCM, apoptosis and proliferation in the CM cluster from scRNA-seq datasets of D10 EBs from control, Lmna−/− and Lmna−/− mES cells after shRNA mediated silencing of Cth. Data are presented as mean ± s.d. Differences in b,f,h were assessed using one-way ANOVA with Tukey correction. Differences in d were assessed with two-sided Fisher’s exact test.
Single-cell transcriptomic analysis further revealed that Cth silencing suppressed the upregulation of dilated cardiomyopathy (DCM)-associated genes such as Ryr2, Myh7, Ldb3 and Tnni3. Additionally, pro-apoptotic gene expression elevated upon Lmna loss was diminished, classical proliferation markers were restored, and cell cycle arrest signatures were normalized (Fig. 6i).
Together, these findings demonstrate that Cth and Cbs silencing rescues the cellular abnormalities observed in Lmna-deficient cardiomyocytes.
CBS levels regulate the balance between histone H3K9 methylation and acetylation in naive pluripotent stem cells
DNA damage repair defects, along with genomic instability attributed to elevated levels of SUV39H1 and H3K9me3, have been demonstrated to contribute to the premature ageing phenotype associated with Lmna p.G609G mutation29,30,48. The methyl group donor S-adenosylmethione (SAM), which is essential for histone methylation8, is generated by the methionine cycle, which is tightly intertwined with cysteine metabolism (Fig. 7a). Indeed, homocysteine can either be converted into ʟ-cystathionine by CBS or recycled into methionine by the 5-methyltetrahydrofolate-homocysteine methyltransferase (MTR). Thus, the loss of CBS in LmnaG609G/G609G mES cells might increase the flux of homocysteine to the methionine cycle, altering histone methylation. We therefore examined whether changes in CBS activity may alter SAM levels and affect the balance between histone methylation and acetylation. Indeed, overexpression of CBS in mES cells decreased SAM levels and SAM:S-adenosylhomocysteine (SAH) ratio, whereas silencing of Cbs resulted in a significant increase in SAM and the SAM:SAH ratio, supporting the hypothesis that CBS reroute metabolic fluxes in mES cells (Fig. 7b and Extended Data Fig. 8a,b). Notably, the elevated CBS levels in Lmna−/− mES cells came along with lower SAM and SAM:SAH ratio, which were restored to control levels upon Cbs silencing (Fig. 7b and Extended Data Fig. 8a). Conversely, progeria cells exhibited notably higher SAM concentrations, which were normalized by CBS overexpression. Furthermore, in LmnaG609G/G609G mES cells, we observed an increase in H3K9me3, alongside with decreased H3K9ac (Fig. 7c), whereas in Lmna−/− mES cells, H3K9me3 levels were lower (Fig. 7d) and H3K9ac levels were higher (Extended Data Fig. 5d).
a, Schematic representation of the homocysteine degradation pathways and the hypothesis: When CBS is depleted in mES cell, cysteine catabolic flux is perturbed. This leads to homocysteine reflux into the methionine cycle, generating increased levels of SAM. b, SAM (S-adenosylmethionine) levels in control, Lmna−/− and Lmna−/− transfected with control or Cbs shRNA as well as control or LmnaG609G/G609G mES cells transfected with control or CBS OE plasmid. n = 4 biological replicates. c, Representative immunoblot analysis of H3K9ac and H3K9me3 in control, LmnaG609G/+ and LmnaG609G/G609G mES cells. d, Representative immunoblot analysis of H3K9me3 in control and Lmna−/− mES cells. e, Representative immunoblot analysis of SUV39H1 levels in chromatin-bound fractions (CBFs) and soluble nuclear extracts (SNEs) of control, CBS OE and CTH OE mES cells. f, Representative immunoblot for SUV39H1 in CBF and H3K9me3 in histone extracts of control and Lmna−/− mES cells transfected with control, Cbs and Cth shRNA. g, Representative immunoblot analysis for SUV39H1 levels in CBF and SNE of control and siRNA mediated Cbs/Cth knockdown mES cells. h, Representative immunoblot analysis of SUV39H1 in CBF of control, LmnaG609G/+ and LmnaG609G/G609G mES cells. i, Representative immunoblot analysis for SUV39H1 in CBF and H3K9me3 in histone extracts of control and LmnaG609G/G609G mES cells transfected with control, CBS or CTH plasmid. j, Representative immunoblot analysis for SUV39H1 in CBF and H3K9me3 in histone extracts of control and Lmna−/− mES cells either not treated or treated with 50 µmol l−1 SAM for 6 h. k, Genome tracks of H3K9me3 ChIP-seq of control (grey), Lmna−/− (red) and LmnaG609G/G609G (blue) mES cells. Differences in b were assessed using one-way ANOVA with Tukey correction.
SUV39H1 and SUV39H2 are the major mammalian histone methyltransferases that catalyse heterochromatic H3K9me3 deposition60,61. Of note, overexpression of CBS, but not CTH, resulted in a decrease in chromatin-bound SUV39H1 (Fig. 7e), consistent with the pivotal role of CBS in metabolic flux. Furthermore, in line with the increased CBS levels following lamin A/C LOF, immunoblotting revealed a reduction of chromatin-bound SUV39H1 in Lmna−/− mES cells compared with control cells, a change reversible by Cbs silencing but not Cth (Fig. 7f). Conversely, the silencing of both Cbs and Cth in mES cells increased chromatin-bound SUV39H1 (Fig. 7g). A similar increase in chromatin-bound SUV39H1 was observed in LmnaG609G/G609G mES cells, consistent with the decreased levels of CBS in these cells (Fig. 7h). Overexpression of CBS, but not CTH, decreased chromatin-bound SUV39H1 and the H3K9me3 catalysed by it in progeroid cells (Fig. 7i). Notably, the levels of SUV39H1 in total nuclear extracts did not differ in either Cth/Cbs double knockdown mES cells (Fig. 7g) or in CBS-overexpressing mES cells (Fig. 7e), suggesting a role for CBS or CBS-regulated SAM in SUV39H1 complex recruitment. To test this, we treated control and Lmna−/− mES cells with SAM, which led to an increase in chromatin-bound SUV39H1 and a corresponding increase of H3K9me3 in both cell lines (Fig. 7j), whereas the levels in the soluble nuclear extract remained unchanged (Extended Data Fig. 8c). Moreover, H3K9me3 ChIP-seq analysis confirmed the decreased levels of H3K9me3 in Lmna−/− mES cells and increased levels in LmnaG609G/G609G mES cells (Fig. 7k).
In summary, our data indicate that CBS-dependent rerouting of cysteine catabolic flux, coupled with rewiring of methionine metabolism and subsequent changes in SAM levels, lead to altered SUV39H1 recruitment to chromatin and subsequent H3K9me3 modifications.
Impact of CBS-dependent rerouting of cysteine catabolic flux on DNA damage repair and cellular senescence
Given our observation that CBS levels play a critical role in regulating metabolic fluxes within the methionine cycle and cysteine metabolism pathways, we next investigated whether re-expression of CBS could potentially restore DNA repair capacity in Lmna p.G609G mES cells. As high levels of CBS induce cellular senescence, we carefully titrated the lentiviruses used to reinstate CBS but not to overexpress it relative to control levels (Extended Data Fig. 8d). This approach proved to be protective, resulting in a significant reduction in DNA damage, as evidenced by staining for γ-H2AX (Fig. 8a). Additionally, it rescued the DNA damage repair defects (Fig. 8b and Extended Data Fig. 8e). In line with this, reducing methionine concentration in the medium to 10–25 μM also led to a notable reduction in H3K9me3 and γ-H2AX (Fig. 8c). Furthermore, CBS re-expression to endogenous levels attenuated SA-β-gal activity, a gold standard of senescence, in Lmna p.G609G mES cells (Fig. 8d,e and Extended Data Fig. 8f). Progeria mES cells also displayed elevated MitoSOX signal, indicative of increased mitochondrial oxidative stress, which was also normalized by CBS re-expression (Fig. 8f). Moreover, many of the genes upregulated in Lmna p.G609G mES cells are signature factors of the senescence-associated secretory phenotype (SASP) (Fig. 8g). Notably, expression of these genes was restored to baseline levels upon re-expression of CBS to physiological levels (Fig. 8h, Extended Data Fig. 8g and Supplementary Table 4). These findings further highlight the critical role of CBS in modulating key hallmarks of the progeroid phenotype.
a, Immunostaining for γ-H2AX (green), CBS (magenta) and DAPI (blue) in control, LmnaG609G/G609G and CBS OE LmnaG609G/G609G mES cells treated with H2O2 for 1 h and allowed to repair in fresh medium for 4 h. b, Quantification of γ-H2AX-positive mES cells after 0 h, 1 h and 4 h of repair. n = 3 biological replicates. c, Immunoblot analysis of γ-H2AX and H3K9me3 in control and LmnaG609G/G609G mES cells treated with H2O2 for 1 h and cultured with the indicated amounts of methionine for 24 h. d,e, SA β-gal staining (d) and percentage of SPIDER SA β-gal+ determined by FACS analysis (e) of control, LmnaG609G/G609G and CBS OE-LmnaG609G/G609G mES cells either not treated (w/o) or treated with H2O2 for 2 h and cultured in fresh medium for 24 h. n = 3 biological replicates. f, FACS analysis (left) and mean MitoSOX Green fluorescence intensity (right) in control, LmnaG609G/G609G, and CBS OE–LmnaG609G/G609G mES cells. n = 3 biological replicates. g, GO analysis of genes upregulated (Padjusted < 0.05) in LmnaG609G/G609G compared with control mES cells and downregulated in CBS OE-LmnaG609G/G609G. Significance was calculated in DAVID using a one-sided Fisher’s exact test. h, Heatmap representation of RNA-seq analysis showing expression of p16, p21 and SASP markers. n = 2 biological replicates. i, qPCR analysis of Cbs, Cth, progerin and total Lmna expression in the hearts of young (3-month-old) and aged (24-month-old) mice. n = 7 mice per group. j,k, Immunostaining of heart sections for lamin A/C (green) (j) or CBS (green) (k) together with cardiac troponin I (magenta) and DAPI (blue) in young (3-month-old) and aged (24-month-old) mice. n = 7 mice per group. l, Relative H3K9me3 levels in young (3-month-old) and aged (24-month-old) hearts normalized to H3 control. n = 7 mice per group. m, SAM-to-SAH ratio in livers of Lmna+/+ and LmnaG609G/G609G mice fed either a control diet or a methionine-restricted (MR) diet60. n,o, RNA-seq analysis of CBS and CTH expression in fibroblasts from young (<30 years) and aged (>70 years) individuals (m) or patients with progeria (GSE113957) (o). FPKM, fragments per kilobase of transcript per million mapped fragments. Differences were assessed using an two-tailed unpaired Student’s t-test (i,l,n,o) and a one-way ANOVA with Tukey correction (b,e,f,m). Data are presented as mean ± s.d. NS, not significant.
Next, we investigated whether similar mechanisms could be relevant during physiological ageing and in progeria patients. Notably, hearts from aged mice exhibited increased expression of progerin transcripts, whereas global Lmna levels remained unchanged (Fig. 8i). In addition, aged cardiomyocytes showed accumulation of lamin A/C at the nuclear periphery and reduced nuclear size (Fig. 8j and Extended Data Fig. 8h). In parallel, we detected a significant reduction in Cbs levels (Fig. 8i,k), increased H3K9me3 (Fig. 8l and Extended Data Fig. 8i) and upregulation of SASP genes (Extended Data Fig. 8j). Of note, analysis of metabolomics data from previous studies demonstrating that methionine restriction extends lifespan in progeroid mice62 revealed that Lmna p.G609G mice display elevated SAM:SAH ratio, which can be reduced by methionine restriction (Fig. 8m). Further, CBS mRNA levels were markedly decreased in old compared with young individuals (Fig. 8n), and progeria patients also displayed reduced CBS expression (Fig. 8o). These findings suggest that accumulation of progerin in aged cardiomyocytes as well as in patients with progeria may disrupt the balance between cysteine metabolism and methionine cycle through direct regulation of CBS levels, thereby contributing to premature ageing.
Taken together, our work uncovered a critical role of the nuclear lamin A/C in regulating metabolic flux within the 1C metabolism to maintain proper cellular fate and function and prevent cellular senescence and ageing (Extended Data Fig. 9).
Discussion
Mounting evidence indicates the crucial role of nuclear lamins in anchoring lineage specific genes to the nuclear periphery and ensuring precise lineage specification and tissue morphogenesis. Here, we uncovered nuclear lamina-mediated epigenetic control beyond LADs, which, exerted by lamin A/C-regulated cysteine flux into acetyl-CoA and histone acetylation is crucial for both proper stem cell fate and differentiation as well as longevity. Lamin A/C is high in naive pluripotent stem cells, where it suppresses metabolic and differentiation-associated genes (Extended Data Fig. 9a). During the transition to the primed state, RA signalling induces a decline in lamin A/C levels, leading to reduced PRC2-mediated H3K27me3 at the promoters of the key cysteine-generating and -metabolizing enzymes Cth and Cbs, thereby enabling SP1 binding and their transcriptional activation. Enhanced de novo cysteine synthesis fosters histone H3 acetylation specifically at lysine 9 and lysine 27, which triggers epigenome remodelling that is critical to support subsequent differentiation and germ layer formation (Extended Data Fig. 9b). Notably, in Lmna−/− mES cells, acetyl groups derived from cysteine were predominantly incorporated into H3K9, H3K27 and H4K12, whereas no such incorporation was detected in wild-type cells. Deciphering how lamin A/C decline during developmental transitions confers specificity toward these residues will be an important direction for future studies. It is noteworthy in this context that the majority of genes associated with the nuclear lamina are transcriptionally inactive and marked by repressive histone modifications, such as H3K27me3 and H3K9me2/3 (refs. 23,63). Thus, decreased SAM levels and increase in acetyl groups’ bioavailability, together with dissociation from the nuclear lamina, might account for the specificity. Numerous studies have highlighted the significance of H3K27me3 and H3K9me2/3 in repressing critical developmental genes and regulating cell fate decisions during embryonic development60. Thus, it is unsurprising that alterations in the balance between methylation and acetylation at these residues, brought about by changes in cysteine catabolic flux, lead to abnormal cell fate and differentiation. Yet, the precise contribution of cysteine and methionine metabolism to epigenetic reprogramming remains a question. Notably, the spatial conversion of cysteine to pyruvate has not been thoroughly investigated. Nuclear acetyl-CoA, a key player in histone acetylation, can be generated by enzymes such as acetyl-CoA synthetase (ACSS) and ATP-citrate lyase, both present in the cytosol and the nucleus. Additionally, recent studies revealed that the mitochondrial pyruvate dehydrogenase complex and TCA cycle-related enzymes translocate into the nucleus where they play important roles in epigenetic reprogramming during zygotic genome activation64 and in primed-to-naive transitions65. Given the importance of nuclear lamins in zygotic genome activation66 and naive pluripotency33, it would be important to study the subcellular localization and function of these metabolic enzymes during lineage transitions. It is also noteworthy to mention that elevated cysteine bioavailability could potentially increase cellular acetyl-CoA levels. This increased acetyl-CoA could freely diffuse through the nuclear envelope67 and target histone residues, potentially influencing histone acetylation, chromatin structure and gene regulation.
In contrast to Lmna−/− mES cells, the dominant, gain-of-function mutation Lmna p.G609G, associated with HGPS and premature ageing, resulted in decreased cysteine availability and catabolic flux. We observed that the rerouting of cysteine catabolic flux, resulting from decreased CBS levels, increased SUV39H1 recruitment to chromatin and raised H3K9me3 levels in progeroid cells. Elevated SUV39H1 and H3K9me3 levels have also been detected in mice lacking the prelamin A-processing metalloprotease, Zmpste24, which largely phenocopy the features of HGPS29,30. Previous studies have shown that depletion of Suv39h1 led to a reduction in H3K9me3 levels, re-established DNA repair ability, decreased mitochondrial ROS production, downregulated SASP gene expression and delayed senescence in progeroid cells29,30. Similarly, in our study re-expression of CBS in Lmna p.G609G mES cells back to control levels restored the DNA repair capacity, mitigated mitochondrial ROS production, reduced SASP genes expression and prevented senescence of progeroid Lmna p.G609G mES cells. This protective effect is likely mediated through its role in limiting SAM availability, thereby reducing SUV39H1 recruitment to chromatin and attenuating SUV39H1-dependent H3K9me3 deposition. In addition, methionine metabolism and thereby SAM availability act as critical regulators of epigenetic remodelling and gene expression during developmental transitions. In human induced pluripotent stem cells, high methionine demand supports elevated levels of histone methylation, including H3K9me3, which helps maintain pluripotency. Depletion of methionine reduces H3K9me3 and promotes mesoderm differentiation8,68. However, compared with differentiated cells, embryonic stem cells contain relatively low levels of H3K9me3-marked heterochromatin, and the progressive accumulation of heterochromatin is thought to contribute to the establishment of cell identity69. Disruption of H3K9me3 through genetic inactivation of its methyltransferases led to de-repression of developmental genes and impaired differentiation69, indicating that stringent regulation of H3K9me3 is crucial for proper cell fate determination. How increased SUV39H1 binding to chromatin and elevated H3K9me3 levels influence cell fate and function in response to changes in lamin A/C levels remains to be further investigated. Notably, overexpression of CTH alone was sufficient to rescue the cell fate and differentiation defects of Lmna p.G609G mES cells, underscoring the central role of cysteine metabolism and CTH-mediated histone acetylation in modulating developmental potential. However, as CTH overexpression also induced CBS upregulation, and vice versa, disentangling the respective contributions of histone methylation and acetylation is challenging. Notably, depletion of both CTH and CBS markedly impaired mesoderm specification and the differentiation of mesodermal derivatives, resembling the defects observed in HGPS, which predominantly affects mesoderm- and mesenchyme-derived lineages42.
Additionally, we observed a significant increase in progerin levels and accumulation of lamin A/C at the nuclear periphery in aged cardiomyocytes, consistent with previous observations in aged fibroblasts70. However, unlike the widely reported global loss of H3K9me3 in aged fibroblasts and mesenchymal stem cells70,71,72, we observed elevated H3K9me3 levels in aged cardiomyocytes. This finding aligns with recent reports showing concomitant increases in H3K9me3 and H3K27me3 levels73 as well as reduced chromatin accessibility in aged cardiomyocytes74. This could be due to the limited CBS levels resulting in elevated SAM availability. Notably, progeroid mice display elevated SAM:SAH ratios, which can be normalized by methionine restriction62, a dietary intervention that extends their lifespan. Intriguingly, dietary restriction more broadly has been shown to confer benefits such as enhanced longevity and stress resistance75. These observations highlight the need to further investigate how alterations in the nuclear lamina intersect with the activity of cysteine-generating and catabolizing enzymes, and to explore their potential roles in promoting longevity and systemic health.
Finally, as we found abnormal cysteine synthesis and catabolism in stem cells carrying LMNA mutations, we suggest that inhibiting or activating this pathway could serve as a new therapeutic strategy in laminopathies. Considering the substantial evidence from epidemiological research and animal studies indicating the profound impact of nutrition before and during pregnancy on the health and overall welfare of offspring76, dietary interventions may hold promise in the prevention of these diseases. On a more general note, our study emphasizes that the reversibility of epigenetic modifications through dietary intervention could counteract perturbations in genes encoding epigenetic modifiers and highlights the need for further studies at the interface of epigenetic alterations and metabolism in development, homeostasis and healthy ageing.
While we have shown that abnormal cysteine synthesis and catabolism during early embryogenesis alters cell fate, function and longevity, and could represent a new therapeutic strategy in laminopathies, several limitations should be considered. Modulation of the transsulfuration pathway impacts the methionine–SAM methylation axis and the folate 1C cycle, both of which are tightly interconnected and essential for proper embryonic development. Given that pregnancy represents a period of highly sensitive epigenetic programming, direct pharmacological targeting of CTH and CBS during gestation carries substantial uncertainty and risk. Safer translational strategies are likely to involve nutritional modulation and selective, time-limited interventions. Notably, similar changes in CTH and CBS were also found in differentiated cells and animal models, indicating that this pathway remains responsive beyond early development and warrants evaluation of its therapeutic potential in adulthood.
Methods
Mouse lines
The Lmna tm1.1Yxz/J line77 was obtained from The Jackson Laboratory and was maintained on a C57BL/6J background. The Lmna tm1.1Yxz/J and C57BL/6J mice were housed in a pathogen-free animal facility under standard conditions with a 12 h light–dark cycle, temperature of 20–25 °C and humidity range of 30–70%. In the ageing studies, C57BL/6J mice aged 3 months and 24 months were designated as young and aged groups, respectively. Embryos were collected at E3.5 and E6.5 from plug-positive mice. Both male and female mice were included in all experiments. All animal experiments were performed according to the regulations issued by the Committee for Animal Rights Protection of the State of Baden-Württemberg (Regierungspraesidium Karlsruhe, Experimental protocol Az. I-25/09 and 35-9185.81/G-17/24).
Mouse ES cell line culture and differentiation
Murine male E14-NKX2-5-EmGFP generated by Hsiao et al.78 and male R1 (ATCC, SCRC-1011) mES cells were cultured and differentiated as described previously7,33. In brief, mES cells were maintained on mitomycin C-treated mouse embryonic fibroblasts in high-glucose DMEM (Thermo Fisher Scientific) supplemented with 15% FBS and 1,000 U ml−1 leukemia inhibitory factor (LIF) (ESGRO, Millipore). For non-directed differentiation, mES cells were dissociated with 0.05% trypsin–EDTA to single cells and EBs were formed in ES cell growth medium without LIF in 15-μl hanging drops (3.3 × 104 cells per ml) for 2 days. EBs were then transferred to low-attachment dishes and collected on day 3–4 (germ layer), day 5–6 (progenitor) and day 8–10 (cardiomyocyte) for analysis.
Naive mES cells were cultured on feeders in either high-glucose DMEM with 15% FBS and 1,000 U ml−1 LIF or knockout DMEM (Thermo Fisher Scientific) with 10% serum replacement and 1,000 U ml−1 LIF. Primed mES cells were generated by culturing naive cells on 0.1% gelatin-coated plates in DMEM/F12 with 20% knockout serum replacement, 10 ng ml−1 Activin A, and 10 ng ml−1 bFGF or in Neurobasal:DMEM/F12 (1:1) supplemented with 2,000 U ml−1 LIF and 10 ng ml−1 BMP4 for 2 days. For methionine or cysteine restriction, mES cells were cultured in high-glucose DMEM lacking methionine and cystine (Thermo Fisher Scientific) supplemented with 15% FBS, 1,000 U ml−1 LIF and indicated amounts of ʟ-cysteine (Sigma-Aldrich, 168149) or ʟ-methionine (Sigma-Aldrich, M5308).
Generation of Lmna −/− and Lmna p.G609G mES cell lines
The generation of Lmna−/−, LmnaG609G/+ and LmnaG609G/G609G murine E14-NKX2-5-EmGFP ES cells has been described elsewhere33. A similar strategy was used for CRISPR/Cas9-mediated gene editing in R1 mES cells. In brief, two gRNAs were employed: Lmna gRNA-1: 5′-CACCGCACTGCTCACGTTCCACCAC-3′, and Lmna gRNA-2: 5′-CACCGAGCTATCAGCACTCTGTTAT-3′. Annealed gRNAs were cloned into PX459 V2.0 (Addgene #62988) and transfected using Lipofectamine 2000. Cells were selected with puromycin (4 µg ml−1) for 48 h, plated on feeders and single colonies were screened by PCR and Sanger sequencing.
Generation of stable knockdown and overexpressing mES cell lines
Female HEK293T cells (ATCC CRL-3216) were cultured in DMEM high glucose with GlutaMAX (Gibco, 61965059) supplemented with 10% FBS. At ~70% confluence, cells were transfected with either shRNA plasmids targeting Cbs (ACACTATCATTGAGCCAACTT), Cth (GCTATATTTGTGTCCAAGGAA) or control (pLKO) for gene knockdown or overexpression plasmids for CBS (TRCN0000473237) and CTH (TRCN0000474979), along with 0.975 µg CMV∆R8.74 packaging plasmid and 0.525 µg VSV.G envelope plasmid using X-tremeGENE (Roche, 6366236001). Viral supernatants were collected 48 h post-transfection and 100,000 mES cells were transduced with 2 ml virus in the presence of 8 µg ml−1 Polybrene (Sigma-Aldrich, TR-1003-G) on 5% poly-HEMA (Sigma-Aldrich, P3932)-coated plates for 12 h. Transduced mES cells were plated on feeders the following day and selected with 10 ng ml−1 puromycin for 48 h. For doxycycline-inducible lamin A overexpression, HEK293T cells were transfected with 1.5 µg pLenti CMV TRE3G Neo GFP-lamin A (Addgene #118709) and mES cells were subsequently transduced and selected with 400 µg ml−1 G418 for 48 h.
Transient gene knockdown or overexpression in mES cell
For transient experiments, mES cells were cultured feeder-free in knockout DMEM supplemented with 10% serum replacement and 2,000 U ml−1 LIF. After 24 h, cells were transfected with 50 nM ON-TARGETplus siRNA targeting Cbs, Cth, Slc1a1, Slc1a5 or Sp1 (Horizon Discovery, SMARTpool) or scrambled control using Lipofectamine RNAiMax (Thermo Fisher). For transient overexpression, 2.5 µg CBS or CTH plasmid (TRC OE library) was transfected using Lipofectamine 2000. Cells were used for hanging drop differentiation or other analyses 48 h post-transfection.
Treatment with small molecules
The mES cells were treated for 72 h with 4 µM UNC1999 (Selleck, S7165), 100 nM Chaetocin (Selleck, S8068) or 5 nM Trichostatin A (Selleck, S1045) and then collected for further analysis. Cells carrying the inducible lamin A construct were treated with 1 µg ml−1 doxycycline (Sigma, D1822) to induce expression.
Quantification of amino acids and biogenic amines
Cells were lysed in ice-cold Triton X-100 buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 20 mM NaF, 10 mM Na4P2O7and 1% Triton X-100) supplemented with protease and phosphatase inhibitors. Lysates were sonicated (10 s, 80% amplitude, Sonoplus mini20) and centrifuged (16,000g, 10 min, 4 °C). Protein concentrations were determined by Bradford assay, and 250 µg protein per sample was used for amino acid profiling. Samples were prepared using the EZ:faast LC–MS free amino acid analysis kit (Phenomenex) with minor modifications; internal standards (10 µl) were added, and sample pH was adjusted to 1.5–6.0. Metabolites were analysed by LC–MS/MS on an Agilent 1290 Infinity LC system with EZ:faast AAA-MS HPLC column coupled to a QTrap 5500 (Sciex) using positive electrospray ionization. Intensities were normalized to internal standards, and data were processed using Analyst v.1.6.2 and MultiQuant v.3.0.
Quantification of tricarboxylic acid cycle metabolites
Cells were lysed as above and 100 µg protein per sample was mixed with 5× methanol and 2.5× isotope-labelled internal standard (water:methanol, 50:50). Samples were evaporated in a vacuum concentrator (30 °C), reconstituted in 50 µl water and analysed by LC–MS/MS. Conditioned medium was mixed 1:1 with internal standard before analysis. Reversed-phase LC separation was performed using a Waters Acquity UPLC HSS T3 column (2.1 × 150 mm, 1.8 µm) at 40 °C with a 10-min gradient (0.15% formic acid in water:acetonitrile), flow rate of 400 µl min−1and injection volume of 2.5 µl. Negative-ion ESI was employed; MRM transitions were normalized to internal standards and calibration curves were generated using authentic standards. Data were analysed using MetaboAnalyst v.5.0.
Metabolite profiling for cysteine flux analyses
Control and Lmna−/− mES cells were cultured feeder-free in knockout DMEM with 10% serum replacement and 2,000 U ml−1 LIF. After 48 h, cells were starved for 1 h in methionine- and cystine-free DMEM/F12 containing 10% serum replacement and 2,000 U ml−1 LIF. Unlabelled ʟ-cysteine or ʟ-cysteine-13C3 (200 µM) was added for 3 h. Cells were washed in cold PBS with 5 µM valproic acid, collected in 85% methanol, sonicated, incubated on ice, and proteins pelleted (12,000g, 10 min, 4 °C). Supernatants were split for TCA flux (800 µl) and amine flux/GSH analysis (200 µl). For amine flux, 700 µl water containing 100 µg ml−1 homotaurine was added; samples were freeze dried, reconstituted in 80 µl boric acid buffer, incubated on ice for 15 min and derivatized with 20 µl AQC reagent (AccQ-Tag, Waters) at 55 °C for 10 min. Samples were centrifuged (12,000g, 10 min, 4 °C) and supernatants were analysed by LC–MS/MS. TCA flux and amine flux analyses were performed on an Agilent 1290 Infinity II Bio LC coupled to a 6495C QQQ. Amine-flux separation used an Agilent ZORBAX Extend RR HD column (2.1 × 150 mm, 1.8 µm) at 30 °C with a 19-min gradient (0.1% formic acid in water/acetonitrile), flow rate of 0.3 ml min−1, injection of 5 µl and positive-ion ESI. TCA flux used an Acquity UPLC HSS T3 column (2.1 × 150 mm, 1.8 µm), 40 °C, 15-min gradient and flow of 0.3 ml min−1. Metabolites and 13C-isotopologues were identified via authentic standards and retention times. Data were processed using Analyst 1.6.2, MultiQuant 3.0 and MetaboAnalyst 5.0.
Quantitative proteomic analysis of histone modifications
Cells were labelled with ʟ-cysteine-13C3 and collected as described in ‘Metabolite profiling for cysteine flux analyses’. Histones were isolated according to the protocol published by Shechter et al.79 and subsequently subjected to acetylomics80 from Creative Proteomics.
ʟ-Cysteine, pyruvate, acetyl-coenzyme A and SAM measurements
Pyruvate levels were determined using a colorimetric pyruvate assay kit (Sigma-Aldrich, MAK332). ʟ-Cysteine and acetyl-CoA were measured with fluorometric assay kits (Sigma-Aldrich, MAK255 and MAK039, respectively). SAM and SAH levels were measured with an SAH ELISA kit (Cell Biolabs, MET-5151) following the manufacturer’s instructions. For nuclear acetyl-CoA, 5 × 106 mES cells were washed with PBS and lysed in 1 ml hypotonic buffer (50 mM Tris-HCl, pH 8.0; 2 mM EDTA, pH 8.0; 0.1% NP-40; and 10% glycerol) on ice for 15 min. Nuclei were pelleted at 960g for 5 min at 4 °C, and resuspended in 100 μl acetyl-CoA assay reagent. Nuclear fractions were sonicated, centrifuged at maximum speed for 15 min at 4 °C and 30 μl of supernatant was used for measurement.
CBS and CTH enzymatic activity assays
The mES cells were dissociated using 0.05% trypsin–EDTA, pre-plated for 1 h and collected by centrifugation. Cells were washed once with cold PBS, lysed in a mild buffer containing 50 mmol l−1 Tris-HCl, pH 8.0, 150 mmol l−1 NaCl, 0.1% v/v NP-40) and treated with 1 mmol l−1 of ʟ-cystathionine for CTH and ʟ-homocysteine for CBS, in the presence of 10 μmol l−1 PLP, following incubation for 30 min at 37 °C. Next, samples were prepared for metabolomics analysis and the respective ʟ-cysteine and ʟ-cystathionine levels were detected through LC–MS/MS.
HDAC enzymatic activity assays
A total of 80,000 mES cells were seeded per well in gelatin-coated 96-well plates and cultured for 2 days in knockout DMEM supplemented with 10% serum replacement and 2,000 U ml−1 LIF. Before HDAC activity measurement, wells were washed with PBS and 100 µl of phenol red-free RPMI was added. For inhibitor control, 1 µl of 300 µM Apicidin (Enzo Life Sciences, BML GR340-0001) was added to designated wells and incubated at 37 °C for 30 min. HDAC class I activity was assessed by adding 5 µl of 1 mM ZLPA substrate (CBZ-Propionyllysine-AMC, GenScript) to each well, followed by incubation at 37 °C for 2 h. Reactions were stopped by adding 50 µl stop solution (PBS, 1.5% Triton X-100, 3 µM Trichostatin A and 0.75 mg ml−1 trypsin) and incubating at 37 °C for 20 min. AMC fluorescence was measured using a PerkinElmer EnSight plate reader (excitation/emission, 360/460 nm) and background signal was subtracted using buffer blanks. Substrate conversion was determined from an AMC standard curve.
Sulfide measurement
The mES cells were dissociated using 0.05% trypsin–EDTA, pre-plated for 1 h, washed with cold PBS and lysed on ice for 1 h in a buffer containing 50 mmol l−1 Tris-HCl (pH 8.0), 150 mmol l−1 NaCl, 1% NP-40 and 1% Triton X-100. Lysates were centrifuged at 20,000g for 10 min at 4 °C, and protein concentration was determined using a BCA assay. Then, 300 µg of protein was incubated with 100 mmol l−1 Tris-HCl (pH 8.0), 50 µM pyridoxal 5′-phosphate hydrate (Sigma-Aldrich, P9255), 10 mmol l−1 ʟ-cysteine, 0.5 mmol l−1 ʟ-homocysteine (Sigma-Aldrich, 69453) and 10 µM fluorescent H2S probe 7-azido-4-methylcoumarin (Sigma-Aldrich, 802409) at 37 °C for 2 h. Fluorescence was measured using a Tecan Infinite 200 Pro plate reader with excitation at 340 nm and emission at 445 nm.
ROS detection
For ROS detection, mES cells were incubated with 1 μM MitoSOX Green (Invitrogen, M36006) for 30 min at 37 °C in a 5% CO2 in HBSS. After three washes with pre-warmed HBSS, cells were analysed by flow cytometry with a GFP channel.
Senescence β-galactosidase staining and SPIDER β-galactosidase assay
To assess senescence, mES cells were either not treated or treated with 200 µM H2O2 for 2 h and cultured in fresh medium for 24 h. mES cells were either directly fixed for senescence β-galactosidase staining using a Senescence β-Galactosidase Staining kit (Cell Signalling Technology, 9860S) or dissociated into single cells by 0.05% trypsin–EDTA and stained for FACS analysis with SPiDER-β-gal reagent (1 μmol l−1, Dojin, SG02-10) for 15 min at 37 °C. After washing with PBS, mES cells were resuspended in 300 µl FACS buffer (0.4% BSA/PBS) and analysed using a BD FACSCanto II (version 8.0.1, firmware version 1.49) flow cytometer with a 488 nm excitation and 638 nm emission wavelength.
Western blotting
Proteins were quantified with Pierce BCA protein assay kit (Pierce Biotechnology, 23225), separated via SDS–PAGE followed by a transfer to a nitrocellulose membrane (Sartorius, 1130641BL). After blocking in 5% skimmed milk/PBST, membranes were incubated with appropriate primary antibodies. The membranes were then washed with PBS/0.1% Tween-20 and incubated with a corresponding secondary antibody for 1 h at room temperature and visualized by Amersham Imager 600. Antibodies used in this study are listed in Supplementary Table 5.
Immunofluorescence staining
For immunostaining of whole-mount embryos, E3.5 and E6.5 embryos were dissected and fixed for 20 min with 4% paraformaldehyde (PFA) in PBS at room temperature. After three washes with 0.1% Tween-20/PBS, each 10 min, the embryos were permeabilized using 0.5% Triton X-100 in PBS for 20 min and then rinsed three times with 0.1% Tween-20/PBS, each 10 min, followed by incubation in blocking buffer 3% BSA/PBS for 2 h. The embryos were further incubated overnight at 4 °C with primary antibodies in 0.5% Triton X-100/ 1% BSA. After three consecutive 10-min washes in 0.1% Tween-20/PBS, embryos were incubated for 3 h with the corresponding secondary antibody in PBS, followed by 4,6-diamidino-2-phenylindole (DAPI) staining.
For immunostaining of cells, cells were fixed with 3.7% paraformaldehyde for 10 min and permeabilized with 0.5% Triton X-100/PBS at room temperature for 10 min, followed by blocking with 3% BSA/PBS for 30 min. The cells were then incubated with the primary antibodies in 0.3% Triton X-100/1% BSA overnight at 4 °C. After three consecutive 5-min washes in PBS, cells were incubated for 1 h with a corresponding secondary antibody in PBS, followed by DAPI staining. Antibodies used in this study are listed in Supplementary Table 5.
Flow cytometry
Cells were washed twice with HBSS, dissociated with 1 mg ml−1 collagenase I at 37 °C for 30 min and blocked in 10% FBS/PBS for 30 min at room temperature. For intracellular markers, 400,000 cells were fixed with 3.7% paraformaldehyde for 30 min, permeabilized with 0.5% saponin/5% FBS/PBS for 15 min on ice, and stained with APC-conjugated anti-troponin T antibody for 2 h. For OCT4, NANOG and lamin A/C, primary antibodies were incubated 2 h, followed by incubation with secondary antibodies for 1 h. Cells were washed and resuspended in FACS buffer for analysis. Apoptosis was assessed using APC annexin V and 7-AAD staining according to the manufacturer’s protocol (BioLegend, 640930). Cardiomyocytes were distinguished by Nkx2-5 GFP expression.
Histone isolation and subcellular fractionation
Cells were washed with cold PBS and lysed with histone extraction buffer (10 mmol l−1 HEPES, 1.5 mmol l−1 MgCl2, 10 mmol l−1 KCl, 1 mmol l−1 dithiothreitol (DTT), 1.5 mmol l−1 phenylmethyl sulfonyl fluoride (PMSF), 0.2 mol l−1 HCl and 1× protease inhibitor cocktail) on ice for 30 min, followed by neutralization with Tris-HCl, pH 9.0. A Subcellular Protein Fractionation kit (Thermo Fisher scientific, 78840) was used to separate cytoplasmic, membrane, nuclear soluble, chromatin-bound and cytoskeletal protein extracts according to the manufacturer’s protocol.
RNA isolation, RT–PCR and real-time PCR
Total RNA was isolated by TRIzol RNA Isolation Reagent (Invitrogen, 15596018). cDNA was synthesized with the High Capacity cDNA Reverse Transcription kit (Applied Biosystems, 4368813). Real-time PCR was performed by the qPCRBIO SyGreen Blue Mix (Nippon, PB20.16-51). Progerin expression in young and aged mouse heart tissue was analysed as previously described81. Real-time PCR primers were designed with Primer3 and synthesized by Sigma-Aldrich. All primers used in this study are listed in Supplementary Table 5.
RNA sequencing and data analysis
Nkx2-5-GFP mES cell lines (shControl, shControl Lmna−/−, shCbs Lmna−/−, shCth Lmna−/−, CBS OE, CTH OE, control, LmnaG609G/+, LmnaG609G/G609Gand CBS OE-LmnaG609G/G609G) were cultured on feeders in DMEM high glucose with 15% FBS and 1,000 U ml−1 LIF. After pre-plating to remove feeders, total RNA was extracted using the RNeasy Plus Universal Mini kit (QIAGEN, 73404). RNA integrity was assessed on a Bioanalyzer 2100 (Agilent). Libraries were prepared and sequenced on a BGISEQ-500 platform. Reads were trimmed with Trimmomatic-0.39 and aligned to the mm10 genome using STAR (−alignIntronMin 20 −alignIntronMax 500,000). Quality control was performed using MultiQC. Read counting was performed with HOMER (analyzeRepeats.pl, –count exons –strand both –noadj) after creating tag directories. Visualization in Integrative Genomics Viewer (IGV) was carried out using merged bam files via BamTools, with bigwig files generated using deepTools (BamCoverage, –bs 20 –smoothLength 40 –normalizeUsing RPKM –e 150). Differential gene expression was analysed with DESeq2 (log2 fold change ≤ −0.58 or ≥0.58, Padjust < 0.05). GO and KEGG pathway analyses were performed with DAVID Bioinformatics Resources 2023. Principal-component analysis, volcano plots and boxplots were generated using custom R scripts.
ChIP-sequencing and data analysis
A total of 1–5 million cells were crosslinked with 1% formaldehyde for 10 min at room temperature and quenched with 125 mM glycine. Cells were washed three times with ice-cold PBS and lysed in cell lysis buffer (50 mM Tris pH 8.0, 2 mM EDTA, 0.1% NP-40, 10% glycerol) with protease and phosphatase inhibitors for 10 min on ice. Nuclei were pelleted and lysed in nuclei lysis buffer (50 mM Tris pH 8.0, 1% SDS and 5 mM EDTA) for 10 min at room temperature. Chromatin was sonicated to 200–500 bp fragments using Bioruptor Plus (45 cycles, 30 s ON/30 s OFF) and centrifuged. Supernatants were diluted 1:10 in ChIP dilution buffer (50 mM Tris pH 8.0, 200 mM NaCl, 5 mM EDTA and 0.5% NP-40) with inhibitors and pre-cleared with BSA-coated Protein G Sepharose beads (GE Healthcare, 17061801) for 1 h at 4 °C. Chromatin was incubated overnight at 4 °C with 1 µg antibodies against H3K9ac (Cell Signalling Technology, 9649s), H3K27ac (Active motif, 39135), H3K9me3 (Abcam, ab8898) or H3K27me3 (Millipore, 07-449), followed by incubation with BSA-coated Protein G beads. Immunoprecipitates were washed sequentially with NaCl wash buffer, LiCl wash buffer and TE buffer. Complexes were eluted with 2% SDS buffer, reverse-crosslinked at 65 °C overnight, treated with proteinase K, and DNA was purified using the ChIP DNA Clean & Concentrator kit (Zymo Research). Chromatin fragment size was verified using a Bioanalyzer 2100 (Agilent) and NGS libraries were prepared with NEBNext Ultra II DNA Library Prep kit (Illumina).
For data analysis, ChIP-seq reads were trimmed with Trimmomatic, aligned to mm10 using Bowtie2, and PCR duplicates removed using Picard MarkDuplicates. BAM files from replicates were merged using BamTools. Normalized coverage tracks (bigWig) were generated using bamCoverage from deepTools (-b 20 -smooth 40 –normalizeUsing RPKM -e 150). Peaks were called using MACS14, and ENCODE mm10 blacklist regions were removed. Genome-wide enrichment profiles were generated using plotProfile from deepTools. Histone mark enrichment over ATAC-seq gain/loss regions (log2 fold change ≥0.58, ≤−0.58; P < 0.05) was visualized using ngs.plot (v.2.47.1) and plotted in R with ggplot2 (v.3.5.2).
For ChIP-seq, input DNA was used for normalization; for ChIP–qPCR, primers in gene desert regions were used to assess background, and results were normalized to input. ChIP–qPCR primers were designed with Primer3 and synthesized by Sigma-Aldrich.
Lamin A/C and SP1 ChIP were performed similarly with 5 million cells using 5 µg lamin A/C (Abcam, ab8984) or 2 µg SP1(Santa Cruz, sc-17824) antibody.
10x Genomics scRNA-sequencing and data analysis
Day 10 EBs were dissociated with Collagenase I as described in the ‘Flow cytometry’ section. Single live cells were sorted using SYTOX Blue Dead Cell Stain (1:2,000, Thermo Fisher Scientific) and diluted to 1,000 cells per µl in 0.4% BSA. A total of 8,000 cells were used for library preparation with the Chromium Next GEM Single-Cell 3’ kit v.3.1 (10x Genomics) and sequenced on a BGISEQ-500 platform. Raw reads were processed using Cell Ranger (v.7.1.0), and filtered feature matrices were integrated using Seurat (v.4.3.0). Low-quality cells expressing fewer than 900 or more than 9,000 genes were excluded. Clustering was performed with FindClusters (resolution = 0.5, random.seed = 1 and algorithm = 1). Uniform Manifold Approximation and Projection and t-stochastic neighbour embedding plots were generated using RunUMAP and RunTSNE (dims = 1:30, n.neighbours = 30). Differential gene expression was visualized with VlnPlot after normalization with NormalizeData.
Hi-C and DamID data analysis
Loess-quantile normalized DamID log2 Dam-laminA:Dam and Dam-laminB1:Dam ratios were obtained from GSE17051 and GSE62685. LADs were identified using circular binary segmentation (CBS) via LADetector with DNAcopy82. DpnII-fragmented genomic bins were used, and neighbouring segments were merged with a 2-kb sliding window; regions <25 kb were excluded. LAD coordinates were lifted from mm9 to mm10 using UCSC LiftOver. Peripheral lamin A LADs (LA/LB1 LADs) were defined by intersecting lamin A and lamin B1 LADs, whereas non-LAD genes overlapped neither. Scripts and the full DamID analysis workflow are available at https://github.com/thereddylab/LADetector. Significant genomic interactions from Hi-C in control and Lmna−/− cells were analysed using HOMER (v.4.2.1) with analyzeHiC (-res 5,000 -window 10,000 -superRes 10,000 -pvalue 0.01)83. Interactions were annotated via annotateInteractions.pl using merged H3K9ac and H3K27ac peaks and TSS (±2 kb) from mm10 RefSeq. Interactions were further compared with gained/lost H3K9ac, H3K27ac ChIP-seq and ATAC-seq peaks using bedtools intersect (v.2.30.0).
Statistics and reproducibility
All experiments were performed at least three times with biologically independent replicates, and n values always refer to biologically independent samples. Randomization was not applicable, as group allocation was determined by mouse genotype or genetic manipulation. Cell culture treatments (compound exposure, gene silencing or overexpression) were applied in parallel to avoid bias. Whenever possible, investigators were blinded during data acquisition. Samples were labelled with sequential numbers during preparation and, in most cases, throughout data collection, image acquisition and metabolite analyses. No animals were excluded from the study. Immunoblot images in Fig. 1g represent more than six independent experiments; those in Fig. 3d,g,i represent three independent experiments, with quantifications provided. Extended Data Fig. 1f shows immunoblots from three individual mES cell clones, with quantifications. Immunoblots in Figs. 5p and 7c–j are representative of at least three independent experiments with similar results. Data are presented as mean ± s.d. No statistical methods were used to predetermine sample sizes; however, chosen sample sizes are consistent with previous studies7,33. Data distribution was assumed to be normal but this was not formally tested. Group differences were evaluated using unpaired two-tailed Student’s t-tests or one-way/two-way analysis of variance (ANOVA) followed by Tukey’s multiple-comparison test, as specified in the figure legends. Statistical analyses and figure preparation were performed using GraphPad Prism v.9.4.1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data generated in this study have been deposited in the Gene Expression Omnibus under accession code GSE248534. The proteomics data have been deposited in the PRIDE repository under accession number PXD071040.
Lamin A DamID (GSE62685)84, lamin B1 DamID (GSE17051)18 ATAC-seq data from Lmna−/−, LmnaG609G/+ and LmnaG609G/G609G mES cell as well as Hi-C and RNA-seq data from Lmna−/− embryonic stem cell (GSE164069)33 were retrieved from previously published studies. Single-cell sequencing data of mouse embryos were retrieved from GSE100597 (ref. 50). ATAC-seq data of naive and primed mES cells were retrieved from E-MTAB-7207 (ref. 85). RNA-seq data of young and aged individuals, as well as patients with progeria were retrieved from GSE113957 (ref. 86). Source data are provided with this paper.
References
Rossant, J. & Tam, P. P. L. New insights into early human development: lessons for stem cell derivation and differentiation. Cell Stem Cell 20, 18–28 (2017).
Van Bortle, K. & Corces, V. G. Spinning the web of cell fate. Cell 152, 1213–1217 (2013).
Wu, J. & Izpisua Belmonte, J. C. Stem cells: a renaissance in human biology research. Cell 165, 1572–1585 (2016).
Wu, J., Ocampo, A. & Belmonte, J. C. I. Cellular metabolism and induced pluripotency. Cell 166, 1371–1385 (2016).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Weinberger, L., Ayyash, M., Novershtern, N. & Hanna, J. H. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 17, 155–169 (2016).
Sperber, H. et al. The metabolome regulates the epigenetic landscape during naive-to-primed human embryonic stem cell transition. Nat. Cell Biol. 17, 1523–1535 (2015).
Shiraki, N. et al. Methionine metabolism regulates maintenance and differentiation of human pluripotent stem cells. Cell Metab. 19, 780–794 (2014).
Zhou, W. et al. HIF1α induced switch from bivalent to exclusively glycolytic metabolism during ESC-to-EpiSC/hESC transition. EMBO J. 31, 2103–2116 (2012).
Moussaieff, A. et al. Glycolysis-mediated changes in acetyl-CoA and histone acetylation control the early differentiation of embryonic stem cells. Cell Metab. 21, 392–402 (2015).
Mathieu, J. et al. Hypoxia-inducible factors have distinct and stage-specific roles during reprogramming of human cells to pluripotency. Cell Stem Cell 14, 592–605 (2014).
TeSlaa, T. et al. α-Ketoglutarate accelerates the initial differentiation of primed human pluripotent stem cells. Cell Metab. 24, 485–493 (2016).
Hwang, I.-Y. et al. Psat1-dependent fluctuations in α-ketoglutarate affect the timing of ESC differentiation. Cell Metab. 24, 494–501 (2016).
Cornacchia, D. et al. Lipid deprivation induces a stable, naive-to-primed intermediate state of pluripotency in human PSCs. Cell Stem Cell 25, 120–136.e10 (2019).
Lu, V. et al. Glutamine-dependent signaling controls pluripotent stem cell fate. Dev. Cell 57, 610–623.e8 (2022).
Zhang, J. et al. LIN28 regulates stem cell metabolism and conversion to primed pluripotency. Cell Stem Cell 19, 66–80 (2016).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Shimi, T. et al. The A- and B-type nuclear lamin networks: microdomains involved in chromatin organization and transcription. Genes Dev. 22, 3409–3421 (2008).
Sullivan, T. et al. Loss of A-type lamin expression compromises nuclear envelope integrity leading to muscular dystrophy. J. Cell Biol. 147, 913–920 (1999).
Vergnes, L., Peterfy, M., Bergo, M. O., Young, S. G. & Reue, K. Lamin B1 is required for mouse development and nuclear integrity. Proc. Natl Acad. Sci. USA 101, 10428–10433 (2004).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Reddy, K., Zullo, J., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Butin-Israeli, V., Adam, S. A. & Goldman, R. D. Regulation of nucleotide excision repair by nuclear lamin b1. PLoS ONE 8, e69169 (2013).
Jenkins, H. et al. Nuclei that lack a lamina accumulate karyophilic proteins and assemble a nuclear matrix. J. Cell Sci. 106, 275–285 (1993).
Moir, R. D., Spann, T. P., Herrmann, H. & Goldman, R. D. Disruption of nuclear lamin organization blocks the elongation phase of DNA replication. J. Cell Biol. 149, 1179–1192 (2000).
Lee, J. et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy. Nature 572, 335–340 (2019).
Liu, B. et al. Genomic instability in laminopathy-based premature aging. Nat. Med. 11, 780–785 (2005).
Liu, B., Wang, Z., Le Zhang, S. G., Zheng, H. & Zhou, Z. Depleting the methyltransferase Suv39h1 improves DNA repair and extends lifespan in a progeria mouse model. Nat. Commun. 4, 1868 (2013).
Liu, G. H. et al. Recapitulation of premature ageing with iPSCs from Hutchinson-Gilford progeria syndrome. Nature 472, 221–225 (2011).
Shah, P. P. et al. Pathogenic LMNA variants disrupt cardiac lamina-chromatin interactions and de-repress alternative fate genes. Cell Stem Cell 28, 938–954.e9 (2021).
Wang, Y. et al. Lamin A/C-dependent chromatin architecture safeguards naive pluripotency to prevent aberrant cardiovascular cell fate and function. Nat. Commun. 13, 6663 (2022).
Bertero, A. et al. Chromatin compartment dynamics in a haploinsufficient model of cardiac laminopathy. J. Cell Biol. 218, 2919–2944 (2019).
Salvarani, N. et al. The K219T-Lamin mutation induces conduction defects through epigenetic inhibition of SCN5A in human cardiac laminopathy. Nat. Commun. 10, 2267 (2019).
Burke, B. & Stewart, C. L. The nuclear lamins: flexibility in function. Nat. Rev. Mol. Cell Biol. 14, 13–24 (2013).
Dittmer, T. A. & Misteli, T. The lamin protein family. Genome Biol. 12, 222 (2011).
Poleshko, A. et al. Genome-nuclear lamina interactions regulate cardiac stem cell lineage restriction. Cell 171, 573–587.e14 (2017).
Bianchi, A. et al. Dysfunctional polycomb transcriptional repression contributes to lamin A/C dependent muscular dystrophy. J. Clin. Invest. 130, 2408–2421 (2020).
Oldenburg, A. et al. A lipodystrophy-causing lamin A mutant alters conformation and epigenetic regulation of the anti-adipogenic MIR335 locus. J. Cell Biol. 216, 2731–2743 (2017).
Wang, Y. & Dobreva, G. Epigenetics in LMNA-related cardiomyopathy. Cells 12, 783 (2023).
Capell, B. C. & Collins, F. S. Human laminopathies: nuclei gone genetically awry. Nat. Rev. Genet. 7, 940–952 (2006).
Hegele, R. A. Lamin mutations come of age. Nat. Med. 9, 644–645 (2003).
Schreiber, K. H. & Kennedy, B. K. When lamins go bad: nuclear structure and disease. Cell 152, 1365–1375 (2013).
Maclean, K. N., Kraus, E. & Kraus, J. P. The dominant role of Sp1 in regulating the cystathionine β-synthase –1a and –1b promoters facilitates potential tissue-specific regulation by Kruppel-like factors. J. Biol. Chem. 279, 8558–8566 (2004).
Yang, G., Pei, Y., Teng, H., Cao, Q. & Wang, R. Specificity protein-1 as a critical regulator of human cystathionine γ-lyase in smooth muscle cells. J. Biol. Chem. 286, 26450–26460 (2011).
Eriksson, M. et al. Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423, 293–298 (2003).
Misteli, T. HGPS-derived iPSCs for the ages. Cell Stem Cell 8, 4–6 (2011).
Kubben, N., Brimacombe, K. R., Donegan, M., Li, Z. & Misteli, T. A high-content imaging-based screening pipeline for the systematic identification of anti-progeroid compounds. Methods 96, 46–58 (2016).
Mohammed, H. et al. Single-cell landscape of transcriptional heterogeneity and cell fate decisions during mouse early gastrulation. Cell Rep. 20, 1215–1228 (2017).
Swift, J. et al. Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341, 1240104 (2013).
Sternberg, A. K., Buck, V. U., Classen-Linke, I. & Leube, R. E. How mechanical forces change the human endometrium during the menstrual cycle in preparation for embryo implantation. Cells 10, 2008 (2021).
Sbodio, J. I., Snyder, S. H. & Paul, B. D. Regulators of the transsulfuration pathway. Br. J. Pharmacol. 176, 583–593 (2019).
Kabil, O., Yadav, V. & Banerjee, R. Heme-dependent metabolite switching regulates H2S synthesis in response to endoplasmic reticulum (ER) stress. J. Biol. Chem. 291, 16418–16423 (2016).
Bibli, S. I. & Fleming, I. Oxidative post-translational modifications: a focus on cysteine S-sulfhydration and the regulation of endothelial fitness. Antioxid. Redox Signal. 35, 1494–1514 (2021).
Bibli, S. I. et al. Mapping the endothelial cell S-sulfhydrome highlights the crucial role of integrin sulfhydration in vascular function. Circulation 143, 935–948 (2021).
Zivanovic, J. et al. Selective persulfide detection reveals evolutionarily conserved antiaging effects of S-sulfhydration. Cell Metab. 30, 1152–1170.e13 (2019).
Cho, S. et al. Mechanosensing by the lamina protects against nuclear rupture, DNA damage, and cell-cycle arrest. Dev. Cell 49, 920–935.e5 (2019).
Puente, B. N. et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157, 565–579 (2014).
Elsherbiny, A. & Dobreva, G. Epigenetic memory of cell fate commitment. Curr. Opin. Cell Biol. 69, 80–87 (2021).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
Bárcena, C. et al. Methionine restriction extends lifespan in progeroid mice and alters lipid and bile acid metabolism. Cell Rep. 24, 2392–2403 (2018).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R. A. & Feinberg, A. P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Nagaraj, R. et al. Nuclear localization of mitochondrial TCA cycle enzymes as a critical step in mammalian zygotic genome activation. Cell 168, 210–223.e11 (2017).
Li, W. et al. Nuclear localization of mitochondrial TCA cycle enzymes modulates pluripotency via histone acetylation. Nat. Commun. 13, 7414 (2022).
Pal, M., Altamirano-Pacheco, L., Schauer, T. & Torres-Padilla, M. E. Reorganization of lamina-associated domains in early mouse embryos is regulated by RNA polymerase II activity. Genes Dev. 37, 901–912 (2023).
Guertin, D. A. & Wellen, K. E. Acetyl-CoA metabolism in cancer. Nat. Rev. Cancer 23, 156–172 (2023).
Sim, E. Z. et al. Methionine metabolism regulates pluripotent stem cell pluripotency and differentiation through zinc mobilization. Cell Rep. 40, 111120 (2022).
Nicetto, D. et al. H3K9me3-heterochromatin loss at protein-coding genes enables developmental lineage specification. Science 363, 294–297 (2019).
Scaffidi, P. & Misteli, T. Lamin A-dependent nuclear defects in human aging. Science 312, 1059–1063 (2006).
Zhang, W. et al. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348, 1160–1163 (2015).
Mrabti, C. et al. Loss of H3K9 trimethylation leads to premature aging. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-4012025/v1 (2024).
Le, C. et al. Dietary glutamine supplementation alleviates age-related cardiac dysfunction by reducing elevated H3K27me3. Preprint at bioRxiv https://doi.org/10.1101/2025.05.16.654624 (2025).
Kirkland, N. J. et al. Age-dependent Lamin changes induce cardiac dysfunction via dysregulation of cardiac transcriptional programs. Nat. Aging 3, 17–33 (2023).
Lee, B. C., Kaya, A. & Gladyshev, V. N. Methionine restriction and life-span control. Ann. NY Acad. Sci. 1363, 116–124 (2016).
Fleming, T. P. et al. Origins of lifetime health around the time of conception: causes and consequences. Lancet 391, 1842–1852 (2018).
Kim, Y. & Zheng, Y. Generation and characterization of a conditional deletion allele for Lmna in mice. Biochem. Biophys. Res. Commun. 440, 8–13 (2013).
Hsiao, E. C. et al. Marking embryonic stem cells with a 2A self-cleaving peptide: a NKX2-5 emerald GFP BAC reporter. PLoS ONE 3, e2532 (2008).
Shechter, D., Dormann, H. L., Allis, C. D. & Hake, S. B. Extraction, purification and analysis of histones. Nat. Protoc. 2, 1445–1457 (2007).
Tang, H. et al. Multiplexed parallel reaction monitoring targeting histone modifications on the QExactive mass spectrometer. Anal. Chem. 86, 5526–5534 (2014).
Santiago-Fernández, O. et al. Development of a CRISPR/Cas9-based therapy for Hutchinson–Gilford progeria syndrome. Nat. Med. 25, 423–426 (2019).
Harr, J. C. et al. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J. Cell Biol. 208, 33–52 (2015).
Cordero, J. et al. Nuclear microRNA 9 mediates G-quadruplex formation and 3D genome organization during TGF-beta-induced transcription. Nat. Commun. 15, 10711 (2024).
Amendola, M. & van Steensel, B. Nuclear lamins are not required for lamina-associated domain organization in mouse embryonic stem cells. EMBO Rep. 16, 610–617 (2015).
Yang, S.-H. et al. ZIC3 controls the transition from naive to primed pluripotency. Cell Rep. 27, 3215–3227.e6 (2019).
Fleischer, J. G. et al. Predicting age from the transcriptome of human dermal fibroblasts. Genome Biol. 19, 221 (2018).
Acknowledgements
We thank S. Uhlig (FlowCore Mannheim), the Preclinical Models and NGS Core Facilities of the Medical Faculty Mannheim at Heidelberg University, and J. Hamel for excellent support. Schematics were created in BioRender.com. G.D. was supported by the CRC 1550 (project A03), the CRC 1366 (project A03) funded by the German Research Foundation (DFG), the German Centre for Cardiovascular Research (DZHK) (project ID 81Z0500202), funded by the Federal Ministry of Education and Research (BMBF) as well as the Baden-Württemberg foundation special programme ‘Angioformatics single cell platform’. S.-I.B. and I.F. were supported by the CRC1366 (project B1, project ID 456687919) and the Cardio-Pulmonary Institute, EXC 2026 (project ID 390649896) funded by the DFG. S.-I.B. was also supported by the DFG Emmy Noether Programme BI 2163/1-1 and the Johanna Quandt Young Academy at Goethe.
Author information
Authors and Affiliations
Contributions
The study was conceived and designed by G.D., Y.W. and S.-I.B. Experiments were performed and analysed by Y.W., H.S., J.W., Y.R., J.C. and M.D. J.H., J.M., A.B., J.B., T.W. and I.F. provided important reagents and intellectual input. NGS-sequencing data analysis was performed by Y.W., Y.R. and J.C. G.D., Y.W. and S.-I.B. wrote the paper. All of the authors contributed to editing of the paper and support the conclusions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Reviewer recognition
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Yanina-Yasmin Pesch and Alfredo Gimenez-Cassina, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Up-regulation of Cbs and Cth upon Lmna loss.
a, Quantification of the distance of Cbs and Cth to nuclear periphery in individual nuclei of Lmna+/+ and Lmna-/- mES cells. Cbs: Lmna+/+ (n = 105) and Lmna-/- (n = 161); Cth: Lmna+/+ (n = 106) and Lmna-/- (n = 103). b, Integrative Genomics Viewer (IGV) tracks of ATAC-seq and RNA-Seq of control (grey), and Lmna-/- mES cells (red) at the Cbs and Cth locus. Significantly increased peaks are highlighted in beige boxes. c, Relative fold changes in ATAC-seq signal at the transcription start site (TSS) and distal regions of Cbs and Cth in Lmna-/- mES cells compared to control mES cells. Differential accessibility was calculated in DESeq2 using two-sided Wald test. n = 3 biological replicates. d, RNA-seq analysis showing RPKM values of Cbs and Cth in Lmna-/- and control mES cells. n = 3 biological replicates. Differential expression was calculated in DESeq2 using two-sided Wald test. e, CTH (left) and CBS (right) enzymatic activity in control and Lmna-/- mES cells. n = 3 biological replicates. Purified CBS and CTH protein served as a positive control. f, g, Immunoblot analysis of key enzymes involved in the one-carbon metabolism in control and Lmna-/- mES cells (f), and quantification of protein relative fold changes (g). n = 3 biological replicates. h, i, Representative immunoblot for SP1 (h) and ATF4 (i) in chromatin-bound fraction (CBF) and total protein extract of control and Lmna-/- mES cells. Lamin B1 and α-Tubulin served as a loading control. j, Relative mRNA expression of Cbs and Cth in WT mES cells treated with DMSO or the Ezh1/2 inhibitor UNC1999. n = 4 biological replicates. k, ChIP–qPCR for H3K27me3 at the Cth and Cbs promoter in control and Lmna-/- mES cells normalized to input. n = 4 biological replicates. l, ChIP–qPCR for H3K27me3 and SP1 in mES cells treated with DMSO or UNC1999. n = 3 biological replicates. Differences in a, e, g, j–l were assessed using an unpaired two-tailed Student’s t-test. Data are presented as mean ± SD.
Extended Data Fig. 2 Progerin suppresses Cbs and Cth in ESCs.
a, Confocal images of immunostainings for lamin A/C (red), lamin B1 (green), and DAPI (nucleus, blue) of Lmna+/+ and LmnaG609G/G609G mES cells. Scale bar, 4 µm. b, Quantification of lamin A/C fluorescence intensity in Lmna+/+ and LmnaG609G/G609G mES cells. n = 82 for Lmna+/+ and n = 71 for LmnaG609G/G609G. Boxplots show median (centre line), 25th-75th percentiles (box), and 1.5× Interquartile Range (whiskers). c, Overlap of genes showing decreased ATAC-Seq signal in LmnaG609G/+ and LmnaG609G/G609G mES cells with genes within LA/LB1 LADs. d, qPCR analysis of Cbs and Cth expression in Lmna+/+, LmnaG609G/+, and LmnaG609G/G609G mES cells. n = 4 biological replicates. e, Quantification of relative fold changes in CBS and CTH protein levels in Lmna+/+, LmnaG609G/+, and LmnaG609G/G609G mES cells. n = 4 biological replicates. f, Representative immunoblot for SP1 in chromatin bound fraction and total protein extract of control and LmnaG609G/G609G mES cells. Lamin B1 and α-Tubulin served as a loading control. g, ChIP–qPCR analysis of SP1 enrichment at Cth and Cbs gene promoter region in control and LmnaG609G/G609G mES cells; GD, gene desert region, used as a negative control. ChIP–qPCR results are normalized to the corresponding input signals. n = 4 biological replicates. h, ChIP-qPCR for H3K27me3 and Lamin A/C at the Cth and Cbs promoter in control and LmnaG609G/G609G mES cells normalized to input. n = 3 biological replicates. Data are presented as mean ± SD. Differences in b, g and h were assessed using an unpaired two-tailed Student’s t-test; in d, and e using one-way ANOVA with Tukey correction. ns: not significant.
Extended Data Fig. 3 Lamin A/C-dependent CTH and CBS activation drives the transition from naive to primed pluripotency.
a, Pathway impact analysis of day 3 (D3) EBs versus naive mES cells. n = 6 biological replicates. Significance was calculated using a one-sided hypergeometric test in MetaboAnalyst 5.0. b, Intracellular methionine (Met), homocysteine (Hcys), cystathionine (Cysta) and cysteine (Cys) levels in mESC (n = 6) and D3 (n = 6), D6 (n = 3) or D10 (n = 3) EBs. c, d, Immunoblot analysis of CBS, CTH, lamin A/C and lamin B1 (c) and intracellular cysteine levels (d) in mES cells cultured in naïve or primed conditions. n = 4 biological replicates. e, Intracellular cysteine levels in H1 hESC cultured in naïve and primed conditions (n = 6)7. f, g, OCT4 immunostaining (f) and percentage of cells highly expressing NANOG (NANOGhigh) determined by FACS analysis of control and Lmna-/- mES cells after silencing with control shRNA or shRNA against Cth or Cbs or of CTH or CBS-overexpressing (OE) mES cells (g). n = 3 biological replicates. Scale bar, 40 µm. h, Heatmap representation of primed genes in RNA-Seq (n = 2) of control and Lmna-/- mES cells after Cth or Cbs silencing. i, FACS analysis (top) and percentage of OCT4high cells (bottom) in mES cells cultured in standard medium, medium depleted of cysteine, or supplemented with AOAA for 48 h. n = 4 biological replicates. j, l, Immunostaining of CBS, CTH, and Lamin A-GFP (j) and fluorescence intensity (l) at 0 h (n = 31, 30 and 142), 6 h (n = 49, 35 and 115), 8 h (n = 40, 30 and 113), 12 h (n = 35, 30 and 67), and 24 h (n = 44, 30 and 79) after doxycycline treatment of mES cells overexpressing pLenti-CMV-TRE3G-Neo-GFP-Lamin A plasmid. Scale bar, 10 µm. k, Relative Cth and Cbs mRNA expression (left) or cysteine levels (right) 6 h after doxycycline treatment. n = 3 biological replicates. m, Percentage of NANOGhigh cells determined by FACS analysis at different time points after doxycycline treatment. n = 3 biological replicates. n, o FACS analysis (n) and percentage of NANOGhigh cells (o) in mES cells overexpressing inducible GFP-Lamin A cultured in naïve conditions or primed conditions for 24 h with or without doxycycline. n = 3 biological replicates. Data in b, d, e, g, i, k, l, m and o represent mean ± SD. Differences in e and k were assessed using an unpaired two-tailed Student’s t-test; in b, d, g, i, l, m and o by one-way ANOVA with Tukey correction. ns: not significant.
Extended Data Fig. 4 Lmna depletion leads to increased cysteine metabolism and the transition from naïve to primed pluripotency in R1 mES cells.
a, b, Immunoblot analysis of lamin A/C in E14 and R1 mES cells (a), and quantification of lamin A/C protein relative fold changes from three biological replicates (b). c, Representative FACS analysis of lamin A/C and NANOG expression in E14 and R1 mESC. d, Immunoblot analysis of CBS, CTH, lamin A/C levels in R1 mES cells cultured in naïve (15% FCS/LIF) or primed (bFGF/Activin A) conditions. e, Relative fold change of intracellular cysteine levels in R1 mES cells cultured under naïve conditions (2i/LIF), as well as in R1 mES cells and epiblast stem cells (EpiSC) cultured under primed conditions (bFGF/Activin A)1. n = 6 biological replicates. f, g, Immunoblot analysis of lamin A/C, CBS and CTH in control and Lmna-/- R1 mES cells (f), and quantification of CBS and CTH protein relative fold changes from three biological replicates (g). h, Relative fold change of intracellular cysteine levels in control and Lmna-/- R1 mES cells. n = 6 biological replicates. i, j Representative histograms showing FACS analysis (i) and percentage of NANOGhigh cells (j) in control and Lmna-/- R1 mES cells. n = 3 biological replicates. k, Phase contrast images showing colony morphology of control and Lmna-/- R1 mES cells. Scale bar, 50 µm. l, qPCR analysis of genes found to be upregulated in E14 Lmna-/- mES cells, measured in control and Lmna-/- R1 mES cells. n = 4 biological replicates. Differences in b, g, h, j and l were assessed using an unpaired two-tailed Student’s t-test; in e were assessed using one-way ANOVA with Tukey correction. Data represent mean ± SD.
Extended Data Fig. 5 Enhanced cysteine metabolism promotes histone acetylation in Lmna mutant mES cells.
a, H2S levels generated through CBS/CTH in control and Lmna-/- mES cells. n = 3 biological replicates. Purified CBS and CTH proteins served as positive control. b, Expression of cysteine transporters in RNA-seq datasets of control and Lmna-/- mES cells. n = 3. RPKM: reads per kilobase of transcript per million mapped reads. Differential expression was calculated in DESeq2 using two-sided Wald test. c, Total (left) and nuclear (right) acetyl-CoA levels of control and Lmna-/- mES cells determined by fluorometric Acetyl-Coenzyme A assay. n = 3 biological replicates. d, Immunoblots analysis of H3K9ac, H3K27ac, H3K56ac and pan-H3ac levels in control and Lmna-/- mES cells (top), and quantification of protein relative fold changes (bottom, n = 3 biological replicates). e, Immunoblot analysis of CBS, CTH, and pan-H3ac levels in WT mES cells after Cbs or Cth single knockdown (top) or knockdown of both Cbs and Cth (bottom). f, Immunoblot analysis of H3K9ac, H3K27ac, and pan-H3ac levels in WT mES cells after 0, 1, 3, 6, 9, and 24 h of AOAA treatment (top), and quantification of protein relative fold changes (bottom, n = 3 biological replicates). g, Immunoblot analysis of whole cell lysates for L-lysine acetylated proteins in control and Lmna-/- mES cells. h, Immunoblots analysis of H3K9ac, H3K27ac, H3K56ac and pan-H3ac levels in control and Lmna-/- R1 mES cells (top), and quantification of protein relative fold changes (bottom, n = 3 biological replicates). i, Quantification of H3K9ac, H3K27ac and pan-H3ac relative fold changes in control, LmnaG609G/G609G, and LmnaG609G/G609G mES cells overexpressing both CBS and CTH. n = 3 biological replicates. j, Immunostaining for H3K9ac (magenta), and Lamin A-GFP (top) and quantification (bottom) of H3K9ac fluorescence intensity at 0 (n = 80), 6 (n = 67), 8 (n = 55), 12 (n = 60), and 24 (n = 66) hours after treatment with doxycycline of WT mES cells overexpressing the pLenti CMV TRE3G Neo GFP-Lamin A plasmid. Scale bar, 10 µm. Data in a–d, f, h, i and j are presented as mean ± SD. Differences in a–d, and h were assessed using an unpaired two-tailed Student’s t-test. Differences in f, i and j were assessed by one-way ANOVA with Tukey correction. ns: not significant.
Extended Data Fig. 6 CTH/CBS upregulation results in gene activation of lamin A/C-dependent genes.
a, Overlap of genes upregulated in Lmna-/- mES cells with genes downregulated in Lmna-/- ESCs after shRNA mediated silencing with shRNA against Cth or Cbs. b, Split violin plots representing the log10 (PRKM + 0.1) for transcripts of genes in LA/LB1 LADs and non-LADs in RNA-seq datasets of control, Lmna-/- mES cells or Lmna-/- mES cells after shRNA mediated silencing with control shRNA or shRNA against Cth or Cbs. RPKM, Reads Per Kilobase per Million mapped reads. c, Overlap of genes upregulated in Lmna-/- mES cells with genes upregulated in CTH OE or CBS OE mES cells. d, Split violin plots representing the log10 (RPKM + 0.1) for transcripts of genes in LA/LB1 LADs and non-LADs in RNA-seq datasets of control, CBS OE and CTH OE mES cells. e, Dot-plot representation of gene ontology analysis of genes upregulated in CTH OE and CBS OE mES cells or genes upregulated in CTH OE or CBS OE and Lmna-/- mES cells. f, Overlap of the genes downregulated in LmnaG609G/G609G mES cells with genes upregulated in CTH OE and CBS OE mES cells. g, Dot-plot representation of gene ontology analysis of genes both downregulated in LmnaG609G/G609G mESC and upregulated in CTH OE or CBS OE mESC. h, qPCR analysis of genes found within LA/LB1 LADs and outside LADs in control, LmnaG609G/G609G mESC as well as LmnaG609G/G609G mES cells after CTH or CBS overexpression. n = 4 biological replicates. Data in h are presented as mean ± SD. In the split violin plots, the centre line represents the median, the upper bound corresponds to the 75th percentile, and the lower bound corresponds to the 25th percentile. Differences in a, c and f were assessed using one-sided hypergeometric test; Differences in d were assessed using an unpaired two-tailed Student’s t-test; Differences in b and h were assessed using one-way ANOVA with Tukey correction. Significance in e and g was generated from DAVID with a one-sided Fisher’s exact test.
Extended Data Fig. 7 Abnormal cysteine metabolism upon lamin A/C mutation induces aberrant cell fate choices.
a, b, qPCR analysis of cardiac progenitor marker genes in D6 EBs (a) and CM marker genes in D10 EBs (b), differentiated from mES cells cultured in medium containing the indicated amounts of cysteine. n = 3 biological replicates. c, Representative FACS analysis of cTnT+ CMs in D10 EBs differentiated from mES cells cultured in medium containing the indicated concentrations of cysteine. n = 3 biological replicates. d, Representative FACS analyses of cTnT+ cardiomyocytes at D10 of differentiation of control and Lmna-/- mES cells after shRNA mediated silencing with control shRNA or shRNA against Cth or Cbs. e, Immunostaining for α-actinin (red) and DAPI (blue) in D10 EBs differentiated from control ESCs and Lmna-/- mES cells either non-treated or treated with AOAA for 48 h before differentiation. Scale bar, 10 µm. f, t-SNE plot of 10x genomics scRNA-Seq datasets of D10 EBs differentiated from control, Lmna-/- and Lmna-/- mES cells after shRNA mediated silencing of Cth. Cell types: CM: cardiomyocyte, ECs: endothelial cell, SMC: smooth muscle cell, Mono/MΦ: Monocytes/ Macrophages, NPCs: neuron progenitor cell, Hepa: hepatocyte, Ery: erythrocyte. g, Violin plots showing differential gene expression in the different cell clusters in D10 EBs differentiated from control, Lmna-/- and Lmna-/- mES cells after shRNA mediated silencing of Cth. Cell types: CM: cardiomyocyte, SMC: smooth muscle cell, Ep: epithelial cell, Hepa: hepatocyte, Ery: erythrocyte, NPCs: neuron progenitor cell. h, i, qPCR analysis of germ layer markers in D4 EBs (h) and CM marker genes in D10 EBs (i) differentiated from control, LmnaG609G/+ and LmnaG609G/G609G ESCs. n = 4 biological replicates. Data in a, b, h and i are presented as mean ± SD. Differences in a, b, h, and i were assessed using one-way ANOVA with Tukey correction. Differences in g were assessed using one-way ANOVA with Tukey’s correction after aggregating single-cell expression data to a pseudo-bulk level.
Extended Data Fig. 8 Perturbated cysteine metabolism results in DNA damage and senescence in Lmna p.G609G mES cells.
a, Ratio of S-adenosylmethionine (SAM) to S-adenosylhomocysteine (SAH) measured in control, Lmna-/- and LmnaG609G/G609G mES cells transfected with control, Cbs shRNA, or CBS OE plasmid. n = 4 biological replicates. b, Immunoblot analysis for CBS and CTH in siRNA mediated Cbs, Cth single or double knockdown mES cells. c, Immunoblot analysis for SUV39H1 in soluble nuclear extracts (SNE) of control and Lmna-/- mES cells either not treated or treated with 50 µmol/L SAM for 6 h. d, Immunoblot analysis of γ-H2AX and CBS levels in control and LmnaG609G/G609G mES cells, and LmnaG609G/G609G mES cells infected with different concentration of lentivirus carrying CBS cDNA. e, Immunostaining for γ-H2AX (red) and DAPI (blue) of control, LmnaG609G/G609G and CBS OE LmnaG609G/G609G mES cells either not treated (w/o) or treated with H2O2 for 1 h and allowed to repair in fresh medium for 0 h and 4 h. Scale bar, 10 µm. f, Representative histograms showing FACS analysis of the percentage of SPIDER SA β-gal+ cells in mES cells either not treated (w/o) or treated with H2O2 for 2 h and cultured in fresh medium for 24 h. g, qPCR analysis of Cdkn2a and SASP marker genes in control and LmnaG609G/G609G mES cells, as well as in LmnaG609G/G609G mES cells overexpressing CBS. n = 4 biological replicates. h, Ratio of peripheral to nucleoplasmic mean lamin A/C fluorescence intensity (left) and nuclear size (right) in young (3-month-old) and aged (24-month-old) mice. Left: n = 51 CMs for each group; Right: young (n = 107) and old (n = 118). i, Immunoblot analysis of H3k9me3 and H3 in protein extracts from young (3-month-old) and aged (24-month-old) mice. n = 7 mice for each group. j, qPCR analysis of Cdkn2a and SASP gene expression in hearts of young (3-month-old) and old (24-month-old) mice. n = 7 mice per group. Data in a, g, h and j are presented as mean ± SD. Differences in h and j were assessed using an unpaired two-tailed Student’s t-test. Differences in a and g were assessed using one-way ANOVA with Tukey correction. ns: not significant.
Extended Data Fig. 9 Working model illustrating the interplay between the nuclear lamina and cysteine metabolism in regulating stem cell fate and longevity.
a, During naïve-to-primed state transition, RA signalling induces a reduction in Lmna expression, thereby enhancing SP1 occupancy on chromatin. Increased SP1 binding promotes the upregulation of the cysteine-generating and -metabolizing enzymes Cbs and Cth. This, in turn, boosts de novo cysteine synthesis, leading to enhanced histone H3 acetylation at lysine 9 and lysine 27. The resulting chromatin changes drive epigenome remodelling that is essential for subsequent differentiation and germ layer formation. b, In naïve pluripotent stem cells, lamin A/C suppresses Cth and Cbs expression, thereby limiting cysteine bioavailability. CTH and CBS are highly upregulated in ESC with lamin A/C loss-of-function (Lmna-/-) or decreased in ESC with lamin A/C dominant, gain-of-function mutation (Lmna p. G609G) leading to either enhanced or reduced cysteine metabolism and rewiring of the methionine cycle. Ultimately, these non-physiological levels of CTH and CBS are responsible for abnormal epigenetic signatures, genomic instability and aberrant cell fate and function.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6 and FACS gating strategy.
Supplementary Table 1 (download XLSX )
LC–MS-based untargeted metabolomics analysis was performed on control and Lmna−/− mES cells, as well as during cardiomyocyte differentiation of WT mES cells.
Supplementary Table 2 (download XLSX )
RNA-seq analysis of Lmna+/+ and Lmna−/− mES cells after shRNA mediated silencing with control shRNA or shRNA against Cth or Cbs or mES cells overexpressing CTH (CTH OE) or CBS (CBS OE).
Supplementary Table 3 (download XLSX )
RNA-seq analysis of Lmna+/+, LmnaG609G/+ and Lmna G609G/G609G mES cells.
Supplementary Table 4 (download XLSX )
RNA-seq analysis of Lmna+/+, LmnaG609G/G609G mES cells and LmnaG609G/G609G after CBS overexpression.
Supplementary Table 5 (download XLSX )
Primers and antibodies used in this study.
Supplementary Table 6 (download XLSX )
Supplementary source data for Supplementary Figs. 1–6.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 1 (download PDF )
Unprocessed western blots for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 3 (download PDF )
Unprocessed western blots for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 5 (download PDF )
Unprocessed western blots for Fig. 5.
Source Data Fig. 6 (download XLSX )
Statistical source data for Fig. 6.
Source Data Fig. 7 (download XLSX )
Statistical source data for Fig. 7.
Source Data Fig. 7 (download PDF )
Unprocessed western blots for Fig. 7.
Source Data Fig. 8 (download XLSX )
Statistical source data for Fig. 8.
Source Data Fig. 8 (download PDF )
Unprocessed western blots for Fig. 8.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots for Extended Data Fig. 8.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Shi, H., Wittig, J. et al. Lamin A/C-regulated cysteine catabolic flux modulates stem cell fate through epigenome reprogramming. Nat Metab 8, 431–453 (2026). https://doi.org/10.1038/s42255-025-01443-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01443-2