Abstract
Gene drives are genetic modifications designed to propagate efficiently through a population. Most applications rely on homologous recombination during sexual reproduction in diploid organisms such as insects, but we recently developed a gene drive in herpesviruses that relies on co-infection of cells by wild-type and engineered viruses. Here, we report on a viral gene drive against human herpes simplex virus 1 (HSV-1) and show that it propagates efficiently in cell culture and during HSV-1 infection in mice. We describe high levels of co-infection and gene drive-mediated recombination in neuronal tissues during herpes encephalitis as the infection progresses from the site of inoculation to the peripheral and central nervous systems. In addition, we show evidence that a superinfecting gene drive virus could recombine with wild-type viruses during latent infection. These findings indicate that HSV-1 achieves high rates of co-infection and recombination during viral infection, a phenomenon that is currently underappreciated. Overall, this study shows that a viral gene drive could spread in vivo during HSV-1 infection, paving the way toward therapeutic applications.
Similar content being viewed by others
Introduction
Herpesviruses are ubiquitous pathogens that establish lifelong infections and persistently infect most of the human population. Their large dsDNA genomes (100–200 kb) replicate in the nucleus and encode hundreds of genes. After primary infection, herpesviruses enter latency and occasionally reactivate, causing recurrent disease. Herpes simplex viruses (HSV) 1 and 2 first infect mucosal surfaces before spreading to the nervous system via axons. The viral genome remains latent in neurons in sensory and autonomic ganglia. Periodic reactivation typically causes lesions in the facial or genital areas, which can be highly painful and stigmatizing. In addition, herpes encephalitis can be life-threatening in newborns and immunocompromised individuals1,2, and HSV-2 is a major risk factor for HIV acquisition3. Antiviral drugs like acyclovir can block viral replication and reduce symptoms, but cannot eradicate the latent reservoir. HSV-1 and 2 lack vaccines or curative treatments and new therapeutic strategies for HSV diseases are critically needed.
During an infection, cells are often co-infected by several virions and therapeutic approaches that rely on viral co-infection have great potential. These viral interference strategies rely on defective viral particles that interfere with the replication of wild-type viruses after co-infection, thus reducing viral levels4. Most viral interference approaches have focused on RNA viruses such as SARS-CoV-2, RSV, Influenza, Zika, Chikungunya, or HIV5,6,7,8,9,10. We recently developed a system of viral interference against herpesviruses, using a technique of genetic engineering known as a gene drive11,12. Gene drives are genetic modifications designed to spread efficiently through a population13,14. They usually rely on CRISPR-mediated homologous recombination during sexual reproduction to efficiently spread a transgene in a population, and applications have been developed in insects to eradicate diseases such as malaria13,14. By contrast, the viral gene drive that we invented spread by the co-infection of cells with wild-type and engineered viruses (Fig. 1a). Specifically, upon co-infection with a gene drive virus expressing CRISPR-Cas9, the wild-type genome is cut and repaired by homologous recombination, producing new recombinant gene drive viruses that progressively replace the wild-type population. In a proof-of-concept study with human cytomegalovirus (hCMV), we previously showed that a viral gene drive propagated efficiently in cell culture in a population of unmodified viruses11. Importantly, an attenuated gene drive virus could spread efficiently until the wild-type population had been eliminated, ultimately reducing viral levels. This represented a new way to engineer herpesviruses for research or therapeutic purposes.
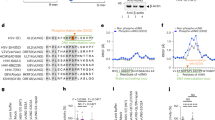
a Gene drive viruses carry Cas9 and a gRNA targeting the same location in a wild-type genome. After coinfection of cells by wild-type (WT) and gene drive (GD) viruses, Cas9 cleaves the wild-type sequence and homology-directed repair –using the gene drive sequence as a repair template– causes the conversion of the wild-type locus into a new gene drive sequence and the formation of new recombinant gene drive viruses (rGD). Artwork was modified from ref. 11, 12. b Modified and unmodified UL37-38 region. The gene drive cassette was inserted between the UL37 and UL38 viral genes and was composed of spCas9 under the control of a CBH promoter followed by the SV40 polyA signal, a CMV promoter driving an mCherry reporter, followed by the beta-globin polyA signal, and a U6-driven gRNA. c Localizations of the gene drive sequence and YFP/CFP reporters on HSV-1 genomes. GD represents a functional gene drive virus, GD-ns carries a non-specific gRNA, and Cas9 is deleted in GD-ΔCas9. UL/US: unique long/short genome segments. d, e Recombination products and examples of viral plaques after cellular co-infection with HSV1-WT expressing YFP and gene drive viruses expressing mCherry and CFP. Representative images from more than n > 10 experiments. Scale bars: 100 μm.
The spread of a viral gene drive relies on cellular co-infection events, which occur at a high frequency in cell culture experiments. While numerical simulations suggest that a viral gene drive could still spread with low co-infection rates11, the frequency of co-infection events during animal or human infections is poorly characterized. In mice, high levels of recombination are observed during herpes simplex encephalitis15,16, and attenuated viruses –which individually are harmless– can complement each other and cause severe disease17,18. Moreover, HSV strains circulating in humans exhibit evidence of extensive recombination19,20,21,22. These indirect observations suggest that cells are often co-infected by several virions during HSV-1 infection. However, whether a viral gene drive could propagate in vivo remains unknown.
In the present study, we tested if a viral gene drive could spread in animal models of herpesvirus infection. HSV-1 is an important human pathogen with established infection protocols in mice, and we developed a gene drive that targeted a neutral region in the HSV-1 genome. This design was not intended to impact viral fitness or limit viral spread, but to investigate the potential of the technology in vivo. Our results show that a gene drive could spread efficiently during acute HSV-1 infection, particularly in neuronal tissues. Using fluorescently labeled viruses, we directly observed high levels of cellular co-infection in the brain. Finally, we show evidence that a superinfecting gene drive virus could recombine with wild-type genomes during latent infection, paving the way toward therapeutic applications.
Results
Design of a gene drive against HSV-1
We aimed to build a gene drive that would not affect viral infectivity and could spread efficiently into the wild-type population. We designed a gene drive targeting an intergenic sequence between the UL37 and UL38 genes, a region known to tolerate the insertion of transgenes with little or no impact on viral replication in cell culture and in vivo23,24. We created a donor plasmid containing homology arms, Cas9 (from Streptococcus pyogenes) under the CBH promoter, an mCherry fluorescent reporter under the CMV promoter, and a U6-driven guide RNA (gRNA) targeting the intergenic UL37-UL38 region (Fig. 1b). Importantly, to prevent self-cleaving, introduction of the gene drive cassette removed the gRNA target sequence from the construct. To create the gene drive virus (GD), Vero cells were co-transfected with purified HSV-1 viral DNA and the gene drive donor plasmid, and mCherry-expressing viruses created by homologous recombination were isolated by three rounds of plaque purification until a pure population was obtained. To follow recombination events between viral genomes, the gene drive virus also carried a cyan fluorescent reporter (CFP) inserted into another neutral region between the US1 and US2 viral genes15,25 (Fig. 1c). Similarly, we built two control viruses with non-functional CRISPR systems, one with a non-specific gRNA (GD-ns) that did not target HSV-1, and one without Cas9 (GD-ΔCas9). Finally, to be used as a wild-type virus with an unmodified UL37-UL38 region, we generated a virus expressing a yellow fluorescent reporter (YFP) from the same US1-US2 region, hereafter referred to as HSV1-WT or simply WT (Fig. 1c). All the viruses described here originated from the highly neurovirulent HSV-1 strain 17 + .
CRISPR technology presents the risk of off-target editing of the host genome. The gRNA targeting the HSV-1 UL37-UL38 intragenic region was designed to minimize off-target editing of the mouse genome. Amplicon sequencing of 14 putative off-target sites after infection of mouse cells with GD or GD-ns confirmed the high specificity of the gRNA. Only one locus showed evidence of off-target editing significantly above background, with a limited mutation rate of 0.02% (Supplementary Fig. 1).
Recombination between gene drive and wild-type genomes can result in four different genome configurations expressing the different fluorescent reporters, which can be followed by plaque assay (Fig. 1d, e). In highly susceptible cell lines such as Vero cells, viral plaques originate from single virions, which allows reconstituting the recombination history of individual viral genomes. Plaques expressing YFP only, or CFP and mCherry together, represent the original WT and gene drive viruses, respectively, while plaques expressing CFP only, or YFP and mCherry together, represent recombination products that exchanged the gene drive cassette.
Gene drive spread in cell culture
First, we tested if our gene drive could spread in vitro and conducted co-infection experiments in cell culture. Some cell lines, in particular Vero cells or other epithelial cells, efficiently restrict co-infection by a mechanism known as superinfection exclusion26,27. However, we found that mouse neuronal N2a cells could sustain high levels of co-infection (Supplementary Fig. 2). Thus, co-infection experiments were conducted in N2a cells, while plaque assays were performed in Vero cells. Importantly, the WT, GD, GD-ns and GD-ΔCas9 viruses individually replicated with similar dynamics in N2a cells, showing that insertion of the gene drive cassette in the UL37-UL38 region did not affect infectivity in cell culture, as expected (Fig. 2a).
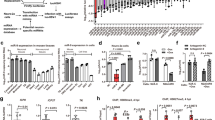
a Viral titers in the supernatant after infection of N2a cells with WT, GD, GD-ns or GD-ΔCas9. Cells were infected with a single virus at MOI = 1. n = 4. b, c Viral titers in the supernatant after co-infection of N2a cells with WT + GD, WT + GD-ns or WT + GD-ΔCas9, with a starting proportion of gene drive virus of 20% (b) or 40% (c). MOI = 1, n = 4. d–g Evolution of the viral population after co-infection with WT + GD, WT + GD-ns or WT + GD-ΔCas9, with a starting proportion of gene drive virus of 20% (d, e) or 40% (f, g). Panels (d and f) show the proportion of viruses expressing mCherry, representing gene drive virus. Panels (e and g) show the proportion of viruses expressing the different fluorophore combinations. Viral titers are expressed in log-transformed PFU (plaque-forming unit) per mL of supernatant. Error bars represent the standard error of the mean (SEM) between biological replicates. n = 4. Source data are provided as a Source Data file.
To test if the gene drive could efficiently spread in the viral population, N2a cells were co-infected with WT + GD, WT + GD-ns, or WT + GD-ΔCas9. Cells were infected at a combined MOI of 1 with an initial proportion of gene drive virus of 20% (Fig. 2b, d, e). Titers and proportion of progeny viruses expressing the different fluorescent markers were measured by plaque assay, from day one to day three post-infection. For each combination tested, the dynamics of total viral replication were indistinguishable from WT-only infection (Fig. 2b). However, the composition of the viral population changed over time. The proportion of viruses expressing mCherry, which represents gene drive viruses, increased from 20% to 85% when cells were co-infected with WT + GD (Fig. 2d). Importantly, YFP-only viruses disappeared and were replaced by recombinant viruses expressing both YFP and mCherry, representing 45% of the final population (Fig. 2e). This indicated that the WT population had been converted to new recombinant gene drive viruses, as anticipated. The proportion of viruses expressing both CFP and mCherry, which represent the original gene drive virus, increased slightly from 20% to 40% (Fig. 2e). By contrast, in the control experiments with GD-ns or GD-ΔCas9, the proportion of mCherry-expressing viruses did not change and remained close to its initial value of 20% (Fig. 2d). In these control experiments, around 10% of viruses expressed both YFP and mCherry at day 3, but this population of recombinant viruses was mirrored by viruses that had lost mCherry and expressed CFP only (Fig. 2e). These represented viruses that had exchanged their YFP and CFP regions in a nonspecific manner. Importantly, CFP-only viruses were not observed after co-infection with WT + GD, highlighting that the efficient incorporation of the gene drive cassette into unmodified genomes is a unilateral and targeted process requiring both Cas9 and a specific gRNA.
To confirm these observations, we repeated these co-infection experiments with an initial proportion of gene drive virus of 40% (Fig. 2c, f, g). With this higher starting point, the gene drive achieved almost complete penetrance and the proportion of mCherry-expressing viruses reached 95% after 3 days, with the population of wild-type viruses expressing YFP-only being converted to recombinant gene drive viruses expressing both YFP and mCherry. As observed above, in the WT + GD-ns or WT + GD-ΔCas9 control experiments, the proportion of mCherry-expressing viruses remained constant at around 40%, and approximately 15% of YFP+mCherry and CFP-only viruses symmetrically appeared by CRISPR-independent recombination. This further showed that a gene drive could spread efficiently in the wild-type population and that the drive was mediated by a functional CRISPR system.
Together, these results indicated that a gene drive could be designed against HSV-1 and spread efficiently in cell culture. Importantly, both during single and co-infections, viral growth of WT and GD viruses followed similar dynamics and the proportion of viruses expressing CFP remained constant (Fig. 2). Thus, the rapid increase of recombinant viruses expressing both mCherry and YFP could not be explained by a higher fitness of the GD virus, but resulted from efficient CRISPR-directed homologous recombination. These results align with the findings of a parallel study28 and expanded on our previous work with hCMV, showing that a viral gene drive could be developed in a second, unrelated, herpesvirus.
Gene drive spread during herpes encephalitis
Next, we tested if the gene drive could spread during acute HSV-1 infection in mice. Previous studies showed that HSV-1 naturally sustains high levels of recombination in the mouse brain during herpes simplex encephalitis15. Thus, we hypothesized that a gene drive could spread efficiently in this context. We used a well-established infection model, where HSV-1 is inoculated intravitreally in the eye and infects the retina and other ocular tissues before propagating to the nervous system via cranial nerves (Fig. 3a). Specifically, HSV-1 travels to the brain via the optic, oculomotor and trigeminal nerves (cranial nerves CN II, III, and V, respectively), either directly or indirectly by first infecting ganglionic neurons of the peripheral nervous system. In particular, HSV-1 infects the trigeminal ganglia (TG) before reaching the brain stem. In the following experiments, a total of 106 plaque-forming units (PFU) were inoculated intravitreally in the left eye, and tissues were collected, dissociated, and analyzed by plaque assay after four days. Five to eight-week-old male and female Balb/c mice were used and we did not observe any differences between sexes. Importantly, individual infections with WT, GD or GD-ns reached similar titers in the eye, the TG and the brain, showing that the gene drive cassette in the UL37-UL38 region did not impact infectivity in vivo (Fig. 3b).
a Infection routes along the optic, oculomotor and trigeminal nerves (cranial nerves II, III and V, respectively) following ocular inoculation of HSV-1. Male and female Balb/c mice were infected with 106 PFU in the left eye. b, c Viral titers after four days in the eye, TG and whole brain after (b) infection with a single virus, n = 5 mice, or (c) with a starting proportion of gene drive virus of 15%, n = 6 mice. d, e Viral population in the eye, TG and whole brain after co-infection with WT + GD or WT + GD-ns, after four days. n = 6. f Proportion of viral genomes with a mutated target site in the brain after four days. n = 3. g Viral titers in the spinal cord and brain after inoculation of WT, WT + GD, or WT + GD-ns in the right hind leg footpad, after 5–7 days. n = 8 for WT and WT + GD, n = 4 for WT + GD-ns. h, i Viral population in the spinal cord and whole brain after co-infection with WT + GD or WT + GD-ns, after 5–7 days. n = 5 for WT + GD, n = 1 for WT + GD-ns. Viral titers are expressed in log-transformed PFU. In panels (b, c and g) black lines indicate the median. n.d.: non-detected. Panels (d, e, f, h, and i) show the average and SEM between biological replicates. Source data are provided as a Source Data file.
To test if the gene drive could efficiently spread in vivo, mice were inoculated intravitreally with WT only, WT + GD, or WT + GD-ns, with an initial proportion of gene drive virus of 15% (Fig. 3c–e). Total viral titers in the eye, TG and brain after four days were indistinguishable between the different conditions, showing that the gene drive did not perturb the overall dynamics of infection (Fig. 3c). The population of gene drive viruses expressing mCherry increased from 15% to 30% in the eye, to 60% in the TG, and 70% in the brain, respectively. We observed a high variation between replicates, with the percentage of gene drive viruses in the brain ranging from 50% to 90%, with a median of 77% (Fig. 3d). Furthermore, and as observed in cell culture, wild-type viruses expressing YFP-only were converted to recombinant gene drive viruses expressing YFP and mCherry, representing 40% of the final population. The proportion of original gene drive viruses expressing CFP and mCherry increased slightly, from 15% to 20% in the TG, and 30% in the brain, respectively (Fig. 3e). Critically, in the control experiment with GD-ns, the proportion of gene drive viruses did not change and remained close to its initial value around 15% in all tissues, with a similar proportion of YFP+mCherry and CFP-only viruses appearing by CRISPR-independent recombination (Fig. 3e). Altogether, this important result showed that the gene drive efficiently spread in the viral population as the infection progressed to the brain, with recombinant gene drive viruses increasing from 15% to 70% in four days.
Gene drive propagation relies on efficient homologous recombination after CRISPR cleavage, but DNA double-strand breaks can also be repaired by non-homologous end joining (NHEJ), resulting in small insertions and deletions that render viruses resistant to the drive12,28. After PCR and Sanger sequencing of infectious viruses isolated from the brain, we found that 20% of the remaining target sites had been mutated by NHEJ (Fig. 3f). Since the gene drive represented 70% of viruses at this point, this result suggested that drive-resistant viruses represented around 6% of the total viral population. This indicated that the drive did not achieve full penetrance after four days, and confirmed the observation made with hCMV that mutagenic repair by NHEJ is infrequent compared to homologous recombination during gene drive spread11,12.
We next tested if our gene drive could spread in another infection model, where HSV-1 is inoculated in the hind leg footpad. In this model, HSV-1 travels to the spinal cord via the sciatic nerve, and finally to the brain. 106 PFU of WT, WT + GD or WT + GD-ns were inoculated in the right hind footpad, with an initial proportion of gene drive virus of 15%. Tissues were collected five to seven days later and analyzed by plaque assay (Fig. 3g–i). Around half the mice did not show any symptoms of infection and had no detectable virus in the spinal cord and the brain, while others developed severe neurological symptoms with very high titers in both tissues (Fig. 3g). In animals with detectable virus, the average proportion of gene drive viruses reached 60% in the brain, with a range between 30% and 80%, and 50% in the spinal cord (Fig. 3h). Once again, wild-type viruses expressing YFP-only had been converted to recombinant gene drive viruses expressing both YFP and mCherry, while the population of viruses expressing CFP and mCherry remained constant (Fig. 3i). Only one control animal infected with WT + GD-ns developed a detectable infection, with mCherry-expressing viruses reaching 35% in the spinal cord. These results confirmed that a gene drive could spread after inoculation via a second route of HSV-1 infection.
Altogether, these findings showed that a gene drive could propagate efficiently in vivo during acute HSV-1 infection in mice, and that the drive was mediated by a functional CRISPR system.
High heterogeneity during gene drive spread in the brain
HSV-1 travels to the brain via different neuronal pathways, and we hypothesized that following gene drive propagation in different regions of the brain could bring novel insight into the dynamics of HSV-1 infection and recombination. After intravitreal inoculation, HSV-1 infects retinal neurons and travels via the optic nerve (CN II) to the hypothalamus and the thalamus –both part of the interbrain– before reaching the cortex through visual pathways. Secondary branches of the optic nerve also connect to the midbrain –the rostral part of the brain stem. After infecting other ocular tissues and specifically the ciliary ganglion, HSV-1 separately reaches the midbrain via the oculomotor nerve (CN III). Finally, HSV-1 travels via the trigeminal nerve (CN V) to the TG and then to the brain stem (Fig. 4a).
a Infection routes following ocular inoculation of HSV-1. Male and female Balb/c mice were co-infected with 106 PFU of WT + GD in the left eye, with a starting proportion of gene drive virus of 15%. b Viral titers over time. Black lines indicate the median. n.d.: non-detected. c, d Proportion of gene drive viruses over time. Data show the average and SEM between biological replicates. e Heatmap summarizing panels (b and c). n = 4 mice for day 2 and 3, n = 6 mice for day 4. Source data are provided as a Source Data file.
To investigate tissue-specific differences in gene drive propagation, Balb/c mice were inoculated intravitreally with 106 PFU of WT + GD, with an initial proportion of gene drive virus of 15%. Viral titers and gene drive-directed recombination were measured by plaque assay in the eye, TG, brain stem, interbrain, cortex and cerebellum at days two to four post-infection (Fig. 4). Viral titers increased progressively throughout the brain, first reaching the interbrain and brain stem after 2 days and then spreading to the cortex and cerebellum (Fig. 4b and summary heatmap in Fig. 4e, upper panel). As described above, the proportion of gene drive viruses remained low in the eye and reached around 60% in the TG (Fig. 4c–e). Interestingly, gene drive levels varied greatly between the different brain regions. The gene drive reached 80% in the brain stem after only two days, and progressively increased to 65% in the interbrain and cerebellum, with a penetrance of up to 90% in some animals. By contrast, gene drive levels remained low in the cortex, at 25% on average. As observed above, in the TG, brain stem and interbrain –but not in the eye and cortex, viruses expressing YFP-only had been converted to recombinant gene drive viruses expressing both YFP and mCherry (Fig. 4d). Together, this showed that gene drive spread in the brain was highly heterogeneous, with almost complete penetrance in some regions and barely any in adjacent ones.
This heterogeneity gives an interesting insight into the dynamic of HSV-1 infection. Gene drive propagation relies on cellular co-infection and one intuitive hypothesis is that higher viral levels would increase the probability that cells are co-infected by several virions, and, thus, that recombination levels should positively correlate with viral titers. However, no correlation between viral titers and recombination was observed (Supplementary Fig. 3). For example, HSV-1 levels were the highest in the eye, but gene drive levels remained the lowest in this organ. Similarly, HSV-1 reached similar titers in the brain stem, interbrain and cortex, but almost no recombination occurred in the cortex while high levels were observed in the brain stem and interbrain (Fig. 4d). This shows that recombination does not simply correlate with viral titers, as could be expected, but is likely explained by other tissue-specific cellular or viral mechanisms.
High levels of cellular co-infection in the TG and the brain
The results described above suggest that cells are frequently co-infected by several virions during HSV-1 infection. Thus, we aimed to directly measure co-infection levels during HSV-1 infection, as this would provide important insight into the basic biology that supports gene drive propagation.
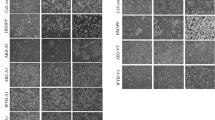
HSV-1 and related viruses expressing fluorescent proteins have long been used to probe neuronal pathways of the visual system29,30,31. To investigate cellular co-infection, Balb/c mice were infected ocularly with equal amounts of three different viruses, expressing either YFP, CFP or RFP from the same US1/US2 locus (Fig. 5a). The fluorescent reporters carried nuclear localization signals and were not incorporated into virions, and, thus, marked infected nuclei. Mice were injected intravitreally with a total of 106 PFU in the left eye and dissected four days later. Fluorescence was observed directly on frozen sections without staining. Strikingly, we observed very high levels of co-infection in the TG (Fig. 5) and different regions of the brain (Fig. 6), with cells often co-expressing two or more fluorophores.
a Balb/c mice were co-infected with equivalent amounts of three viruses expressing YFP, CFP and RFP, respectively, with a total of 106 PFU in the left eye. b. YFP and CFP cellular intensity after machine learning-assisted cell segmentation of TG sections. Datapoints represent individual cells and were colored by converting YFP and CFP signals into the CYMK color space. 4035 cells were detected, originating from 53 images and n = 4 mice. c Percentage of infected cells expressing YFP, CFP, or both. n = 4 mice. d Percentage of infected cells expressing one or two fluorescent markers. n = 4 mice. e, f Representative images of TG sections from four biological replicates, highlighting high levels of co-infection. Arrows indicate cells co-expressing YFP, CFP and RFP together. Scale bars: 100 μm. Panels (c and d) show the average and standard deviation (SD) between biological replicates. Source data are provided in Supplementary data 2 and as a Source Data file.
a Images of brain sections were collected in three regions in the thalamus, midbrain and brain stem after ocular infection. b YFP, CFP and RFP cellular intensity after machine learning-assisted cell segmentation of brain sections. Datapoints were colored by converting YFP, CFP and RFP signals into the CYMK color space. 10,028 cells were detected, originating from 95 images and n = 3 mice. c Percentage of infected cells expressing one, two, or three fluorescent markers, both in the whole brain and in specific subregions. n = 3 mice. Data show the average and SD between biological replicates. d Representative images of the brain in the thalamus (sections S1), midbrain (sections S2) and brain stem (section S3), from three biological replicates. e–h Representative images and summary of co-infection patterns in specific subregions. LGN: lateral geniculate nucleus; SC: superior colliculus; EW: Edinger–Westphal nucleus; TGN: trigeminal nerve nuclei. Scale bars: 100 μm. Source data are provided in Supplementary data 2 and as a Source Data file.
In the TG, YFP and CFP were expressed at high levels, but the RFP signal was weaker and rarely above background. Thus, for the TG, we restricted our analysis to YFP and CFP. We observed widespread expression distributed over the entire length of the TG on some sections, or more tightly localized clusters in others (Fig. 5e, Supplementary Fig. 4e). Areas with the strongest signal likely localized to the neuron-rich ophthalmic division of the TG, but the widely disseminated expression indicated that HSV-1 had spread to other areas. Importantly, a majority of infected cells appeared to express both YFP and CFP, indicating extensive cellular co-infection in the TG (Fig. 5e). To quantify co-infection precisely, we used machine learning to automatically segment cells and measure YFP and CFP intensity on TG sections (Fig. 5b–d, Supplementary Fig. 4a-d). A total of 4035 cells was detected, with a clear separation between YFP and CFP signals and high consistency between replicates. We found that an average of 52% of cells expressed both YFP and CFP, with a range of 45% to 60% between replicates. Since RFP was excluded from this analysis, this was probably an underestimate of the co-infection frequency. In fact, in the few images with strong RFP signal, we detected numerous cells co-expressing RFP, YFP and/or CFP, with instances of cells expressing all three markers (Fig. 5f, Supplementary Fig. 4e). Together, this showed that more than 50% of cells were co-infected by two or more virions in the TG.
We then repeated this analysis in the brain (Fig. 6, Supplementary Figs. 5–10). We detected strong fluorescence along well-characterized routes through the optic, oculomotor and trigeminal nerves (Fig. 6d, Supplementary Fig. 5). In particular, fluorescence was observed: 1) in the lateral geniculate nucleus (LGN) in the thalamus and the superior colliculus (SC) in the midbrain, where most axons of the optic nerve terminate; 2) in the Edinger–Westphal nucleus (EW), one of the two nuclei of the oculomotor nerve in the midbrain; and 3) throughout the hindbrain, likely corresponding to trigeminal nerve nuclei (TGN). Fluorescence was also detected in other areas associated with visual pathways such as the optic tract, olivary pretectal nucleus, suprachiasmatic nuclei or visual cortex (Supplementary Fig. 5). Additionally, disseminated fluorescence was detected over wide areas not easily associated with the visual system, with important variation between replicates. This likely corresponded to the secondary or tertiary spread of HSV-1 throughout the brain. Images were collected in three main regions in the thalamus, midbrain, and hindbrain (Fig. 6a). This time, RFP had a stronger signal and was included in the analysis. After machine learning-assisted segmentation, a total of 10,028 cells were analyzed over three brains. RFP was observed in 10-15% of infected cells, while YFP and CFP were detected in equivalent proportions in 40–60% of cells (Supplementary Fig. 6). When analyzing all images together, we observed a high level of co-infection. In total, 29% of cells expressed two colors, and 5% of cells had three colors, with results highly consistent between replicates (Fig. 6b, c and Supplementary Fig. 6). This confirmed the high level of co-infection found in the TG, with more than 34% of cells co-infected by two or more virions in the whole brain.
Next, we focused our analysis on the brain regions associated with primary HSV-1 spread, namely the LGN, SC, EW, and TGN. Interestingly, the LGN and SC, where the optic nerve terminates, had relatively low co-infection levels, around 20%. By contrast, the EW and TGN, where the oculomotor and trigeminal nerves terminate, respectively, had much higher co-infection levels, around 40% (Fig. 6c). This visually correlated with very different infection patterns. In the LGN and SC, well-separated and tight foci expressing only one color were observed, with co-infected cells at the boundaries (Fig. 6e, f closeups and additional examples in Supplementary Figs. 7 and 8). This was reminiscent of viral plaques in cell culture and may suggest clonal spread from a single infected cell. By contrast, in the EW and TGN, infected cells were broadly disseminated, with few infected cells touching each other. The different colors were uniformly distributed and cells with one, two, or three colors were observed without evidence of spatial clustering (Fig. 6g, h, Supplementary Figs. 9 and 10). Together with the high heterogeneity observed during gene drive spread (Fig. 4), these observations suggested that co-infection dynamics vary significantly depending on the viral propagation route.
Finally, we analyzed co-infection levels in the eye (Fig. 7, Supplementary Fig. 11). At this late stage of infection, HSV-1 replicated at very high titer and the eye retained poor physical integrity. Additionally, high background autofluorescence in the CFP and RFP channels made the interpretation of co-infection patterns challenging. Despite these caveats, we detected widespread infection in most ocular tissues, and in particular the retina and cornea (Fig. 7b, Supplementary Fig. 11a). In the retina, fluorescent reporters appeared physically separated, forming non-overlapping stripes spanning the width of the retina (Fig. 7c, Supplementary Fig. 11a). This suggested that HSV-1 spread from the upper to the deeper layers of the retina with limited amounts of co-infection. By contrast, we observed high levels of co-infection in a small posterior ocular structure that we identified as the ciliary ganglion (Fig. 7d–f, Supplementary Fig. 11b–g). Co-infection levels in the ciliary ganglion were similar to those observed in the TG, with more than 50% of cells infected by two or more virions, suggesting that high levels of co-infection may be favored in all ganglia.
a Following intravitreal inoculation, HSV-1 principally infects the retina and other ocular tissues such as the cornea, before invading the peripheral and central nervous system. In particular, HSV-1 infects the ciliary ganglion (CG) before propagating to the midbrain through the oculomotor nerve. b Representative image of an infected eye, showing infected cells in the retina, cornea and CG. The boxed area is shown in panel (c). c Representative image of the retina. Despite the high background fluorescence in the CFP and RFP channels, viruses expressing YFP, CFP and RFP appear restricted to non-overlapping areas. d Representative image of the ciliary ganglion (CG), showing high levels of co-infection. The CG was identified as a small structure close to the optic nerve in the posterior region of the eye, with clear fluorescent signals similar to other neuronal tissues in the TG or the brain. e YFP, CFP and RFP intensity in the CG after machine learning-assisted cell segmentation. 222 cells originating from 4 images and n = 2 mice. f Percentage of infected cells expressing one, two, or three fluorescent markers in the CG. n = 2. Scale bars: 100 μm. In panels (b, c, and d,) images are representative images from two biological replicates. Source data are provided in Supplementary data 2 and as a Source Data file.
Altogether, this analysis revealed that neuronal tissues sustain high levels of co-infection during HSV-1 infection, with more than 50% and 40% of cells infected with two or more viruses in peripheral ganglia and specific brain regions, respectively. The differences in co-infection patterns between tissues and brain regions are intriguing and will form the basis of future investigations.
Gene drive spread during latent infection
Our results showed that a gene drive could spread efficiently during acute infection. Next, we investigated if a gene drive could also spread in the context of a latent infection, as this would open important avenues for therapeutic interventions. After primary oro-facial infection, HSV-1 typically establishes latency in the TG and other peripheral ganglia. Following reactivation, HSV-1 travels back to the mucosal surface, causing lesions or shedding asymptomatically. In the following experiments, we used an ocular model of latent infection and drug-induced reactivation in mice to test if a gene drive virus –administered at a later time point, thus “superinfecting”– could target and recombine with latent HSV1-WT (Figs. 8 and 9).
a Experimental outline: Swiss-Webster mice were infected with 105 PFU of HSV1-WT on both eyes after corneal scarification. Four weeks later, mice were superinfected with 107 PFU of GD or GD-ns on both eyes, after corneal scarification. Another four weeks later, latent HSV-1 was reactivated twice with JQ1, two weeks apart. n = 14 mice per group. b Titer and number of shedding events in eye swabs on days 1–3 following JQ1 treatment, by qPCR. Shedding events from the same mouse are connected by a line. c Genotyping of positive eye swabs from five mice, using two duplex ddPCR assays. The first assay detected and quantified mCherry levels. The second assay distinguished between YFP and CFP. n = 8. d Number and proportion of TG and mice with detectable CFP. e Latent viral load in the TG by duplex ddPCR, detecting mCherry, YFP, CFP markers, or all HSV sequences. n = 28. f Proportion of CFP and mCherry in the TG. n = 28. Titers are expressed in log-transformed copies per swab, or per million cells after normalization with mouse RPP30 levels. Black lines indicate the median. n.d.: non-detected. Source data are provided as a Source Data file.
C57Bl/6 mice were infected with 106 PFU of HSV1-WT on both eyes after corneal scarification. Four weeks later, mice were superinfected with 107 PFU of GD or GD-ns on both eyes, after corneal scarification. After another four weeks, latent HSV-1 was reactivated three times, two weeks apart, with JQ1 and Buparlisib. n = 26 for GD, n = 18 for GD-ns. a Titer and number of shedding events in eye swabs on days 1–3 following JQ1 treatment, by qPCR. Shedding events from the same mouse are connected by a line. b Genotyping of positive eye swabs, detecting mCherry, YFP and CFP markers. Light red indicates low mCherry levels, less than 5% of the total swab titer. Swab genotypes were assessed by ddPCR and confirmed by qPCR for low-titer swabs. Details are shown in Supplementary Fig. 14. c Number and proportion of TG and mice with detectable CFP marker from GD/GD-ns. d, e Latent viral load and proportion in the TG. n = 52 for GD, n = 36 for GD-ns. Black lines indicate the median. f mCherry as a function of CFP. Data was fitted either to the best possible line or to the identity line, and the fits were compared using the extra sum-of-squares F test. GD datapoints were significantly higher than the identity line (p < 0.0001), suggesting gene drive propagation in the TG. Same data as panel (d), excluding non-detected samples. n = 38 for GD, n = 23 for GD-ns. g Log2 fold-change between the proportion of latent mCherry and CFP, using data from panel (e). Samples with low levels of less than 0.5% were excluded. Black lines show the average. Asterisks summarize the results of two-sided Welch’s t-test (p = 0.0053). n = 33 for GD, n = 20 for GD-ns. Titers are expressed in log-transformed copies per swab, or per million cells after normalization with mouse RPP30. n.d.: non-detected. Source data are provided as a Source Data file.
To establish a latent infection, Swiss-Webster mice were infected ocularly with HSV1-WT, with 105 PFU in both eyes after corneal scarification. Four weeks later, animals were inoculated with GD or control GD-ns after corneal scarification (Fig. 8a, Supplementary Fig. 10a, b). Immune responses induced by the primary infection limit the spread of a superinfecting virus, allowing mice to be safely inoculated with 107 PFU per eye of GD or GD-ns while being transiently immunosuppressed with the glucocorticoid dexamethasone. After four more weeks and once the second infection had resolved, latent HSV-1 –originating from the primary, secondary infection, or both– was reactivated twice by treating animals with the bromodomain inhibitor JQ1. Mice do not naturally reactivate HSV-1, but JQ1 induction reproducibly induces viral shedding32,33. Eye swabs collected on days 1-3 following JQ1 treatment were screened for viral DNA by qPCR. We observed overall low shedding rates. A total of 10 shedding events were detected, with some animals shedding on consecutive days (Fig. 8b). Gene drive-directed recombination was measured in positive swabs using two duplex digital droplet (dd)PCR assays, that distinguish between the YFP, CFP and mCherry markers of WT and gene drive viruses and quantify the proportion of gene drive recombinants. Eight out of ten positive swabs could be successfully genotyped, with the gene drive sequence detected in three of them (Fig. 8c, Supplementary Fig. 12g). Critically, one of the reactivated viruses (from mouse #24) was a recombinant gene drive virus carrying YFP and mCherry markers, whereas two others carried CFP and mCherry, representing the original gene drive virus. Despite the limited number of shedding events, this showed that GD and GD-ns could successfully reach the latent reservoir and later reactivate, with one recombination event detected.
Next, both TG from each animal were collected and latent viral loads were measured by duplex ddPCR. The CFP marker originating from the superinfecting GD or GD-ns viruses could be detected in around 40% of TG, with 60% of mice having detectable CFP in at least one TG (Fig. 8d). When detected, GD and GD-ns viral loads were approximately two orders of magnitude lower than WT, as measured by the respective titers of YFP and CFP/mCherry (Fig. 8e). Overall, GD and GD-ns represented between 0 and 60% of the total latent viral load (average around 5%, median at 0%), with most detected samples ranging from 1% to 10% (Fig. 8f). This low proportion contrasted with the relatively high frequency of gene drive sequences detected in reactivated swabs (3/8, or 37%), suggesting that the superinfecting GD/GD-ns viruses could successfully reactivate and shed despite representing only a small proportion of the latent reservoir.
Swiss Webster mice are highly susceptible to HSV-1 infection. During primary infection, mice exhibited moderate to severe symptoms, with often extensive facial lesions. As a result, most animals had residual scar tissue on their eyes once the primary infection had resolved. Symptoms and final eye scarification were scored during the primary infection, and mice separated into groups with homogenous symptoms and eye scores before superinfection with GD or GD-ns (Supplementary Fig. 12c, d). Importantly, at the end of the experiment, GD and GD-ns were detected almost exclusively in the TG of mice with perfect eyes, with even low levels of scarification preventing successful superinfection (Supplementary Fig. 12e). HSV-1 typically does not cause long-lasting scars in humans, and this residual scar tissue represented an unfortunate confounding factor in our mouse model.
To alleviate this effect, we repeated the experiment in C57Bl/6 mice (Fig. 9). C57Bl/6 are more resistant to HSV-1 infection and typically experience minimal symptoms. C57Bl/6 mice were infected ocularly with HSV1-WT, with 106 PFU in both eyes after corneal scarification. Compared to Swiss Webster and despite the higher dose, mice exhibited limited symptoms and reduced mortality during primary infection. Around 20% of mice had residual scar tissues, usually on only one eye (Supplementary Fig. 13a–d). Four weeks later, mice were superinfected with 107 PFU per eye of GD or GD-ns, while being transiently immunosuppressed with dexamethasone and tacrolimus. Then, another four weeks later, mice were injected with JQ1 and Buparlisib to reactivate latent HSV-1. Buparlisib is a phosphoinositide 3-kinase inhibitor and evidence suggested that it could improve HSV-1 reactivation (34,35 and Supplementary Fig. 13e). Reactivation rates ranged from 0 to 33% (Fig. 9a). Most events were close to the detection limit (around 100 copies/swab) and we could successfully obtain viral genotypes from 17 out of 22 reactivation events (Fig. 9b, Supplementary Fig. 14). Nine out of 17 swabs (53%) were either recombinant viruses carrying mCherry and YFP, or original gene drive viruses with mCherry and CFP. In addition, two swabs were genotyped as predominantly WT (from mice #25 and #33), but had detectable amounts of mCherry, representing less than 5% of the total titer (Fig. 9b, Supplementary Fig. 14). In one positive swab with a low viral titer (mouse #45), both YFP and CFP markers, originating from the same US1/US2 locus, were detected, suggesting a mix of reactivated viruses. Importantly, we detected recombinants expressing YFP and mCherry in mice superinfected with either GD (1/9) or GD-ns (3/8), showing that both viruses could recombine with HSV1-WT. Because of the overall low number of reactivation events, further statistical analysis could not be conducted. In summary, we found that gene drive viruses represented more than 50% of reactivation events, with several examples of recombination with HSV1-WT.
Next, TG were dissected and latent viral loads were measured by duplex ddPCR. GD and GD-ns viruses could be detected in around 79% and 66% of TG, respectively, with more than 90% of mice having detectable GD or GD-ns in at least one TG (Fig. 9c). GD and GD-ns titers were one to two orders of magnitude lower than WT (Fig. 9d), and represented a small percentage of the total latent viral load, ranging from 0.5% to 50% for most samples (average at 10% and median around 1%, Fig. 9e). We investigated whether we could detect evidence of gene drive spread in the TG. In particular, we determined whether mCherry was significantly overrepresented compared to the CFP baseline, as the gene drive cassette containing mCherry potentially recombined with HSV1-WT and increased in frequency. Of note, we observed no correlation between the viral loads in the left and right TG collected from the same animals (Pearson correlation, r2 = 0.11), which allowed us to treat all samples as independent replicates in these analyzes (Supplementary Fig. 15a). When plotting the titers of mCherry versus CFP, we observed that GD datapoints were significantly above the identity line (p < 0.0001), while GD-ns datapoints were mostly situated along the line and did not significantly deviate from it (Fig. 9f, statistical analysis in Supplementary Fig. 15b). This suggested that the gene drive had recombined with wild-type viruses and increased in frequency in the TG of GD-superinfected mice. To quantify this enrichment, we calculated the fold change between the proportion of mCherry and CFP in the TG (Fig. 9g). On average, in the GD-superinfected samples, we found a 70% increase in the relative proportion of mCherry compared to CFP, which was significantly higher than the control samples with GD-ns where no enrichment was observed (p = 0.0053, t-test). In the most extreme cases, the gene drive had spread more than 10-fold over the CFP baseline, for example increasing from 9% to 90% in one sample, or from 1.3% to 10% in another (Supplementary Fig. 15c). Together, this analysis showed a limited but statistically significant spread of the gene drive in the latent wild-type population.
In summary, we found that a superinfecting gene drive virus could reach the latent reservoir and spread in the wild-type population. The immune response induced by the primary infection likely limited the spread of the second infection and the drive did not reach full penetrance. However, in both Swiss-Webster and C57Bl/6 mice, GD and GD-ns were detected in more than 50% of shedding events after JQ1 reactivation. This indicated that the superinfecting GD and GD-ns viruses could successfully reactivate and shed despite representing only a small proportion of the latent reservoir.
Discussion
In this manuscript, we designed a gene drive against HSV-1 and showed that it could spread efficiently in cell culture and in vivo. In particular, we observed high levels of co-infection and gene drive-directed recombination in neuronal tissues during herpes encephalitis in mice. Furthermore, we found evidence that a superinfecting gene drive virus could recombine with HSV1-WT during latent infection. Altogether, this work presented an important proof-of-concept and showed that a gene drive could spread in vivo during acute and latent infection.
Our cell culture results recapitulated our previous work with hCMV and aligned with the findings of a recent study11,28. This showed that a gene drive could be designed in a second herpesvirus with very different infection dynamics. This suggests that viral gene drives could be developed in a wide variety of herpesviruses, which significantly expands the potential of the technology.
In co-infection experiments, both in cell culture and in vivo during acute infection, wild-type viruses expressing YFP-only were efficiently converted to recombinant gene drive viruses expressing YFP and mCherry, in a CRISPR-dependant manner. Interestingly, the proportion of original gene drive viruses expressing CFP and mCherry also increased slightly, from 20% to 40% in cell culture, and from 15% to 30% in the brain, respectively (Figs. 2e, 3e). This increase was observed only in experiments with GD and not in control experiments with GD-ns. Since WT, GD and GD-ns replicated with similar dynamics (Figs. 2a, 3b), this cannot be explained by a competitive advantage of one virus over the next. We had made a similar observation in our earlier work with hCMV11, suggesting that this small increase in the population of the original gene drive virus may be a constitutive feature of gene drive propagation. It may reflect complex intracellular processes occurring during viral recombination that were imperfectly captured by the three-color system used throughout our studies (Supplementary Fig. 16).
The spread of a viral gene drive relies on the co-infection of cells by wild-type and engineered viruses. High co-infection levels can easily be achieved in cell culture, either by using cell lines naturally susceptible to co-infection such as N2a cells, or by infecting cells at a high MOI to bypass restriction mechanisms28. Indirect observations in animal studies and recombination patterns in HSV strains circulating in humans indicate that co-infection events take place in vivo15,16,17,18,19,20,21,22. However, whether these co-infection events occur at a high frequency was unknown. Here, we showed that a gene drive could spread efficiently during acute infection in mice, with the level of gene drive-directed recombination increasing from 15% to 80% in some brain regions after only four days. Using fluorescent reporters, we directly observed high levels of co-infection in peripheral ganglia and the brain, with more than 50% of cells infected by two or more virions in some regions. These findings revealed that HSV-1 achieves high rates of co-infection and recombination during viral spread. Furthermore, we showed that co-infection and recombination levels varied depending on the cranial nerves used to travel to the brain. For example, co-infection and recombination were low in regions accessed by the optic nerve and much higher in regions associated with the oculomotor and trigeminal nerves (Figs. 4–7). The oculomotor and trigeminal nerves transit through the ciliary and trigeminal ganglia, respectively, and we hypothesize that these ganglia could serve as convergence zones for the virus, leading to high levels of co-infection and gene drive propagation in regions downstream of the ganglia, such as the brain stem. Conversely, the low level of gene drive recombination in the eye correlated with low levels of co-infection in the retina, and the low level of recombination in the cortex correlated with limited co-infection upstream in the viral propagation route. Propagation to the cortex originates in the retina and transits through the optic nerve and the thalamus. Co-infection was limited in the retina and the thalamus, suggesting that this route of infection may present few opportunities for co-infection and recombination. Together, this suggests that the heterogeneity in gene drive propagation reflects different levels of co-infection depending on the infection route, and that co-infection frequency is the main factor limiting gene drive spread.
Herpes encephalitis is a severe but rare condition, and HSV disease is more often characterized by chronic lesions on the facial and genital area. We showed that a gene drive virus could reach the latent reservoir and spread in a limited manner in the wild-type population (Figs. 8 and 9). This limited penetrance was likely caused by immune responses against HSV-1 induced by the primary infection, thus restricting the propagation of the superinfecting gene drive virus during latent infection. However, gene drive viruses represented around half of the shedding events after drug-induced reactivation, despite these viruses representing a much smaller proportion of the latent viral load. This finding has important implications. It suggests that shedding viruses originated from only a small fraction of the latent reservoir and that a superinfecting gene drive could efficiently reach the physiologically relevant neurons that seed viral reactivation. Mice do not shed HSV-1 spontaneously and our study was limited by the low reactivation rates. Further studies in animals that better recapitulate human disease, such as rabbits or guinea pigs, will be necessary to thoroughly investigate the potential of a gene drive during chronic infection.
This research represents an important milestone toward therapeutic applications. Our future work will focus on designing gene drives that can limit infectivity and reduce disease severity. Interestingly, we noticed that the titer of reactivated gene drive viruses was significantly lower than the titer of wild-type ones (Supplementary Fig. 14c). Insertion of the gene drive sequence did not affect infectivity in cell culture or during acute infection. However, this unexpected observation suggested that the gene drive may have hampered reactivation or shedding. This could occur either directly if insertion of the gene drive had genetically impaired HSV-1, or indirectly through immune or other host-mediated effects. Similar approaches, where the gene drive virus can reach the latent reservoir efficiently but then suppress reactivation, will form the basis of future strategies. Together, our work may pave the way toward new therapeutics for HSV diseases.
As a final note, the development of gene drives in mosquitoes and other insects has generated important ecological and biosafety concerns14,36. Our approach followed the guidelines established by the NIH and the National Academy of Science37,38,39. In the future, the risks and benefits of viral gene drives will need to be properly addressed and discussed with the scientific community.
Methods
These studies and all procedures were approved by the Institutional Biosafety Committees and Institutional Animal Care and Use Committees of the Buck Institute and Fred Hutchinson Cancer Center. Our research followed the guidelines on gene drive research established by the NIH and the National Academy of Science37,38,39.
Cells and viruses
African green monkey epithelial Vero cells and murine neuroblastoma N2a cells were obtained from the ATCC and cultured in DMEM (Corning, Corning, NY, USA) supplemented with 10% FBS (Sigma-Aldrich, St-Louis, MO, USA). Viral infections and plaque assays were performed using DMEM supplemented with 2% FBS. Cells were maintained at 37 °C in a 5% CO2 humidified incubator and frequently tested negative for mycoplasma contamination.
Unmodified HSV-1 strain 17+ and HSV1-CFP expressing cyan fluorescent protein mTurquoise2 from the US1/2 locus15 were provided by Matthew P Taylor (Montana State University, USA). Viruses generated for this study were made by modifying HSV-1 and HSV1-CFP, as described below. To prepare viral stocks for cell culture experiments, Vero cells in 15 cm dishes were infected for one hour at MOI = 0.01, and kept in culture for 48 hours or until 100% cytopathic effect was observed. Cells and supernatant were scraped out of the plate, sonicated three times at maximum power with a probe sonicator, and debris pelleted away by centrifugation (500 × g rpm, 10 minutes, 4 °C). Media containing viruses was collected in single-use aliquots and titers measured by plaque assay.
For high-titer and high-purity viral stocks used for animal experiments, Vero cells in 15 cm dishes were infected for one hour at MOI = 0.01, and kept in culture for 48 hours or until 100% cytopathic effect was observed. Supernatants and cells were collected, and cells were pelleted by centrifugation (500 × g, 5 minutes, 4 °C). Supernatants were collected in clean tubes and reserved for later. Cell pellets were resuspended in a small volume of culture media and cell-bound virions were released by two cycles of freeze-thaw in dry ice. Debris were pelleted again and the supernatant containing released virions was combined with the supernatant reserved earlier. Virions were then pelleted by ultracentrifugation (22,000 rpm, 90 min, 4 °C, Beckman-Colter rotor SW28) on a 5-mL cushion of 30% sucrose. Supernatants were discarded, and virions were resuspended in PBS containing 2% BSA. Single-use aliquots were prepared and titers were measured by plaque assay.
Co-infection experiments were performed in 12-well plates by co-infecting N2a cells with HSV1-WT and gene drive viruses for 1 h, with a total MOI of 1, before replacing inoculum with 1 mL of fresh medium. 100uL of supernatant was collected at regular intervals and analyzed by plaque assay.
Cloning and generation of recombinant viruses
A donor plasmid containing the gene drive cassette against the HSV-1 UL37-38 intergenic region (GD and derivatives) was generated by serial modifications of the GD-mCherry donor plasmid used in our previous study11. All modifications were carried out by Gibson assembly (NEB, Ipswich, MA, USA), using PCR products from other plasmids or synthesized DNA fragments (GeneArt String fragments, ThermoFisher, USA). The final GD donor plasmid included homology arms for the UL37-38 region, the CBH promoter driving SpCas9 followed by the SV40 polyA terminator, the CMV promoter driving an mCherry reporter followed by the beta-globin polyA signal, and the U6 promoter controlling gRNA expression. The functional GD plasmid carried a gRNA targeting the UL37-38 region (ACGGGATGCCGGGACTTAAG), while the non-specific GD-ns control targeted a sequence absent in HSV-1 (ACATCGCGGTCGCGCGTCGG). GD-ΔCas9 donor construct was subsequently generated by removing SpCas9 by digestion and ligation. Donor constructs to insert CMV-driven yellow (YFP) or red (RFP) fluorescent protein reporters into the US1/US2 locus were built similarly, by replacing mTurquoise with mCitrine2 or mScarlet2 in a donor plasmid for the US1/US2 region, respectively (pGL002, from ref. 15). Of note, the YFP, CFP and RFP reporters carried a nuclear localization signal.
To build recombinant viruses, 1.5 million Vero cells were co-transfected with linearized donor plasmids and purified HSV-1 strain 17+ or HSV1-CFP viral DNA. Viral DNA was purified from infected cells by HIRT DNA extraction, as described previously11. Transfection was performed by Nucleofection (Lonza, Basel, Switzerland) and cells were plated in a single 6-well. After 2–4 days, mCherry-expressing viral plaques were isolated and purified by several rounds of serial dilutions and plaque purification. Purity and absence of unmodified viruses were assayed by PCR and Sanger sequencing after DNA extraction (DNeasy kit, Qiagen, Germantown, MD, USA). Viral stocks were produced as specified above and titered by plaque assay.
Plaque assay
Plaque assays were performed either directly from cell culture supernatants or from frozen mouse tissues. To release infectious virions from tissues, frozen samples were resuspended in cell culture media and disrupted using a gentle tissue homogenizer (Pellet Pestle, Fisher Scientific, USA). Samples were sonicated three times at maximum power with a probe sonicator, and debris pelleted away by centrifugation (500 × g, 10 minutes, 4 °C). Volumes were adjusted to a final volume of 1 mL, and titers were measured by plaque assay.
Viral titers and recombination levels were determined by plaque assay with 10-fold serial dilutions. Confluent Vero cells in 24-well plates were incubated for 1 h with 100uL of inoculum, and overlaid with 1 mL of complete media containing 1% methylcellulose, prepared using DMEM powder (ThermoFisher, USA), methylcellulose (Sigma-Aldrich, USA) and 2% FBS. After two or three days, fluorescent plaques expressing YFP, CFP and/or mCherry were manually counted using a Nikon Eclipse Ti2 inverted microscope. Every viral plaque was analyzed on YFP, CFP and red channels. 5–100 plaques were counted per well, and each data point was the average of 3–4 technical replicates (i.e., 3–4 different wells). Images of fluorescent viral plaques were acquired with an EVOS automated microscope (ThermoFisher, USA), and adjusted for contrast and exposure with ImageJ (v2.14.0/1.54 f).
The deconvolution of Sanger sequencing in Fig. 3f was performed using Synthego ICE online tools (https://ice.synthego.com).
gRNA design and amplicon sequencing of putative off-target sites
The gRNA targeting the HSV-1 UL37-UL38 intragenic region was designed to minimize potential off-target editing of the mouse genome, using the online design tool CRISPOR v5.240 (http://crispor.gi.ucsc.edu/). The list of potential off-target sites, none with less than three mismatches, was downloaded and ranked according to their “Cutting Frequency Determination” (CFD) specificity score41. The 14 sites with CFD scores > 0.1 were analyzed by amplicon sequencing. N2a cells were infected with GD and GD-ns at MOI = 1 and cells collected after 3 days. DNA was extracted using Qiagen DNeasy kit. PCR amplicons, ranging from 180 to 300 bp for the 14 sites, were amplified using Phusion Plus high-fidelity polymerase (ThermoFisher, USA) on a Biorad Thermocycler. Amplicons were barcoded and libraries prepared using Illumina DNA Prep Kit (Illumina, USA). Barcoded amplicons were sequenced on an Illumina Nextseq 2000 using 2 × 150 read format. Demultiplexing was performed using BCLConvert (Illumina) to generate raw fastqs. Reads were trimmed and aligned on the amplicon sequences using BWA. Mutation rates at the target site were analyzed using software developed previously42 and available on Github https://github.com/proychou/TargetedMutagenesis). The list and location of off-target sites, PCR primers, amplicon sequences and a summary of the sequencing results are presented in Supplementary data 1.
Mouse experiments
All animal procedures were approved by the Institutional Animal Care and Use Committee of the Buck Institute and Fred Hutchinson Cancer Center, under protocol numbers A10252 and 1865, respectively. This study was carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health (“The Guide”). Standard housing, diet, bedding, enrichment, and light/dark cycles were implemented under animal biosafety level 2 (ABSL2) containment.
Acute infection after intravitreal inoculation
Acute infections were performed at the Buck Institute. Male and female Balb/c mice were purchased from Charles River Laboratories and bred in house. Between five and eight weeks-old animals were infected by intravitreal injection in the left eye, as described previously43,44. Briefly, mice were anesthetized by intraperitoneal injection of ketamine (100 mg/kg) and xylazine (10 mg/kg) and laid prone under a stereo microscope. The left eye was treated with a thin layer of veterinary ophthalmic ointment and the sclera was exposed using ophthalmic forceps. 2 μl of HSV-1 stock containing 106 pfu was injected slowly in the intravitreal space, using a 5uL Hamilton syringe and a 30 gauge needle. In the days following infection, mice were treated with sustained-release buprenorphine to minimize pain (Ethiqa XR, Fidelis Animal Health, North Brunswick, NJ, USA). Animals were humanely euthanized after two to four days. For plaque assay analysis, tissues were collected and snap-freezed in liquid nitrogen.
Latent infection after corneal scarification
Latent infections were performed at the Fred Hutch Cancer Center, using female Swiss-Webster or C57bl/6 mice five to six weeks old purchased from Taconic (Germantown, NY, USA). Mice were anaesthetized by intraperitoneal injection of ketamine (100 mg/kg) and xylazine (10 mg/kg) and laid under a stereo microscope. Mice corneas were lightly scarified using a 28-gauge needle, and 4uL of viral inoculum was dispensed on both eyes. Swiss-Webster and C57bl/6 mice were infected with 105 and 106 PFU, respectively. Following inoculation, ophthalmic drops of local analgesic (Diclofenac) were deposited on both eyes, and the analgesic Meloxicam was added to the drinking water ad libitum for 1-5 days following infection. From five to fifteen days following primary infection, symptoms of infection were reported and scored using an in-house scoring system. Mice experiencing severe symptoms were humanely euthanized. Once the infection had fully resolved, final eye scarification levels were scored in both eyes and averaged, using the following scores: 0: perfect eye; 1: lightly damaged and/or cloudy cornea, 2: scar tissue covering a small portion of the eye; 3: scar tissue covering most of the eye; 4: extremely bad-looking eye, fully blind.
The second infection with GD and GD-ns was performed the same way four weeks after the primary infection, with 107 PFU per eye. Mice were transiently immunosuppressed with dexamethasone (Fig. 8 in Swiss-Webster) or with dexamethasone and tacrolimus (Fig. 9 in C57bl/6). Tacrolimus and Dexamethasone were diluted in the drinking water and administrated ad libitum from one day before to seven days after infection. Drug concentration was calculated according to the average mice weight, considering that mice drink around 5 mL per day, in order to reach a dose of 1 mg/kg/day and 2 mg/kg/day for dexamethasone and tacrolimus, respectively. For example, for an average mouse weight of 25 g, dexamethasone and tacrolimus were diluted at 5 mg/L and 10 mg/L, respectively. Of note, drugs were diluted in medidrop sucralose (ClearH20, Westbrook, ME, USA) instead of regular drinking water, to make tacrolimus more palatable to mice.
HSV reactivation
HSV reactivation was performed by intraperitoneal injection of JQ1 (MedChemExpress HY-13030, NJ, USA) at a dose of 50 mg/kg, and, when indicated, Buparlisib (MedChemExpress HY-70063, NJ, USA) at a dose of 20 mg/kg. JQ1 and Buparlisib were prepared from stock solutions (10x at 50 mg/mL and 100x at 200 mg/mL, in DMSO, respectively) by dilution in a vehicle solution of 10% w/v 2-hydroxypropyl-β-cyclodextrin (Sigma-Aldrich, St-Louis, MO, USA) in PBS. In the C57bl/6 experiments (Fig. 9), mice were transiently immunosuppressed with dexamethasone and tacrolimus in the drinking water, at the concentrations indicated above, from one day before to three days after JQ1/Buparlisib injection. Mouse eyes were gently swabbed with cotton swabs moistened with PBS, on day one to three following injection. Swabs were collected into vials containing 1 ml of digestion buffer (KCL, Tris HCl pH8.0, EDTA, Igepal CA-630) and stored at 4 °C before DNA extraction.
HSV quantification of viral loads in swabs and tissues
HSV quantification of eye swabs by qPCR
DNA was extracted from 200 μl of swab digestion buffer using QiaAmp 96 DNA Blood Kits (Qiagen, Germantown, MD, USA) and eluted into 100 μl AE buffer (Qiagen, Germantown, MD, USA). 10 μl of eluted DNA was used to set up 30 μl real-time Taqman quantitative PCR reactions, using QuantiTect multiplex PCR mix (Qiagen, Germantown, MD, USA), using the following PCR cycling conditions: 1 cycle at 50 °C for 2 minutes, 1 cycle at 95 °C for 15 minutes, and 45 cycles of 94 °C for 1 minute and 60 °C for 1 minute. Exo internal control was spiked into each PCR reaction to monitor inhibition. A negative result was accepted only if the internal control was positive with a cycle threshold (CT) within 3 cycles of the Exo CT of no template controls. Primers and probes have been described previously45 and are provided in Supplementary Table 1. Swabs positive for HSV were then further analyzed by ddPCR, as described below.
Duplex digital droplet PCR (ddPCR) of tissues and swabs
Total genomic DNA was isolated from ganglionic tissues using the DNeasy Blood and tissues kit (Qiagen, Germantown, MD, USA) and eluted in 60 μl of EB buffer, per the manufacturer’s protocol.
Quantification of the YFP, CFP and mCherry markers, as well as total HSV viral load was measured with two separate duplex ddPCR, using 10uL of eluted DNA. ddPCR was performed using the QX200 Droplet Digital PCR System and ddPCR Supermix for Probes (No dUTP) from Biorad (Hercules, CA, USA), following the manufacturer’s instructions. Primers were used at a final concentration of 900 nM and probes at 250 nM (Supplementary Table 1). Primers and probes were ordered from IDT (Coralville, IA, USA), using IDT custom PrimeTime ZEN double-quenched qPCR probes, with FAM and HEX fluorescent dyes. The first duplex assay used two sets of primers/probes to quantify mCherry (HEX probe) and HSV UL38 gene (FAM probe). UL38 primers/probe set was located in the UL38 viral gene and recognized both wild-type and gene drive genomes. The second duplex assay distinguished between YFP and CFP, using one set of common primers amplifying both markers and YFP and CFP-specific probes with FAM and HEX dyes, respectively. Primer specificity and sensitivity were validated on plasmid DNA before use in mouse samples. A limit of detection of three copies per reaction was used throughout the study, except in Fig. 8d–f, where a cutoff of 10 was applied to mCherry to account for a small PCR contamination. Final titers were normalized and expressed in log-transformed copies per million cells (MCells). Cell numbers in tissue samples were quantified by ddPCR using a mouse-specific RPP30 primer/probe set42.
The duplex assays allowed us to determine the proportion of latent mCherry and CFP, using absolute values measured in the same PCR reaction, thus, limiting technical variation. Proportions were calculated as follows:
Swab genotyping
Positive swabs identified by qPCR were analyzed using the duplex ddPCR assays described above. Samples expressing YFP only with no detectable CFP and mCherry were categorized as wild-type. Swabs expressing mCherry at the same level as UL38 were categorized as gene drives. Swabs expressing both CFP and mCherry represented the original GD/GD-ns, while swabs expressing both YFP and mCherry represented recombinants (Supplementary Figs. 12g, 14). Some swabs expressed mCherry one to two orders of magnitude lower than HSV and were genotyped as wild-type but with detectable amounts of mCherry. For low-titer swabs, the genotype was further confirmed by duplex qPCR using the same primers.
Brain and TG imaging and image analysis
Tissue processing and image collection
Balb/c mice were infected ocularly with equivalent amounts of three viruses expressing either YFP, CFP or RFP from the US1-US2 locus. A total of 106 PFU was inoculated intravitreally in the left eye. Of note, the fluorescent proteins carry nuclear localization signals. They are expressed in infected cells and are not incorporated into virions. Four days after infection, mice were injected intraperitoneally with a terminal dose of euthanasia solution containing Sodium Pentobarbitol (Euthasol). Once unresponsive, mice were subjected to thoracotomy and transcardially perfused with PBS followed by 4% Paraformaldehyde-Lysine-Periodate solution (PLP) through the aorta to fix tissues46. Brains, TG, and eyes were dissected, fixed in PLP overnight, and transferred to a 20% sucrose solution for 24 hours, and finally to a 30% sucrose solution for at least 24 hours for cryo-protection. All tissues were stored in 30% sucrose before processing. This protocol was a courtesy of J.P. Card44.
TG and eyes were embedded in OCT and serial sections of 15 µm made using a Cryostat (Zeiss) at -20 °C. TG sections were mounted on subbing solution-treated slides and polymerizing mounting media containing DAPI (Vectashield, Vector Labs, Burlingame, CA, USA) was added before coverslipping. Brains were immobilized in 30% sucrose on a freezing microtome stage set to -18 °C (Physitemp, Clifton, NJ). Serial coronal sections at 30 µm on a horizontal sliding microtome (AO Optical) were collected. Brain sections were binned into six parallel groups. One bin was arranged and mounted on slides before counterstaining with polymerizing mounting media containing DAPI and coverslipping. Epifluorescence imaging was performed on a Nikon Ti-Eclipse (Nikon Instruments, Melville, NY, USA) inverted microscope equipped with a SpectraX LED (Lumencor, Beaverton, OR, USA) excitation module and fast-switching emission filter wheels (Prior Scientific, Rockland, MA, USA). Fluorescence imaging used paired excitation/emission filters and dichroic mirrors for DAPI, CFP, YFP and TRITC (Chroma Technology Corp., Bellow Falls, VT, USA). All images were acquired with an iXon 896 EM-CCD (Andor Technology LTD, Belfast, NI, USA) camera using NIS-Elements software. Image tiles with 4x and 10x Phase objectives were acquired to assess fluorescent protein expression across sectioned tissues. Specific regions were imaged with the 20x ELWD to acquire detailed localization and fluorescent protein expression images for subsequent data analysis.
Image analysis
From the five animals originally infected, one animal was not included as very few infected cells could be detected in the brain and TG, suggesting that the infection had failed. Furthermore, one brain and two eyes from the remaining four mice was irremediably damaged during processing. Thus, the analysis was conducted on four TG, three brains and two eyes.
Using ImageJ (v2.14.0/1.54 f), Nd2 images acquired with Nikon NIS-Element software were batch converted into tiff files using an ImageJ macro (modified from https://github.com/singingstars/). During the conversion process, brightfield and DAPI channels were discarded, and ImageJ background subtraction was performed on the YFP, CFP and RFP channels. An additional grayscale channel was created, composed of the maximum projection of the YFP, CFP and RFP channels. This composite channel contained every cell irrespective of the original colour and was used for segmentation (Supplementary Fig. 4a). Machine learning-assisted segmentation was performed using an online analysis tool from www.biodock.ai (Biodock, AI Software Platform. Biodock 2023). The software was trained to recognize cells on the grey channel using a few training images, and segmentation was then run on the entire dataset. Around 3–4% of cells with aberrant area or eccentricity were discarded, and average signal intensity was measured in the original YFP, CFP and RFP channels for each detected cell. Of note, for the TG, this analysis was performed using only the YFP and CFP channels. Data was then further processed and plotted using R (RStudio v2023.09.1 + 494). For YFP and CFP, the intensity was simply log10 converted. Because RFP had a higher background and different intensity ranges across images, RFP intensity was first scaled across images, and then log10 converted. Stringent intensity thresholds were applied on the three channels and used to quantify cells infected with one, two, or three viruses, with around 5% of cells below thresholds being discarded (Supplementary Fig. 6b). Thresholds were chosen stringently to unequivocally identify co-infected cells, and are reported in Supplementary Figs. 4c and 6c, 11e. For visualization and plotting, signal intensities in the YFP, CFP and RFP channels were converted into CYMK color space. Source data are provided in Supplementary data 2.
Representative images shown in the manuscript were minimally adjusted for contrast and exposure using ImageJ. Some images were rotated or flipped horizontally to consistently present the brain in a caudal direction, with the left hemisphere on the right side.
Statistics and reproducibility
Experiments were carried out in multiple replicates. Investigators were blinded when performing plaque assays, collecting swabs, and analyzing DNA samples. No data was excluded, except when indicated in the main text, methods or figure legends. Statistical analyzes were performed using GraphPad Prism version 10.1.1 for macOS (GraphPad Software, USA, www.graphpad.com). Statistical tests and their results are described in the text and figure legends. Images of co-infected tissues presented in Figs. 5–7 are representative images from four TG, three brains, and two eyes, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary files. Amplicon Sequencing data have been deposited in the Short Read Archive with BioProject accession no. PRJNA1128239. Plasmids, viruses, and other reagents developed in this study are available upon request and subject to standard material transfer agreements with the Buck Institute and Fred Hutch Cancer Center. Source data are provided with this paper.
Code availability
CRISPR off-target analysis was performed using software developed previously42 and available on Github https://github.com/proychou/TargetedMutagenesis).
References
Berrington, W. R. et al. Clinical correlates of herpes simplex virus viremia among hospitalized adults. Clin. Infect. Dis. 49, 1295–1301 (2009).
Brown, E. L. et al. Effect of maternal herpes simplex virus (HSV) serostatus and HSV type on risk of neonatal herpes. Acta Obstet. Gynecol. Scand. 86, 523–529 (2007).
Freeman, E. E. et al. Herpes simplex virus 2 infection increases HIV acquisition in men and women: systematic review and meta-analysis of longitudinal studies. AIDS 20, 73–83 (2006).
Tanner, E. J., Kirkegaard, K. A. & Weinberger, L. S. Exploiting genetic interference for antiviral therapy. PLoS Genet 12, e1005986 (2016).
Chaturvedi, S. et al. Identification of a therapeutic interfering particle—a single-dose SARS-CoV-2 antiviral intervention with a high barrier to resistance. Cell 184, 6022–6036 (2021).
Dimmock, N. J., Rainsford, E. W., Scott, P. D. & Marriott, A. C. Influenza virus protecting RNA: an effective prophylactic and therapeutic antiviral. J. Virol. 82, 8570–8578 (2008).
Mercado-López, X. et al. Highly immunostimulatory RNA derived from a Sendai virus defective viral genome. Vaccine 31, 5713–5721 (2013).
Rezelj, V. V. et al. Defective viral genomes as therapeutic interfering particles against flavivirus infection in mammalian and mosquito hosts. Nat. Commun. 12, 2290 (2021).
Levi, L. I. et al. Defective viral genomes from chikungunya virus are broad-spectrum antivirals and prevent virus dissemination in mosquitoes. PLoS Pathog. 17, e1009110 (2021).
Pitchai, F. N. N. et al. Engineered deletions of HIV replicate conditionally to reduce disease in nonhuman primates. Science 385, eadn5866 (2024).
Walter, M. & Verdin, E. Viral gene drive in herpesviruses. Nat. Commun. 11, 4884 (2020).
Walter, M., Perrone, R. & Verdin, E. Targeting conserved sequences circumvents the evolution of resistance in a viral gene drive against human cytomegalovirus. J. Virol. 95, e0080221 (2021).
Esvelt, K. M., Smidler, A. L., Catteruccia, F. & Church, G. M. Concerning RNA-guided gene drives for the alteration of wild populations. Elife 3, e03401 (2014).
Bier, E. Gene drives gaining speed. Nat. Rev. Genet. 23, 5–22 (2022).
Law, G. A., Herr, A. E., Cwick, J. P. & Taylor, M. P. A new approach to assessing HSV-1 recombination during intercellular spread. Viruses 10, 220 (2018).
Card, J. P. et al. A dual infection pseudorabies virus conditional reporter approach to identify projections to collateralized neurons in complex neural circuits. PLoS One 6, e21141 (2011).
Javier, R. T., Sedarati, F. & Stevens, J. G. Two avirulent herpes simplex viruses generate lethal recombinants in vivo. Science 234, 746–748 (1986).
Nishiyama, Y., Kimura, H. & Daikoku, T. Complementary lethal invasion of the central nervous system by nonneuroinvasive herpes simplex virus types 1 and 2. J. Virol. 65, 4520–4524 (1991).
Bowden, R., Sakaoka, H., Donnelly, P. & Ward, R. High recombination rate in herpes simplex virus type 1 natural populations suggests significant co-infection1. Infect. Genet. Evol. 4, 115–123 (2004).
Casto, A. M. et al. Large, stable, contemporary interspecies recombination events in circulating human herpes simplex viruses. J. Infect. Dis. 221, 1271–1279 (2020).
Koelle, D. M. et al. Worldwide circulation of HSV-2 × HSV-1 recombinant strains. Sci. Rep. 7, 44084 (2017).
Liljeqvist, J. Å., Tunbäck, P. & Norberg, P. Asymptomatically shed recombinant herpes simplex virus type 1 strains detected in saliva. J. Gen. Virol. 90, 559–566 (2009).
Gierasch, W. W. et al. Construction and characterization of bacterial artificial chromosomes containing HSV-1 strains 17 and KOS. J. Virol. Methods 135, 197–206 (2006).
Taylor, M. P., Kobiler, O. & Enquist, L. W. Alphaherpesvirus axon-to-cell spread involves limited virion transmission. Proc. Natl Acad. Sci. USA 109, 17046–17051 (2012).
Morimoto, T., Arii, J., Akashi, H. & Kawaguchi, Y. Identification of multiple sites suitable for insertion of foreign genes in herpes simplex virus genomes. Microbiol. Immunol. 53, 155–161 (2009).
Criddle, A., Thornburg, T., Kochetkova, I., DePartee, M. & Taylor, M. P. gD-Independent superinfection exclusion of alphaherpesviruses. J. Virol. 90, 4049–4058 (2016).
Cwick, J. P. et al. Superinfection exclusion of alphaherpesviruses interferes with virion trafficking. Microbiol Spectr. 10, e0068422 (2022).
Yao, Q. et al. Un1Cas12f1 and Cas9 gene drive in HSV1: viruses that ‘infect’ viruses. Preprint at bioRxiv. https://doi.org/10.1101/2023.12.04.569968 (2023).
Wojaczynski, G. J., Engel, E. A., Steren, K. E., Enquist, L. W. & Patrick Card, J. The neuroinvasive profiles of H129 (herpes simplex virus type 1) recombinants with putative anterograde-only transneuronal spread properties. Brain Struct. Funct. 220, 1395–1420 (2015).
Card, J. P. & Enquist, L. W. Transneuronal circuit analysis with pseudorabies viruses. Curr. Protoc. Neurosci. 68, 1.5.1-39 (2014).
Banfield, B. W., Kaufman, J. D., Randall, J. A. & Pickard, G. E. Development of pseudorabies virus strains expressing red fluorescent proteins: new tools for multisynaptic labeling applications. J. Virol. 77, 10106–10112 (2003).
Aubert, M. et al. Gene editing for latent herpes simplex virus infection reduces viral load and shedding in vivo. Nat. Commun. 15, 4018 (2024).
Alfonso-Dunn, R. et al. Transcriptional elongation of HSV immediate early genes by the super elongation complex drives lytic infection and reactivation from latency. Cell Host Microbe 21, 507–517.e5 (2017).
Cliffe, A. R. et al. Neuronal stress pathway mediating a histone methyl/phospho switch is required for herpes simplex virus reactivation. Cell Host Microbe 18, 649–658 (2015).
de Gooijer, M. C. et al. Buparlisib is a brain penetrable pan-PI3K inhibitor. Sci. Rep. 8, 10784 (2018).
Brossard, D., Belluck, P., Gould, F. & Wirz, C. D. Promises and perils of gene drives: navigating the communication of complex, post-normal science. Proc. Natl Acad. Sci. USA 116, 7692–7697 (2019).
Emerson, C., James, S., Littler, K. & Randazzo, F. Principles for gene drive research. Science 358, 1135–1136 (2017).
National Academies of Sciences, E. A. M. Gene Drives on the Horizon. (National Academies Press, Washington, D.C., 2016).
NOT-OD-24-093: Notice of revisions to the NIH guidelines for research involving recombinant or synthetic nucleic acid molecules. https://grants.nih.gov/grants/guide/notice-files/NOT-OD-24-093.html.
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Aubert, M. et al. Gene editing and elimination of latent herpes simplex virus in vivo. Nat. Commun. 11, 4148 (2020).
Card, J. P., Whealy, M. E., Robbins, A. K., Moore, R. Y. & Enquist, L. W. Two alpha-herpesvirus strains are transported differentially in the rodent visual system. Neuron 6, 957–969 (1991).
Card, J. P. & Enquist, L. W. Use and visualization of neuroanatomical viral transneuronal tracers. in Visualization Techniques 225–268 (Humana Press, Totowa, NJ, 2012).
Jerome, K. R., Huang, M.-L., Wald, A., Selke, S. & Corey, L. Quantitative stability of DNA after extended storage of clinical specimens as determined by real-time PCR. J. Clin. Microbiol. 40, 2609–2611 (2002).
McLean, I. W. & Nakane, P. K. Periodate-lysine-paraformaldehyde fixative. a new fixation for immunoelectron microscopy. J. Histochem. Cytochem. 22, 1077–1083 (1974).
Acknowledgements
We thank ophthalmologist Koji Kitazawa (Buck Institute and Kyoto Prefectural University of Medicine) for training MW with intravitreal injections. We thank members of the Verdin and Jerome labs for technical and conceptual help. This study was supported by NIH grant R21AI178255 and through institutional support from the Buck Institute for Research on Aging and the Fred Hutch Cancer Center. In particular, MW received funding from the VIDD faculty initiative award from the Fred Hutch Cancer Center.
Author information
Authors and Affiliations
Contributions
M.W. designed the study with input from E.V., K.R.J., and M.P.T. M.W. performed all cell culture experiments. M.W. and R.R. wrote the IACUC protocol at the Buck Institute. M.W., R.R., A.K.H., P.A.M., M.A.L., D.E.S., and M.A. contributed to mouse husbandry and mouse experiments. L.M.K., L.S., and T.K.S. processed samples at the University of Washington Virology Laboratory. M.W. and M.P.T. performed the analysis of co-infection in the brain. P.R. contributed to the off-target analysis. M.W., M.P.T., K.R.J., and E.V. analyzed the data. M.W. wrote the manuscript with input from all authors. M.W., E.V., and K.R.J. supervised and funded the project.
Corresponding authors
Ethics declarations
Competing interests
A patent application describing the use of a gene drive in DNA viruses has been filed by the Buck Institute for Research on Aging (Application number 17054760, inventor: M.W.). The authors declare no further competing interests.
Peer review
Peer review information
Nature Communications thanks Rongtuan Lin and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Walter, M., Haick, A.K., Riley, R. et al. Viral gene drive spread during herpes simplex virus 1 infection in mice. Nat Commun 15, 8161 (2024). https://doi.org/10.1038/s41467-024-52395-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52395-2
This article is cited by
-
Partial inhibition of viral replication machinery enhances recombination in herpes simplex viruses
Biology Direct (2025)
-
Gene drive-based population suppression in the malaria vector Anopheles stephensi
Nature Communications (2025)
-
Exploiting social traits for clinical applications in bacteria and viruses
npj Antimicrobials and Resistance (2025)
-
Cas12f1 gene drives propagate efficiently in herpesviruses and induce minimal resistance
Genome Biology (2024)