Abstract
High-throughput sequencing technologies generate a vast number of DNA sequence reads simultaneously, which are subsequently analysed using the information contained within these fragmented reads. The assessment of sequencing technology relies on information efficiency, which measures the amount of information entropy produced per sequencing reaction cycle. Here we propose a fuzzy sequencing strategy that exhibits information efficiency more than twice that of currently prevailing cyclic reversible terminator sequencing methods. To validate our approach, we develop a fully functional and high-throughput fuzzy sequencer. This sequencer implements an efficient fluorogenic sequencing-by-synthesis chemistry and we test it across various application scenarios, including copy-number variation detection, non-invasive prenatal testing, transcriptome profiling, mutation genotyping and metagenomic profiling. Our findings demonstrate that the fuzzy sequencing strategy outperforms existing methods in terms of information efficiency and delivers accurate resequencing results with faster turnaround times.
Similar content being viewed by others
Main
High-throughput next-generation sequencing (NGS)1,2 has been extensively employed for precise quantification of nucleic acids at a whole-genome, exome or transcriptome scale3,4. Among the primary applications of NGS is resequencing, which encompasses copy-number variation (CNV) assessment5,6, transcriptome profiling (RNA sequencing (RNA-seq))7,8 and non-invasive prenatal testing (NIPT)9 that rely on counting sequence-based DNA fragments. The accuracy of counting largely depends on the effectiveness of extracting information from DNA fragments and accurately mapping it onto reference sequences. Although mapping algorithms continue to evolve, the sequencing steps themselves have remained largely unchanged for several years.
NGS relies on the incorporation of nucleotides to encode information, and the efficiency of information encoding varies across different systems. The majority of high-throughput DNA sequencers use sequencing-by-synthesis (SBS) chemistries, with signal generation schemes falling into two major variations: single-nucleotide addition (SNA) or cyclic reversible terminator (CRT)10,11. We define the intrinsic information efficiency of a sequencer as the information entropy per cycle of the sequencing signals. According to this definition, SNA chemistry produces natural DNA duplex but suffers from relatively low intrinsic information efficiency, averaging at 0.67 nt per cycle or 1.33 bits per cycle for encoding random sequences. In contrast, multistep CRT chemistry, although producing scar-containing duplex, offers 2 bits per cycle intrinsic information efficiency with an exact 1 nt per cycle extension rate (Extended Data Fig. 1a). Enhancing the information entropy encoding efficiency becomes a clear avenue for improving sequencing performance.
While the information entropy embedded in the nucleotide sequence plays a decisive role in mapping a read to a reference, mapping accuracy is also influenced by the corresponding region of the reference. One example is the mate-pair reads which, compared with conventional paired-end reads, are more mappable because of larger reference target size instead of higher information entropy. This effect becomes particularly evident in genomes with a high proportion of repeat sequences. From the perspective of the sequencer, we define the extrinsic information efficiency as the information entropy per cycle of the sequenced DNA fragment, or two times the read length per cycle. This reference-side extrinsic information is equally important but often overlooked. This oversight is primarily due to prevalent DNA sequencers providing explicit sequences of four bases, where the intrinsic information efficiency is equal to the extrinsic information efficiency, despite the differences in efficiency between SNA and CRT. Enhancing the extrinsic information efficiency could also lead to improvements in sequencing technology.
Decoupling the intrinsic and extrinsic information entropy within the same read suggests a change in sequencing read formatting. When aligning a DNA fragment sequence to reference sequences, the alignment process relies not on the string format, but rather on the information entropy contained within it. The reference genome is typically represented by the conventional and natural four base letters (A, C, G and T), but the read itself does not necessarily need to adhere to the same formatting. As long as the information is sufficiently rich for unique alignment, alternative formatting can be employed. Although sequencing by oligonucleotide ligation and detection is being phased out in the market due to low-efficiency chemistry and very short read lengths, it demonstrates the possibility of sequencing with alternate formatting through a colour–space strategy12.
From a sequencing strategy standpoint, achieving higher information efficiency (>2 bits per cycle) for both intrinsic and extrinsic information entropy could benefit the entire sequencing field. In this report, we introduce a fuzzy SBS strategy that sacrifices explicit sequencing information in exchange for improved information encoding performance. Implemented as a fully functional and high-throughput fuzzy sequencer, this strategy shows consistent results across various application scenarios.
Results
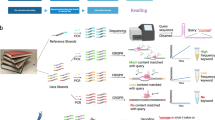
The fuzzy SBS sequencing strategy is based on fluorogenic sequencing chemistry using terminal phosphate-labelled fluorogenic nucleotides13,14,15 (TPLFN, Fig. 1a) we reported earlier. In this chemistry, the fluorophore Tokyo green (TG)-labelled TPLFN can turn from a dark state to a light state after catalysis by the Bst polymerase and alkaline phosphatase, and its fluorescence intensity indicates the number of the incorporated nucleotides (Fig. 1b). We constructed DNA fragments into a library with two specific end adapters for clonal amplification on hydrogel microbeads carrying amplification primers. The DNA fragments hybridized on the beads were then amplified into clones through emulsion PCR16 or surface-tethered recombinase polymerase amplification (RPA) (Fig. 1c). Each clone typically contained ~30,000 copies of DNA for the SBS reaction. After annealing with the sequencing primer, the DNA fragments could be effectively sequenced through a conventional SNA strategy (Fig. 1d).
a, Chemical structure of the TPLFN used in fuzzy sequencing. b, Schematic of the fluorogenic sequencing chemistry adopted by fuzzy sequencing. CIP, calf intestinal alkaline phosphatase. c, Schematic of the fuzzy sequencing pipeline. The DNA templates are amplified on the beads and loaded into the wells of the sequencing chip. After hybridization by the sequencing primers, the DNA templates are primed for sequencing. d–f, Schematic of the nucleotide flowgrams used in single-nucleotide addition (d), monochromatic double-nucleotide addition (BitSeq; e) and dichromatic double-nucleotide addition (SuperBitSeq; f).
Fuzzy SBS can be implemented using different flowgrams (Fig. 1e,f and Extended Data Fig. 1b), allowing the introduction of nucleotides beyond the limitation of a single type per cycle (see Supplementary Text 1). Briefly, ‘BitSeq’ employs a double-nucleotide addition flowgram, where a combination of two nucleotides is alternately added in each cycle. For instance, K (a mix of G and T) is added during odd cycles, while M (a mix of A and C) is added during even cycles (see Fig. 1e). BitSeq generates fuzzy sequences since it is not possible to distinguish the signal associated with the incorporation of the two nucleotides in the mix. Similarly, TritSeq employs a triple-nucleotide addition flowgram, where a combination of three nucleotides is alternately added in each cycle (Extended Data Fig. 1b). All of these fuzzy SBS approaches can achieve intrinsic and/or extrinsic information efficiency beyond 2 bits per cycle, effectively reducing the cycle number required to achieve a specific read length and ultimately leading to faster turnaround speed (Supplementary Table 1). In this paper, the focus is primarily on BitSeq and its dichromatic form SuperBitSeq as they strike a reasonable balance between information efficiency and signal dynamic range per reaction cycle.
We constructed a laboratory prototype sequencer for conducting high-throughput BitSeq (Fig. 2a,b). The bit sequencer comprises a microfluidic flowcell for implanting clonally amplified DNA fragments for sequencing, and an imaging system for signal acquisition (Extended Data Fig. 2a–g). We implemented fluorogenic SBS chemistry to naturally align with the BitSeq flowgram13,14,15,17.
a, Design of a prototype for high-throughput fluorogenic fuzzy sequencing. b, Photograph of a sequencing module, which consists of a flowcell connected to a manifold for liquid routing and placed on a thermoelectric Peltier device for temperature control. c, Photograph of a sequencing flowcell, which is made of a glass slide and a FOP, with a flow channel in between with shape defined by laser-cut double-sided adhesive. IPA, isopropanol. d, Scanning electron microscopy image of the microwell array made by selective etching of FOP. These femtolitre microwells are placed inside the flow chamber. e, Size of a microwell. f, Running procedure for one sequencing cycle. First, the microbeads with clonal amplified templates for sequencing are introduced into the microwells and tethered on the inner surface of the microwells. Then, the reaction buffer is primed into the flow chamber and all microwells are sealed by oil flowing into the chamber to separate each microwell from cross-talk. The SBS reaction is triggered by elevation of the temperature. The fluorogenic product is proportional to the bases that are incorporated during each reaction cycle. When the reaction is finished, the flowcell is cooled to take fluorescence images. g, One typical field of view from 171 tiles per cycle and the procedure of fluorescence intensity extraction. The microwells that contain beads generate fluorescence after reaction of each cycle, and higher intensity indicates longer degenerate polymer length (DPL) in that reaction cycle. Each microwell is addressable and indexed to produce an intensity series that can be later deduced into bit sequences. h, Read length distribution of bit sequences. i, Error rate of BitSeq.
The microfluidic flowcell (Fig. 2c) contains 28 million 30-femtolitre microwells (Fig. 2d,e). Each cycle of the SBS reaction can extend one or more bases and produce a corresponding amount of fluorophore. The microwells can be sealed to prevent signal cross-talk due to fluorophore diffusion. These microwells were fabricated by selective wet-etching of fibre-optic plates (FOPs), allowing the fluorogenic signal to propagate through the fibre waveguide to the outer surface of the flowcell for imaging (Fig. 2f). The inner surface of the microwells was made hydrophilic, while the adjoining surface between microwells was topologically coated to be hydrophobic, enabling the highly parallel sealing of the SBS reactions using oil (Extended Data Fig. 2h–k).
The oil-based microwell sealing is robust, reversible and compatible with fluorogenic SBS reaction conditions. In each cycle of the fluorogenic SBS reaction, we first filled the microwells with reaction buffer containing polymerase and two fluorogenic unnatural nucleotides at a low temperature of 4 °C. We then sealed the microwells by steadily flowing in fluorinated oil (Fig. 2f). Next, we quickly raised the temperature of the flowcell to 65 °C to initiate the synthesis reaction, followed by cooling down to 25 °C for fluorescence image acquisition. After that, we removed the sealing by flowing in isopropanol and performed an aqueous wash to reset the reaction conditions for the next cycle. The acquired images were aligned to identify each microwell, which was then registered for tracking throughout all reaction cycles (Fig. 2g).
Defective regions resulting from local sealing or unsealing failures were eliminated before further processing (Extended Data Fig. 3a). Empty microwells and microwells with multiple beads were identified and filtered out of further analysis. The fluorescence signal of each monoclonal microwell was extracted from the images after noise reduction and background correction. Due to the asynchronization of the molecular reaction within each clone, the intensity–cycle series could not be directly converted into bit sequences without a meticulous dephasing process. To supervise the dephasing algorithm, we spiked a standard DNA library, constructed from the lambda phage genome, into the samples to be sequenced together. As a result, the sequenced reads could be easily classified into two categories: standard dots (SD) and library dots (LD), on the basis of the characteristic signal profiles of the first few cycles. We employed the dual-base flowgram dephasing model to fit the SD signals and obtain the parameters18, which were then applied to LD signals to construct a flux matrix for dephasing correction and to convert the signals into bit sequences.
We initially tested our prototype sequencer on a lambda phage genomic library (Extended Data Fig. 3c and Supplementary Table 2). After 61 cycles of repeated sealing and sequencing, more than 90% of the chip area remained unaffected by stains (Extended Data Figs. 3b and 4h,m). In a single run, we generated 6,578,364 reads with an average length of 112.2 bp (Fig. 2h and Extended Data Fig. 4a–c). To map the BitSeq signals, we encoded them as sequence reads that could be aligned using conventional mappers such as Burrows–Wheeler Aligner (BWA)19,20 and Bowtie2 (Supplementary Text 2 and Tables 3–6)21,22. By mapping the bit sequences back to the reference genome, we successfully aligned 99.11% of the reads, which had an average length of 111.6 bp. The error rate of BitSeq was found to be 0.39% in the first 10 bp and 0.62% in the 130–140-bp region, which are acceptably low. This indicates the feasibility of accurate read mapping and DNA fragment counting (Fig. 2i).
Using the same aligning strategy, we then evaluated the performance of the BitSeq approach in four widely used resequencing applications: CNV identification, NIPT, transcriptomic analysis (RNA-seq) and metagenomics sequencing (mNGS).
We sequenced two genomic DNA samples collected from a normal male and a male patient with Down syndrome at low coverage (0.1×). The sequencing coverage across the whole genome was mostly uniform for both samples (Extended Data Fig. 5a–d) except for the sex chromosomes which were half of that of autosomes. The additional copy of chromosome 21 in the patient with Down syndrome was accurately counted through BitSeq, and validated by an Ion Torrent sequencer.
Besides aneuploidies, BitSeq can also faithfully detect the clinically relevant small-size CNVs. We used BitSeq to identify the 2.9-Mb deletion at Chr22.q11.21 that is associated with DiGeorge syndrome from a patient, and two CNVs (20.8-Mb gain at Chr 11.q and 10.0-Mb gain at Chr22.q11) from another patient with developmental delay (Fig. 3a), with high concordance with other conventional sequencing approaches. Notably, BitSeq showed a lower median of the absolute values of all pairwise differences (MAPD) than CRT when truncated to read lengths with the same sequencing cycles, indicating more precise CNV determination (Fig. 3b).
a, CNV of a patient with DiGeorge syndrome (left) and a patient with developmental delay (right). b, BitSeq shows lower MAPD than CRT under the same sequencing cycles. MAPD for CRT under 25 cycles is missing because such short reads cannot be mapped uniquely to the genome. c, Z-scores of normal and mock trisomy NIPT ccfDNA samples. d, Z-scores of single-blinded NIPT ccfDNA samples. Red dashed lines in c and d indicate the threshold of 3. P values were determined using two-sided Student’s t-test without adjustments for multiple comparisons. e,f, Microbe abundance comparison between BitSeq and Illumina by metagenomic sequencing (e) and targeted metagenomic sequencing (f). g, TPM comparison of mEF (blue) and mES (red) cell lines between BitSeq and commercial sequencers. E. avium, Enterococcus avium; E. faecium, Enterococcus faecium; S. thermophilus, Streptococcus thermophilus; C. innocuum, Clostridium innocuum; S. suis, Streptococcus suis; HAdV, human adenovirus; E. gilvus, Enterococcus gilvus; E. lactis, Enterococcus lactis; E. faecalis, Enterococcus faecalis; E. casseliflavus, Enterococcus casseliflavus; P. taiwanensis, Pseudolabrys taiwanensis; P. sp., Pseudolabrys species; B. sp., Bradyrhizobium species; R. sp, Rhodoplanes species; B. erythrophlei, Bradyrhizobium erythrophlei; B. diazoefficiens, Bradyrhizobium diazoefficiens; M. sp, Mesorhizobium species; L. mirabilis, Lautropia mirabilis.
We then tested the effectiveness of BitSeq on NIPT, a widely used resequencing application in prenatal diagnostics that requires high accuracy of DNA counting. We started the test with mock samples prepared by spiking fragmented genomic DNA from patients with trisomy into circulating cell-free DNA (ccfDNA) of an unpregnant woman at different ratios. These 12 mock NIPT samples had trisomy in Chr21, Chr18 and Chr13, and the mixing proportions were 10, 5, 3.5 and 2.5%. We also collected 28 NIPT ccfDNA samples from pregnant women with normal fetuses. We applied BitSeq to all the samples and used 24 out of the 28 normal samples as the control set (the other 4 used as the negative test set) to calculate the Z-score for each NIPT sample (Fig. 3c). Using Z = 3 as the cut-off, BitSeq shows comparable power to identify the positive trisomy NIPT samples when the mock fetal DNA fraction is ≥3.5%. We further tested 10 single-blinded true NIPT samples, 5 of which tested positive based on the same cut-off using BitSeq (Fig. 3d). These results were confirmed by parallel tests using Ion Torrent sequencers as well as the clinical records (Extended Data Figs. 5e and 6).
Combined with metagenomic sequencing, BitSeq may identify pathogens in acute infections in a timely manner. We tested the capability of BitSeq in metagenomics by sequencing one anal swab and one throat swab from a previously reported paediatric patient suffering from multi-organ abscesses23. The 10 species types with the highest abundance identified by BitSeq and Illumina sequencer are identical. Their abundances, ranging from <0.1% to >95%, are also close (Fig. 3e). In addition, using a custom panel, we also tested BitSeq on targeted metagenomics by sequencing bronchoalveolar lavage fluid samples from 14 patients with community-acquired pneumonia (CAP). The species types as well as their normalized read number reflecting their relative abundance, are also consistent (Fig. 3f).
We next verified that BitSeq could be used for RNA-seq. Using mouse embryonic fibroblasts (mEF) and embryonic stem (mES) cells, BitSeq could provide almost identical results as Ion Torrent or Illumina sequencers. The number of genes detected was comparable and the gene expression levels were correlated with high linearity between these approaches (Fig. 3g and Extended Data Fig. 5f,g). We also checked the Gene Ontology (GO) terms of genes only detected by one sequencer and found no specific preferences in between (Extended Data Fig. 5h–j).
All these experiments show that BitSeq is fully capable for those applications based on DNA fragment counting. We expect that other resequencing approaches that do not require single-base resolution but focus on pattern alterations, such as bisulfite sequencing for DNA methylation24,25,26 or Hi-C for chromatin topology analysis27,28, are also possible with BitSeq. Due to its high information and long extension length per cycle as well as the fast fluorogenic sequencing chemistry, BitSeq would enable more-efficient DNA fragment counting that is valuable in both basic science research and clinical testing.
We further considered the possibility to extend the fuzzy sequencing strategy from BitSeq to SuperBitSeq by labelling the two different nucleotides in each reaction cycle with distinguishable fluorophores (Fig. 1f). SuperBitSeq possesses the same extrinsic information efficiency as BitSeq, but even higher intrinsic information efficiency (Extended Data Fig. 1d).
Similar to BitSeq, we also designed an encoding strategy for SuperBitSeq that is compatible with prevailing short read mappers such as BWA (Fig. 4a). We conducted in silico analysis to validate the alignment accuracy of the fuzzy SBS encoding strategy. For our analysis, we randomly selected 1 × 107 positions from the human reference genome and 1 × 106 positions from the Arabidopsis reference genome. We simulated the BitSeq, SuperBitSeq and CRT sequencing from these positions by extending the nucleotide strings with certain cycles (Extended Data Fig. 7a), and then aligned the simulated reads back to their respective reference genomes. We observe that reads generated with more sequencing cycles provided greater information, resulting in a higher unique mapping rate (UMR) for both fuzzy sequencing and CRT approaches. Notably, SuperBitSeq consistently exhibited the highest UMR (Fig. 4b, Extended Data Fig. 7b and Supplementary Tables 7–58). In addition, the UMR of BitSeq with the M-K form was consistently higher than that of CRT and BitSeq with the R-Y form. This preference for the M-K form may be attributed to the occurrence of transition base substitutions following gene duplication during evolution29.
a, Encoding strategy of SuperBitSeq in M-K form as an example. b, Simulated UMR of DNA reads by fuzzy sequencing and CRT for different genomes. For mapping the specific proportion of the genome, both BitSeq and SuperBitSeq need much shorter minimally required sequencing cycles, and this advantage can also be reflected by the reaction cycle reduction factor (RCRF) that is defined by the ratio of minimally required cycles between technologies. c, Distinguishable and indistinguishable SNV types of BitSeq and SuperBitSeq. d, Ratio of indistinguishable SNV by BitSeq and SuperBitSeq. e, Freqencies of extended base number of different flowgrams. f, Fractal of encoded SuperBitSeq signals. g, Fluorescence intensities (top) and their dephasing-corrected signals (bottom) of a single-template SuperBitSeq experiment. Black dots represent the signal of each replicate. Bars represent the mean signal of replicates. Yellow dots indicate the theoretical values where the corresponding mean signal is erroneous.
The elevated information efficiency enables the detection of substantially more single-nucleotide variants (SNVs) that cannot be identified through BitSeq (Fig. 4c). For example, in BitSeq, the A > C/G > T mutations cannot be identified in the M-K form, or the A > G/C > T mutations in the R-Y form. In SuperBitSeq, they can all be well captured, except for only specific indistinguishable base swap events such as the AC > CA in the M-K form. To evaluate the potential of SuperBitSeq for single-nucleotide polymorphism (SNP)/SNV detection, we filtered The Single Nucleotide Polymorphism Database (build 155)30 and ClinVar (20210908)31 to exclude items with N, and obtained 1,297,977,577 and 1,009,591 items, respectively. About 17–52% of known human SNPs/SNVs cannot be distinguished by BitSeq, whereas the non-detection rates are ~0.01% or even lower for SuperBitSeq (Fig. 4d). This means that SuperBitSeq can de facto detect almost all known SNPs/SNVs. In each reaction cycle, the fluorescence signal released by extended bases can be split into two channels, resulting in reduced requirement for signal detection dynamic range and enhanced accuracy in detecting long copolymers (Fig. 4e and Extended Data Fig. 7c). Interestingly, we also found that the encoded superbit sequences can form a fractal when we mapped the points in a square (Fig. 4f and Supplementary Text 4).
To implement SuperBitSeq experimentally, we synthesized a fluorophore, Peking orange (PO), which was designed to exhibit excellent fluorogenic properties (on/off ratio ~2,000) when terminally labelled on the tetraphosphate nucleotides (PO-TPLFNs, or PO-deoxynucleotide tetraphosphate; Fig. 1a and Extended Data Fig. 8a–j). PO was used to pair with TG for labelling two different nucleotides in each reaction cycle. PO and TG have similarly high fluorescence quantum efficiency but distinct excitation and emission spectra to avoid signal cross-talk (Extended Data Fig. 8k). PO-deoxynucleotide tetraphosphate has great photo and thermal stability, and showed excellent signal-to-base linearity upon nucleotide extension reaction (Extended Data Fig. 8i–n). The same dephasing algorithm we optimized for TG-deoxynucleotide tetraphosphate substrates could be seamlessly adopted to PO-deoxynucleotide tetraphosphate when sequencing a single template (Extended Data Fig. 9a–d). According to our previous in silico simulation, dephasing correction for SuperBitSeq could be built upon the algorithm of BitSeq by separating two channels for independent correction with the same dephasing parameters18. We also experimentally confirmed this strategy using the same single template for SuperBitSeq (Fig. 4g and Extended Data Fig. 9e,f).
We added a fluorescence channel to the bit sequencer and converted it to the superbit sequencer (Fig. 5a and Extended Data Fig. 10a), while the image processing and signal extraction pipeline was largely adapted from before. We demonstrated the SNV detectability of SuperBitSeq by sequencing the G719S and T790M mutations of the egfr gene (Fig. 5b). The signal acquired from 9 imaging tiles (0.67 × 0.67 mm2 each, 212,874 reads in total) could be clearly clustered into 4 groups (Fig. 5c,d), and each group represented one of the DNA templates with single nucleotide difference. With dephasing correction, the signals of these clusters clearly unveiled the notable difference at the specific reaction cycle, which is associated with the SNVs we targeted.
a, A merged fluorescence image of green (TG-labelled T or G) and red (PO-labelled C or A) channels from which the signal intensities are extracted. The image was associated with Cycle 6 in this specific experiment and yellow dots represent the co-existence of green and red signal. The signal series of four different DNA templates are shown on the right. b, Violin plot of fluorescence intensity and dephasing-corrected signals of four different DNA templates. The blue boxes show the distinguishable signals that differentiate the SNVs using SuperBitSeq. c,d, The four different DNA templates can be distinguished by the difference of fluorescence intensities (c) or dephasing-corrected signals (d) in Cycle 5–6. e, Error rate of SuperBitSeq in SARS-CoV-2 samples. f, Allele frequencies in SARS-CoV-2 samples detected by SuperBitSeq and an Illumina sequencer.
While BitSeq can only determine the pathogen species and abundances in metagenomic sequencing, SuperBitSeq allows fast pathogen genotyping and origin tracing, which is valuable in epidemic outbreaks. We sequenced 3 SARS-CoV-2 samples from a previously reported epidemiology study32 for 26 cycles using SuperBitSeq33. The total reaction time is only 42.6 min and we got ~200,000 reads. After mapping the SuperBitSeq reads to the reference genome, we restored the SNV information by separating the read and the reference into two semi-sequences, aligning individually and then merging together (Extended Data Fig. 10b–d and Supplementary Text 5). The experimental data demonstrated low error rate (~1%; Fig. 5e) and an average read length of 43 bp (Extended Data Fig. 10a). The SNVs called by SuperBitSeq are consistent with those called by an Illumina sequencer (Fig. 5f).
Discussion
We present a fuzzy sequencing strategy with over twice the information efficiency of current CRT sequencing. We developed a high-throughput fuzzy sequencer using efficient fluorogenic SBS chemistry. After testing in multiple applications such as CNV detection, it was proven that this strategy provides accurate resequencing results with faster turnaround times.
One reasonable extension of our information-richer SuperBitSeq is to combine three rounds of SuperBitSeq with orthogonal dual-base mixes for error-correction code (ECC) sequencing, which may surpass the previously demonstrated high-accuracy ECC. Our in silico simulation showed that compared with traditional ECC, error correction using three rounds of superbit sequences has lower false-negative rate in detecting potential sequencing errors under the same noise level (Extended Data Fig. 10f). Both BitSeq and SuperBitSeq are specific forms of the fuzzy SBS strategy that encode DNA with high information efficiency for sequencing. More advanced forms such as TritSeq or SuperTritSeq are definitely worthy of more expectation. However, the signal dynamic range of TritSeq would be too large to practice for fluorogenic chemistry, although TritSeq would generate longer extension length and higher information efficiency than BitSeq. One of the limitations of BitSeq is its incompatibility with prevailing sequencing platforms such as Illumina’s NovaSeq X or other similar instruments. The fuzzy SBS strategy relies on fluorogenic sequencing chemistry, in which the fluorophore is cleaved off right after nucleotide incorporation. If fuzzy SBS is performed in the flowcells in other sequencers without microwell sealing, the fluorophore will diffuse, causing signal mixing. Fuzzy SBS uses the microwell chip with reversible sealing functions to confine reactions and prevent signal mixing. This microwell chip has potential applications beyond fuzzy sequencing, including digital PCR and single-cell transcriptomics where reaction isolation is crucial.
There are three major challenges that have not been well addressed in this work. First, the fundamental mathematical structure of the fuzzy coding of DNA is not yet fully understood. For example, after transforming the bit sequences into binary fraction numbers and then converting into decimal fraction numbers, every infinite long DNA sequence can be mapped to a point in the square region [0,1) × [0,1) (Fig. 4f) and formed fractal patterns for both SuperBitSeq with M-K and W-S forms. These two fractal patterns have identical Hausdorff dimension of ~1.7716, although the nature of it is still unknown (Supplementary Text 4). Second, currently available fragment mapping algorithms and software, such as BWA or Bowtie2, were designed for recognizing natural base format but not the fuzzy sequences. Although we have circumvented this problem by renaming the bit strings to be compatible with current software, a more-general mapping tool that directly handles the nature of fuzzy sequence is still highly desirable. Last, further optimizations are still required to reduce the overall operational time and cost. Although cycle number reduction by fuzzy sequencing leads to reduced reaction time and reagent cost, multiple steps such as template amplification, fluidics and imaging still take up a large proportion of the time required, and the throughput should be further increased to reduce the average cost.
Methods
Ethics and sample collection
Peripheral blood DNA samples of patients with congenital heart disease (CHD) were obtained from a previous CHD study34. Sample usage was approved by the Medical Ethics Committee of Tsinghua University. Peripheral blood plasma cfDNA samples of pregnant women were obtained from a previous prenatal diagnostic study35,36. Sample usage was approved by the Medical Ethics Committee of Beijing Obstetrics and Gynecology Hospital and Tsinghua University. The anal swab and throat swab samples were obtained from a previous case study23,37. Sample usage was approved by the Medical Ethics Committee of the Institute of Pathogen Biology, Chinese Academy of Medical Sciences and Peking Union Medical College. The collection and usage of bronchoalveolar lavage fluid samples of patients with CAP was approved by the Medical Ethics Committee of Peking University Third Hospital37. All samples were deidentified before use in this study.
DNA extraction and library preparation
Genomic DNA sequencing library preparation
The lambda phage genomic DNA was obtained from New England Biolabs (NEB)38. Genomic DNA (50 ng) in 130 μl TE buffer was fragmented to ~180 bp using sonication (Covaris S220) with the following parameter setting: 175 W peak incident power, 10% duty factor, 200 cycles per burst, 300 s treatment time. Libraries were constructed from fragmented genomic DNA using NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB) following manufacturer protocol. Standard index PCR primers were replaced by custom dual-unique indexing primers for Illumina sequencing, or Adaptor A and P1 (synthesized by Thermo Fisher; Extended Data Fig. 3c and Supplementary Table 2) for Ion Torrent and BitSeq.
ccfDNA extraction
Blood samples (10 ml) were centrifuged at 1,600g for 10 min at 4 °C within 8 h after collection. The extracted plasma was centrifuged again at 16,000g for 10 min at 4 °C to further remove cellular components, before being stored at −80 °C. ccfDNA from the plasma was extracted using the CWhipro Circulating Nucleic Acid kit (CW Biotech) following manufacturer instruction. The quantity and quality of the ccfDNA was checked using a Qubit 3.0 fluorometer (Thermo Fisher) and a 2100 Bioanalyzer (Agilent).
Mock trisomy ccfDNA preparation
Genomic DNA from the patient with trisomy was sheared using Covaris S220. The 150–250-bp part of the fragmented DNA was gel purified. The mock trisomy ccfDNA was prepared by mixing fragmented trisomy genomic DNA with ccfDNA extracted from non-pregnant female volunteers. The mixing ratios were validated through droplet digital PCR.
ccfDNA sequencing library preparation
ccfDNA (2–5 ng) was end repaired in a total of 60 μl mix containing 1× T4 polynucleotide kinase buffer (TIANGEN), 0.167 mM dNTP mix (NEB), 3 U T4 DNA polymerase (TIANGEN), 10 U T4 polynucleotide kinase (TIANGEN), 5 U Klenow fragment (TIANGEN) and sterile water at 25 °C for 25 min, 70 °C for 10 min and held in 4 °C. The end-repaired DNA was then ligated in a total of 100 μl mix with 1× T4 DNA ligase buffer (TIANGEN), 3,600 U T4 DNA ligase (TIANGEN), 40 U Bst DNA polymerase (TIANGEN), 50 nM Adaptor A and P1 at 25 °C for 15 min, 65 °C for 5 min and held at 4 °C. The ligated DNA was cleaned up using 0.9× volume of Agencourt AMPure XP beads (Beckman Coulter). The library was sent to Beijing Jinnuo Ruijie Gene Science and Technology for Ion Torrent sequencing. This library was also used for BitSeq.
RNA extraction and reverse transcription
Total RNA was extracted from 106 feeder-free mouse ES cells or mEF cells separately using RNeasy Micro kit (Qiagen) following manufacturer instructions. On-column DNase digestion was performed to remove genomic DNA. The quantity and quality of the RNA was checked using a Qubit RNA HS Assay kit (Invitrogen) and a 2100 Bioanalyzer. Messenger RNA was isolated and fragmented from 1 μg of extracted total RNA using the NEBNext Poly(A) mRNA Magnetic Isolation Module, followed by reverse transcription using NEBNext Ultra II RNA First and Second Strand Synthesis Module according to manufacturer instructions. Libraries were constructed from 10 ng of double-strand cDNA obtained above using NEBNext Ultra II DNA Library Prep Kit for Illumina following manufacturer protocol. Standard index PCR primers were replaced by custom dual-unique indexing primers for Illumina sequencing, or Adaptor A and P1 for Ion Torrent and BitSeq. After 10 cycles of PCR amplification, the final libraries were quantified using a Qubit dsDNA HS Assay kit (Invitrogen) and Agilent Fragment Analyzer39.
Targeted metagenomics sequencing
Briefly, 600 µl of bronchoalveolar lavage fluid isolated from each enrolled patient was collected and mixed with lysozyme and glass beads. DNA extraction was performed by using the TIANamp Micro DNA kit (TIANGEN). The libraries were prepared using the targeted metagenomics sequencing panel of Cygnus Biosciences following manufacturer protocol.
Sequencer design
A ×10 microscope objective (NA 0.45, CFI Plan Apo Lambda, Nikon) and a scientific complementary metal–oxide semiconductor sensor (Flash 4.0, Hamamatsu) were used for image acquisition in the sequencer prototype. A high-power blue LED with wavelength of ~460 nm (CBT-90-B-TE, Luminus) was used for excitation. Both excitation (482 ± 20 nm) and emission (536 ± 40 nm) bandpass filters were purchased from Semrock. The x-axis translation stage (MSMF012, Panasonic) had 2 μm precision, taking ~1 s to move 1.33 mm at each step. The y-axis translation stage (IKO) had 1 μm precision and 65 mm moving range, taking ~300 ms to move 1.33 mm at each step. We imaged 171 tiles (9 × 19) in each reaction cycle to cover the whole reaction chamber. Reagent flow was driven by a syringe pump (Cavro XC, Tecan) and a series of solenoid valves (LVM10R6, SMC). The temperature of the reaction was controlled by a Peltier device (TE Tech), with a water-cooling system connected to the heat sink. We used LabVIEW (National Instruments) to control the whole prototype.
Synthesis of Peking orange
Synthesis of (2-bromo-5-methoxyphenyl)(3-methoxyphenyl)methanol
The chemical structure is shown as 1a in Extended Data Fig. 8a. In a 250 ml round-bottom flask (flame dried) equipped with a constant pressure dropping funnel, 80 ml anhydrous tetrahydrofuran solution of m-methoxyphenyl magnesium chloride (1.2 eq) was added and cooled to −40 °C with stirring. Then 20 ml anhydrous tetrahydrofuran solution of 3-methoxy-o-bromobenzaldehyde (10.8 g) was added dropwise to the reaction flask through dropping funnel while being kept at −40 °C with stirring. After addition, the reaction was kept for 2–6 h, monitored by thin-layer chromatography (TLC) and stopped when the raw material, 3-methoxy-o-bromobenzaldehyde, disappeared. Saturated ammonium chloride aqueous solution (20 ml) was added to quench the reaction. The reaction mixture was subjected to a rotary evaporator to remove most of the solvent, and then extracted twice with dichloromethane (100 ml each). After separation, the organic phases were combined, washed with saturated brine, dried over Na2SO4, concentrated by a rotary evaporator and then purified by silica gel column chromatography with developing solvent: the product (petroleum ether/ethyl acetate ~10/1, retention factor (Rf) ~0.3) was collected to get 15 g of colourless liquid 1a with 93% yield.
1H NMR (CDCl3, 500 MHz): δ 7.39 (d, J = 10 Hz, 1H, Ar-H), 7.23 (t, J = 10 Hz, 1H, Ar-H), 7.13 (d, J = 5 Hz, 1H, Ar-H), 6.96 (m, 1H, Ar-H), 6.81–6.79 (m, 1H, Ar-H), 6.96 (m, 1H, Ar-H), 6.70 (dd, J = 10 Hz, 5 Hz, 1H, Ar-H), 6.08 (d,1H, CH), 3.77 (s, 3H), 3.75 (s, 3H).
13C NMR (126 MHz, CDCl3): δ 159.70, 159.29, 143.76, 143.48, 133.41, 129.51, 119.33, 115.12, 114.01, 113.11, 113.05, 112.80, 74.63, 55.49, 55.23.
HRMS: Calculated for C15H15BrO3Na (M+Na), 345.0097. Found, m/z 345.0096.
Synthesis of (2-bromo-5-methoxyphenyl)(3-methoxyphenyl) methanone
The chemical structure is shown as 1b in Extended Data Fig. 8a. In 80 ml of dichloromethane in a 250 ml flask, 6.5 g of 1a was dissolved, added with 10 g of pyridinium chlorochromate (PCC) and 10 g celite, and the mixture was kept for 2–5 h under rapid stirring at room temperature. The reaction was monitored by TLC until the raw materials disappeared. The reaction solution was filtered through a short column pad with celite and washed with dichloromethane. The filtrate was collected, concentrated on a rotary evaporator and subjected to silica gel column chromatography. Developing solvent: petroleum ether/ethyl acetate ~10/1, Rf ~0.6. A 6.1 g light yellow solid was obtained after concentration, with 95% yield.
1H NMR (CDCl3, 500 MHz): δ 7.50 (d, J = 10 Hz, 1H, Ar-H), 7.45 (m, 1H, Ar-H), 7.35 (d, J = 10 Hz, 1H, Ar-H), 7.31–7.29 (m, 1H, Ar-H), 7.16–7.14 (m, 1H, Ar-H), 6.91–6.88 (m, 1H, Ar-H), 6.87 (d, J = 5 Hz, 1H, Ar-H), 3.85 (s, 3H), 3.79 (s, 3H).
13C NMR (126 MHz, CDCl3): δ195.40, 159.92, 158.74, 141.48, 137.29, 133.93, 129.61, 123.52, 120.47, 117.36, 114.18, 113.76, 109.69, 77.05, 55.65, 55.49.
HRMS: Calculated for C15H14BrO3 (M + H), 321.0121. Found, m/z 321.0120.
Synthesis of 1-bromo-4-methoxy-2-(2-(3-methoxyphenyl)propan-2-yl) benzene
The chemical structure is shown as 1c in Extended Data Fig. 8a. Dichloromethane (70 ml) was added to a dry 250 ml round-bottom flask and then cooled to −40 °C. Then, TiCl4 (9 ml) and dimethyl zinc solution (80 ml of 1 M toluene solution) were slowly added with stirring under the protection of argon. After stirring at 40 °C for 15 min, 30 ml of dichloromethane solution of 1b (6.1 g) was added dropwise to the above solution and kept at −40 °C for 3 h. After that, the temperature was slowly raised to 0 °C and the reaction was continued for 5–10 h. The reaction was monitored by TLC and stopped when raw material 1b disappeared. The brown reaction solution was quenched by pouring into crushed ice. The mixture was extracted with dichloromethane, dried over Na2SO4, concentrated on a rotary evaporator and purified by silica gel column chromatography with developing solvent: petroleum ether/ethyl acetate ~15/1, Rf ~0.6 product was collected and concentrated to obtain 5.5 g colourless liquid 1c with yield of 87%.
1H NMR (CDCl3, 500 MHz): δ 7.38 (d, J = 10 Hz, 1H, Ar-H), 7.21 (d, J = 5 Hz, 1H, Ar-H), 7.18–7.14 (m, 1H, Ar-H), 6.87–6.81 (m, 1H, Ar-H), 6.72–6.69 (m, 1H, Ar-H), 6.63 (dd, J = 10 Hz, 5 Hz, 1H, Ar-H), 3.82 (s, 3H), 3.74 (s, 3H), 1.73 (s, 6H).
13C NMR (126 MHz, CDCl3): δ 159.51, 158.70, 151.23, 148.67, 135.90, 133.52, 128.94, 119.03, 115.78, 113.10, 111.93, 109.87, 55.38, 55.14, 44.88, 30.16, 13.29.
HRMS: Calculated for C17H20BrO2 (M + H), 335.0642. Found, m/z, 335.0641.
Synthesis of 1-bromo-4-methoxy-2-(2-(3-methoxyphenyl)propan-2-yl) benzene
The chemical structure is shown as 1d in Extended Data Fig. 8a. Compound 1c (3 g) in 30 ml anhydrous tetrahydrofuran was added into a dry 100 ml round-bottom flask and cooled to −78 °C under argon. t-Butyllithium n-hexane solution (~9–10 mmol) was added ‘with extreme caution’ using a well-dried syringe. The solution was kept with constant stirring at −78 °C to ~−60 °C for 1 h. Then 10 ml anhydrous tetrahydrofuran solution of o-methyl benzaldehyde (11.5 mmol) was slowly added to the reaction at −78 °C with a syringe pump. After the addition was completed, the temperature was allowed to increase slowly with stirring for 2–6 h and monitored by TLC. The reaction was quenched by adding 10 ml of saturated ammonium chloride solution. After removing most of the solvent by rotary evaporator, the residue mixture was extracted three times with dichloromethane (100 ml each), the organic phase was collected, dried over Na2SO4 and concentrated to obtain the crude compound 1d, which was directly used in the next reaction step.
Synthesis of 1-bromo-4-methoxy-2-(2-(3-methoxyphenyl)propan-2-yl) benzene
The chemical structure is shown as 1e in Extended Data Fig. 8a. The compound 1d from the previous step was dissolved in 40 ml of dichloromethane and 4 g of PCC was added under rapid stirring. The reaction was kept for 2–4 h at room temperature and monitored by TLC until the raw material disappeared. The reaction solution was filtered through a short column pad with celite aid and washed with dichloromethane. The filtrate was collected, concentrated on a rotary evaporator and subjected to silica gel column chromatography. Developing solvent: petroleum ether/ethyl acetate ~7/1, Rf ~0.5 product was collected. After concentration, 2.9 g of yellowish solid was obtained as crude compound 1e, which was directly used in the next reaction step without further purification needed.
Synthesis of 1-bromo-4-methoxy-2-(2-(3-methoxyphenyl)propan-2-yl) benzene
The chemical structure is shown as 1 in Extended Data Fig. 8a. Compound 1e (2.9 g) from the above step was dissolved in 40 ml anhydrous dichloromethane under ice-water-bath cooling, and boron tribromide (2–5 eq) was added dropwise (with extreme caution). The reaction was kept for ~2–5 h after the addition was completed. Then, 20 ml of ice water was carefully added to quench the reaction with stirring for 30 min. After that, the reaction was adjusted to pH ~7.0 with saturated aqueous NaHCO3, extracted twice with 150 ml of dichloromethane, and the organic phase washed with water and saturated brine, and dried over anhydrous Na2SO4. The organic phase was concentrated using a rotary evaporator and then drained using a vacuum oil pump under reduced pressure to obtain a slightly yellow oil. Then, 5 ml of methanesulfonic acid was added to the oil and the mixture heated to ~80–100 °C with stirring. The reaction was stopped and cooled to room temperature after 1 h of heating. The reactant was poured into crushed ice while stirring, the precipitated solid was collected by filtration, washed with water and dried under vacuum to obtain ~2.2 g of crude product of the target product 1. The crude product was separated and purified by silica gel column chromatography to obtain 1.8 g of orange-red solid PO with yield of 72%.
1H NMR (CDCl3, 500 MHz): δ 7.32–7.29 (m, 1H, Ar-H), 7.24–7.18 (m, 2H, Ar-H), 7.02 (d, J = 5 Hz, 1H, Ar-H), 6.95 (s, 1H, Ar-H), 6.94 (s, 1H, Ar-H), 6.86 (d, J = 10 Hz, 1H, Ar-H), 6.50 (dd, J = 10 Hz, 5 Hz, 2H, Ar-H), 1.94 (s, 3H), 15.2 (s, 3H), 1.48 (s, 3H).
13C NMR (126 MHz, CDCl3): δ 174.51, 157.24, 155.28, 136.62, 135.96, 130.24, 129.10, 128.84, 125.72, 122.37, 120.66, 118.89, 114.02, 46.98, 40.15, 32.50, 31.93, 19.52.
HRMS: Calculated for C23H19O2 (M−H), 327.1386. Found, m/z 327.1390.
Chip preparation and sequencing
Etching of FOPs
FOPs were fabricated by Guangzhou Honsun Opto-Electronic. The etchant was prepared by adding 85 ml concentrated hydrochloric acid and 20 ml hydrogen peroxide to 895 ml water. One face of the FOP chip was covered with a piece of adhesive tape, while the other face was exposed to etchant. Etching was operated at 25 °C for 2 h, then the chip was rinsed with water and air dried. A thin layer (~100–400 nm) of silica was deposited on the etched FOP by ion-beam deposition.
Silane deposition
The FOP chips were first cleaned using air plasma, and then put into a vacuum oven for hydrophobic modification under argon atmosphere and trichloro(1H, 1H, 2H, 2H-perfluorooctyl)silane vapour under 100 °C and 100 Pa for 1 h. Then, the chips were cooled to room temperature and transferred to another vacuum oven under argon atmosphere and (3-mercaptopropyl)tri-methoxysilane vapour at 100 °C and 100 Pa for 30 min. Finally, the chips were placed on a hot stage at 130 °C to age the silane modification.
Polyethylene glycol and streptavidin coating
The silane-coated chips were rinsed with isopropanol and then 1× PBS buffer. Maleimide–polyethylene glycol–biotin (0.45 mM, M.W. 5,000, Laysan Bio) solution was injected into the chip, incubated for 30 min and washed with 10 ml MilliQ water. Then, 100 μg ml−1 streptavidin (Sigma Aldrich) in 1× PBS buffer was injected into the chip, incubated for 10 min and washed with 10 ml 1× PBS buffer.
Preparation and loading sequencing beads
The Ion Sphere Particles (ISPs) were prepared by emulsion PCR and enrichment following the Ion Torrent OneTouch2 protocol (Thermo Fisher). Then, 100 μl ISP solution dispersed in 1× terminal deoxynucleotidyl transferase buffer was treated with terminal deoxynucleotidyl transferase solution and 10 μM biotin-16-deoxyuridine triphosphate (NEB) at 37 °C for 1 h, followed by addition of 100 μM dideoxynucleotide triphosphate at 37 °C for 4 h to further block all free 3’ DNA terminals. After that, the ISP solution was diluted to 0.3× by 1× PBS and injected into the sequencing chip. The chip was centrifuged under 1,000g for 10 min and washed with wash buffer to remove ISPs that were not firmly immobilized on the chip.
Sequencing
Sequencing was done by sequentially introducing dual-base reaction mix (the dual-base flowgram) as we have previously described15. The sequencing reaction mix contains Bst polymerase, calf intestinal alkaline phosphatase (NEB), MnCl2 (Sigma Aldrich) and two types of fluorogenic nucleotide to provide specific degenerate combinations. In each sequencing cycle, the chip was filled with the sequencing reaction mix at 4 °C and then sealed with fluorinated oil (FC-40, 3 M); the sequencing-by-synthesis reaction was triggered by heating the sealed chip to 65 °C for 30 s. Then, the chip was cooled to 25 °C to stop the reaction and to perform tiling-image acquisition. The chip was then washed with isopropanol and wash buffer at the end of each reaction cycle. The sequencing operation was controlled by a LabVIEW programme.
Base calling
The microwells on the FOP were imaged as multiple bright dots with different intensities in the fluorescent images. For each fluorescent image, the microwells were recognized as local maxima by image morphological dilation. Then, the images were upscaled to increase the resolution. The microwell positions were refined by the weighted average of pixels around the local maxima on the resized image. The intensities of the microwells were extracted using the sum of pixels around the refined positions on the resized image. The extracted intensities were further corrected by subtracting intensities of neighbouring dark microwells. Light stains and dark stains were identified as large areas with over-bright or over-dark intensities, and microwells from stained regions were discarded in the subsequent steps (Extended Data Fig. 3a). Images of the same tile in different cycles were registered through the positioning markers on the bright-field image (Extended Data Fig. 2k).
We plotted a histogram for microwell intensities extracted from each cycle (Extended Data Fig. 4d). Two thresholds were identified in the histogram to separate the first peak as the dark dots, the second peak as the monoclonal bright dots and the rest as polyclonal bright dots. Only monoclonal bright dots were retained for further processing.
We mixed two kinds of DNA samples in each run: the standard dots (SD) and library dots (LD). All SDs have known identical sequences, while LDs are from the sample to be sequenced and have variant sequences. All LDs have an identical starting sequence GTAGCC (the key sequence in Extended Data Fig. 3c), which is different from that of SDs. We calculated the Pearson’s correlation coefficients between intensities of Cycle 1–6 and the degenerate polymer length (DPL) of LD/SD (Extended Data Fig. 4d,e). Dots with correlation >0.9 with the LD DPL were identified as LD, and dots with correlation >0.95 with the SD DPL were identified as SD. Dots that were neither LD nor SD were identified as error dots (ED).
The SD intensities f were corrected from dephasing using our previous algorithm15,18. Briefly, we constructed a flux matrix T from the lead (ε1, ε2), lag (λ1, λ2) and SD DPL h to transform h to a dephased signal s, where ε1 and λ1 are for odd cycles and ε2 and λ2 are for even cycles. Then, the predicted intensities were \({f}^{* }={a\cdot }\left(1-b\right){t\cdot s}+{c\cdot }{s}_{1}+{d\cdot }{s}_{2}\), where a is the unit signal, b is the decay coefficient, c is hydrolysis1 (hydrolysis of odd cycles), d is hydrolysis2 (hydrolysis of even cycles), s1 is 1 in odd cycles and 0 otherwise, s2 is 1 in even cycles and 0 otherwise. We minimized the loss function \(y={{\rm{||}}\;f-{f}^{* }{\rm{||}}}^{2}+{r}^{2}[{\left({\varepsilon }_{1}-{\varepsilon }_{2}\right)}^{2}+{\left({\lambda }_{1}-{\lambda }_{2}\right)}^{2}]\) to fit all the parameters \(({\varepsilon }_{1},{\varepsilon }_{2},{\lambda }_{1},{\lambda }_{2},a,b,c,d)\). The regularization coefficient r was set to 109. The corrected signal was obtained by reverse transformation of intensity f to DPL h using the fitted parameters.
The LD intensities were first normalized to the known DPLs of their key sequence and then corrected from dephasing using the mean dephasing coefficients obtained from SD. The corrected DPL of LD were rounded to their nearest integers and then transformed to bit sequences.
The dephasing correction of SuperBitSeq is similar to that of BitSeq, except that the signals from the two channels were processed separately, according to our previous simulation results18.
Simulation
Mapping the simulated error-free bit sequences to the genome
To test the accuracy of the mapping of bit and superbit sequences, we simulated the mapping of error-free sequences to three genomes: Homo sapiens (GRCh38, or hg38), Arabidopsis thaliana (TAIR10.1) and a simulated random genome. The simulated genome has 3 × 109 bases equally distributed in 20 mock chromosomes (150 Mb each) since BWA can only process chromosomes <2 Gb. The simulated genome fully constituted base A, C, G and T without any ambiguous bases such as N. Each base had equal occurrence probability of 0.25 and the base type in each site of the genome was independent from any other sites. Sites (1 × 107, 1 × 106 and 1 × 107) were uniformly sampled from the GRCh38, TAIR10.1 and the simulated genome, respectively. Regions with ambiguous bases (marked as N in the reference genome) were omitted in the sampling. For each site, by retrieving sequences with different lengths, we simulated sequencing reads under the MK, RY, WS and CRT flowgram with 5–150 cycles (Extended Data Fig. 7a). BitSeq and SuperBitSeq flowgrams were named with a heading ‘b’ and ‘s’, respectively. For example, bMK stands for BitSeq in MK flowgram and sRY stands for SuperBitSeq in RY flowgram. This nomenclature is consistent throughout this paper. No errors were added to these sequences. Sequencing signals were encoded as both bit and superbit sequences according to the method described in ‘Coding strategy compatible with the prevalent mapping software’ in Supplementary Text 2. These sequences were mapped to their corresponding reference genomes using Bowtie2, BWA-MEM or BWA-SW. We calculated the percentage of the following kinds of sequences from the Binary Alignment/Map files (Supplementary Tables 7–24 and 47–52):
-
(1)
Total mapping: mapped to the reference genome by the software;
-
(2)
Unique mapping: no XS tag, or the value of XS tag is less than that of the AS tag;
-
(3)
Q20 mapping: the mapping quality is ≥20;
-
(4)
Q30 mapping: the mapping quality is ≥30;
-
(5)
Correct mapping: the distance between the mapped site and the sampling site is ≤5 bp;
-
(6)
Unique and correct mapping: both unique and correct mapping.
Mapping the simulated erroneous sequences to the human genome
To test the accuracy of the mapping of bit sequences with a few errors, 1 × 106 sites were uniformly sampled from the hg38 genome. Regions with ambiguous bases were omitted in the sampling. For each site, we also simulated sequencing reads under the MK and RY flowgram with 15–150 cycles, as in the error-free sequence simulation. Erroneous sequences under the CRT flowgram were not simulated because they have different error profiles compared with bit sequences (more substitutions than indels). To simulate errors in bit sequences, we used a matrix \({P}_{n\times n}\) to describe the error pattern, in which Pij is the probability that DPL i is sequenced as j. \({P}_{n\times n}\) was generated as:
Then, we normalized P such that the sum of each row is 1. n was set to 50, an integer sufficiently greater than most DPLs. λ was set to 447.6, 222.6, 110.1, 53.8 and 25.7, corresponding to error rates of 0.25%, 0.5%, 1%, 2% and 4%, respectively. In the simulation, DPLs were transformed from the sampled bit sequences, then modified according to the probability given by the matrix P. For example, the probability that DPL i remains unchanged is \({P}_{i,i}\), and the probability that DPL i is erroneously sequenced as i + 1 is \({P}_{i,i+1}\) and so on. The modified DPLs were transformed back to bit sequences and mapped to the reference genome using Bowtie2, BWA-MEM or BWA-SW. The percentages of total mapping, unique mapping, Q20 mapping, Q30 mapping, correct mapping, and unique and correct mapping reads (Supplementary Tables 25–42) were calculated using the same method as in the previous section for mapping the simulated error-free sequences.
Mapping simulated random sequences to the human genome
As a negative control, random sequences were used to validate that non-genome bit sequences would not be erroneously mapped onto the genome. We generated 1 × 106 random DNA sequences of 1 kb. The probability of each of the 4 kinds of base is 0.25 and each site was generated independently from each other. For each random sequence, by retrieving subsequences with different lengths, we simulated sequencing reads under the MK, RY and CRT flowgram with 5–50 cycles, as in the error-free sequence simulation. These sequences were mapped to their corresponding genomes using Bowtie2, BWA-MEM or BWA-SW. The percentages of total mapping, unique mapping, Q20 mapping and Q30 mapping reads were calculated (Supplementary Tables 43–46) using the same method as previously described. Correct mapping or unique and correct mapping reads were not calculated since random sequences should not have been mapped.
Whole-genome screening of faithful mapping sites of the human genome
We defined faithful mapping as:
-
(1)
the sequence being mapped to where it was generated (correct mapping);
-
(2)
the mapping quality was ≥20 (Q20 mapping);
-
(3)
no XS tag, or the value of the AS tag minus that of the XS tag was ≥30 (unique mapping).
We enumerated every 50, 70 and 100 bp sequences on the GRCh38. For example, in the case of 50 bp, the sequences were from sites of 1–50, 2–51, 3–52 and so on until the end of each chromosome. These sequences were transformed into MK and RY bit sequences, and mapped to their corresponding reference genome along with their original DNA sequences using Bowtie2 and BWA-MEM. We selected sites where the generated sequences can be faithfully mapped as the faithful mapping sites. The number of faithful mapping sites of different read lengths, and software are provided in Supplementary Tables 3–8.
Comparison of error-correction capabilities
One round bit sequence or superbit sequence encodes 50% or 84% information of the 4-base DNA sequence, and 3 orthogonal rounds (MK, RY, WS) of them provide information redundancy that can be used to correct potential sequencing errors. To compare their error-correction capability, we performed the following simulation:
-
(1)
Simulate 1 × 107 100-bp random DNA sequence reads (denoted as A);
-
(2)
Calculate their theoretical three-round sequencing signals S;
-
(3)
Add white noise with standard deviation σ = C × S + 0.05 to S (C stands for coefficient of variation (c.v.)); then round to integer D; the 0.05 in σ is to disturb the zero signals in SuperBitSeq;
-
(4)
Use the error-correction principle to check potential sequencing errors in D;
-
(5)
If no error is found, mark D as legal;
-
(6)
For legal signals, decode the signal as a new sequence read B;
-
(7)
If B is not a substring of A starting from the beginning, mark B as false negative (sequencing error not detected);
-
(8)
Calculate the ratio of legal signals and false-negative sequences of BitSeq and SuperBitSeq under different c.v.
Due to the reduced dynamic range and greater information entropy, error correction of SuperBitSeq has higher legal signal rates and lower false-negative rates than BitSeq under the same noise c.v. The results are illustrated in Extended Data Fig. 10f.
Bioinformatic analysis
Analysis of lambda phage genomic DNA sequencing
The bit sequences from the lambda phage genomic DNA were mapped to its corresponding encoded reference genome using BWA-MEM (v.0.7.14) with the following parameters: -B 5 -O 3 -E 3. Only reads with mapping quality ≥20 were used for calculating the error rate, using the method described in ref. 40. Soft-clipped bases were omitted during the calculation.
Identification of CNVs
The bioinformatic pipeline for CNV detection is as follows:
-
(1)
Generating Browser Extensible Data files. We first screened for faithful mapping sites (see section ‘Whole-genome screening of faithful mapping sites of the human genome’). We retained faithful mapping sites that occurred consecutively for over 200 bp and discarded the remaining sporadic faithful mapping sites. We then transformed the retained sites to BED form files. Two different BED files were generated for normal 4-base DNA sequences and bit sequences.
-
(2)
Binning. The reference genome was divided into multiple bins in which every bin contained exactly L faithful mapping sites recorded in the BED files. If there were less than L sites when binning at the end of a chromosome, this bin was discarded. The binning resolution L was 2 Mb for identification of aneuploidies and 100 kb for small CNVs.
-
(3)
Filtering. Sequencing reads were mapped to the reference genome using the default settings of BWA-MEM (v.0.7.14), then transformed to the BAM format and sorted using Samtools (v.1.10). Then, we retained reads that:
-
(a)
were mapped to regions recorded in the BED files;
-
(b)
had mapping quality ≥20;
-
(c)
contained no XS tag in the alignment result, or the value of the AS tag minus the XS tag was ≥5.
-
(a)
-
(4)
Counting. We counted the number Ni of reads mapped to each bin of the reference genome as the sequencing depth.
-
(5)
GC correction. Supposing the GC content of each bin is gi, we used the locally weighted polynomial regression (LOWESS) to fit the curve of \(\log \left({N}_{i}\right)\) with respect to gi. Supposing the fitted values are fi, we calculated \({x}_{i}=\exp \left(\log \left({N}_{i}\right)-{f}_{i}\right)\) as the sequencing depth after GC correction.
-
(6)
Normalization. We calculated the mean value \(\bar{x}\) of \({x}_{i}\) on all autosomes, and calculated \({x}_{i}^{{\prime} }=2{x}_{i}/\bar{x}\) as the normalized depth of each bin.
-
(7)
CNV identification. We used the circular binary segmentation algorithm of the DNAcopy package41 in the R language to calculate the segment mean mi from xi. The segment mean mi is the copy number in this bin.
Non-invasive prenatal testing
We first used the same pipeline as the CNV identification to get the normalized depth \({x}_{i}^{{\prime} }\) using a binning resolution of L = 2 Mb. Then, we calculated the representative value of Chr21 of each sample as
We calculated the mean \({\mu }_{{\rm{chr}}21}\) and standard deviation \({\sigma }_{{\rm{chr}}21}\) of the Chr21 representative values of all 24 negative controls, and the Chr21 Z-score of a sample s to be detected was calculated as:
The Z-scores for Chr18 and Chr13 were calculated in the same manner. A Z-score >3 was tested as trisomy positive, otherwise negative.
The gender of each sample was classified by calculating the mean depth of ChrY as:
In the calculation of \({d}_{{\rm{chrY}}}^{s}\), we omitted the second bin of ChrY for both BitSeq and conventional 4-base DNA sequencing, because this bin is akin to the centromere and its depth varies substantially.
RNA-seq
For conventional 4-base DNA sequences, the transcripts per kilobase million (TPM) of each gene were directly obtained using Salmon (v.1.4.0)42. For bit sequences, we first used the default settings of BWA-MEM to map the reads to the corresponding reference genome, retained reads with <2 errors and an alignment read length ≥70 bp, then used Salmon v.1.4.0 to obtain the TPM.
Metagenomics sequencing
For the untargeted metagenomics sequencing, the raw sequencing data were filtered using fastp v.0.23.2 to remove reads with length <50 bp or mean base quality <20, and trim reads >100 bp to 100 bp. The data were further deduplicated using seqkit v.2.3.0 and reads with >25 bp homopolymers were removed. The filtered data were mapped to the human genome GRCh38 (MK encoded) using hs-blastn v.0.0.5 with default parameters, and the unmapped reads were again mapped to RefSeq (downloaded in May 2022, MK encoded) using hs-blastn with the parameter ‘-evalue 1e-30 -outfmt 6 -dust no -word_size 50’. For each read, only the query with the highest score was retained for species identification. For targeted metagenomics sequencing, the sequences of targeted regions of each species were compiled to form the reference sequences. The sequencing reads were mapped to these reference sequences using BWA-MEM (v.0.7.17-r1188). The read numbers mapped to each species were counted using a bespoke Python script and normalized to the total read number with unit in 100,000 reads.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive43 at the National Genomics Data Center44, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA, CRA024244, CRA024246, CRA024247, CRA024248; GSA-Human, HRA010915) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human.
Code availability
The code demonstrating how to process fuzzy sequences using BitSeq and SuperBitSeq is publicly available at https://github.com/wxzhou/FuzzySequencer (ref. 45).
References
Levy, S. E. & Myers, R. M. Advancements in next-generation sequencing. Annu. Rev. Genomics Hum. Genet. 17, 95–115 (2016).
Shendure, J. et al. DNA sequencing at 40: past, present and future. Nature 550, 345–353 (2017).
Manolio, T. A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Li, R. et al. A body map of somatic mutagenesis in morphologically normal human tissues. Nature 597, 398–403 (2021).
Li, X. et al. Improvement in the risk assessment of oral leukoplakia through morphology-related copy number analysis. Sci. China Life Sci. 64, 1379–1391 (2021).
Zarrei, M., MacDonald, J. R., Merico, D. & Scherer, S. W. A copy number variation map of the human genome. Nat. Rev. Genet. 16, 172–183 (2015).
Wu, A. R., Wang, J., Streets, A. M. & Huang, Y. Single-cell transcriptional analysis. Annu. Rev. Anal. Chem. 10, 439–462 (2017).
Di, L. et al. RNA sequencing by direct tagmentation of RNA/DNA hybrids. Proc. Natl Acad. Sci. USA 117, 2886–2893 (2020).
Wong, F. C. K. & Lo, Y. M. D. Prenatal diagnosis innovation: genome sequencing of maternal plasma. Annu. Rev. Med. 67, 2.1–2.14 (2016).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Metzker, M. L. Sequencing technologies – the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Valouev, A. et al. A high-resolution, nucleosome position map of C. elegans reveals a lack of universal sequence-dictated positioning. Genome Res. 18, 1051–1063 (2008).
Sims, P. A., Greenleaf, W. J., Duan, H. & Xie, X. S. Fluorogenic DNA sequencing in PDMS microreactors. Nat. Methods 8, 575–580 (2011).
Chen, Z. et al. Fluorogenic sequencing using halogen-fluorescein-labeled nucleotides. ChemBioChem 16, 1153–1157 (2015).
Chen, Z. et al. Highly accurate fluorogenic DNA sequencing with information theory-based error correction. Nat. Biotechnol. 35, 1170–1178 (2017).
Li, M. et al. BEAMing up for detection and quantification of rare sequence variants. Nat. Methods 3, 95–97 (2006).
Sood, A. et al. Terminal phosphate-labeled nucleotides with improved substrate properties for homogeneous nucleic acid assays. J. Am. Chem. Soc. 127, 2394–2395 (2005).
Zhou, W. et al. A virtual sequencer reveals the dephasing patterns in error-correction code DNA sequencing. Natl. Sci. Rev. 8, nwaa227 (2021).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie2. Nat. Methods 9, 357–359 (2012).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Wu, M. et al. Pathogenesis diagnosis of a pediatric patient suffering from multi-organ abscesses. Diagn. Pathol. 18, 85 (2023).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Li, X. et al. The comprehensive DNA methylation landscape of hematopoietic stem cell development. Cell Discov. 7, 86 (2021).
Liu, M. C. et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann. Oncol. 31, 745–759 (2020).
Luo, X. et al. 3D genome of macaque fetal brain reveals evolutionary innovations during primate corticogenesis. Cell 184, 723–740 (2021).
Ding, B. et al. Noncoding loci without epigenomic signals can be essential for maintaining global chromatin organization and cell viability. Sci. Adv. 7, eabi6020 (2021).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2017).
Song, S. et al. Genomic epidemiology of SARS-CoV-2 in Pakistan. Genomics Proteomics Bioinformatics 19, 727–740 (2021).
Huang, Y. Covid-19 of Fuzzy-seq. CRA024248. GSA https://ngdc.cncb.ac.cn/gsa/browse/CRA024248 (2025).
Du, M. et al. Unique dual indexing PCR reduces chimeric contamination and improves mutation detection in cell-free DNA of pregnant women. Talanta 217, 121035 (2020).
Wang, Y. et al. Cell-free DNA screening for sex chromosome aneuploidies by non-invasive prenatal testing in maternal plasma. Mol. Cytogenet. 13, 10 (2020).
Huang, Y. A fuzzy sequencer for fast DNA fragment counting and genotyping. HRA010915. GSA-Human https://ngdc.cncb.ac.cn/gsa-human/browse/HRA010915 (2025).
Huang, Y. Metagenomic sequencing of Fuzzy-seq. CRA024246. GSA https://ngdc.cncb.ac.cn/gsa/browse/CRA024246 (2025).
Huang, Y. Lambda phage sequencing of Fuzzy-seq. CRA024247. GSA https://ngdc.cncb.ac.cn/gsa/browse/CRA024247 (2025).
Huang, Y. RNA-seq of mouse cell lines. CRA024244. GSA https://ngdc.cncb.ac.cn/gsa/browse/CRA024244 (2025).
Loman, N. J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Seshan, V. E. & Olshen, A. DNAcopy: DNA copy number data analysis. R package version 1.70.0 (2022).
Patro, R. et al. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Chen, T. et al. The Genome Sequence Archive Family: toward explosive data growth and diverse data types. Genomics Proteomics Bioinformics 19, 578–583 (2021).
CNCB-NGDC Members and Partners. Database resources of the National Genomics Data Center, China National Center for Bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Zhou, W. Demo code for fuzzy sequencing. Zenodo https://doi.org/10.5281/zenodo.15615588 (2025).
Acknowledgements
We thank Y. Shang, W. Li, L. Du, X. Wu, B. Feng, J. Huang, R. Du, N. Wang and W. Wang for experimental and analytical assistance. Funding was partially provided by National Natural Science Foundation of China Grants T2225005 (J.W.) and T2188102 (Y.H.), Beijing National Laboratory for Molecular Sciences Grant BNLMS-CXTD-202401 (Y.H.), Beijing Municipal Science and Technology Commission Grants Z201100005320016 (Y.H.), Z211100003321006 (Y.H.) and Z221100007022003 (Y.H.), National Health Commission of China Grant 2023ZD0519900 (J.W.), and the Beijing Advanced Innovation Center for Genomics.
Author information
Authors and Affiliations
Contributions
Y.H., Z.C. and W.Z. conceptualized the project. W.Z., L.K., S.Q., H.D., C.L., R.Z., L.S., Y.X., W.Y., Y.W., L.D., Y.P., M.L., L.R. and C.Y. conducted experiments. W.Z., L.K., S.Q., Z.L., M.T., L.L., M.D., X.Z., J.W., Z.C. and Y.H. performed data analysis. W.Z., L.K., J.W., Z.C. and Y.H. wrote and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
W.Z., L.K., S.Q., H.D., Z.C. and Y.H. were listed as inventors of fuzzy sequencing in patent applications CN111575355B, CN111667882B and CN108165616B, all filed in China in December 2016. H.D., Z.C. and Y.H. are founders, and R.Z., L.L. and L.S. are employees, of Cygnus Biosciences. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic of commercial high-throughput sequencing chemistries.
(a) Schematic of Cyclic reversible terminator (CRT) used by Illumina and MGI. (b) Schematic of triple-nucleotide addition (TritSeq). (c) Nucleotide substrates used in different sequencing chemistries. The TPLFN is used in fluorogenic sequencing, that is, error-correction code (ECC) sequencing, BitSeq and SuperBitSeq. (d) The information efficiency of different sequencing technologies. Compared with SNA or CRT, BitSeq and SuperBitSeq show higher information efficiency that can be illustrated as the product of intrinsic and extrinsic components.
Extended Data Fig. 2 The instrument and chip design.
(a) The ‘black-box’ prototype with major components including the camera, the imaging optics, the motion stages equipped with sequencing modules, as well as the liquid delivery. (b) Inside the ‘black-box’. Two sequencing modules sit on the moving stages. Above one of the sequencing module is the microscopic objective lens mounted on a piezo focusing adjustment stage. The manifolds are used for connection between the flow cell and the fluidic system (the selector and solenoid valves are shown in the photo). (c) The graphical user interface of the sequencer operating software (written in LabVIEW). (d) The schematic of the structure and fabrication of the flow-cell chip. (e) The fully assembled flow-cell chip. (f) The flow-cell chip and its holder. (g) The fluid flowing process in each reaction period that contains two reaction cycles. (h) The bleaching test to confirm the effectiveness of sealing of the microreactors. Top: Field-of-view (FOV) where fluorophores were sealed inside the microwells by FC-40. Middle: Using the pinhole to selectively photo-bleach part of the FOV, the fluorescent intensity is lower in the exposed area. Bottom: After several minutes, the margin of the exposed area remains sharp and clear, indicating the sealing is successful so that non-bleached fluorophores in the unexposed area cannot diffuse into the bleached area. (i) When sealing fails, the margin of exposed area is blurred, and the intensity of the exposed area recovers as a result of fluorophore diffusion. (j) A typical example of light stains (blue) and dark stains (red). (k) A typical example of bright-field image with tetris-like positioning markers.
Extended Data Fig. 3 The data processing pipeline and library structure.
(a) The schematic of the data processing pipeline. Defects are filtered and the intensity of each microwell at each reaction cycle is extracted and further filtered to keep monoclones for dephasing correction. The correction parameters are determined by fitting the signal to the standard dots (SD) which have been spiked into the sequencing library. (b) >90% of the area remains stain-free after 61 cycles of sequencing. (c) The structure of the DNA library. The 8-bp index sequence is written as I’s and the inserted part is written as N. The library is modified from ion torrent’s library by adding an extended sequence to enhance the melting temperature of the sequencing primer and a key sequence for LD identification. The underlined TCAG is used in ion torrent for library identification and signal normalization.
Extended Data Fig. 4 The base-calling results.
(a–c) Spatial distribution of light stains (a), dark stains (b) and both (c) in one sequencing experiment. The beads within these stains are excluded from further analysis. (d) Schematic of how dots were selected according to their intensities. The three peaks were identified as dark dots, monoclonal and polyclonal, respectively. (e) LD and SD were identified through the Pearson’s correlation coefficient ρ between the raw intensities of each dot and the starting DPL of LD/SD. (f, g) Signal distribution of SD (f) and LD (g) in each cycle. Top: intensities normalized by dividing by the intensities of Cyc 2. Bottom: DPLs corrected from dephasing. Black: intensity distributions. Yellow dots: Ideal DPLs equal to the rounded mean intensity. Red dots: Ideal DPLs not equal to the rounded mean intensity. The SD all had the same sequence and were shown as single peaks in all cycles. The first 7 cycles of LD were from the key region, showing as a single peak. The latter cycles of LD were from the inserted random region, showing as multiple peaks. (h–m) Spatial distribution of loading rate (the percentage of nonempty wells) per tile (h), number of light dots (i), number of LD (j), unit signals (k), number of SD (l), number of ED (m). In i-m, numbers were counted in 8×8 regions that each tile was divided into. (n) The distributions of signal parameters estimated from SD. (o) The spatial distribution of number of errors per read in SD. (p) The distribution of number of errors per read in SD. (q) Error rate of SD. The error rate by base is the error rate by cycle divided by the corresponding DPL. (r) The percentage of reads with zero or one error in the first n cycles.
Extended Data Fig. 5 Additional comparison of BitSeq with commercial sequencers in resequencing.
(a, b) The normalized sequencing depth of a normal male adult by BitSeq (a) and Ion Torrent (b). (c, d) The normalized sequencing depth of a Trisomy 21 patient by BitSeq (c) and Ion Torrent (d). (e) The average sequencing depth of ChrY of each sample. These values by both BitSeq and ion torrent differ substantially between male and female fetuses. (f) Number of genes detected using different thresholds for RNA-seq data of mES and MEF cells. (g) Venn diagram of TPM > 1 genes. (h, j) Gene Ontology analysis of genes that were only detected by each sequencer in RNA-seq.
Extended Data Fig. 6 The normalized sequencing depth of 53 patients involved in the NIPT study.
X axis: Genome sites. Green dots: BitSeq. Orange dots: Ion Torrent. The dark and light gray backgrounds indicate the chromosomes.
Extended Data Fig. 7 Mapping capabilities of BitSeq and SuperBitSeq.
(a) How error-free sequences were simulated. Analogous simulation method was adopted by SuperBitSeq. CRT: cyclic reversible terminator (the sequencing chemistry used by Illumina and MGI sequencers). (b) Longer read length or more information enable more accurate genome mapping. (c) Polymer distributions of genomes of lambda phage (top left), Escherichia coli (top right) and Saccharomyces cerevisiae (bottom left).
Extended Data Fig. 8 Chemical synthesis and characterization of Peking Orange (PO).
(a) The synthetic route of the fluorophore PO. (b–i) NMR of intermediate products. (b, c) Chemical 1a. (d, e) Chemical 1b. (f, g) Chemical 1c. (h, i) Chemical PO. (bdfh) 1H-NMR. (cegi) 13C-NMR. (j) HRMS of PO. (k) Excitation and emission spectra of PO. (l) The fluorescence emission of PO is pH sensitive. The pink shade indicates the standard deviation. (m) Fluorogenic kinetics of PO-dN4P extending multiple bases. (n) The sequencing reaction kinetics using PO-dN4P and TG-dN4P as substrates.
Extended Data Fig. 9 Single-template sequencing results.
(a–d) BitSeq using PO. (e, f) SuperBitSeq using PO and TG. Top of each panel: fluorescent intensities. Bottom of each panel: dephasing-corrected signals. Black dots represent signal of each replicates. Bars represent the mean signal of replicates. Yellow dots indicate the theoretical values where the corresponding mean signal is erroneous. a-d share the same legend and e-f share the same legend.
Extended Data Fig. 10 High-throughput SuperBitSeq results.
(a) Parameter and read length distributions of dephasing correction in SuperBitSeq. (b–d) SNV reconstruction in SuperBitSeq. (b) Site map is defined as the positions of encoded bases in the original genome and is recorded during SuperBitSeq encoding. (c) In some cases SNV information may be altered after SuperBitSeq encoding. (d) Schematic of SNV reconstruction. The sequencing read and its matched reference sequence are separated into two semi-sequences, respectively. One semi-sequence is composed of A and G, which are both labeled with the green fluorophore Tokyo Green. The other semi-sequence is composed of C and T, which are both labeled with the red fluorophore Peking Orange. The semi-sequences are locally aligned individually, then the two alignments are merged together by sorting the site map in ascending order. (e) Normalized depth of the 3 SARS-CoV-2 samples. Bin size: 200 bp. (f) Error-correction capability of three rounds of bit sequences or superbit sequences.
Supplementary information
Supplementary Information (download PDF )
Supplementary Texts 1–5.
Supplementary Tables (download XLSX )
Supplementary Tables 1–59.
Supplementary Video 1 (download MP4 )
The fluorescence imaging process. After each cycle of reaction, the flowcell was cooled down to room temperature and then the imaging process started. The fluorescence images were taken in tiles, and in every cycle, we took 171 (19 × 9) images to cover the whole flowcell.
Supplementary Video 2 (download MP4 )
A small field of view demonstrating the cyclic reaction and fluorogenic signals. The total cycle number is 69 for this experiment, and the images drifted due to motion errors and thermal cycles. Through image registration, every microwell is indexed, two of which are labelled by red and green circles in the video. The signal intensity varies in each well through the cyclic sequencing reaction, representing the different lengths of elongation during each reaction cycle. Such intensity series can be quantified and then subjected to image processing and other correction procedures to generate the bit sequences.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, W., Kang, L., Qiao, S. et al. A fuzzy sequencer for rapid DNA fragment counting and genotyping. Nat. Biomed. Eng 9, 2068–2082 (2025). https://doi.org/10.1038/s41551-025-01430-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41551-025-01430-8