Abstract
Metabolic-dysfunction-associated steatohepatitis (MASH) is the leading cause of chronic liver disease, but an incomplete understanding of MASH-induced liver fibrosis has limited therapeutic options. Here we show that hepatocyte caspase-8 drives MASH fibrosis through an apoptosis-independent mechanism. Hepatic caspase-8 expression correlates with liver fibrosis in both human and experimental MASH, and hepatocyte-specific caspase-8 deletion in male mice with MASH suppressed liver fibrosis and hepatic stellate cell (HSC) activation without affecting hepatocyte apoptosis. Mechanistic studies showed that a caspase-8–YY1 pathway in hepatocytes induces secretory meteorin (Metrn), which activates HSCs via a c-Kit–STAT3 pathway. Meteorin expression was increased in human and male mouse MASH livers and decreased by deletion of hepatocyte caspase-8 in MASH mice and human and mouse primary hepatocytes. Genetic restoration of hepatocyte meteorin in hepatocyte-caspase-8-deleted MASH mice restored HSC activation and liver fibrosis while silencing hepatocyte meteorin lowered liver fibrosis. These findings reveal a therapeutically targetable pathway promoting MASH fibrosis involving a non-apoptotic function of caspase-8 and a newly discovered HSC activator, meteorin.
Similar content being viewed by others
Main
MASH (formerly non-alcoholic steatohepatitis (NASH)) is a progressive form of metabolic-dysfunction-associated liver disease (MASLD; formerly non-alcoholic fatty liver disease (NAFLD)) characterized by liver steatosis, inflammation and fibrosis, with the potential to progress to cirrhosis and liver failure1,2. MASH is emerging as the leading cause of liver disease3, but therapeutic options are limited. This is largely attributable to the incomplete understanding of how steatosis progresses to liver fibrosis, which is the main contributor to both liver- and non-liver-related mortality in MASH4,5. The primary mechanism of liver fibrosis in MASH involves the activation of collagen-producing hepatic stellate cells (HSCs), facilitated by hepatocyte–HSC cross-talk2,6.
Caspase-8, an apoptosis-inducing protease, has been identified as a possible contributor to MASH progression. Apoptosis is a feature of MASH7, and in a multi-ancestry genome-wide association study, a variant at rs10201587, located in the intron of CASP8, was associated with both increased CASP8 expression in the liver8 and chronically elevated plasma alanine transaminase (ALT) levels (P < 5 × 10−8)9, a marker of liver injury. However, in a clinical MASH trial testing the efficacy of emricasan, a drug that blocks the proapoptotic protease activity of many caspases, including caspase-8, liver fibrosis was worse in the drug-treated cohort10. Moreover, two studies investigating the deletion of hepatocyte caspase-8 in a weight-loss model of fatty liver disease (the methionine-choline-deficient model) came to opposite conclusions regarding the role of caspase-8 in liver injury, inflammation and fibrosis11,12. In one study, liver disease was mitigated in the knockout cohort, suggesting that caspase-8 is pathogenic, and there was a decrease in liver cell apoptosis11. In the other study, the knockout cohort experienced worse liver disease, suggesting that caspase-8 is protective, and there was an increase in the expression of RIP3, a pro-inflammatory signalling molecule that is negatively regulated by caspase-8 (ref. 12). A third study reached conclusions similar to those of the first study, but the investigators used an unusual hepatocyte JNK1- and JNK2-deficient model that promotes liver cell death13. Thus, the roles and mechanisms of caspase-8 in MASH, particularly MASH fibrosis, in humans and experimental models remain unclear.
To address this gap, we undertook a study using several bona fide mouse MASH and steatohepatitis models in which human-relevant diets were used to induce metabolic liver disease, including early fibrosis. Moreover, we interrogated human MASH liver specimens for evidence of pathways revealed by our mouse studies. Our findings show that the non-apoptosis-inducing form of hepatocyte caspase-8 is increased in human and mouse MASH liver and, on the basis of findings from hepatocyte-specific knockout studies, caspase-8 promotes HSC activation and MASH fibrosis. The mechanism does not involve the apoptosis-inducing function of caspase-8. Rather, elevated caspase-8 in hepatocytes in MASH leads to Ying Yang 1 (YY1)-mediated induction of a gene, Metrn, encoding meteorin, which has never before been implicated in liver function or pathology. In the setting of MASH, meteorin is secreted from hepatocytes in a caspase-8–YY1-dependent manner and then activates HSCs. Notably, we showed evidence of this pathway in human MASH liver, and the non-apoptotic role of caspase-8 in this pathway is consistent with the aforementioned emricasan trial, in which inhibiting the cell-death function of caspases did not improve liver fibrosis in people with MASH10. These findings herein reveal a heretofore unknown and therapeutically targetable pathway that contributes to MASH fibrosis.
Results
Hepatocyte caspase-8 deletion suppresses fibrotic MASH but not apoptosis
Before conducting causation studies in experimental MASH, we analysed CASP8 expression in human MASH versus control liver using datasets from the Gene Expression Omnibus. We found that the average level of CASP8 mRNA expression was substantially higher in the livers of individuals with histologically confirmed MASH than in the livers of lean, healthy control individuals (Extended Data Fig. 1a). To complement these findings, we show that the expression of full-length caspase-8 protein was higher in the livers of individuals with MASH than in healthy controls (Extended Data Fig. 1b). We then turned to two models of experimental MASH, in which mice were fed a fructose, palmitate and cholesterol (FPC) diet14 or an American-lifestyle-induced obesity syndrome diet (ALIOS)15. We also used a well-characterized steatohepatitis model in which mice are fed a high-fat, choline-deficient l-amino-acid-defined diet (HF-CDAA)16. In all three models, full-length caspase-8, not the apoptotic form p18, was markedly higher in the livers of mice fed the MASH/steatohepatitis diets than in those fed the chow diet. We also observed a modest increase in the cleaved form of caspase-8 (p43) and an increase in Casp8 mRNA levels, but not the cleaved form of caspase-8 (p18), in the FPC model (Extended Data Fig. 1c–e). Moreover, caspase immunostaining revealed an increase in hepatocyte caspase-8 in human MASH versus normal livers, (Extended Data Fig. 1f), and caspase-8 expression was elevated in hepatocytes isolated from human MASH livers versus normal livers (Extended Data Fig. 1g).
To determine whether this increase in caspase-8 plays a role in the progression of steatosis to MASH, we injected AAV8-TBG-Cre into Casp8fl/fl mice with pre-MASH hepatosteatosis, which had been fed an FPC diet for 8 weeks to specifically delete hepatocyte caspase-8 (ref. 14); Casp8fl/fl mice injected with AAV8-TBG-LacZ served as the control. The mice were then maintained on the FPC diet for an additional 8 weeks to allow for the development of MASH and early fibrosis (Fig. 1a). In our previous work, we documented that in 16-week-old FPC-fed mice, liver fibrosis, markers of HSC activation, pro-fibrotic gene expression and plasma ALT levels are higher than in chow-fed control mice14. For example, Sirius Red staining is approximately fourfold higher than that in the livers of chow-fed mice, and Tgfb1 and Acta2 levels are approximately five- to sixfold higher14. We confirmed successful caspase-8 knockout in the livers of the AAV8-TBG-Cre-treated mice (H-Casp8-KO) (Fig. 1b), and there were no significant differences in body weight, liver to body weight ratio, blood glucose levels or liver triglyceride content between the two groups (Extended Data Fig. 2a–d). Most notaly, the Sirius-red-positive area was smaller in the livers of H-Casp8-KO (Cre) than in those of control (LacZ) mice (Fig. 1c, top images), indicating that liver fibrosis decreased. There was also a modest decrease in the number of mononuclear cells in the KO livers (Fig. 1c, bottom images), suggesting a reduction in inflammation. Immunofluorescence staining revealed that the α-smooth muscle actin (α-SMA)-positive area was reduced, indicating fewer HSCs and F4/80-positive macrophages in the KO livers, providing further evidence of decreased fibrosis and inflammation (Fig. 1d,e). Furthermore, the livers of the H-Casp8-KO mice ehibited lower expression levels of mRNAs associated with HSC activation, fibrosis and inflammation (Fig. 1f), and plasma ALT concentration was also lower in the KO mice (Fig. 1g). However, liver neutrophil numbers were similar between the two groups (Extended Data Fig. 2e). Importantly, the percentages of cleaved-caspase-3-positive (apoptotic) and TUNEL-positive hepatocytes were not significantly different between the two groups (Fig. 1h,i). As expected, no cleaved caspase-3-positive (apoptotic) cells were detected in the livers of normal, chow-fed mice (Extended Data Fig. 2f). These data, along with the finding that liver expression of the full-length (non-activated) form of caspase-8 correlates with MASH in humans and mice, led us to hypothesize that hepatocyte caspase-8 promoted MASH progression through a non-apoptotic mechanism, which we will further explore below.
a–i, Male Casp8fl/fl mice were fed the FPC diet for 8 weeks to induce steatosis. The mice were then injected with AAV8-TBG-Cre or AAV8-TBG-LacZ control and continued on the FPC diet for 8 more weeks. a, The experimental set-up. b, Immunoblot of caspase-8 (Casp8) in liver, with β-actin as the loading control. c, Staining of liver sections for Sirius red (top; scale bar, 500 μm) and H&E (bottom; scale bar, 100 μm), with quantification of the Sirius-red-positive area (arrows) and hepatic mononuclear cells. Data are presented as mean ± s.e.m.; *P = 0.019, **P = 0.0021, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. d, α-SMA immunofluorescence (red) and quantification. DAPI counterstain (blue) for nuclei is shown. Scale bar, 200 μm. Data are mean ± s.e.m.; ****P < 0.0001, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. e, F4/80 immunofluorescence (red; arrows) and quantification. DAPI counterstain (blue) is shown. Scale bar, 200 μm. Data are mean ± s.e.m.; **P = 0.0038, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. f, Liver Tgfb1, Spp1, Acta2 and Mcp1 mRNA levels. Data are mean ± s.e.m.; *P = 0.024, **P = 0.0011 (Spp1), ***P = 0.0005 (Acta2), ****P < 0.0001, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. g, Plasma ALT levels. Data are mean ± s.e.m.; **P = 0.007, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. h, Cleaved caspase-3 (cl-Casp3) immunofluorescence (red; arrows) and quantification. DAPI counterstain (blue) is shown. Scale bar, 200 μm. Data are mean ± s.e.m.; n.s., non-significant, unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8. i, TUNEL staining (red; arrows) and quantification; DAPI counterstain (blue) is shown. Scale bar, 50 μm. Data are mean ± s.e.m.; unpaired two-tailed Student’s t-test; LacZ, n = 8; Cre, n = 8.
We then tested the effect of H-Casp8-KO in HF-CDAA-fed mice, in which the development of steatohepatitis is more rapid and associated with a higher incidence of hepatocyte apoptosis than in the FPC model16. As with the FPC model, a previous study documented that liver fibrosis, markers of HSC activation, pro-fibrotic gene expression and plasma ALT levels are higher in HF-CDAA-fed mice than in chow-fed control mice14. For example, in that study, the Sirius-red-positive area was approximately tenfold higher in HF-CDAA-fed mice livers than in those of chow-fed mice; Tgfb1 and Acta2 expression levels were also elevated three- to fivefold14, consistent with our Sirius red staining and Tgfb1 expression data (Extended Data Fig. 2g,h). Casp8fl/fl mice were fed the HF-CDAA diet for 4 weeks, injected with AAV8-TBG-Cre (or AAV8-TBG-LacZ) and continued on the diet for an additional 4 weeks (Fig. 2a). Caspase-8 deletion in the Cre-treated livers was confirmed (Fig. 2b), and the two cohorts had similar body and liver weights, blood glucose levels, liver triglyceride content and plasma ALT levels (Extended Data Fig. 2i–m). As with the FPC model, H-Casp8-KO led to decreases in liver fibrosis and inflammation (Fig. 2c–e and Extended Data Fig. 2n), although total numbers of liver T cells were similar (Extended Data Fig. 2o). Despite the relatively high level of hepatocyte apoptosis in HF-CDAA-fed mice, the percentage of apoptotic hepatocytes and DNA double-strand breaks was not significantly different between the two groups (Fig. 2f and Extended Data Fig. 2p,q). These combined data provide evidence that hepatocyte caspase-8 is increased in MASH and contributes to MASH fibrosis independently of apoptosis changes.
Male Casp8fl/fl mice were fed the HF-CDAA diet for 4 weeks to induce steatosis. The mice were then injected with AAV8-TBG-Cre or control AAV8-TBG-GFP and continued on the HF-CDAA diet for 4 more weeks. a, The experimental scheme. b, Immunoblot of caspase-8 in liver, with β-actin as the loading control. c, Staining of liver sections for Sirius red (arrows), with quantification. Scale bar, 200 μm. Data are presented as mean ± s.e.m.; ****P < 0.0001, unpaired two-tailed Student’s t-test; GFP, n = 7; Cre, n = 7. d, COL1A1 immunohistochemistry and quantification. Scale bar, 200 μm. Data are mean ± s.e.m.; ***P = 0.0002, unpaired two-tailed Student’s t-test; GFP, n = 7; Cre, n = 7. e, α-SMA immunofluorescence (red) and quantification. DAPI counterstain (blue) for nuclei is shown. Scale bar, 200 μm. Data are mean ± s.e.m.; **P = 0.0013, unpaired two-tailed Student’s t-test; GFP, n = 7; Cre, n = 7. f, Cleaved caspase-3 immunofluorescence (red; arrows) and quantification; DAPI counterstain (blue) is shown. Scale bar, 200 μm. Data are mean ± s.e.m.; unpaired two-tailed Student’s t-test; GFP, n = 7; Cre, n = 7.
Meteorin links caspase-8 to HSC activation
We next sought to explore potential mechanisms linking hepatocyte caspase-8 to the progression of liver fibrosis in MASH. Activated HSCs are the main source of collagen-producing myofibroblasts in fibrotic liver disease and play a key role in MASH fibrosis2,6,17. To look for a link between hepatocyte caspase-8 and HSC activation, we began with an ex vivo model14 in which primary mouse or human HSCs were incubated with conditioned media (CM) from mouse or human primary hepatocytes transfected with Casp8-targeted small interfering RNA (siCasp8) or control siRNA. Consistent with the in vivo data, levels of markers of HSC activation were reduced in HSCs incubated with CM, from Casp8-silenced AML12 mouse hepatocytes, THLE2 human hepatocytes or primary human hepatocytes compared with control CM (Fig. 3a, and Extended Data Fig. 3a–d). Further, human spheroid data showed a reduction in fibrotic gene expression in caspase-8-silenced spheroids (Extended Data Fig. 3e). Consistent with the idea that caspase-8 promotes HSC activation independently of its apoptotic function (described above), we found that treatment of hepatocytes with the caspase-8 inhibitor IETD, which blocks apoptosis18, did not impede hepatocyte CM from activating HSCs (Extended Data Fig. 3f). To further support this idea, we transfected Casp8-silenced hepatocytes with wild-type caspase-8 versus catalytically inactive caspase-8-C360A, which cannot induce apoptosis19. We then tested the ability of CM from these cells to activate HSCs ex vivo. As before, the HSCs were less activated by the CM of Casp8-silenced hepatocytes than by the CM of control hepatocytes, and CM-induced HSC activation was restored equally by transfecting the Casp8-silenced hepatocytes with wild-type or mutant caspase-8-C360A (Extended Data Fig. 3g). Finally, heat treatment of the CM before incubation with the HSCs blocked CM-induced HSC activation and abrogated the difference between siCasp8 and control conditions, suggesting at least one secreted protein was responsible for hepatocyte caspase-8-dependent activation of HSCs (Extended Data Fig. 3h).
a, AML12 mouse hepatocytes were transfected with scrambled RNA (Scr) or siCasp8. After 24 h, the media were changed to DMEM with 0.1% FBS, and after an additional 24 h, the CM were transferred to quiescent primary mouse HSCs. After 72 h,Timp1, Spp1 and Col1a1 mRNA levels were analysed in the HSCs. Data are presented as mean ± s.e.m.; *P = 0.012, *P = 0.0001, ****P < 0.0001; two-way ANOVA; n = 4 per group. b, The scheme for screening caspase-8-dependent secretory genes and proteins in hepatocytes that can activate HSCs. c, Volcano plot from the RNA-seq data; false discovery rate (FDR)-adjusted P value < 0.05 (two-tailed t-test) was classified as differentially expressed. d, Heat plot for the RNA-seq data. e, Venn diagram plot from the RNA-seq and LC–MS/MS data, integrated with a database of secretory proteins; the asterisk indicates the region of candidate genes. f, mRNA levels of candidate genes in the livers of the mice in Fig. 1a–i. Data are mean ± s.e.m.; *P = 0.029 (Ccl7), *P = 0.014 (Ccl9), **P = 0.0012, two-way analysis of variance (ANOVA); LacZ, n = 4, Cre, n = 4. g, Immunoblot of meteorin in the livers of control or hepatocyte caspase-8-knockout mice fed the FPC diet for 16 weeks, with quantification. Data are mean ± s.e.m.; ***P = 0.0005, unpaired two-tailed Student’s t-test; LacZ, n = 4; Cre, n = 4. h, Immunoblot of meteorin in the livers of control or hepatocyte caspase-8 knockout mice fed the HF-CDAA diet for 8 weeks, with quantification. Data are mean ± s.e.m.; **P = 0.0022, unpaired two-tailed Student’s t-test; GFP, n = 4; Cre, n = 4. i, In Scr- and siCasp8-treated AML12 cells, caspase-8 and meteorin (Metrn) levels were analysed by immunoblot. j, In Scr- and siCASP8-treated primary human hepatocytes, levels of CASP8 and METRN mRNA and caspase-8 and meteorin protein were analysed. Data are mean ± s.e.m.; ***P= 0.0005, ****P < 0.0001, two-way ANOVA; n = 3 per group.
To identify candidate secretory proteins that were caspase-8-dependent and potential activators of HSCs, we integrated the following three data sets: liquid chromatography–tandem mass spectrometry (LC–MS/MS) of conditioned media from the cells, to identify secretory proteins that were diminished in Casp8-silenced AML12 mouse hepatocytes (Fig. 3b); RNA sequencing (RNA-seq) to identify genes encoding secretory proteins that were diminished in Casp8-silenced AML12 mouse hepatocytes (Fig. 3c,d); and a database of secretory proteins. On the basis of overlapping proteins and genes in all three analyses (Fig. 3e), we selected 11 genes for quantitative PCR (qPCR) confirmation in the H-Casp8-KO FPC-MASH livers to determine whether their expression decreased when hepatocytes were deficient in caspase-8. Among these genes, Metrn, encoding the protein meteorin, emerged as a candidate (Fig. 3f). Further investigation revealed that levels of the meteorin protein were decreased in the livers of H-Casp8-KO versus control MASH mice (Fig. 3g,h). Likewise, Casp8 silencing in primary mouse hepatocytes lowered meteorin protein levels (Fig. 3i). CASP8 silencing also reduced levels of METRN mRNA and meteorin protein in primary human hepatocytes (Fig. 3j) and THLE2 human hepatocytes (Extended Data Fig. 3i).
Given that there were no reports on the role of meteorin in liver disease or fibrosis, we conducted an initial proof-of-concept test to determine whether recombinant meteorin could activate primary HSCs ex vivo. We found that exposing primary mouse HSCs and primary human HSCs isolated from either normal or MASH liver to recombinant meteorin caused increases in levels of mRNAs associated with HSC activation, for example Timp1, Col1a1 and Spp1 (Fig. 4a). Next, returning to the above conditioned-medium-transfer experiment, we transfected Metrn into Casp8-silenced hepatocytes to restore Metrn levels to those of the to Scr control (Fig. 4b), followed by transfer of the conditioned medium to HSCs. The data show that HSC activation by conditioned medium from Casp8-silenced hepatocytes was restored partially (Timp1) or fully (Spp1) after the hepatocytes were transfected with Metrn (Fig. 4c). Similarly, the conditioned medium from METRN-silenced human hepatocytes led to less HSC activation than did the control hepatocyte medium (Fig. 4d and Extended Data Fig. 4a). Further, treatment of AML12 hepatocytes with the pan-caspase inhibitor ZVAD or the caspase-8 inhibitor IETD (above), which successfully blocked Jo2/Fas-mediated caspase-3 and caspase-8 cleavage, did not impede hepatocyte conditioned medium from inducing Metrn expression (Extended Data Fig. 4b). As further evidence that the proteolytic domain of caspase-8 is not involved, we transfected CASP8-silenced primary human hepatocytes with plasmids containing the amino-terminal death-effector domains (DEDs) of caspase-8 but lacking the carboxy-terminal p18/p10 proteolytic domain19, finding that the DEDs could restore METRN expression (Extended Data Fig. 4c). Notably, caspase-10 also contains a DED domain, and we observed increased levels of full-length caspase-10 protein in human MASH livers (Extended Data Fig. 4d). Similarly, caspase-10 was elevated in isolated human MASH hepatocytes (Extended Data Fig. 4e). However, CASP10 silencing in human hepatocytes did not alter METRN mRNA expression levels (Extended Data Fig. 4f). Thus, caspase-8-induced HSC activation and meteorin expression are independent of the pro-apoptotic protease activity of caspase-8, setting the stage for meteorin-causation experiments in MASH mice, described below.
a, Primary mouse HSCs (left), primary (1o) human HSCs from a healthy liver (middle) and primary HSCs from the livers of individuals with MASH (right) were treated with 200 ng ml−1 recombinant meteorin (r-meteorin) or vehicle for 48 h. The HSCs were then assayed for fibrotic gene expression by qPCR. Data are presented as mean ± s.e.m. Left, *P = 0.013, ****P < 0.0001; middle, *P = 0.012 (TIMP1), *P = 0.013 (SPP1), ***P = 0.0004; right, *P = 0.015, **P = 0.006, ****P < 0.0001, two-way ANOVA; mouse HSCs, n = 4 per group, human HSCs, n = 3 per group. b, AML12 cells were transfected with Scr siRNa or siCasp8 and with GFP or Metrn plasmid, as indicated. After 24 h, the conditioned media were changed to DMEM with 0.1% FBS, and after a further 24 h, the AML12 cells were assayed for meteorin and caspase-8 protein. c, Conditioned media from the AML12 cells in b were transferred to quiescent HSCs. After 72 h, the HSCs were assayed for Timp1 and Spp1 mRNA. Data are mean ± s.e.m.; *P = 0.012, ****P < 0.0001, one-way ANOVA; n = 4 per group. d, THLE2 cells were transfected with Scr or siMETRN. After 24 h, the media from THLE2 were transferred to quiescent primary mouse HSCs. After 72 h, Timp1 and Spp1 mRNA levels in the HSCs were assayed. Data are mean ± s.e.m.; *P = 0.014, ****P < 0.0001, two-way ANOVA; n = 4 per group. e, Primary mouse HSCs were treated with 400 ng ml−1 r-meteorin or vehicle for 48 h. The levels of phosphorylated (p-Y705) and total Stat3 in the HSCs were then assessed using immunoblot. f, Primary mouse HSCs were transfected with scrambled (Scr), siKit or siStat3. After 24 h, the cells were treated with vehicle or r-meteorin. After another 48 h, Timp1 and Spp1 mRNA levels were measured in the HSCs. Data are mean ± s.e.m.; **P = 0.01, ***P = 0.0007, ****P < 0.0001, one-way ANOVA; n = 4 per group.
We next sought to elucidate the signalling pathway involved in meteorin-induced HSC activation. For this purpose, we focused on previous work showing that STAT3 can drive the pro-fibrotic activation of HSCs20,21 and that STAT3 activation in the liver correlates with MASH fibrosis in humans22. We then reasoned that meteorin might activate STAT3 in HSCs, given that meteorin-like protein, which has ~45% homology to meteorin, can activate the c-KIT receptor23, and c-KIT is a known activator of STAT3 (ref. 24). In this context, we found that treatment of primary mouse HSCs with recombinant meteorin markedly increased Y705 phosphorylation of STAT3 (Fig. 4e), a measure of STAT3 activation. Second, both siKit and siStat3 decreased meteorin’s ability to boost Timp1 and Spp1 levels in primary mouse HSCs (Fig. 4f and Extended Data Fig. 4g). These data provide initial ex vivo support for the idea that meteorin activates HSCs through a c-KIT–STAT3 pathway.
Finally, as an initial step towards understanding how caspase-8 might increase Metrn levels, we performed chromatin immunoprecipitation (ChIP) analysis, which revealed that Casp8 silencing reduced Pol-II binding to the Metrn promoter in AML12 hepatocytes (Extended Data Fig. 4h), suggesting that caspase-8 induces Metrn transcription. We then conducted an in silico analysis of the Metrn promoter and found a binding site for the transcription factor YY1, consistent with a report studying YY1-mediated induction of Metrn in hypoxic haematopoietic stem or progenitor cells25. Most importantly, YY1 silencing lowered Metrn levels in AML12 hepatocytes and METRN in primary human hepatocytes (Fig. 5a). Similarly, YY1 silencing blocked the ability of DEDs to induce Metrn in Casp8-silenced hepatocytes (Fig. 5b). Further, using YY1 ChIP analysis, we showed reduced YY1 binding to the Metrn promoter upon Casp8 silencing (Fig. 5c). Silencing Casp8 reduced nuclear YY1 levels in mouse AML12 hepatocytes and primary human hepatocytes, as determined by immunofluorescence microscopy (Extended Data Fig. 5a–c). Similarly, in FPC-fed HC-Casp8 knockout livers, nuclear YY1 was substantially reduced, as shown by immunohistochemistry (Extended Data Fig. 5d). Moreover, immunofluorescence microscopic analysis revealed that nuclear YY1 expression was increased in HSA+ hepatocytes in human MASH liver versus normal liver. By contrast, levels of nuclear YY1 in non-hepatocytes (HSA−) were similar in MASH and control livers (Fig. 5d). Similar data in humans and FPC-fed mice were obtained using immunohistochemistry, based on the morphology of hepatocytes (parenchymal cells) versus non-hepatocytes (Extended Data Fig. 5e and Fig. 5e). Finally, a previous single-nucleus RNA-seq study of the livers of mice with ALIOS-diet-induced MASH identified five hepatocyte subclusters, termed pericentral (PC), intermediate (Int), periportal (PP), mNASH-Hep1 and mNASH-Hep2, with the latter two subclusters highly enriched in MASH versus normal liver and having distinct gene signatures associated with MASH progression15. We found that the expression of Casp8 and Yy1 was enriched in the mNASH-Hep1 and mNASH-Hep2 subclusters versus the other three subclusters (Fig. 5f). These combined data provide support for the following hypothesis: caspase-8, which is elevated in MASH, leads to YY1-mediated induction of meteorin, which is secreted and activates HSCs to promote liver fibrosis.
a, qPCR was used to assess YY1 and METRN expression in Scr- and siYy1-treated AML12 cells (left) or primary human hepatocytes (right). Data are presented as mean ± s.e.m.; *P = 0.017 (Metrn), *P = 0.011 (YY1), *P = 0.020 (METRN); ***P = 0.0007, two-way ANOVA; n = 3 per group. b, AML12 hepatocytes treated with Scr, siCasp8 or siYy1 were transfected with constructs expressing GFP control or DED domains. After 48 h, the Metrn mRNA level was measured. Data are mean ± s.e.m.; *P = 0.039, **P = 0055, ***P = 0.0006, one-way ANOVA; n = 3 per group. c, Hepatocyte nuclear extracts from AML12 cells treated with siCasp8 or Scr siRNA were subjected to YY1 ChIP analysis using anti-YY1 or IgG control. The promoter region sequence was amplified by qPCR and normalized to the values obtained from the input DNA. Data are mean ± s.e.m.; **P = 0.0045, one-way ANOVA; n = 4 per group. d, YY1 immunofluorescence in liver sections of normal and MASH human livers; scale bar, 50 μm. Arrows, an example of a YY1+HSA+ hepatocyte in each sample, highlighting the higher staining intensity of nuclear YY1 in MASH liver, which is quantified as mean fluorescence intensity (MFI) in the graph. Arrowheads, YY1+HSA− cells. The MFI for nuclear YY1 is presented as mean ± s.e.m.; ***P = 0.0002, unpaired two-tailed Student’s t-test; n = 6 for normal and n = 7 for MASH. a.u., arbitrary unit. e, Immunohistochemical analysis of YY1 protein in the livers of FPC diet-fed mice. Scale bar, 50 μm. Arrows, parenchymal cells (hepatocytes); arrowheads, mononuclear cells. Integrated density (IntDen) data for nuclear YY1 in parenchymal and mononuclear cells are presented as mean ± s.e.m.; **P = 0.0075, unpaired two-tailed Student’s t-test; chow, n = 4; FPC, n = 4. f, snRNA-seq data from 9-month ALIOS-diet-fed mouse liver were analysed for Casp8 and Yy1 expression across five hepatocyte clusters. Data are mean ± s.e.m. The P values in the graphs refer to the difference between groups 4–5 (n = 1,052) versus groups 1–3 (n = 4,882) by unpaired two-tailed Student’s t-test.
Meteorin promotes MASH fibrosis
To investigate the role of liver meteorin in MASH, we assayed METRN mRNA and meteorin protein levels in human MASH versus normal liver and probed a public bulk RNA-seq human dataset. Both our data and the RNA-seq dataset showed increased METRN mRNA levels in MASH liver, and we also showed an increase in meteorin protein (Fig. 6a,b). Elevated levels of meteorin protein were also observed in the livers of FPC-fed, HF-CDAA-fed and ALIOS-diet-fed mice with MASH (Fig. 6c) but not in the steatotic livers of 8-week-old FPC-fed mice or humans with simple steatosis (Fig. 6d and Extended Data Fig. 5f). On the basis of immunofluorescence staining of human MASH liver and immunoblot analysis of isolated MASH hepatocytes, the increase in meteorin in MASH was determined to occur in hepatocytes (Fig. 6e,f). These data are consistent with the finding that the two proteins upstream of meteorin, caspase-8 and YY1, were not increased in the livers of mice with simple steatosis (Extended Data Fig. 5g,h). We next addressed causation by determining whether genetic restoration of meteorin in hepatocytes could restore liver fibrosis in H-Casp8-KO FPC-MASH mice (above). We administered AAV8-TBG-Cre with or without AAV8-TBG-Metrn (or AAV8-TBG-GFP control) to Casp8fl/fl mice after 8 weeks of the FPC diet and then analysed the mice after they were fed an FPC for 16 weeks (Fig. 7a). As before, we achieved robust caspase-8 deletion with AAV8-TBG-Cre, with a concomitant decrease in meteorin, and AAV8-TBG-Metrn restored meteorin to levels comparable to those in control livers (Fig. 7b). There were no significant differences in body weight, liver to body weight ratio or blood glucose levels among the three groups of mice (Extended Data Fig. 6a–c). Also as before, the Sirius-red-positive area and number of mononuclear cells were decreased by H-Casp8-KO, but, most notably, these changes were abrogated in the H-Casp8-KO mice treated with AAV8-TBG-Metrn (Fig. 7c). Analyses of fibrotic and inflammatory mRNA expression, COLA1- and F4/80-positive areas and plasma ALT further confirmed that restoration of meteorin in hepatocytes abrogated the beneficial effects of H-Casp8-KO on MASH (Fig. 7d–f). These findings provide in vivo causation evidence that hepatocyte caspase-8 promotes MASH by inducing meteorin.
a, METRN mRNA and meteorin protein were assayed in the livers of individuals with or without MASH. Data are presented as mean ± s.e.m.; *P = 0.029, unpaired two-tailed Student’s t-test; normal, n = 6; MASH, n = 6. b, METRN mRNA levels in livers of healthy individuals and those with MASLD, based on the RNA-seq dataset GSE126848. Data are mean ± s.e.m.; *P = 0.012, unpaired two-tailed Student’s t-test; healthy, n = 14; MASLD, n = 31. a.u., arbitrary units. c, Meteorin protein was increased in the livers of mice fed the FPC diet for 16 weeks, the HF-CDAA diet for 8 weeks or the ALIOS diet for 36 weeks. d, Meteorin immunoblot and quantification of liver extracts from C57BL/6J mice fed chow or the FPC diet for 8 or 16 weeks. Data are mean ± s.e.m.; **P = 0.0061, one-way ANOVA; chow, n = 3, 8 weeks FPC, n = 4, 16 weeks FPC, n = 4. e, Meteorin immunofluorescence in liver sections of normal and MASH human livers. Scale bar, 50 μm. Arrows, an example of a meteorin+HSA+ hepatocyte in each sample, highlighting the higher staining intensity of meteorin in MASH liver, which is quantified as the MFI. Arrowheads, meteorin+HSA− cells. The MFI for hepatocyte meteorin is presented as mean ± s.e.m.; ****P < 0.0001, unpaired two-tailed Student’s t-test; normal, n = 4; MASH, n = 4. f, Meteorin protein was assayed in primary hepatocytes from people with or without MASH.
a–f, Male Casp8fl/fl mice were fed the FPC diet for 8 weeks and then injected with AAV8-TBG-Cre (KO), AAV8-TBG-Cre/Metrn (KO+Metrn) or AAV8-TBG-GFP (control) and continued on the FPC diet for an additional 8 weeks. a, The experimental scheme. b, Immunoblot of caspase-8 and meteorin in liver, with β-actin as loading control. Data are presented as mean ± s.e.m.; ****P < 0.0001, one-way ANOVA; control: n = 5, KO: n = 5, KO+Metrn: n = 5. c, Staining of liver sections for Sirius red (top; scale bar, 200 μm) and H&E (bottom; scale bar, 200 μm), with quantification of Sirius red-positive area and hepatic mononuclear cells. Data are mean ± s.e.m.; **P = 0.0023 (Sirius red), **P = 0.0011 (mononuclear cells), ***P = 0.0003, one-way ANOVA; n = 7 per group. d, Liver mRNA levels of Timp1 (*P = 0.013), Spp1 (*P = 0.0004), Col1a1 (**P = 0.0047), Tnfa (*P = 0.034), Mcp1 (*P = 0.042) and Emr1 (*P = 0.040). Data are mean ± s.e.m.; one-way ANOVA; n = 7 per group. e, COL1A1 and F4/80 immunohistochemistry, with quantification; scale bar, 200 μm. Data are mean ± s.e.m.; *P = 0.030 (COL1A1), ***P = 0.0003; *P = 0.031 (F4/80: con versus KO), *P = 0.033 (F4/80: KO versus KO+Metrn), one-way ANOVA; con, n = 7; KO, n = 7; KO+Metrn, n = 7. f, Plasma ALT levels. Data are mean ± s.e.m.; **P = 0.0024, ***P = 0.0003, one-way ANOVA; con, n = 7; KO, n = 7; KO+Metrn, n = 7.
Silencing hepatocyte meteorin in MASH reverses liver fibrosis
Given that therapies that silence genes specifically in hepatocytes, for example using GalNAc-siRNA, are now used in people to treat various diseases26, we sought to conduct a proof-of-concept study in experimental MASH to determine whether hepatocyte-specific deletion of meteorin could block the progression from early MASH to more advanced MASH, particularly lessening liver fibrosis. For this purpose, we fed mice an FPC diet for 16 weeks (basal), followed by treatment with hepatocyte-specific AAV8-H1-shMetrn or a control vector and then continuation of the diet for an additional 12 weeks (Fig. 8a). As anticipated, Metrn mRNA and meteorin protein levels were reduced in the livers of mice treated with shMetrn (Fig. 8b). The three groups of mice had similar body weights, liver to body weight ratios, blood glucose levels and liver triglyceride content (Extended Data Fig. 6d–g). After 28 weeks of FPC feeding, liver fibrosis and inflammation were increased compared with that in the 16-week basal livers, but shMetrn treatment during the progression period resulted in regression to the 16-week level (Fig. 8c). A similar pattern was observed for the endpoints of fibrosis- and inflammation-related mRNAs, COL1A1- and F4/80-positive areas, and plasma ALT levels (Fig. 8d–f). However, the total number of T cells was comparable between the two groups (Extended Data Fig. 6h). These data complement the findings from the meteorin-restoration model described in the previous section by showing the pro-fibrotic nature of hepatocyte meteorin in MASH, and they provide proof-of-concept evidence that silencing hepatocyte meteorin in early MASH can reverse the disease, particularly MASH fibrosis.
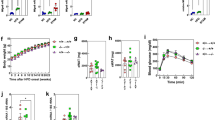
Male C57BL/6J mice were fed an FPC diet for 16 weeks and then injected with AAV8-H1-shMetrn (shMetrn) or AAV8-H1-Scr (control) and continued on the FPC diet for an additional 12 weeks. a, The experimental scheme. b. Liver Metrn mRNA levels and immunoblot of meteorin protein, with β-actin as the loading control (right). Data are presented as mean ± s.e.m.; *P = 0.016, one-way ANOVA; basal, n = 8, con, n = 7, shMetrn, n = 7. c, Staining of liver sections for Sirius red (top; scale bar, 200 μm) and H&E (bottom; scale bar, 200 μm), with quantification of the Sirius-red-positive area and hepatic mononuclear cells. Data are mean ± s.e.m.; **P = 0.0023 (Sirius red: con versus shMetrn), ***P = 0.0003, **P = 0.0033 (mononuclear cells: basal versus con), **P = 0.0077 (mononuclear cells: con versus shMetrn); one-way ANOVA; basal, n = 8, con, n = 6, shMetrn, n = 7. d, Liver mRNA levels of Timp1 (***P = 0.0009), Spp1 (***P = 0.0002), Col1a1 (*P = 0.030), Col1a2 (*P = 0.012), Tgfb1 (**P = 0.0033), Tnfa (*P = 0.011), Mcp1 (**P = 0.004) and Emr1 (****P < 0.0001). Data are mean ± s.e.m.; one-way ANOVA; basal, n = 8, con, n = 7, shMetrn, n = 7. e, COL1A1 and F4/80 immunohistochemistry, with quantification; scale bar, 200 μm. Data are mean ± s.e.m.; *P = 0.022; **P = 0.0036 (COL1A1: basal versus con), **P = 0.0016 (COL1A1: con versus shMetrn), **P = 0.0046 (F4/80), one-way ANOVA; basal, n = 8, con, n = 7, shMetrn, n = 7. f, Plasma ALT levels. Data are mean ± s.e.m.; *P = 0.013, **P = 0.0021, one-way ANOVA; basal, n = 8, con, n = 7, shMetrn, n = 7.
Discussion
The data herein provide evidence for a hepatocyte–HSC cross-talk pathway that contributes to early MASH fibrosis: caspase-8, through a non-apoptosis function involving DED domains, leads to YY1-mediated induction of meteorin, a secretory protein that we show activates HSCs through a c-Kit–STAT3 pathway. The relevance of these findings to humans is supported by our analyses of human MASH liver and our data from primary human hepatocytes and HSCs. Moreover, prior genetic data have linked an intronic CASP8 variant and elevated CASP8 expression to chronically elevated ALT levels9, although future studies will be needed to determine whether this variant is associated with MASH fibrosis, and vice versa for possible loss-of-function CASP8 variants. Additionally, our evidence that the pathway is independent of apoptosis, including the finding that the pathway can be activated by the non-proteolytic DED domains of caspase-8, is consistent with a clinical trial showing that pharmacologic inhibition of the pro-apoptotic activity of caspases does not block MASH fibrosis10.
As expected for a previously unknown pathway, several fundamental questions remain to be explored. One is how caspase-8 expression in hepatocytes is increased in MASH. CASP8 mRNA and caspase-8 protein levels are increased in early MASH but not steatosis. These data suggest that an early stage in the transition from steatosis to MASH could lead to CASP8 induction in hepatocytes, which then further amplifies MASH progression through the YY1–meteorin pathway described here. In general, CASP8 induction could involve either a hepatocyte-autonomous process or signals from another cell, for example, monocyte-derived early MASH macrophages. One interesting possibility arises from the accumulation of cholesterol in hepatocytes during the early stages of the steatosis to MASH transition27. We showed that the trafficking of plasma membrane cholesterol into cholesterol-enriched hepatocytes triggers the upregulation of the gene regulator TAZ in early MASH, inducing the secretion of an HSC activator, Indian hedgehog (Ihh)28. A previous study has shown that Casp8 levels were increased in the livers of fat-fed steatotic mice lacking the cholesterol-transport enzyme StarD5, associated with an increase in liver cholesterol29. Other possibilities include cell-autonomous or cell cross-talk pathways that increase transcription factors for CASP8, for example IRF1 and IRF3 (refs. 30,31), and/or pathways that promote CASP8 mRNA stability mediated by the ELAV1–HuR pathway31.
Another interesting question is how caspase-8 leads to an increase in nuclear YY1 and YY1-mediated induction of Metrn. Previous work examining non-apoptosis functions of caspase-8 has focused on its roles in inflammation, immune regulation and cell differentiation32, but our finding that caspase-8, through its DED domains, leads to transcription-factor-mediated gene induction represents a new area that is ripe for future investigation. Our data suggest that the process involves the nuclear localization of YY1. Previous work has shown that YY1 shuttling from the cytoplasm to the nucleus can be triggered by post-translational modifications of YY1 and/or interaction with YY1-binding proteins33. Thus, it is possible that caspase-8 interacts directly with YY1 or with an intermediary YY1-binding protein, leading to one or more YY1 alterations that promote its nuclear localization. Another issue is whether other YY1-inducible genes in addition to Metrn are upregulated downstream of caspase-8 in early MASH and, if so, how this might affect MASH progression. Interestingly, a recent study implicated hepatic YY1-mediated repression of farnesoid X receptor in high-fat diet-induced hepatosteatosis34, so YY1 might play roles in both early (steatotic) and later (fibrotic) stages of MASLD.
Meteorin has not previously been implicated in either physiologic or pathophysiologic liver processes. Most studies have focused on its role in the central nervous system, with functions including astrocyte to radial glia differentiation and axonal extension in sensory ganglia neurons35. Other functions of meteorin include its role in Nodal expression during mesendoderm development36 and angiogenesis37. Our idea to look at YY1 as the link between caspase-8 and Metrn induction came from a study showing that YY1, together with hypoxia-induced factor 1α, in hypoxic bone marrow macrophages induces the expression of meteorin, which restrains the mobilization of haematopoietic stem and progenitor cells through an oxidative stress pathway25. Whether meteorin secreted by hepatocytes in early MASH has MASH-promoting functions in addition to HSC activation remains to be determined.
Our findings have potentially important therapeutic implications, based on two key points: first, liver fibrosis is a major contributor to both liver- and non-liver-related mortality in MASH, with limited therapeutic options4,5; second, therapeutic siRNAs that silence pathogenic genes exclusively in hepatocytes are in clinical use26. Thus, although caspase-8 has been identified as a therapeutic target in MASH11, the new mechanistic findings here favour the approach of silencing caspase-8 in hepatocytes, for example, using therapeutic GalNAc-siCASP8, rather than blocking its pro-apoptotic enzymatic activity, which is consistent with the failure of the aforementioned emricasan trial10. Moreover, we have identified a more specific target, meteorin, whose therapeutic silencing could\ also be considered as a new strategy to help prevent the progression of fibrotic MASH.
Methods
Animal studies
The research was conducted in accordance with all applicable ethical regulations. Male wild-type C57BL/6J mice (no. 000664, 11–12 weeks old) and Casp8fl/fl mice (no. 027002, on C57BL/6J background, PIMD: 16148088) were obtained from Jackson Laboratory and acclimated for 1 week at the Columbia University Irving Medical Center Institute of Comparative Medicine before being randomly assigned to experimental groups. To induce MASH, the mice were fed either a fructose–palmitate diet with 1.25% cholesterol (FPC diet, Envigo, TD.160785) for 16 weeks to induce MASH14 or a high-fat choline-deficient l-amino acid-defined (HF-CDAA, Research diet, A06071302) diet16 for 8 weeks. The control chow diet was obtained from LabDiet (catalogue no. 5053). Casp8fl/fl mice were injected through tail vein with AAV8-TBG-Cre, AAV8-TBG-Metrn, AAV8-TBG-LacZ or control AAV8-TBG-GFP (2 × 10¹¹ genome copies per mouse) at either 8 weeks (FPC) or 4 weeks (HF-CDAA) after the MASH diet was initiated. Additionally, AAV8-H1-scrambled RNA or AAV8-H1-shMetrn (2 × 10¹¹ genome copies per mouse) was administered through tail vein 16 weeks after starting the FPC diet, with the mice remaining on the diet for another 12 weeks. Mice were housed in standard cages at 22 °C under a 12-h light/dark cycle in a barrier facility. All procedures were approved by the Institutional Animal Care and Use Committee at Columbia University and conducted in accordance with institutional guidelines (protocol no. AC-AABL5573).
Human liver specimens and human primary liver cells
De-identified human liver specimens were obtained from the Liver Tissue Cell Distribution System at the University of Minnesota. Both male and female specimens were collected at the time of liver transplantation and preserved as frozen samples. Written informed consent was obtained from all donors for tissue samples. Diagnostic information is provided in Supplementary Table 1. Phenotypic and pathological assessments were performed by medical physicians and pathologists affiliated with the Liver Tissue Cell Distribution System. The study protocol was approved by the Institutional Review Board (IRB) at Columbia University Irving Medical Center. Diseased (MASH, METAVIR Score = F4); adult human liver cells were obtained from the Human Synthetic Liver Biology Core at the Pittsburgh Liver Research Center and the Center for Transcriptional Medicine. The Institutional Review Board at the University of Pittsburgh has approved our protocol and given the Not Human Research Determination (IRB no. STUDY24020093). Hepatocytes were isolated using a three-step collagenase digestion technique, as previously described38. Cell viability was assessed after isolation using trypan blue exclusion, and only cell preparations with viability >80% were used for the analysis.
Human primary hepatocytes were sourced from the Liver Tissue Cell Distribution System at the University of Pittsburgh, and human primary HSCs were isolated as described39. All participants provided consent to the use of these specimens, and the protocol was reviewed and approved by local IRBs.
Reagents and kits
Adeno-associated virus subtype 8 (AAV8)-shRNA targeting murine Metrn was generated by annealing complementary oligonucleotides (5′-CACCA GAACGTATGACCGGCAATCTATCAAGAG TAGATTGCCGGTCATACGTTC-3′ and 5′-AAAAGAACGTATGACCGGCAATCTACTCTTGATAGATTGCCGGTCATACGTTCT-3′). These oligonucleotides were ligated into the self-complementary (sc) AAV8-RSV-GFP-H1 vector, following a previously described protocol14. AAV8-TBG-Metrn and AAV8-H1-shRNAs were amplified and packaged into AAV particles by PackGene Biotech. The following reagents were used in the study: AAV8 containing hepatocyte-specific TBG-Cre recombinase (AAV-TBG-Cre, 107787-AAV8) and the control vector AAV8-TBG-LacZ (105534-AAV8) from Addgene; mouse Metrn plasmid (no. 215466) from Origene; caspase inhibitor Z-VAD-FMK (G7231) from Promega; caspase-8 inhibitor Z-IETD-FMK (FMK007) from R&D Systems; human METRN recombinant protein (TP761464) from Origene; mouse METRN recombinant protein (OPCD05218) from Aviva; WT Casp8 (no. 11817) and C360A (no. 11818) plasmids from Addgene; plasmids encoding the death effector domains of caspase-8 as described19; Jo2 antibody (anti-mouse CD95, no. 554254) from BD Biosciences; ChIP kit (no. 9003) from Cell Signaling Technology; ALT assay kit (no. A526-120) from TECO Diagnostics; liver TG kit (10010303) from Cayman.
Cell culture and conditioned media transfer
AML12 mouse hepatocytes were obtained from ATCC (CRL-2254) and cultured in DMEM/F12 medium (Life Technologies, no. 11320) supplemented with 10% (vol/vol) heat-inactivated FBS (Gibco, no. 16140-071) and 1× penicillin–streptomycin solution (Corning, 30-002-Cl). Primary mouse hepatocytes were isolated from 10-week-old wild-type C57BL/6J mice, as previously described14, and cultured in DMEM with 10% FBS before being treated as described in the figure legends. Primary mouse HSCs were isolated from 5- to 6-month-old BALB/C mice, as described previously14. In brief, after cannulation of the inferior vena cava, the portal vein was cut, allowing retrograde step-wise perfusion with solutions containing protease (Sigma Aldrich, P5147) and collagenase D (Roche, no. 11088866001). The perfusates were subjected to 9.7% Nycodenz (Accurate Chemical, no. 1002424) gradient centrifugation to isolate the HSCs, which were then plated in tissue culture dishes and used the next day. Primary human HSCs from deidentified donor livers that were declined for transplantation for various reasons were obtained by Lifesharing Organ Procurement Organization under IRB 171883XX. Livers were graded by a pathologist using a double-blinded method and identified as normal or MASH and used for human HSC isolation through perfusion with pronase and collagenase D perfusion followed by gradient centrifugation, as described39. Enriched human HSCs were stained for desmin and glial fibrillary acidic protein (GFAP) and had a purity of >95%. After various treatments, the hepatocytes or HSCs were collected in Laemmli sample buffer (Bio-Rad, no. 1610737) with 2-mercaptoethanol (Bio-Rad, no. 161-0710) for immunoblotting or in RNA lysis buffer (Qiagen, no. 79216) for mRNA quantification. For conditioned-medium-transfer experiments, AML12 cells were cultured in DMEM with 0.1% FBS and incubated for 24 h. The medium was then transferred to HSCs that had been pre-incubated in DMEM with 0.1% FBS for 8 h. After 72 h, the HSCs were analysed for mRNA expression.
Human liver spheroids
Primary human hepatocytes and non-parenchymal cells were co-incubated to form spheroids, as previously described40. In brief, fresh hepatocytes and non-parenchymal cells obtained from patients undergoing liver resection at the University of Pittsburgh were seeded into 96-well Akura Spheroid Microplates (inSphero) at a ratio of 4:1 (hepatocytes:non-parenchymal cells; 2,000 viable cells per well) in 70 μl of William’s E medium supplemented with 2 mM l-glutamine (Thermo Fisher), 1× penicillin–streptomycin solution (Corning, no. 30-002-Cl), 100 nM dexamethasone (Thermo Fisher), ITS X-100 (Thermo Fisher) and 10% fetal bovine serum (FBS, Thermo Fisher), The cells were cultured to form spheroids at 37 °C in a humidified incubator with 5% CO2. On day 5 after seeding, and every 2–3 days thereafter, the spheroids were refreshed with the medium described above but without FBS. The spheroids were treated with siCASP8 or scrambled siRNA on day 7 and collected on day 12 after seeding.
siRNA-mediated gene silencing and transfection
Scrambled siRNA control and gene-targeting siRNAs were transfected into AML12 cells or primary hepatocytes using Lipofectamine RNAiMAX (Life Technologies) at a final concentration of 40 nM siRNA in 24-well plates, following the manufacturer’s protocol. In brief, 2 × 105 cells at 30–40% confluence were incubated for 18 h with 0.5 ml of culture medium containing 1.5 μl Lipofectamine RNAiMAX and 40 pmol siRNA. The siRNA sequences are provided in Supplementary Table 2. Plasmid transfection into AML12 cells was conducted using Lipofectamine LTX Reagent with PLUS Reagent (Life Technologies, no. 15338100). For each well in a 24-well plate, 2 μl LTX, 0.5 μl PLUS reagent and 0.5 μg plasmid DNA were added when the cells reached 30–40% confluence. After overnight incubation, the cells were returned to normal culture medium.
Blood and plasma analyses
Blood glucose levels were measured using a glucose meter (One Touch Ultra, LifeScan) in mice that were deprived of food for 5 h with free access to water. Plasma ALT levels were measured using commercially available kits, as detailed in the ‘Reagents and kits’ section, and the data were collected using SoftMax Pro Software v5.4.1.
Histopathological analysis
Liver fibrosis was evaluated by measuring the area stained with Picrosirius red (Sirius) (Polysciences, no. 24901). Inflammatory cells in H&E-stained liver section images were quantified by counting the number of mononuclear cells per field at ×20 magnification. For immunohistochemistry, paraffin sections were rehydrated and subjected to antigen retrieval by placing them in a pressure cooker with Antigen Unmasking Solution (H-3300, Vector) for 10 min. The slides were then treated with 3% hydrogen peroxide for 10 min and subsequently blocked with Serum-Free Protein Block (X0909, Dako) for 30 min. Sections were incubated overnight with a mouse primary antibodies against anti-MPO (AF3667, R&D Systems, 1:100 dilution), anti-CD3 (ab16669, Abcam, 1:100 dilution), anti-pH2AX (no. 9718, Cell Signaling, 1:100 dilution), anti-COL1A1 (no. 72026, Cell Signaling, 1:500 dilution), anti-F4/80 (no. 70076, Cell Signaling, 1:500 dilution) or anti-CD3 (ab16669, Abcam, 1:100 dilution) and secondary antibody (SignalStain Boost IHC Rabbit, Cell Signaling, no. 8114), then developed using a DAB substrate kit (no. 8059, Cell Signaling). For various staining parameters, computerized image analysis (ImageJ) was used to quantify the stained area, utilizing the same threshold settings across all analyses. For each section, for each mouse, we quantified six randomly selected fields.
Immunofluorescence microscopy
AML12 or primary cultured hepatocytes were fixed in 4% paraformaldehyde for 10 min at room temperature, rinsed with PBS and stained overnight with a primary antibody at 4 °C. On the following day, the cells were washed and incubated with a secondary antibody and DAPI before being viewed in PBS using fluorescence microscopy. Paraffin sections were rehydrated and subjected to antigen retrieval; they were first placed in a pressure cooker with antigen unmasking solution (H-3300, Vector) for 10 min, followed by blocking with serum. Sections were then labelled overnight with antibodies against α-smooth muscle actin (α-SMA) (C6198, Sigma, 1:300 dilution), cl-Casp3 (no. 9664, Cell Signaling, 1:100 dilution), F4/80 (no. 70076, Cell Signaling, 1:500 dilution), human serum albumin (HSA) (ab190706, Abcam, 1:100 dilution), anti-caspase-8 (no. 8873, Cell Signaling, 1:100 dilution), anti-meteorin (OAAI00402, Aviva, 1:100 dilution), or anti-YY1 (no. 46395, Cell Signaling, 1:500 dilution). The sections were then incubated for 1 h with a fluorophore-conjugated secondary antibody (Invitrogen, Alexa-Fluor-555-conjugated donkey anti-rabbit or Alexa-Fluor-488-conjugated goat anti-mouse, 1:500), mounted with a DAPI-containing mounting medium (P36935, Life Technologies) and viewed using Olympus DP74 fluorescence microscopy with cellSens software. ImageJ2 (version 2.14.0) was used to quantify the stained area, applying consistent threshold settings for each analysis. Six randomly selected fields were quantified per section per mouse for image analyses. TUNEL staining was performed using a kit from Roche (no. 12156792910).
Immunoblotting
Liver proteins were extracted using RIPA buffer (Thermo, no. 89900), and the protein concentration was determined with a BCA assay (Thermo, no. 23227). Proteins were separated by electrophoresis on 4–20% Tris gels (Life Technologies, EC60285) and transferred to nitrocellulose membranes (Bio-Rad, no. 1620115). The membranes were blocked for 30 min at room temperature in Tris-buffered saline with 0.1% Tween 20 (TBST) containing 5% (wt/vol) non-fat milk and then incubated overnight at 4 °C with primary antibodies diluted 1:1000 in the same buffer: anti-caspase-8 (no. 4790, Cell Signaling), anti-cl-Casp8 (no. 9429, Cell Signaling), anti-YY1 (no. 46395, Cell Signaling), anti-Metrn (OACD05531, Aviva), anti-cl-Casp3 (no. 9661, Cell Signaling), anti-phospho-Stat3 (no. 9145, Cell Signaling), anti-Stat3 (no. 9139, Cell Signaling), anti-caspase-10 (no. 14311-1-AP, Proteintech), anti-β-actin (no. 5125, Cell Signaling) or anti-GAPDH (no. 3683, Cell Signaling). Protein bands were detected using horseradish-peroxidase-conjugated secondary antibodies (donkey anti-rabbit-HRP, Jackson ImmunoResearch, 711-035-152 or anti-rat IgG HRP-linked antibody, Cell Signaling no. 7077, both at 1:5,000 dilution) and SuperSignal West Pico enhanced chemiluminescent solution (Thermo, no. 34080). For cultured cells, lysis was performed in Laemmli sample buffer (Bio-Rad, no. 161-0737) containing 5% 2-mercaptoethanol, followed by heating at 100 °C for 5 min. The samples were then electrophoresed and immunoblotted as described above.
Chromatin Immunoprecipitation assay
The ChIP assay was conducted following the instructions provided in the SimpleChIP kit from Cell Signaling Technology (no. 9003). In brief, AML12 hepatocytes cultured in a 15-cm dish were incubated with formaldehyde to cross-link proteins to DNA, and the reaction was neutralized by the addition of glycine. The cells were then scraped into cold PBS containing a protease inhibitor cocktail (PIC) and centrifuged, and the supernatant fraction was discarded. The cell pellet was lysed to release the nuclei, and nuclease was added to digest the DNA into fragments ranging from 150 to 900 bp. After halting the digestion, the nuclei were pelleted and sonicated to disrupt the nuclear membrane, followed by centrifugation to remove the pellet. The resulting supernatant fraction, which was enriched with cross-linked chromatin, underwent immunoprecipitation using antibodies from Cell Signaling (YY1, no. 46395, 1:200 dilution; Rpb1, no. 14958, 1:100 dilution; IgG, no. 3900, 1:100 or 1:200 dilution). The suspensions were rotated overnight, after which protein G magnetic beads were added. After washing the beads, the DNA was eluted and further purified. The presence of YY1 and Pol II in the Metrn promoter region was quantified by qPCR and expressed relative to the input genomic DNA. The sequences of the primers used for the ChIP–qPCR assays are detailed in Supplementary Table 3.
Quantitative RT–qPCR
Total RNA was extracted from liver tissue or cultured hepatocytes using the RNeasy kit (Qiagen, no. 74106). The quality and concentration of the RNA were evaluated by measuring absorbance at 260 and 280 nm with a Thermo Scientific NanoDrop spectrophotometer. cDNA synthesis was performed using 1 μg of total RNA with oligo (dT) primers and Superscript II (Invitrogen). Quantitative PCR (qPCR) was conducted on a 7500 Real-Time PCR system (Applied Biosystems 7500 software v2.3) using SYBR Green Master Mix (Life Technologies, no. 4367659). The primer sequences utilized for the qPCR are provided in Supplementary Table 3.
RNA-seq and data analysis
Total mRNA was extracted from AML12 hepatocytes treated with either siCasp8 or scramble RNA using the RNeasy Mini Kit (Qiagen, no. 74104). Libraries were constructed utilizing Illumina TruSeq chemistry and sequenced on the Illumina NovaSeq 6000 at the Columbia Genome Center. For data analysis, FASTQ files were processed using Salmon v1.3.0 to quantify transcript abundance, using an index based on GRCm39 GENCODE VM27 (genome browser, IGV: https://genome.ucsc.edu/cgi-bin/hgTrackUi?db=mm39&g=wgEncodeGencodeVM27)41. The resulting counts were aggregated to the gene level with tximport v1.20.0 (ref. 42). Differential expression analysis was performed using DESeq2 v1.32.0 (ref. 43), with genes exhibiting an absolute fold change of >1.5 and a false discovery rate (FDR)-adjusted P value < 0.05 classified as differentially expressed (DE). The DESeq2 v1.32.0 output was scored and ranked using Approximate Posterior Estimation for Generalized Linear Model (apeglm)44, applying the ranking metric of ‘−log10 P-value multiplied by the sign of log-transformed fold-change’45. The list of DE genes was used to generate a Venn diagram on the website bioinformatics.psb.ugent.be/webtools/Venn. Additionally, the other two datasets for the Venn diagram included a secreted protein list from VerSeDa (genomics.cicbiogune.es/VerSeDa/index.php)46 and proteomics data from cell culture supernatant analysis (see below).
Generic proteome preparation from cell culture supernatants
AML12 cells treated with either scramble or siCasp8 were cultured in three T150 flasks for each condition. After reaching 80–90% confluence, the cells were washed three times with warm PBS and incubated for 1–2 h in pre-warmed serum-free medium to remove residual serum proteins. This medium was discarded and replaced with fresh 37 °C serum-free, phenol-red-free medium (20 ml). The cells were incubated for 24 h, after which the conditioned medium was collected on ice. Protease inhibitors and EDTA were added, and the medium was clarified by centrifugation, filtered through a 0.2-μm filter and concentrated by precipitation using 15% trichloroacetic acid (TCA) on ice for 4 h. The samples were then centrifuged at >9,000g for 15 min at 4 °C, the supernatant fraction was decanted, and freezer-cold (−20 °C) 100% acetone (equal to the original volume) was added. This was followed by centrifugation and decanting, and then the pellet was washed three times. The final pellet was resuspended in 200 mM HEPES (pH 8) at a protein concentration of approximately 1–2 mg ml−1. For global quantitative proteomic analysis, dia-PASEF (parallel accumulation serial fragmentation)-based proteomics was used. In brief, the samples were denatured in sodium deoxycholate (SDC) buffer (1% SDC, 100 mM Tris-HCl, pH 8.5) and heated for 15 min at 60 °C with shaking at 1,000 r.p.m. Protein reduction and alkylation of cysteines were performed with 10 mM TCEP and 40 mM CAA at 45 °C for 10 min. The samples were then sonicated in a water bath, cooled to room temperature and digested overnight using a Lys-C/trypsin mix at a 1:50 ratio (µg enzyme to µg protein) at 37 °C with shaking at 1,400 r.p.m. The resulting peptides were acidified by adding 1% trifluoroacetic acid (TFA), vortexed and subjected to StageTip clean-up using styrene-divinylbenzene-reverse phase sulfonate (SDB-RPS). The peptides were dried in a speed-vac and resuspended in 10 µl of liquid chromatography buffer (3% acetonitrile/0.1% formic acid). Peptide concentrations were determined using a NanoDrop, and 200 ng of each sample was used for dia-PASEF analysis on a Trapped Ion Mobility Spectrometry-Time of Flight (timsTOF) Pro mass spectrometer.
Liquid chromatography with tandem mass spectrometry
Peptides were separated over an 87-min gradient at a flow rate of 400 nanoliters min−1 on a reversed-phase C18 column with an integrated CaptiveSpray Emitter (25 cm × 75 µm, 1.6 µm, IonOpticks). Mobile phase A consisted of 0.1% formic acid in water, and mobile phase B consisted of 0.1% formic acid in acetonitrile. The gradient for phase B increased linearly from 2% to 23% over 70 min, followed by an increase to 35% over 10 min, and then to 80% before re-equilibration. The timsTOF Pro was operated in dia-PASEF mode, with data acquired across 32 × 26 Th isolation windows from m/z 400 to 1,200. To optimize the MS1 cycle time in dia-PASEF, the repetitions were set to 2 in the 16-scan scheme. The collision energy was ramped linearly according to mobility, from 59 eV at 1/K0 = 1.6 Vs cm−2 to 20 eV at 1/K0 = 0.6 Vs cm−2. The acquired dia-PASEF raw data were searched using a library-free workflow in the DIA-NN search engine, utilizing the UniProtKB/Swiss-Prot mouse database. Default settings for the library-free algorithm were used, with match-between-runs (MBR) enabled. A maximum of one missed cleavage for trypsin was allowed, and the maximum variable modification was set to one. Carbamidomethylation was specified as the fixed modification, and protein N-terminal methionine excision, methionine oxidation and N-terminal acetylation were set as variable modifications. The peptide length range was set from 7 to 30 amino acids, with precursor charge ranges of 1 to 4, precursor m/z ranges of 300 to 1,800 and fragment ion m/z ranges of 200 to 1,800. The false discovery rates (FDRs) for protein and peptide levels were set to 1%, with MS1 and MS2 mass tolerances automatically determined by DIA-NN.
Quantification and statistical analysis
Statistical analyses were carried out using GraphPad Prism (version 9.4). Data were tested for normality using the Shapiro–Wilk test. Data that passed the normality test were analysed using the two-tailed Student’s t-test for comparison of two groups or one- or two-way analysis of variance (ANOVA) with Bonferroni post hoc analysis for comparison of more than two groups. Data are shown as mean ± s.e.m., and differences were considered statistically significant at P < 0.05. No statistical methods were used to pre-determine sample sizes, but our sample sizes are similar to those reported in previous publications14,28. No animals or data points were excluded from the analyses. Data collection and analysis were not performed blind to the conditions of the experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data associated with this study are present in the paper or the Supplementary Information. The raw RNA-seq data have been deposited at the National Center for Biotechnology Information Gene Expression Omnibus (NCBI GEO) under accession number GSE280901. The raw LC–MS/MS data have been deposited in the PRIDE database under accession number PXD057540. All the databases and datasets used in the study, along with appropriately accessible links, are provided in the manuscript. All data supporting the figures in this paper are provided in the source data published with the article. Source data are provided with this paper.
Code availability
No original code was created for this study.
References
Wree, A., Broderick, L., Canbay, A., Hoffman, H. M. & Feldstein, A. E. From NAFLD to NASH to cirrhosis-new insights into disease mechanisms. Nat. Rev. Gastroenterol. Hepatol. 10, 627–636 (2013).
Loomba, R., Friedman, S. L. & Shulman, G. I. Mechanisms and disease consequences of nonalcoholic fatty liver disease. Cell 184, 2537–2564 (2021).
Younossi, Z. M., Kalligeros, M. & Henry, L. Epidemiology of metabolic dysfunction-associated steatotic liver disease. Clin. Mol. Hepatol. S32–S50 (2024).
Angulo, P. et al. Liver fibrosis, but no other histologic features, is associated with long-term outcomes of patients with nonalcoholic fatty liver disease. Gastroenterology 149, 389–397 (2015).
Sanyal, A. J. et al. Prospective study of outcomes in adults with nonalcoholic fatty liver disease. N. Engl. J. Med. 385, 1559–1569 (2021).
Schwabe, R. F., Tabas, I. & Pajvani, U. B. Mechanisms of fibrosis development in nonalcoholic steatohepatitis. Gastroenterology 158, 1913–1928 (2020).
Schwabe, R. F. & Luedde, T. Apoptosis and necroptosis in the liver: a matter of life and death. Nat. Rev. Gastroenterol. Hepatol. 15, 738–752 (2018).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Vujkovic, M. et al. A multiancestry genome-wide association study of unexplained chronic ALT elevation as a proxy for nonalcoholic fatty liver disease with histological and radiological validation. Nat. Genet. 54, 761–771 (2022).
Harrison, S. A. et al. A randomized, placebo-controlled trial of emricasan in patients with NASH and F1-F3 fibrosis. J. Hepatol. 72, 816–827 (2020).
Hatting, M. et al. Hepatocyte caspase-8 is an essential modulator of steatohepatitis in rodents. Hepatology 57, 2189–2201 (2013).
Gautheron, J. et al. A positive feedback loop between RIP3 and JNK controls non-alcoholic steatohepatitis. EMBO Mol. Med. 6, 1062–1074 (2014).
Volkert, I. et al. Selective genetic inactivation of Caspase 8 in hepatocytes ameliorates progression of MASH following Jnk deficiency. Hepatology https://doi.org/10.1097/hep.0000000000001286 (2025).
Wang, X. et al. Hepatocyte TAZ/WWTR1 promotes inflammation and fibrosis in nonalcoholic steatohepatitis. Cell Metab. 24, 848–862 (2016).
Xiao, Y. et al. Hepatocytes demarcated by EphB2 contribute to the progression of nonalcoholic steatohepatitis. Sci. Transl. Med. 15, eadc9653 (2023).
Wei, G. et al. Comparison of murine steatohepatitis models identifies a dietary intervention with robust fibrosis, ductular reaction, and rapid progression to cirrhosis and cancer. Am. J. Physiol. Gastrointest. Liver Physiol. 318, G174–G188 (2020).
Mederacke, I. et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat. Commun. 4, 2823 (2013).
Sauerwald, T. M., Oyler, G. A. & Betenbaugh, M. J. Study of caspase inhibitors for limiting death in mammalian cell culture. Biotechnol. Bioeng. 81, 329–340 (2003).
Mielgo, A. et al. The death effector domains of caspase-8 induce terminal differentiation. PLoS ONE 4, e7879 (2009).
Liu, Y. et al. Transforming growth factor-beta (TGF-beta)-mediated connective tissue growth factor (CTGF) expression in hepatic stellate cells requires Stat3 signaling activation. J. Biol. Chem. 288, 30708–30719 (2013).
Meng, F. et al. Interleukin-17 signaling in inflammatory, Kupffer cells, and hepatic stellate cells exacerbates liver fibrosis in mice. Gastroenterology 143, 765–776 (2012).
Jiao, J. et al. Spatial molecular and cellular determinants of STAT3 activation in liver fibrosis progression in non-alcoholic fatty liver disease. JHEP Rep. 5, 100628 (2023).
Reboll, M. R. et al. Meteorin-like promotes heart repair through endothelial KIT receptor tyrosine kinase. Science 376, 1343–1347 (2022).
Ning, Z. Q., Li, J. & Arceci, R. J. Signal transducer and activator of transcription 3 activation is required for Asp(816) mutant c-Kit-mediated cytokine-independent survival and proliferation in human leukemia cells. Blood 97, 3559–3567 (2001).
Dai, Y. W. et al. Meteorin links the bone marrow hypoxic state to hematopoietic stem/progenitor cell mobilization. Cell Rep. 40, 111361 (2022).
Qin, Z. X. et al. GalNac–siRNA conjugate delivery technology promotes the treatment of typical chronic liver diseases. Expert Opin. Drug Deliv. 22, 455–469 (2025).
Horn, C. L., Morales, A. L., Savard, C., Farrell, G. C. & Ioannou, G. N. Role of cholesterol-associated steatohepatitis in the development of NASH. Hepatol. Commun. 6, 12–35 (2022).
Wang, X. et al. Cholesterol stabilizes TAZ in hepatocytes to promote experimental non-alcoholic steatohepatitis. Cell Metab. 31, 969–986 (2020).
Kakiyama, G. et al. StarD5 levels of expression correlate with onset and progression of steatosis and liver fibrosis. Am. J. Physiol. Gastrointest. Liver Physiol. 326, G747–G761 (2024).
Liu, J. et al. Toll-like receptor-4 signalling in the progression of non-alcoholic fatty liver disease induced by high-fat and high-fructose diet in mice. Clin. Exp. Pharm. Physiol. 41, 482–488 (2014).
Deng, B. et al. TNF and IFNgamma-induced cell death requires IRF1 and ELAVL1 to promote CASP8 expression. J. Cell Biol. 223, e202305026 (2024).
Orning, P. & Lien, E. Multiple roles of caspase-8 in cell death, inflammation, and innate immunity. J. Leukoc. Biol. 109, 121–141 (2021).
Verheul, T. C. J., van Hijfte, L., Perenthaler, E. & Barakat, T. S. The why of YY1: mechanisms of transcriptional regulation by Yin Yang 1. Front. Cell Dev. Biol. 8, 592164 (2020).
Lu, Y. et al. Yin Yang 1 promotes hepatic steatosis through repression of farnesoid X receptor in obese mice. Gut 63, 170–178 (2014).
Nishino, J. et al. Meteorin: a secreted protein that regulates glial cell differentiation and promotes axonal extension. EMBO J. 23, 1998–2008 (2004).
Kim, Y. Y. et al. Meteorin regulates mesendoderm development by enhancing nodal expression. PLoS ONE 9, e88811 (2014).
Park, J. A. et al. Meteorin regulates angiogenesis at the gliovascular interface. Glia 56, 247–258 (2008).
Ortiz, K. et al. Human hepatocellular response in cholestatic liver diseases. Organogenesis 19, 2247576 (2023).
Liu, X., Brenner, D. A. & Kisseleva, T. Human hepatic stellate cells: isolation and characterization. Methods Mol. Biol. 2669, 221–232 (2023).
Moore, M. P. et al. Low MBOAT7 expression, a genetic risk for MASH, promotes a profibrotic pathway involving hepatocyte TAZ upregulation. Hepatology 81, 576–590 (2025).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhu, A., Ibrahim, J. G. & Love, M. I. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics 35, 2084–2092 (2019).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Cortazar, A. R., Oguiza, J. A., Aransay, A. M. & Lavín, J. L. VerSeDa: vertebrate secretome database. Database 2017, baw171 (2017).
Acknowledgements
This work was supported by NIH grants DK133694 (I.T. and D.S.) and DK125573 (M.A.L.) and by American Heart Association training grant 827529 (Y.X.). Normal human liver samples and primary hepatocytes were obtained from the Human Liver Tissue & Hepatocytes Research Resource (HLTHRR) programme funded by NIH grants R24DK139775 (A.S.G.). Diseased (MASH) human liver samples and primary hepatocytes were obtained from the Human Synthetic Liver Biology Core (HSLBC), partially supported by NIH grant 1P30DK120531 Pittsburgh Liver Research Center (PLRC) and NIH grants DK099257, DK135606 (A.S.G.). We acknowledge T. Austin for facilitating the distribution of the hepatocytes. Support was also provided by the Digestive and Liver Disease Research Center of Columbia University Irving Medical Center, funded by 5P30DK132710. Samples for histological analysis were prepared in the Molecular Pathology Shared Resource of the Herbert Irving Comprehensive Cancer Center at Columbia University, supported by NIH/NCI grant P30-CA013696. Primary human HSCs were provided by Lifesharing Organ Procurement Organization.
Author information
Authors and Affiliations
Contributions
X.W. and I.T. conceived the study and wrote the manuscript, with all other authors providing comments and editing. X.W., M.P.M. and Y.X. conducted the experiments. S.K. and D.S. provided information regarding the genetic premise of the study. T.K. gave advice and reagents for the human HSC experiments. L.A.P.F., Z.H. and A.S.G. gave advice and reagents for the human hepatocyte experiments. D.G.S. was instrumental in the studies focused on the DED domains of caspase-8. Y.X., J.Z., and M.A.L. provided scRNA-seq data and insight related to the ALIOS model.
Corresponding authors
Ethics declarations
Competing interests
X.W. and I.T., through Columbia University, have filed a provisional patent application related to the therapeutic implications of the findings in this study. The authors declare no other competing interests.
Peer review
Peer review information
Nature Metabolism thanks Ekihiro Seki and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Revati Dewal, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Extended information for experiments in which hepatocyte caspase-8 was deleted in MASH mice, data related to Fig. 1.
a, CASP8 mRNA in the livers of healthy and MASLD human subjects, based on RNAseq GEO dataset 126848. Data are presented as mean ± SEM; *p=0.022, unpaired two-tailed t-test; normal, n=14, MASH, n=16. b, Immunoblot of full-length caspase-8 (Casp8) in normal and MASH human livers, with β-actin as loading control. c, Immunoblot of full-length caspase-8 and p43 cleaved caspase-8 (cl-Casp8), p18 cleaved caspase-8, and Casp8 mRNA, in chow- and FPC-fed mouse livers, with GAPDH as loading control. Data are presented as mean ± SEM; n=6; *p=0.017, unpaired two-tailed t-test. d, Immunoblot of full-length caspase-8 and cleaved caspase-8 in chow- and HF-CDAA-fed mouse livers, with GAPDH as loading control. e, Immunoblot of full-length caspase-8 in chow- and ALIOS-fed mouse livers, with β-actin as loading control. Data are presented as mean ± SEM; unpaired two-tailed t-test; GFP, n=8; Cre, n=8. f, Normal and MASH human livers were analyzed for caspase-8 immunofluorescence (red), human serum albumin (HSA) immunofluorescence (green), and DAPI immunofluorescence (blue). Arrows, an example of a caspase-8+ (CASP8+) HSA+ hepatocyte in each sample, highlighting the higher staining intensity of caspase-8 in MASH liver, which is quantified as MFI in the graph. Arrowheads, CASP8+ HSA− cells. Bar, 50 mm. Data are presented as mean ± SEM; n=4; ***p=0.0003, n.s., non-significant, unpaired two-tailed t-test. g, Immunoblot of full-length caspase-8 in primary human hepatocytes isolated from normal and MASH livers.
Extended Data Fig. 2 Extended information for experiments in which hepatocyte caspase-8 was deleted in MASH mice, data related to Figs. 1 and 2.
a-e, Male Casp8fl/fl mice were fed the FPC diet for 8 weeks to induce steatosis. The mice were then injected with AAV8-TBG-Cre or AAV8-TBG-LacZ control and continued on the FPC diet for an additional 8 weeks. Shown are body weight, percent liver weight of body weight, blood glucose 3 h after food withdrawal, liver triglyceride (TG) and myeloperoxidase (MPO) immunohistochemistry. Data are presented as mean ± SEM; n.s., non-significant, unpaired two-tailed t-test; LacZ, n=8; Cre, n=8. Bar, 200 mm for panel e. f, Cleaved caspase-3 (cl-Caspase 3) immunofluorescence with DAPI counterstain (blue) for nuclei in mice fed a chow diet or FPC diet for 16 weeks; bar, 50 mm. Data are presented as mean ± SEM; **p=0.005, unpaired two-tailed t-test; chow, n=3; FPC, n=3. g, Sirius red staining in mice fed a chow diet or the HF-CDAA diet for 8 weeks; bar, 200 mm. Data are presented as mean ± SEM; ****p<0.0001, unpaired two-tailed t-test; chow, n=5; HF-CDAA, n=5. h, Tgfb1 mRNA expression mice fed a chow diet or the HF-CDAA diet for 2, 4, or 8 weeks. Data are presented as mean ± SEM; ****p<0.0001, one-way ANOVA; chow, n=6; 2wk, n=5; 4wk, n=4; 8wk, n=5. (i-q) Male Casp8fl/fl mice were fed the HF-CDAA diet for 4 weeks to induce steatosis. The mice were then injected with AAV8-TBG-Cre or control AAV8-TBG-GFP and continued on the HF-CDAA diet for an additional 4 weeks. Data are presented as mean ± SEM; unpaired two-tailed t-test; GFP, n=7; Cre, n=7. i-m, Body weight, percent liver weight of body weight, blood glucose 3 h after food withdrawal, liver triglyceride (TG), and plasma ALT. n-q, The livers were analyzed for H&E staining, CD3 immunohistochemistry, TUNEL immunofluorescence (red), and p-H2AX immunohistochemistry. In p, DAPI counterstain for nuclei (blue) is shown in the bottom panels. Bars, 200 mm for panels n, o, and q and 50 mm for panel p. For panels i-q, data are presented as mean ± SEM; **p=0.0041; n.s., non-significant, unpaired t-test; GFP, n=7; Cre, n=7.
Extended Data Fig. 3 Extended information on hepatocyte caspase-8-induced HSC activation, related to Fig. 3.
a, Immunoblot of caspase-8 in AML12 cells treated with scrambled RNA (Scr) or siCasp8. b, CASP8 mRNA in THLE2 cells treated with scrambled RNA (Scr) or siCASP8. Data are presented as mean ± SEM; ***p=0.0002, unpaired two-tailed t-test; n=6 per group. c, THLE2 human HCs were transfected with scrambled RNA (Scr) or siCasp8. After 24 h, the media were changed to DMEM/0.1% FBS, and after an additional 24 h, the conditioned media from the cells was transferred to quiescent primary mouse HSCs. After 72 h, the HSCs were assayed for Timp1, Spp1, and Col1a1 mRNA. Data are presented as mean ± SEM; *p=0.017, **p=0.0066, two-way ANOVA; n=4 per group. d, Control or heat-inactivated (HI; 100 °C x 5 min) CM from Scr- or siCasp8-treated AML12 HCs was added to quiescent primary mouse HSCs as in panel a. After 72 h, the HSCs were assayed for the indicated mRNAs. Data are presented as mean ± SEM; ***p=0.0004, ****p<0.0001, n.s., non-significant, two-way ANOVA; n=4 per group. e, Left, CASP8 mRNA in primary human hepatocytes treated with scrambled RNA (Scr) or siCASP8. Data are presented as mean ± SEM; ****p<0.0001, unpaired two-tailed t-test; n=4 per group. Right, Primary human hepatocytes (HCs) were transfected with scrambled RNA (Scr) or siCasp8. After 24 h, the media were changed to DMEM/0.1% FBS, and after an additional 24 h, the conditioned media (CM) were transferred to human primary HSCs. After 48 h, the HSCs were assayed for TIMP1 and SPP1 mRNA. Data are presented as mean ± SEM; *p=0.024 (TIMP1), *p=0.023 (SPP1: grey vs. white), *p=0.016 (SPP1: hash vs. grey), **p = 0.0020, two-way ANOVA; n=4 per group. f, Human primary hepatic 3D spheroids treated with scrambled RNA (Scr) or siCASP8 (n=3 per group). Left, CASP8 mRNA. Data are presented as mean ± SEM; *p=0.036, unpaired two-tailed t-test. Right, The indicated mRNAs were assayed, Data are presented as mean ± SEM; *p=0.025 (SPP1), *p=0.012 (COL1A1), *p=0.049 (COL3A1), two-way ANOVA. g, AML12 mouse hepatocytes cultured in DMEM/0.1% FBS were treated ± IETD. After 24 h, the conditioned media (CM) were transferred to quiescent primary mouse HSCs, with one set of control HSCs receiving no CM (HSC only). After 72 h, the HSCs were assayed for Timp1 and Col1a1 mRNA. Data are presented as mean ± SEM; n.s., non-significant, one-way ANOVA; n=4 per group. h, Scr- or siCasp8-treated AML12 cells were transfected with GFP control, wild-type Casp8, or C360A-mutant Casp8. After 24 h, the media were changed to DMEM/0.1% FBS, and after another 24 h, the CM from the hepatocytes were transferred to quiescent primary mouse HSCs. After 72 h, the HSCs were assayed for Timp1 and Col1a1 mRNA. Data are presented as mean ± SEM; *p=0.012 (Timp1), **p=0.0055 (Timp1), *p=0.011 (Col1a1: GFP vs. CASP8), *p=0.033 (Col1a1: GFP vs. C360A), one-way ANOVA; n=4 per group. i, Scr- or siCASP8-treated THLE2 cells were assayed for CASP8 and METRN mRNA and for caspase-8 and meteorin protein. Data are presented as mean ± SEM; **p=0.0015 (CASP8 and METRN), unpaired two-tailed t-test; n=6 per group.
Extended Data Fig. 4 Extended information on the role hepatocyte caspase-8-induced meteorin in HSC activation, related to Fig. 4.
a, THLE2 cells were treated with scrambled RNA (Scr) or siMETRN. After 24 h, the media were changed to DMEM/0.1% FBS, and after an additional 24 h, the THLE2 cells were assayed for METRN mRNA and meteorin protein. Data are presented as mean ± SEM; ***p=0.0004, unpaired two-tailed t-test; n=3 per group. b, Upper, AML12 cells were treated 24 h with ZVAD or ITED, and Metrn mRNA was measured. Lower, AML12 cells were treated for 24 h with or without 5 mg/ml Jo2 to induce apoptosis in the presence of vehicle (DMSO) or 20 mm ZVAD or 20 mm ITED. The cleaved forms of caspase-3 (cl-Casp3) and caspase-8 (cl-Casp8), which are a measure of their activity in apoptosis, were assayed by immunoblot. Data are presented as mean ± SEM; n.s., no significant difference, unpaired two-tailed t-test; n=3 per group. c, Scr- or siCASP8-treated primary human hepatocytes were transfected with constructs expressing GFP control or DEDs. After 48 h, the cells were assayed for mRNAs encoding METRN and DED domains. Data are presented as mean ± SEM; *p=0.022, ***p=0.0007, ****p<0.0001, one-way ANOVA; n=3 per group. d, Immunoblot of caspase-10 (CASP10) in normal and MASH human livers, with β-actin as loading control. *p=0.029, unpaired two-tailed t-test; n=3 per group. e, Immunoblot and quantification of caspase-10 in primary human hepatocytes from normal and MASH livers. Data are presented as mean ± SEM; *p=0.026, unpaired two-tailed t-test; n=3. f, CASP10 and METRN mRNA in primary human hepatocytes treated with scrambled RNA (Scr) or siCASP10. Data are presented as mean ± SEM; *p=0.019, unpaired two-tailed t-test; n=4 per group. g, Primary mouse HSCs were transfected with scrambled RNA (Scr), siKit, or siStat3. After 48 h, Kit and Stat3 mRNA were measured. Data are presented as mean ± SEM; *p=0.024 (kit) *p=0.029 (Stat3), unpaired two-tailed t-test; left, n=3 per group, right, n=4 per group. h, Hepatocyte nuclear extracts from AML12 cells treated with scrambled RNA (Scr) or siCasp8 were subjected to Pol-II ChIP using anti-Pol-II or IgG control. The Metrn promoter region sequence was amplified by qPCR and normalized to the values obtained from input DNA. Data are presented as mean ± SEM; ***p=0.0003, one-way ANOVA; n=4 per group.
Extended Data Fig. 5 Extended information on the role of YY1 in Metrn induction, data related to Figs. 5 and 6.
a, Scr- or siCasp8-treated AML12 hepatocytes were immunostained for YY1 and then quantified for the percentage of cells with nuclear YY1. Bar, 200 mm. Data are presented as mean ± SEM; ***p=0.0003, unpaired two-tailed t-test; n=4 per group. b, Scr- or siYY1-treated AML12 hepatocytes were immunostained for YY1. Bar, 200 mm. c, Scr- or siCASP8-treated primary human hepatocytes were immunostained for YY1 and then quantified for the percentage of cells with nuclear YY1. Bar, 200 mm. Data are presented as mean ± SEM; *p=0.022, unpaired two-tailed t-test; n=3 per group. d, YY1 immunohistochemistry of liver sections of control (con) or hepatocyte caspase-8 knockout mice (KO) fed the FPC diet (related to Fig. 1a–i) and quantification; bar, 200 mm. Integrated density (IntDen) data for nuclear YY1 in parenchymal are presented as mean ± SEM; ****p<0.0001, unpaired two-tailed t-test; con, n=6; KO, n=6. e, YY1 immunohistochemistry of liver sections of normal and MASH human liver; bar, 50 mm. Arrows, parenchymal cells (hepatocytes). Integrated density (IntDen) data for nuclear YY1 in parenchymal cells are presented as mean ± SEM; ***p=0.0002, unpaired two-tailed t-test; normal, n=6; MASH, n=7. f, Meteorin immunoblot of liver extracts of normal and steatotic human liver. g, Casp8 immunoblot of liver extracts of chow- and 8 weeks FPC-fed liver. h, Immunohistochemistry and quantification for HC-like nuclear YY1 (arrows) in the livers of 8 weeks FPC diet-fed mice. Integrated density (IntDen) data for nuclear YY1 in parenchymal are presented as mean ± SEM; n.s., non-significant, unpaired two-tailed t-test; chow, n = 6; 8 weeks FPC, n = 6.
Extended Data Fig. 6 Extended information on the role of hepatocyte meteorin in experimental MASH, data related to Figs. 7 and 8.
a-c, Body weight, percent liver weight of body weight, and blood glucose 3 h after food withdrawal of the three groups of mice in Fig. 7a–f. d-h, Body weight, percent liver weight of body weight, blood glucose 3 h after food withdrawal, liver TG, and immunohistochemistry (bar, 200 mm) and quantification for CD3+ cells of the two groups of mice in Fig. 8. None of the values are significantly different by one-way ANOVA (panels a-f) or unpaired two-tailed t-test (panels g-h), basal, n=7; con, n=7; shMetrn, n=7.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–4.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 6 (download XLSX )
Statistical source data for Fig. 6.
Source Data Fig. 7 (download XLSX )
Statistical source data for Fig. 7.
Source Data Fig. 8 (download XLSX )
Statistical source data for Fig. 8.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig.2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data for Extended DataFig. 5.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Fig. 6.
Source Data Uncropped Gels (download PDF )
Uncropped gels.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Moore, M.P., Shi, H. et al. A non-apoptotic caspase-8–meteorin pathway in hepatocytes promotes MASH fibrosis. Nat Metab 7, 2067–2082 (2025). https://doi.org/10.1038/s42255-025-01355-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01355-1
This article is cited by
-
Modulation of metabolic, inflammatory, fibrotic, and cell death pathways by resmetirom in metabolic dysfunction-associated steatohepatitis (MASH): a transcriptomic profiling study
Acta Pharmacologica Sinica (2026)
-
Shooting for the stars: caspase-8–meteorin in MASH and fibrosis
Nature Metabolism (2025)