Abstract
The decline in ovarian function with age affects fertility and is associated with increased risk of age-related diseases, including osteoporosis and dementia. Notably, earlier menopause is linked to shorter lifespan, yet the molecular mechanisms underlying ovarian aging remain poorly understood. Recent evidence suggests the gut microbiota may influence ovarian health. Here we show that ovarian aging is associated with distinct gut microbial profiles in female mice and that the gut microbiome can directly influence ovarian health. Using fecal microbiota transplantation from young or estropausal female mice, we demonstrate that heterochronic microbiota transfer remodels the ovarian transcriptome, reduces inflammation-related gene expression and induces transcriptional features consistent with ovarian rejuvenation. These molecular changes are accompanied by enhanced ovarian health and increased fertility. Integrating metagenomics-based causal mediation analyses with serum untargeted metabolomics, we identify candidate microbial species and metabolites that may contribute to the observed effects. Our findings reveal a direct link between the gut microbiota and ovarian health.
Similar content being viewed by others
Main
The ovarian follicle pool is established before birth and is progressively depleted over time, culminating in menopause1,2,3. Menopause onset is associated with an elevated risk for many age-related conditions (for example, osteoporosis and dementia)4,5,6. Moreover, later age at menopause is a good predictor of extended lifespan7,8. While the profound effects of menopause on aging are well documented, the factors that contribute to ovarian aging and strategies for its prevention remain poorly understood.
During reproductive years, the hypothalamus–pituitary–ovarian axis regulates the production and release of hormones necessary for ovulation and menstruation9; however, as menopause progresses, the ovaries become less responsive, ultimately leading to characteristic hormonal imbalances10: increased circulating levels of follicle-stimulating hormone (FSH) and luteinizing hormone (LH) and decreased levels of anti-Müllerian hormone (AMH) and Inhibin A (INHBA)11,12,13. These hormonal adjustments are hallmarks of menopause and highlight diminished ovarian function10. Of note, this disrupted feedback may drive menopausal symptoms and health risks associated with this transition14,15. For instance, recent studies have shown that brain FSH signaling increases Alzheimer’s disease susceptibility and FSH blockade improves cognition in an Alzheimer’s mouse model14,16.
The adult human gut microbiome is composed of approximately 1013 to 1014 microorganisms17,18, composed of bacteria, viruses and fungi19. Perturbations in the gut microbiota composition can lead to disease development and/or progression20,21 (for example, diabetes and depression). Reported health effects of fecal microbiota transplantation (FMT) from aged donors to young recipients are inconsistent22,23. For example, one study demonstrated that FMT from aged to young mice led to accelerated brain, retina and gut aging in recipients22. In contrast, another study found that FMT from aged mice to prepubescent germ-free mice led to enhanced hippocampal neurogenesis and intestinal growth23. These findings underscore the microbiome’s complex, multifaceted effects on organ systems throughout life.
Patients with premature ovarian insufficiency (POI) have significantly altered gut microbial profiles24. Notably, gut microbiome alterations in patients with POI were associated with changes in FSH, LH and AMH serum levels24. Similarly, gut microbiome composition has been implicated in polycystic ovary syndrome (PCOS), a disorder characterized by hyperandrogenism and polycystic ovaries25. Specifically, FMT of samples from patients with PCOS led to ovarian dysfunction in otherwise healthy recipient mice25. Notably, 17β-estradiol levels are significantly lowered in germ-free female mice26. Moreover, transcriptome analysis of liver sexual development marker genes and histological analysis of ovarian follicles in germ-free female mice indicated that sexual maturation is perturbed in microbiota-depleted mice26. Of note, the gut microbiota can modulate serum estrogen levels (through the ‘estrobolome’), further highlighting a role of the gut microbiota in female reproductive health27. The gut microbiota regulates estrogenic action through β-glucuronidase secretion, which promotes estrogen bioavailability27,28,29. Gut microbiota dysbiosis can impair this process, leading to reduced circulating estrogens and potentially contributing to the development of estrogen-modulated conditions27. Collectively, these findings highlight a substantial crosstalk between the gut microbiota and ovarian tissue, suggesting a pivotal role for microbial communities in ovarian health.
In this study, we analyzed the gut microbial profiles of young and estropausal (post-reproductive phase in rodents, analogous to human menopause) female mice via 16S rRNA amplicon sequencing and explored how these distinct microbial communities affect ovarian health through FMT in young recipients. RNA-seq analyses revealed that FMT leads to profound changes in ovarian transcriptomes. Of note, heterochronic FMT of estropausal microbiota to young female recipients led to a reduction in ovarian inflammation-related gene expression and dampened expression of genes upregulated in aging ovaries. Consistently, estropausal FMT recipients showed improved ovarian function, supporting a noteworthy finding whereby old gut microbiota is beneficial to ovarian health. Last, causal mediation analyses identified specific microbial genera and species that might drive observed ovarian transcriptomic changes. Overall, our study shows that gut microbiota manipulation can directly promote ovarian health.
Results
Young and estropausal female mice show distinct fecal microbial profiles
To investigate the potential link between changes in the gut microbiome and ovarian aging in mice, we assessed (1) the ovarian health state and (2) microbial profiles of young female (4 months) and naturally aged estropausal female (20 months) C57BL/6JNia mice30 (Fig. 1a). To quantify overall ovarian health, we developed a composite ‘ovarian health index,’ modeled after the frailty index31 (Extended Data Fig. 1a). Our goal in constructing this index was to create a standardized, integrative tool that would enable comparative analysis of ovarian function across studies. The index combines ovarian follicle counts (Extended Data Fig. 1b and Methods) and serum levels of AMH, FSH and INHBA, using a three-tier scoring system based on the median young and estropausal values as reference points. Specifically, values beyond the young median are scored as 3, values between the medians of the young and estropausal groups are scored as 2, and values beyond the estropausal median are scored as 1 (Extended Data Fig. 1a). Given that ovarian follicle count and serum levels of AMH, FSH and INHBA are widely recognized as indicators of ovarian health32,33,34,35, we combined these parameters to form an objective evaluation of ovarian function (Methods). To promote broader use of the ovarian health index, we developed an online portal, allowing the community to analyze their own data (https://alanxu-usc.shinyapps.io/Ovarian_Health_Index_Calculator/).
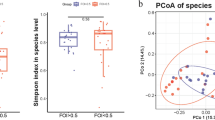
a, Schematic diagram of the experimental setup of female aging cohorts. b, Boxplot of combined follicle count of young and estropausal mice (P value ~ 0.0065). c, Boxplot of serum concentrations of AMH (P value ~ 0.0079). d, Boxplot of ovarian health index of young and estropausal mice (P value ~ 0.0117). e, PCA result of CLR-transformed and batch-corrected ASV counts from young and estropausal female mice. Animals from four independent cohorts (C1–C4), n = 19–20 per group (variation due to animal death before experiment; n = 20 for young female mice; n = 19 for estropausal female mice). f, Boxplots of observed features (P value ~ 9.1 × 10−4) and Shannon entropy (P value ~ 9.7 × 10−6) indices of young and estropausal female mice. Medians from young female mice for each cohort were used to normalize indices per cohort. g, Differential abundance analysis results of microbial genera of young versus estropausal female mice using ALDEx2. h, Functional abundance prediction analysis of young versus estropausal female mice using PICRUSt2. i, Boxplot of β-glucuronidase activity from PICRUSt2 (P value ~ 0.3759). Medians from young female mice for each cohort were used to normalize indices per cohort. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for b–d,f,i; Benjamini–Hochberg-corrected P value of Wilcoxon tests are reported for g and h. C, cohort.
Consistent with previous reports30,36, we observed a significant reduction in ovarian follicle counts and serum levels of AMH and INHBA, together with increased FSH levels in estropausal mice (Fig. 1b,c and Extended Data Fig. 1c–e). Furthermore, the estropausal group exhibited a significantly lower ovarian health index compared to the young group (P value ~ 0.0117; Fig. 1d), providing an internal consistency check that supports the use of our index as a measure of ovarian health.
We collected fecal samples from four independent cohorts of mice and performed 16S rRNA amplicon sequencing to compare the gut microbial profiles of the young and estropausal female mice. 16S rRNA amplicon sequencing allows identification of bacteria at the genus level by amplifying and sequencing variable, species-specific regions of the 16S rRNA gene (for example, V3–V4 regions)37. Quantification and analysis of variable region sequences, or amplicon sequence variants (ASVs), can provide important insights into microbial communities38.
Our analysis focused on both β diversity, which assesses differences between communities, and α diversity, which looks at within-community diversity39,40. The β diversity was evaluated using the Bray–Curtis dissimilarity and Jaccard index (Extended Data Fig. 2a). The analysis of Bray–Curtis dissimilarity and Jaccard index, conducted by cohort, revealed a distinct separation between the young and estropausal groups (Extended Data Fig. 2a). Moreover, the results of the principal component analysis (PCA), applied to center log-ratio (CLR)-transformed batch-corrected ASV counts, highlighted clear clustering between the young and estropausal female mice across cohorts (Fig. 1e). Batch correction was performed to account for technical variability across cohorts41. α diversity measures the diversity within a single microbial community, focusing on the variety and abundance of ASVs in a specific environment36. The estropausal female mice exhibited significantly higher scores in both observed features and Shannon entropy compared to young female mice (Fig. 1f). Our results suggest that the estropausal female mice have a higher biodiversity and increased species richness. This pattern of increased α diversity with age in female mice aligns with previous findings42; however, in humans, α diversity shows variable trends with age, with increases and decreases reported in different studies43,44. A recent study reported elevated α diversity in PCOS model mice45.
We next asked which specific microbial genera changed in abundance in young versus estropausal mice across cohorts (Fig. 1g and Supplementary Figs. 1a and 2a). Our analysis identified three genera that were more (‘UP in EF’) and six that were less abundant (‘DOWN in EF’) in estropausal female microbiota (Fig. 1g and Supplementary Figs. 1a and 2a; absolute effect size >1, adjusted P value < 0.05). Most differentially abundant genera belonged to the Lachnospiraceae family. Previous studies reported relative decreases in Lachnospiraceae microbial species with age in humans46,47. While these findings suggest an association between gut microbial shifts and ovarian aging, additional studies will be needed to determine how these genera are linked to ovarian health. Additionally, we conducted a predictive analysis of functional consequences of microbial abundance changes using PICRUSt2 (ref. 48) (Fig. 1h). This analysis suggested an increase in metabolic pathways such as ketogluconate metabolism and the superpathway of glycerol degradation to 1,3-propanediol, whereas pathways including the superpathway of fatty acid biosynthesis initiation and phospholipases were predicted to decrease in estropausal microbiota (Fig. 1h). These predicted changes, including an increase in ketoglutarate and alterations in lipid metabolism, have been previously associated with ovarian cancers, underscoring the potential relevance of our findings to ovarian health49,50. To further assess whether estrobolome-associated pathways might be affected with age, we examined changes in β-glucuronidase activity in female mice predicted using PICRUSt2, which revealed a nonsignificant trend of decreased β-glucuronidase activity with age (P value ~ 0.3759; Fig. 1i). Our results collectively underscore the pronounced differences in the fecal microbial profiles between young and estropausal female mice.
Chemically induced premature ovarian failure model mice show distinct fecal microbial profiles
The 4-vinylcyclohexene diepoxide (VCD)-injected mouse model is a follicle-deplete, ovary-intact animal model which recapitulates many aspects of human ovarian aging51,52. VCD accelerates atresia by causing selective loss of primary and primordial follicles, while leaving the rest of the ovarian structure intact51,52. We assessed the ovarian health index and fecal microbial profiles of vehicle- and VCD-exposed mice (CTL and VCD, respectively; Extended Data Fig. 3a). In brief, 4-month-old C57BL/6NTac female mice were subjected to daily intraperitoneal injections of either vehicle (safflower oil) or VCD (160 mg kg−1 day−1) for 15 consecutive days, then rested for 100 days53 (Extended Data Fig. 3a). To determine whether the observed effects on microbial profiles in female mice were a direct consequence of VCD-induced ovarian dysfunction rather than off-target effects of VCD, we also generated data from matched male controls.
For female mice, we validated ovarian follicle depletion by histology and alterations in AMH, FSH and INHBA levels by serum measurements (Extended Data Fig. 3b–e). As expected, VCD-treated female mice had reduced ovarian health index (P value ~ 0.0104; Extended Data Fig. 3f).
We conducted 16S rRNA V3–V4 amplicon sequencing on vehicle- and VCD-treated animals. PCA of CLR-transformed ASV counts, along with Bray–Curtis dissimilarity and Jaccard index analyses, showed separation between the vehicle-treated (CTL) and VCD-exposed female mice (Extended Data Fig. 4a, b). The α diversity indices, on the other hand, revealed no significant differences between groups (Extended Data Fig. 4c). These findings suggest that the overall diversity within the group’s communities may seem similar, but they may be differences in the species present, their relative abundances and functions. Of note, analysis in male mice showed no changes in β diversity (Extended Data Fig. 4a, b) nor in α diversity between groups (Extended Data Fig. 4d). These findings support the notion that observed effects in female mice are causally induced by ovarian dysfunction and not VCD off-target effects. Consistently, body weight changes from pre-injection to killing did not significantly differ between groups in either sex, suggesting that weight changes are unlikely to account for observed microbial shifts (Extended Data Fig. 4e).
Our analysis revealed differentially abundant genera from CTL versus VCD groups (Extended Data Fig. 4f and Supplementary Fig. 3a; absolute effect size >1, adjusted P value < 0.05). We detected several genera from the Lachnospiraceae family, yet none of the unique ASV IDs overlapped with those identified in estropausal female mice. We utilized PICRUSt2 to predict functional abundance between the CTL and VCD groups (Extended Data Fig. 4g). Notably, we detected downregulation of the phospholipase pathway in the VCD group, consistent with observations in estropausal female mice (Fig. 1h and Extended Data Fig. 4g). To note, no difference in predicted β-glucuronidase activity between the groups was detected (P value ~ 0.8413; Extended Data Fig. 4h). This lack of change may reflect the fact that VCD induces ovarian failure without substantially altering the taxa contributing to β-glucuronidase activity. Alternatively, estrobolome gut microbes may require longer-term physiological changes, as seen in natural aging, to manifest detectable shifts in functional potential. Our findings suggest that physiological ovarian aging and chemically induced ovarian failure lead to substantially different outcomes on the gut microbiota.
Young and aged male mice show distinct fecal microbial profiles
We also characterized the fecal microbial profiles of young (4 months) and naturally aged (20 months) C57BL/JNia male mice using 16S rRNA amplicon sequencing in three independent mouse cohorts, as above (Extended Data Fig. 5a). Comparisons between female and male aging mice, as well as VCD-exposed mice, aimed at identifying differences in microbial profiles attributable to general versus ovarian aging. PCA of CLR-transformed ASV counts, coupled with Bray–Curtis dissimilarity and Jaccard index analysis, showed a distinct divergence between young versus aged males (Extended Data Fig. 5b, c); however, this separation was less pronounced than what we observed in female mice (Fig. 1e). The α diversity indices indicated no significant changes in aging males (Extended Data Fig. 5d), consistent with a previous study54. Through differential abundance and functional abundance prediction analyses, we identified several genera and functional terms that varied significantly in aging males (Extended Data Fig. 5e, f and Supplementary Fig. 4). Of note, some functional terms were consistent between female and male aging cohorts. Specifically, the L-lysine biosynthesis I pathway was upregulated, whereas the pyridoxal 5-phosphate biosynthesis I pathway and the superpathway of pyridoxal 5-phosphate biosynthesis were downregulated with aging across sexes (Fig. 1h and Extended Data Fig. 5f). These results suggest that these pathways may play roles in the general aging processes, which can be elucidated with future research.
Gut microbiota manipulation can induce changes in the ovarian transcriptome
We conducted FMT experiments to directly assess how gut microbiota alterations after estropause may influence ovarian health and function. Young adult female mice (4 months) were subjected to antibiotics/antimycotic (Ab/Am) treatment to eliminate their initial microbiome, and then received fecal microbiota grafts from either young female (FMT-YF) or estropausal female (FMT-EF) mice (Fig. 2a). Subsequently, the ovaries were collected for bulk RNA-seq analysis (Fig. 2a). Based on previous research that has shown adverse effects of gut microbiota from donors with ovarian dysfunction (for example, PCOS25), we initially hypothesized that FMT from estropausal mice would promote estropause-like phenotypes in recipients.
a, Schematic diagram of the experimental setup of FMT cohort. Fecal microbiota from either young (4 months) or estropausal (20 months) female mice were transplanted into young (4 months) female recipient mice. FMT recipient groups are referred to as FMT-YF (receiving young donor microbiota) and FMT-EF (receiving estropausal donor microbiota). b, PCA result of RNA expression between young and estropausal female FMT recipient mice (FMT-YF and FMT-EF, respectively). c,d, Top GSEA enriched terms from Gene Ontology (GO) Biological Process (BP) (c) and GO Molecular Function (MF) (d). Only up to the top ten most-upregulated and top ten most-downregulated gene sets are plotted for readability. e, Heat map of normalized expression of cytokine genes from ovarian RNA-seq data. f, Representative images of ovarian immunohistochemistry (IHC) against IL-6. Scale bar, 50 µm. g, Quantification of IL-6 IHC data (P value ~ 0.0439). Diaminobenzidine (DAB) signal intensity was normalized to the percentage of total tissue area (hematoxylin signal). For IHC quantification, one data point from the FMT-EF group was excluded based on Grubbs’ test identifying it as a significant outlier (P value < 0.05). Boxplot shows the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the IQR. Individual datapoints are shown. Significance in a nonparametric two-sided Wilcoxon rank-sum test is reported.
PCA and multidimensional scaling (MDS) analyses of the ovarian RNA-seq dataset showed clear separation between the FMT-YF and FMT-EF groups (Fig. 2b and Extended Data Fig. 6a). Additionally, we detected 2,131 differentially expressed genes between the FMT-YF and FMT-EF groups (false discovery rate (FDR) < 5%; Extended Data Fig. 6b). We investigated pathways that were distinctively regulated between the FMT-YF and FMT-EF groups. Gene set enrichment analysis (GSEA)55 results highlighted that immune regulation-related pathways were significantly downregulated with FMT-EF (Fig. 2c, d and Extended Data Fig. 6c). To evaluate whether these changes resulted from changes in immune cell infiltration, we leveraged a publicly available ovarian single-cell RNA-seq dataset56 to conduct deconvolution and infer immune cell proportions within our samples. Two distinct tools, CSCDRNA57 and Granulator (https://github.com/Novartis/granulator), predicted no significant differences in immune cell proportions between FMT-YF and FMT-EF ovaries (Extended Data Fig. 6d), consistent with the notion that observed transcriptional changes are cell autonomous.
Next, we examined expression profiles of detected genes encoding pro- and anti-inflammatory cytokines in our ovarian RNA-seq data58,59. Consistent with GSEA, our analysis showed overall decreased proinflammatory cytokine expression and increased anti-inflammatory cytokine expression in FMT-EF versus FMT-YF ovaries (Fig. 2e). To validate these findings at the protein level, we performed immunohistochemistry for interleukin (IL)-6, a well-established proinflammatory cytokine implicated in ovarian function60. Consistently, IL-6 levels were significantly reduced in the ovaries of FMT-EF mice (P value ~ 0.0439; Fig. 2f, g), further supporting the conclusion that FMT-EF leads to reduced inflammatory signaling in the ovary.
GSEA also revealed terms associated with chromatin structure and dynamics, including kinetochore, chromatin silencing complex, chromosome, centromeric region and spindle pole (Extended Data Fig. 6c). These findings suggest that transcriptional changes in the ovaries encompass chromatin organization and chromosome dynamics, processes critical for maintaining oocyte quality61,62. Together, these results indicate that pathways associated with both immune regulation and chromatin dynamics distinguish FMT-YF and FMT-EF ovaries.
To identify transcription factors (TFs) potentially driving FMT-driven transcriptomic alterations, we utilized Gene Transcription Regulation Database (GTRD)63 with the GSEA paradigm. With a lenient FDR < 10% threshold, we identified two TFs showing trends of increased activity and eight trends of decreased activity in FMT-EF ovaries (Extended Data Fig. 6e). We identified publicly available ChIP-seq datasets for the trending TFs via the Cistrome Data Browser64,65, using corresponding bona fide peaks to determine potential enrichment by GSEA. Our analysis showed significant negative enrichment scores of Ncoa1 and Usp7 target genes within the FMT-EF group (P value ~ 0.0138 and ~1 × 10−4, respectively; Extended Data Fig. 6f). Notably, we noted that Ncoa1 and Usp7 expression levels remained consistent across FMT-YF and FMT-EF groups (Extended Data Fig. 6g), suggesting post-transcriptional or post-translational regulation. While little is known on the roles of Ncoa1 and Usp7 in the ovary, Ncoa1 function is important in egg production traits in Shaobo hens66 and P5091, an inhibitor of Usp7, inhibits ovarian cancer cell proliferation67. Together, our data suggest that Ncoa1 and Usp7 represent potential regulators of ovarian function.
Heterochronic FMT may have rejuvenating effects on the ovarian transcriptome of the recipients
Cumulative studies have indicated that aging leads to a proinflammatory shift in the ovary, adversely affecting ovarian function and oocyte quality68,69. Based on our unbiased GSEA, we next asked whether FMT-EF-driven changes in the ovarian transcriptome could reflect a more general trend of transcriptional rejuvenation. For this purpose, we analyzed two publicly available mouse ovarian aging bulk RNA-seq datasets70,71 to identify gene sets up- or downregulated with ovarian aging (‘UP with ovarian aging’ and ‘DOWN with ovarian aging’; Fig. 3a), and used these gene sets for GSEA. Of note (and contrary to expectations based on previously reported observations at the somatic level for heterochronic FMT), our findings revealed a significant negative enrichment score for genes that were upregulated with ovarian aging (P value ~ 0.0046; Fig. 3b), suggesting that the ovarian transcriptome of the heterochronic FMT recipients (FMT-EF group) displays ‘younger’ transcriptional profiles. As an orthogonal line of evidence, we next asked whether cellular senescence (as measured by the highly curated senescent marker gene set, SenMayo72) was significantly impacted by FMT-EF. Of note, our results showed a significant downregulation of the SenMayo gene set in the FMT-EF group, consistent with overall ‘rejuvenation’ of the ovarian transcriptome upon FMT-EF (P value ~ 0.0382; Fig. 3c).
a, Schematic diagram of the analysis setup of public ovarian aging bulk RNA-seq datasets, CRA003645 and PRJNA1002222, from young (2–3 months) and estropausal (12 months) female mice. b,c, GSEA enrichment score plots of genes that are upregulated with ovarian aging (‘UP with ovarian aging’, left, P value ~ 0.0046) and downregulated with ovarian aging (‘DOWN with ovarian aging, right, P value ~ 0.305) (b) and SenMayo gene set (P value ~ 0.0382) (c). In both b and c, the black vertical lines indicate the positions of the genes from the gene set within the ranked list of genes. The curve represents the running enrichment score (ES), with the peak indicating the point of maximum enrichment for the gene set. The color scale at the bottom of the plot reflects the ranking metric, where red represents upregulated genes and blue represents downregulated genes. Benjamini–Hochberg-adjusted P values derived from permutation-based GSEA statistics are reported. d, Boxplots of DESeq2 normalized log2 counts of Cd38, Cdkn1a and Il1a (P values ~ 0.0111, ~0.2086 and ~0.0041, respectively). Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× IQR. Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for d. DGE, differential gene expression.
A recently published study highlighted the role of nicotinamide adenine dinucleotide (NAD) metabolism in ovarian aging, showing that increased NADase CD38 expression accelerated ovarian aging71. CD38 deletion or pharmacological inhibition preserved fertility and follicular reserves in aged mice71. Thus, we examined the expression of Cd38 as well as inflammation markers from that study, Cdkn1a and Il1a (Fig. 3d). We observed reduced Cd38 expression in the FMT-EF group compared to the FMT-YF group, along with decreased expression of the inflammation markers (Fig. 3d). These findings suggest a potential involvement of the CD38–NAD pathway in the observed phenotypes of the FMT-EF group.
FMT-EF mice show improved ovarian health
As we observed transcriptomic remodeling consistent with a more youthful ovarian state in the FMT-EF group, we next asked whether FMT-EF led to enhanced ovarian function using a two-pronged approach (Fig. 4a): (1) calculating the ovarian health index, and (2) evaluating effective fertility using litter size and time to first pregnancy after mating73 (Fig. 4).
a, Schematic diagram of the experimental setup. b, Boxplot of ovarian health index of FMT-YF and FMT-EF mice (P value ~ 0.0255). Ovarian health index was calculated for two independent cohorts (n = 5 or 7 per group and 10 or 14 per cohort). c, Boxplot of pup counts of first litters of young and estropausal female mice FMT recipient mice (FMT-YF and FMT-EF, respectively; P value ~ 0.1726). Pup counts were measured from two independent cohorts (n = 8 or 10 per group and 16 or 20 per cohort). d, Reproductive success rate analysis result shown as percentages of nulliparous between FMT-YF and FMT-EF (P value ~ 0.002). Markers on the x axis indicate the median latency to breeding for each group. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× IQR. Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for b and c, and a log-rank test for d.
Consistent with our predictions, FMT-EF mice exhibited significantly higher ovarian health indexes than FMT-YF mice (P value ~ 0.0255; Fig. 4b and Extended Data Fig. 7a–d). Notably, for the FMT cohorts, FSH levels were measured using two different methods during the study due to an update implemented by the core facility conducting the assays. To ensure consistency and comparability across cohorts, we applied an assay-specific correction (Methods) to ensure that FSH levels were directly comparable (Extended Data Fig. 7d).
We also evaluated the endpoint of ovarian function (fertility) through litter size and reproductive success. There was a suggestive trend for increased litter size in FMT-EF mice (P value ~ 0.1726; Fig. 4c). In addition, FMT-EF mice remained nulliparous for a significantly shorter amount of time after male pairing (P value ~ 0.002; Fig. 4d). The median time to birth was 23 days for FMT-YF mice and 21.5 days for FMT-EF mice, corresponding to a modest but substantial 6.7% decrease in latency to first litter. To determine whether the inclusion of 0 pup count data influenced our findings, we reanalyzed our data after excluding animals with 0 pup counts. Of note, the latency data remained statistically significant (P value ~ 0.0159 without 0 pup counts versus P value ~ 0.002 with all data). Together, our findings are consistent with improved/rejuvenated ovarian function in response to FMT-EF.
Notably, no significant differences were observed in body weight between FMT-YF and FMT-EF groups after Ab/Am treatment (Extended Data Fig. 7e and Supplementary Table 1). Estrous cycle staging on the day of killing also revealed no significant differences between groups (Supplementary Table 2).
FMT-YF and FMT-EF mice show distinct microbial profiles
We next verified that fecal transplants from young versus estropausal female mice led to distinct microbial profiles in recipients. For this purpose, we conducted 16S rRNA V3–V4 amplicon sequencing on the gut microbiota of FMT-YF and FMT-EF mice (Extended Data Fig. 8).
PCA of CLR-transformed ASV counts, complemented by Bray–Curtis dissimilarity and Jaccard index analyses, showed a clear segregation between the FMT-YF and FMT-EF groups (Extended Data Fig. 8a, b). While no significant differences were noted across α diversity indices (Extended Data Fig. 8c), several microbial genera showed significant differential abundance between the FMT-YF and FMT-EF groups (absolute effect size >1, adjusted P value < 0.05; Extended Data Fig. 8d and Supplementary Fig. 5). Notably, two genera with increased abundance in FMT-EF recipients were also elevated in estropausal donors (Bacteria;Firmicutes;Clostridia;Clostridiales;Ruminococcaceae and Bacteria;Firmicutes;Clostridia; Clostridiales;Lachnospiraceae;Coprococcus 2;uncultured organism; Extended Data Fig. 8e). These results indicate that FMT from young or estropausal female mice changes the microbial landscape of recipients. Other studies also reported the selective enrichment of specific microbial species from donor microbiota in recipients22,74, which may be attributable to variations in intestinal microenvironments that allow the proliferation of certain species.
We used PICRUSt2 to predict functional abundance differences between the FMT-YF and FMT-EF groups (Extended Data Fig. 8f). We identified only significantly upregulated (but not downregulated) functional groups in the FMT-EF group. Notably, there was a notable presence of terms related to the menaquinol biosynthesis pathway, including those involved in 1,4-dihydroxy-6-naphthoate biosynthesis II, 1,4-dihydroxy-6-naphthoate biosynthesis I, and the superpathway of menaquinol-8 biosynthesis II (Extended Data Fig. 8f). Menaquinols, the active hydroquinone forms of vitamin K, also known as menaquinones (MKs), facilitate the carboxylation of vitamin K-dependent proteins, which are crucial for blood clotting, bone health and heart health75.
To enable us to pinpoint specific microbial species, we next performed shotgun metagenomics on the same cohort (Fig. 5a). Consistent with the 16S amplicon data, PCA of CLR-transformed data and β diversity analysis revealed distinct clustering between the FMT-YF and FMT-EF groups (Fig. 5b, c), and no significant differences were observed in α diversity (Fig. 5d). Shotgun metagenomics analysis identified 168 differentially abundant microbial species (FDR < 5%; Fig. 5e, f and Supplementary Fig. 6), confirming distinct FMT-YF and FMT-EF microbial compositions. Notably, microbial species previously reported to be differentially enriched in the fecal microbiota of patients with PCOS, such as Bacteroides xylanisolvens, Bacteroides thetaiotaomicron, Parabacteroides distasonis and Bacteroides stercoris25, were significantly depleted in the FMT-EF group.
a, Schematic diagram of the experimental setup. b, PCA result of CLR-transformed and batch-corrected counts from young and estropausal female mice FMT recipient mice (FMT-YF and FMT-EF, respectively). c, Principal coordinate analysis results of Bray–Curtis dissimilarity (left) and Jaccard (right) indices. d, Boxplots of observed features (P value ~ 0.389) and Shannon entropy (P value ~ 0.818) indices of FMT-YF and FMT-EF mice. e, Volcano plot of differential abundance analysis results of microbial species of FMT-YF versus FMT-EF mice using ALDEx2. f, Bubble plot of top ten up- and downregulated microbial species in FMT-EF. g, Functional abundance analysis of FMT-YF versus FMT-EF mice using HUMAnN3 (q-value < 0.1). h, Boxplot of β-glucuronidase activity from HUMAnN3 (P value ~ 0.317). Median from FMT-YF group was used to normalize indices. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× IQR. Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for d and h, and Benjamini–Hochberg-corrected P values of Wilcoxon tests are reported for f.
Functional abundance analysis using HUMAnN3 (ref. 76) revealed upregulation of pathways related to menaquinol biosynthesis, including PWY-7371: 1,4-dihydroxy-6-naphthoate biosynthesis II, consistent with predictions from the 16S amplicon analysis (Fig. 5g and Extended Data Fig. 8f). Intriguingly, we detected terms associated with the NAD biosynthesis pathway (for example, PWY−7761: NAD salvage pathway II (PNC IV cycle), PYRIDNUCSYN−PWY: NAD de novo biosynthesis I (from aspartate)), which aligns with our ovarian RNA-seq data showing altered Cd38 expression in FMT-EF ovaries (Fig. 3d).
Given the stability of menaquinones and their primary synthesis by gut microbiota, we further evaluated their levels in stool samples (MK6-MK13) to explore potential systemic effects post-absorption, as they contribute substantially to the host’s vitamin K status; however, no significant differences in MK concentrations in the stool samples of FMT-YF and FMT-EF animals were observed (Extended Data Fig. 9a,b), although this may result from high animal-to-animal variability and/or high vitamin K content of standard chow (‘Discussion’).
We also examined predicted β-glucuronidase activity in the FMT cohort using both HUMAnN3 (shotgun metagenomics-based) and PICRUSt2 (16S amplicon-based) analyses (Fig. 5h and Extended Data Fig. 8g). In both datasets, we observed nonsignificant increasing trends in predicted β-glucuronidase activity in the FMT-EF group compared to the FMT-YF group, with a stronger effect being detected in the amplicon data (P value ~ 0.317 for shotgun metagenomics; P value ~ 0.0513 for 16S amplicon).
FMT-YF and FMT-EF mice show distinct serum metabolomic profiles
It is well established that microbiota-derived molecules can enter the bloodstream and influence health77. Additionally, plasma metabolite levels can be affected by aging78. To assess whether gut microbial profile shifts in FMT animals induced changes in circulating metabolites, we performed untargeted metabolomics on serum from two independent cohorts (Fig. 6a and Supplementary Table 3). First, we performed PCA of identified metabolites. Although nonlinear batch effects were present and could not be fully corrected, clear separation was evident within each batch (Fig. 6b). Subsequent linear discriminant analysis (LDA) supported distinct metabolomic profiles of the two groups (Fig. 6c). Thus, we extracted metabolites that changed consistently across cohorts and selected the top 15 based on LDA loadings (Fig. 6d, e and Supplementary Fig. 7a).
a, Schematic diagram of the experimental setup. b, PCA plots of identified serum metabolites from two independent FMT cohorts. c, LDA projection scatter-plot displaying LDA1 scores for the FMT-YF and FMT-EF groups. d, Top 15 metabolites contributing to LDA-based separation, ranked by their positive and negative contributions to the LDA axis. e, Boxplots of 3,4-dihydroxyphenylpropionic acid (3,4-DHPPA, P value ~ 0.07923) and lactamide (P value ~ 0.09084) serum levels between FMT-YF and FMT-EF groups. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× IQR. Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for e.
Among identified metabolites, 3,4-dihydroxyphenylpropionic acid (3,4-DHPPA) and lactamide were significantly decreased in FMT-EF animals at FDR < 0.1 (Fig. 6e). 3,4-DHPPA, a microbial metabolite derived from polyphenol metabolism, has been linked to lipid metabolism and broader metabolic regulation79. Given the critical role of lipid metabolism in hormonal balance and ovarian function, its reduction in FMT-EF animals may reflect metabolic adaptations that contribute to improved ovarian health. Additionally, while the direct role of lactamide in ovarian function is unclear, microbial-derived lactic acid and its derivatives have been implicated in gut microbiota-driven regulation of reproductive health80. For instance, lactic acid bacteria have been shown to influence sex hormone-related gut microbiota composition, leading to beneficial effects on ovarian physiology in preclinical models80. The observed decrease in lactamide levels may indicate broader microbiome-mediated metabolic changes supporting ovarian health.
In addition to significant changes, palmitoylcarnitine, a key metabolite involved in fatty acid β-oxidation and mitochondrial energy production81, was elevated in FMT-EF animals according to LDA loadings (Fig. 6d). This suggests enhanced mitochondrial efficiency and metabolic flexibility, which are essential for oocyte and follicular health82. Furthermore, palmitoylcarnitine is involved in immune modulation83, and increased levels may indicate a shift supporting both energy homeostasis and an anti-inflammatory environment. This aligns with our bulk RNA-seq analysis, which revealed a downregulation of inflammatory pathways in FMT-EF ovaries (Fig. 2c–e).
Conversely, LDA loadings suggested that serum β-estradiol levels are decreased in FMT-EF mice (Fig. 6d). Based on improved ovarian function in FMT-EF mice, this may reflect a more physiologically regulated hormonal environment, rather than a pathological decline. Estradiol plays a critical role in ovarian function, lipid metabolism and mitochondrial activity84, and its levels may be influenced by gut microbiota-driven metabolism27. Notably, predicted β-glucuronidase activity was elevated in the FMT-EF group compared to FMT-YF (Fig. 5h and Extended Data Fig. 8g), despite the lower basal serum estradiol levels. This suggests that increased β-glucuronidase activity may contribute to altered local or systemic estrogen metabolism without necessarily raising basal circulating estradiol, potentially supporting tissue-level hormonal signaling in a more regulated manner. The gut microbiome is known to modulate steroid hormone metabolism27, and FMT from estropausal donors may have contributed to a systemic reprogramming of lipid and hormonal pathways that promote ovarian resilience.
Together, these findings suggest that FMT from estropausal donors induces distinct metabolic shifts that may contribute to ovarian health observed improvements.
Causal mediation analyses reveal microbial species potentially influencing ovarian transcriptome
Last, we performed causal mediation analyses to characterize microbial species that may directly influence ovarian transcriptome remodeling between FMT-YF and FMT-EF mice (Fig. 7a). From the features exhibiting significant abundance variations between groups, we identified four microbial species (Bacteroides zhangwenhongii, Shigella flexneri, Bacteroides stercoris and Bacteroides graminisolvens) with a significant mediation effect (Fig. 7b). Of note, Bacteroides stercoris was previously highlighted as a key contributor to the distinct fecal microbiota profiles between healthy individuals and those with PCOS25. Similarly, we identified four ASV IDs (850e0d58c1aa35238538564a8010e0ff, e4f32cdc3407edb8423447421078fdae, 55ebdcf041fb6b8b4632993423011372 and d4840c45eb37593108d020fb6a63f8e2) with a significant mediation effect on ovarian transcriptome changes (Extended Data Fig. 10a, b).
a, Graphical representation of causal mediation analysis. b, Causal mediation analysis results of shotgun metagenomics counts that showed significant mediation effects on changes in the ovarian transcriptome in FMT recipient mice. Bars represent model-derived mediation estimates (average causal mediation effect, average direct effect and total effect). Vertical lines denote 95% nonparametric bootstrap CIs. Statistical significance is denoted as NS, not significant; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. c, Heat map of Spearman correlation analysis between CLR-transformed shotgun metagenomics count data and serum hormone quantification data. #P < 0.1; *P < 0.05. d, Summary diagram. The estropausal female fecal microbiota exhibits increased levels of commensal microbes and metabolites through compensatory mechanisms. As a result, fecal transplantation of estropausal microbiota enhances ovarian health measures. Potential mechanisms, including NAD and vitamin K synthesis pathways, should be further examined. ACME, average causal mediated effect; ADE, average direct effect.
Finally, we performed Spearman correlation analysis using shotgun metagenomics and hormone quantification data (Fig. 7c). This analysis revealed significant correlations between multiple microbial species and hormone levels, including Shigella flexneri, Parabacteroides merdae and Porphyromonas sp. oral taxon 275, suggesting these species as potential candidates influencing ovarian function. These findings provide a strong foundation for future studies aimed at experimentally validating the roles of these microbial species in modulating ovarian function and hormone levels.
Discussion
In this study, we examined the fecal microbial profiles of three distinct groups: (1) young and estropausal female mice, (2) mice with chemically induced premature ovarian failure and their control counterparts, and (3) young and aged male mice. We identified functional pathways that may be unique to ovarian aging (phospholipases pathway) and some common to aging (L-lysine biosynthesis I, pyridoxal 5-phosphate biosynthesis I and superpathway of pyridoxal 5-phosphate biosynthesis). Furthermore, through FMT, we discovered that age-related alterations in the gut microbiota can substantially impact ovarian phenotypes. We also identified specific microbial species as potential mediators of the observed effects on ovarian function. Further studies will be needed to assess whether these species play a mechanistic role in ovarian health modulation. The vitamin K2 and CD38-NAD pathways emerged as potential mechanisms to explore for investigating the underlying causes of the observed phenotypes in our FMT experiments (Fig. 7d). Future investigations focusing on these species hold considerable promise for crafting new therapeutic strategies aimed at mitigating or reversing ovarian aging.
Our analysis of ovarian RNA-seq data from the FMT recipients revealed significant changes in the ovarian transcriptome following alterations in the gut microbiota. While existing research underscores the broad effects of the gut microbiome on host health, evidence detailing its direct effects on the transcriptome of distal tissues remains sparse. For instance, earlier research demonstrated that mice raised in a germ-free environment exhibit a markedly distinct liver transcriptome compared to their conventionally raised counterparts26. To the best of our knowledge, a direct influence of gut microbiota on ovarian transcriptomes has not yet been documented. Our analysis not only uncovered notable variations in the ovarian transcriptome of FMT recipients (reduced inflammatory pathway with FMT-EF), but we also characterized functional impact of phenotypic alterations (improved fertility, ovarian health). Collectively, these results offer substantial evidence of gut microbiota’s direct role in influencing ovarian function. Furthermore, the microbial species and genera and functional pathways we identified serve as valuable assets for future research aimed at investigating therapeutic strategies.
Based on previous research showing detrimental effects of gut microbiota from donors with ovarian dysfunction25, we initially hypothesized that FMT from estropausal mice would induce estropause-like phenotypes in recipients. However, we observed rejuvenated ovarian transcriptomes and improvements in measures of ovarian health and fertility. These unexpected advantageous effects could be attributed to compensatory mechanisms. For instance, FSH, required for follicle maturation and ovulation, is elevated in post-menopausal humans and estropausal mice2,85,86. After reproductive senescence, the ovaries no longer produce estrogen or mature eggs, so the pituitary compensates by producing more FSH in an attempt to stimulate nonresponsive ovaries30. Similarly, it is conceivable that the post-reproductive gut microbiota could become enriched with microbes and/or metabolites that support ovarian health. Cumulative studies have highlighted the role of gut microbiota in compensatory processes. For instance, the gut microbiota can influence peptide levels that regulate energy balance in the hypothalamus, promoting decreased body fat87. Furthermore, patients with mild cognitive impairment show increased abundance of microbes producing metabolites critical for nervous system’s function (such as serotonin, acetylcholine, histamine and dopamine)88. Therefore, when young adult female mice received FMT from estropausal female mice, it is possible that the enriched gut microbial environment contributed to the manifestation of younger or more reproductively active phenotypes in recipients. Consistently, we observed elevated predicted β-glucuronidase activity in FMT-EF recipients compared to FMT-YF, in contrast to the decline in β-glucuronidase activity seen with natural aging. This enzymatic function, linked to the estrobolome and estrogen metabolism, may represent a transmissible functional adaptation of the donor microbiota that supports hormonal and metabolic regulation despite lower basal serum estradiol levels.
Although our analysis of functional abundance from both shotgun metagenomics and 16S rRNA amplicon sequencing datasets predicted potential shifts in the biosynthesis pathway of MKs in the FMT mice, we observed no significant differences in the stool concentrations of MKs between the FMT-YF and FMT-EF groups. This lack of significant difference might be attributed to the vitamin K content of the standard University of Southern California (USC) vivarium chow (LabDiet, PicoLab Rodent Diet 20). Specifically, the high vitamin K dietary content could have saturated the system, thus masking subtle variations in microbial MK production. Such nuances are typically more pronounced under conditions of a low vitamin K diet89. Vitamin K status influences MK production because gut bacteria require vitamin K substrates to synthesize MKs89. In our study, the presence of vitamin K3 (menadione) in the diet, which is converted into MK4 in the body, likely contributed to the high basal MK levels observed in the fecal matter of recipients. While there have been limited studies, MKs have demonstrated potential benefits for ovarian health in specific contexts90,91 (for example, ovarian cancers and PCOS). Additionally, studies have highlighted anti-inflammatory properties of MKs92. Given our observations of decreased inflammatory function and enhanced ovarian function in the FMT-EF mice, coupled with documented benefits of MKs, MKs may be therapeutic candidates for improving ovarian health and function; however, further research into the direct effects of MKs on ovarian function is needed.
In our study, while we detected certain similarities in the microbial profiles of aging female mice and males, our analysis primarily revealed divergences between sexes. Specifically, while aging was associated with significant changes in α diversity in female mice, such changes were not observed in males. Moreover, our analyses of differential abundance of microbial genera and predictions of functional abundance revealed substantial differences in the gut microbial profiles and their functions between aging female and male mice. Indeed, extensive research has established that aging manifests differently in females and males, a phenomenon known as sex dimorphism93. Particularly, susceptibility to age-related diseases associated (for example, Alzheimer’s, obesity) varies significantly between sexes94,95,96. Furthermore, differences in female vs. male lifespan are consistently observed across species97,98. In agreement with our findings, sex differences in the aging gut microbiome have previously been documented99,100. While the limited number of existing studies suggest a possible influence of sex hormones on the observed sex differences in the gut microbial profiles101,102,103, these findings do not negate potential effects of nonhormonal factors, such as those related to sex chromosomes. Given the substantial sex divergence highlighted in our study, it is imperative to consider sex as a contributing factor in gut microbiome research, not only to deepen our understanding, but also to effectively address the specific differences.
CD38, a primary NADase in mammalian tissues, plays a pivotal role in regulating NAD+ levels104. Emerging evidence implicates the CD38–NAD axis in aging, with increased CD38 expression contributing to age-related declines in NAD+ levels, impairing mitochondrial function and cellular metabolism105,106. In the context of ovarian aging, studies have shown that CD38 overexpression accelerates follicular depletion and diminishes ovarian function, whereas CD38 inhibition preserves follicle reserves and fertility in aged mice71. Here, we observed reduced Cd38 expression in the ovaries of FMT-EF mice, together with downregulation of inflammatory markers (Fig. 3d). Functional predictions from metagenomics further supported these findings, revealing enrichment of pathways related to NAD biosynthesis in the FMT-EF group (Fig. 5g). Specifically, pathways such as NAD salvage pathway II (PNC IV cycle) and NAD de novo biosynthesis I (from aspartate) were upregulated (Fig. 5g). These results suggest that the gut microbiota established through estropausal FMT may suppress CD38 activity, influencing ovarian NAD metabolism and creating a more favorable ovarian environment. This aligns with previous research linking CD38-mediated NAD+ depletion to inflammation and cellular senescence, processes that are detrimental to ovarian function107,108.
Although the potential role of systemic NAD+ levels is of interest, NAD+ was not detected in our untargeted serum metabolomics data. This is consistent with previous reports showing that NAD+ is primarily an intracellular metabolite and typically absent from serum unless released through cell lysis artefacts109. Moreover, standard extraction and LC–MS protocols often fail to detect NAD+ in serum without specialized methods110,111. Future studies using approaches optimized for NAD+ detection in whole blood (with hematocrit correction) or examining NAD+ levels and related metabolites directly in ovarian tissue will be important to determine whether systemic NAD+ levels are altered in response to FMT.
Together, these findings indicate that modulation of the CD38-NAD pathway by gut microbiota may alleviate ovarian inflammation and promote a more youthful transcriptomic profile, contributing to enhanced ovarian health in FMT-EF mice.
One potential caveat in our chemically induced ovarian failure model is that VCD, while selectively depleting ovarian follicles, is a systemically administered agent and could affect other biological systems; however, studies have shown that VCD’s effects are confined to ovarian tissue, and impact on other organ systems have not been reported112,113. To further eliminate off-target effects, we conducted control experiments in male mice, and no significant differences in gut microbial β diversity indices (Bray–Curtis dissimilarity and Jaccard index) were observed. These findings suggest that observed microbial changes in female mice are most likely driven by ovarian dysfunction rather than direct effects of VCD. To further assess whether metabolic changes might explain the observed microbial differences, we analyzed body weight changes from before VCD treatment to euthanasia. We found no significant differences in percent weight change between control and VCD-treated animals in either female or male mice (P = 1.0 for both sexes). While we did not perform detailed metabolic profiling, the lack of significant weight change in either sex suggests that the microbiota shifts observed in female are unlikely to be explained by body weight alone.
It is possible that the improved fertility outcomes in the FMT-EF group, compared to the FMT-YF group, may reflect differences in how aged and young donor microbiota recolonize the gut following antibiotics treatment. For example, the aged microbiota might better adapt to recolonization, reducing the likelihood of artificially poor fertility outcomes in the recipients. As germ-free mice have been shown to exhibit reduced fertility114, differences in recolonization dynamics are an important factor to consider when interpreting these results. We note that the difference in median time to first litter between FMT groups was modest. Thus, future studies with larger cohorts or older animals will be important to confirm these observations (including once fertility starts to wane with age) and better define the contribution of donor microbiota to fertility outcomes. Nonetheless, FMT-driven microbiota changes resulted in significant changes in ovarian health, detected in transcriptomic profiles, fertility outcomes, and health indices in FMT-YF versus FMT-EF mice. A recent study has also reported that the gut microbiota can modulate ovarian function, further supporting the validity of our findings115.
Another limitation involves the use of different C57BL/6 strains across experiments (C57BL/6J and C57BL/6N), which are known to differ genetically, including in the Nnt gene116. The Nnt mutation in the C57BL/6J strain may influence mitochondrial redox biology and age-related metabolic traits116. While all within-experiment comparisons were conducted using the same strain to maintain internal consistency, this difference should be considered when interpreting cross-model comparisons and generalizing findings. Finally, another limitation involves the use of PICRUSt2, which predicts functional abundances based on marker gene sequences. While PICRUSt2 provides insights into potential functional pathways affected by microbiota changes, it is a predictive tool and does not directly measure functional abundances. Similarly, HUMAnN3 relies on alignment to known reference genomes and protein databases to estimate functional profiles from metagenomic data. As a reference-dependent tool, HUMAnN3 can only detect gene families and pathways represented in existing databases, potentially missing new or divergent sample-specific functions. Future studies will require additional assays to validate and expand upon our predictions.
In summary, we performed a comparative analysis of microbial profiles among aging female and male mice, and a chemically induced menopause model mice to characterize microbial genera that are unique to general aging versus ovarian aging. Our work highlights potential microbial species and genera, and functional pathways that may play key roles in modulating ovarian function.
Methods
Mouse husbandry
All animals were treated in accordance with the Guide for Care and Use of Laboratory Animals and all animals were housed under specific-pathogen-free conditions. All experimental procedures were approved by the USC’s Institutional Animal Care and Use Committee (IACUC) and are in accordance with institutional and national guidelines. Samples were derived from animals on approved IACUC protocol number 20770 and 21212.
For the ‘aging cohort,’ female and male C57BL/6JNia mice (4- and 20-month-old animals) were obtained from the National Institute on Aging (NIA) colony at Charles River Laboratories. For the ‘VCD cohort’ and ‘fecal microbiota transplantation (FMT) cohort,’ female and male C57BL/6NTac mice (3.5-month-old animals) were ordered from Taconic Biosciences. All animals were acclimated at the specific-pathogen-free animal facility at USC for 2 weeks before any processing. All animals were fed PicoLab Rodent Diet 20 (LabDiet, 5053). The facility is on a 12-h light–dark cycle and animal housing rooms are maintained at 72 °F and 30% humidity. All animals were killed between 8:00 and 11:00, by CO2 asphyxiation followed by cervical dislocation.
For the ‘VCD cohort,’ animals were subjected to daily intraperitoneal injections of either vehicle (safflower oil) or VCD (at a dosage of 160 mg kg−1 day−1) for 15 consecutive days. All injections were performed between 8:00 and 10:00 to minimize variability due to circadian effects. Lethality was observed in male VCD-injected mice (four out of ten males in the experimental group within 16 days of the start of injections). Of note all lethality cases occurred in males that had to be single-housed before injections due to fighting and wounds. Effects may have been aggravated due to lack of social interactions and/or temperature regulation.
Ovarian follicle counts
For each animal, the right or left ovary was randomly assigned for histological analysis. Ovaries were fixed in Bouin’s solution (Sigma, HT10132) for 24 h at room temperature and transferred to 70% ethanol for storage. Paraffin embedding, tissue sectioning and staining with hematoxylin and eosin (H&E) were performed by the USC Norris Comprehensive Cancer Center Translational Pathology Core Facility. H&E staining slides were imaged on the Keyence BZ-X All-in-One Fluorescence Microscope platform.
Follicles were classified based on established morphological criteria117,118,119,120,121. Primordial follicles contained an oocyte surrounded by a partial or complete layer of squamous granulosa cells. Primary follicles displayed a single layer of cuboidal granulosa cells, while intermediate follicles exhibiting both squamous and cuboidal cells were classified as primary if cuboidal cells predominated. Secondary follicles contained more than one layer of granulosa cells without visible follicular fluid. Antral follicles contained one or more cavities of follicular fluid. Corpus lutea were identified as highly vascularized structures composed of large, pale-staining granulosa lutein cells centrally and smaller, darker-staining theca lutein cells peripherally. A representative H&E image illustrating the classification of each follicle stage is provided in Extended Data Fig. 1b.
Three sections per ovary were used to count the number of primordial, primary, secondary and antral follicles and corpus luteum by three blinded observers. Median counts across the three observers were used for data analysis. Statistical significance between groups was assessed using the nonparametric Wilcoxon rank-sum test.
Quantification of serum AMH, FSH and Inhibin A concentrations
After killing, blood was collected directly from the heart. Blood was allowed to clot at room temperature for 1 h, and serum was separated using MiniCollect Serum Tube (Greiner, 450472), then stored at −80 °C until further processing. Quantification of serum AMH (Rat and Mouse Anti-Müllerian hormone (AMH) ELISA kit, Ansh Labs, AL-113), FSH (Millipore Pituitary Panel Multiplex kit, RPT86K and Ultra-Sensitive Mouse & Rat FSH, UVA Ligand Core, in-house122), and Inhibin A (Inhibin A ELISA kit, Ansh Labs, AL-161) levels was performed by the University of Virginia Center for Research in Reproduction Ligand Assay and Analysis Core. All serum hormone measurements were performed in a blinded manner. Standard normalized values were provided by the core. Statistical significance between groups was assessed using the nonparametric Wilcoxon rank-sum test.
For FSH quantification, two different kits were used during the course of the study, necessitating correction of the dataset generated by the ultra-sensitive kit. To ensure consistency, we generated data using both kits with the same serum samples and devised a polynomial model to fit the relationship between the two datasets (R2 = 0.90). The trained polynomial model was then applied to correct the data generated by the Ultra-Sensitive Mouse & Rat FSH assay. This correction process was specifically applied to the data for the FMT cohort. The raw data used to train the model have been uploaded to GitHub (https://github.com/BenayounLaboratory/Ovarian_Aging_Microbiome).
Ovarian health index calculation
The ovarian health index was calculated by integrating two key components: (1) ovarian hormone levels (AMH, FSH, and Inhibin A) and (2) follicle counts (combined counts of primordial, primary, secondary, antral follicles and corpus luteum). A three-tier scoring system was used for each parameter. Values beyond the young medians were assigned a score of 3, values between the young and estropausal medians were assigned a score of 2, and values beyond the estropausal medians were assigned a score of 1. The average hormone score (calculated from the AMH, FSH and Inhibin A scores) was then combined with the follicle score in a 1:1 ratio to generate the overall ovarian health index. This combined score was subsequently scaled to a 0–100 range to improve interpretability and enable intuitive comparisons across samples and experimental groups.
Statistical significance between groups was assessed using the Wilcoxon rank-sum test. Scripts used to calculate the ovarian health index are available on the Benayoun laboratory GitHub at https://github.com/BenayounLaboratory/Ovarian_Aging_Microbiome.
DNA extraction for 16S rRNA amplicon and shotgun metagenomics analyses
Fecal matter for 16S rRNA V3–V4 amplicon and shotgun metagenomics analyses were collected aseptically after killing from the colon. Samples were immediately snap-frozen in sterile tubes and stored at −80 °C. DNA extraction was performed using QIAamp Fast DNA Stool Mini kit (QIAGEN, 51604), following the manufacturer’s protocol.
V3–V4 region-targeted amplification and sequencing were performed by Novogene. Amplification primers used were CCTAYGGGRBGCASCAG (FWD_341F) and GGACTACNVGGGTWTCTAAT (REV_806R). Amplicon libraries were prepared using an in-house protocol and sequenced on an Illumina NovaSeq 6000 platform. Shotgun metagenomics library preparation and sequencing were performed by Novogene on the Illumina NovaSeq X machine. NEBNext Ultra II DNA Library Prep Kit-PCR-free (New England Biolabs, E7410) was used for library preparation. De-multiplexed FASTQ files were provided by Novogene for data analysis.
16S rRNA V3–V4 amplicon sequencing data analysis
Data pre-processing, quality control and denoising
Data pre-processing, quality control and denoising were performed within the QIIME2 (v.2023.7)123 platform. FASTQ files were imported to QIIME2 and denoised using DADA2 (ref. 124). Forward and reverse reads were truncated at 226 bp and 224 bp, respectively, to obtain reads with quality scores above 25 and to ensure coverage of the V3–V4 region (–p-trunc-len-f 226 –p-trunc-len-r 224).
Batch correction and PCA of amplicon sequence variants
Previous studies have shown that effectiveness of batch correction tools is dataset dependent41. For batch correction, a previously described benchmarking pipeline41 was used to select the most effective batch correction tool for our dataset. The removeBatchEffect function from limma125 (v.3.50.1), Combat from sva126 (v.3.42.0) and FAbatch from bapred127 (v.1.1) were benchmarked. Denoised feature tables were imported to R using qiime2R package (v.0.99.6; https://github.com/jbisanz/qiime2R). Then, features with low counts (<0.01% of all the counts) were removed and filtered counts were CLR-transformed. Based on PCA results (described below) and variance calculations, FA-batch-corrected values were chosen for downstream analyses.
PCA of amplicon sequence variants
For PCA, the pca function from mixOmics128 (v.6.18.1) was used in R. For female and male aging cohorts, PCA was performed on FA-batch-corrected values following CLR transformation. For VCD and FMT cohorts, CLR-transformed values were used for PCA. PCA was applied to CLR-transformed data, which were not suitable for dissimilarity-based analyses such as Bray–Curtis due to the loss of proportional structure post-transformation. To note, FA-batch correction could be not applied to the data from the VCD and FMT cohorts, as they were obtained from single experimental batches.
β and α diversity analyses
For β and α diversity analyses, phylogenetic trees were generated using the ‘qiime phylogeny align-to-tree-mafft-fasttree’ command. Rooted phylogenetic trees were used to run the ‘qiime diversity core-metrics-phylogenetic’ command for β and α diversity analyses. Diversity analysis results were imported to R using qiime2R package (v.0.99.6; https://github.com/jbisanz/qiime2R) for further plot generation and significance tests. Bray–Curtis dissimilarity and Jaccard index principal-coordinate analysis results were used to generate β diversity plots using ggplot2 (v.3.4.2). To ensure comparability across different cohorts, we normalized α diversity metrics within each cohort by dividing the values of each sample by the median value of the corresponding metric (for example, observed features and Shannon entropy) within that cohort (for example, CTL versus VCD and FMT-YF versus FMT-EF). The statistical significance of differences in α diversity between groups was determined using the Wilcoxon rank-sum test.
Differential abundance analysis of microbial genera
Differential abundance analysis of microbial genera was performed using the ALDEx2 (ref. 129) plugin within the QIIME2 (ref. 123) platform. The taxonomic classifier was generated using the SILVA130 (132 release) reference. Reference reads were extracted for CCTAYGGGRBGCASCAG (FWD_341F) and GGACTACNVGGGTWTCTAAT (REV_806R) primer set and a classifier was trained using the ‘qiime feature-classifier extract-reads’ and ‘qiime feature-classifier fit-classifier-naive-bayes’ commands, respectively. Differential abundance analysis results and taxonomic classifier were imported to R using qiime2R (https://github.com/jbisanz/qiime2R) for comparative analysis and plot generation. For female and male aging cohorts, differential abundance analysis results for each cohort for each sex was combined based on the directionality of the differential abundance measures, identifying features that consistently exhibited either an increase or a decrease across all cohorts for each sex. Using Fisher’s method and Benjamini–Hochberg correction, P values for each feature’s differential abundance were combined and corrected, respectively. Additionally, the mean of effect size across cohorts for each feature was used for analysis. For combined female and aging cohorts, as well as VCD and FMT cohorts, features with an adjusted P value < 0.05 and an absolute effect size >1 were identified as significantly different in abundance.
Functional abundance prediction analysis
Functional abundance prediction analysis was performed via the PICRUSt2 (ref. 48) plugin within the QIIME2 (ref. 123) platform using default parameters. PICRUSt2 results were imported to R using qiime2R. For differential abundance and statistical significance analyses, ALDEx2 (ref. 129) was used (500 Monte Carlo samples, Wilcoxon test for statistical testing, effect size calculation enabled and the interquartile log ratio (iqlr) for denominator calculation). To combine independent cohorts for female and male aging cohorts, Fisher’s method and Benjamini–Hochberg multiple testing correction were used to combine and correct the P values. Mean values of effect sizes were calculated for each feature. Adjusted P values < 0.05 and absolute effect size >1 were used as criteria to filter functional pathways that were significantly different in abundance.
For β-glucuronidase activity analysis, functional abundance data corresponding to EC:3.2.1.31 was extracted from PICRUSt2 output. To ensure comparability across different cohorts, β-glucuronidase abundance values for each sample were normalized by dividing by the median value of the corresponding controls within that cohort (young females of aging cohort, vehicle controls of VCD cohort and FMT-YF of FMT cohort). The Wilcoxon rank-sum test was then used to assess statistical significance.
Shotgun metagenomics analysis
Data quality control and pre-processing
Paired-end 150-bp reads were pre-processed to remove sequencing adapters and low-quality reads using Trimmomatic (v.0.39)131. Trimmed reads were then mapped to the mouse reference genome (mm10) using Bowtie2 (v.2.4.4)132 to remove host DNA contamination. Unmapped reads from paired-end files were extracted using the –un-conc-gz option in Bowtie2 and used for subsequent analyses. Quality-controlled and host-depleted reads were then processed for taxonomic classification and functional profiling.
Taxonomic classification and abundance estimation
Kraken2 (v.2.1.2)133 was used for taxonomic classification of unmapped reads. Reads were classified against a pre-built standard Kraken2 database containing bacterial, archaeal and viral sequences. Classification outputs were summarized using Bracken (v.2.7)134 to refine species- and genus-level abundance estimates. Bracken was configured with a read length of 150 bp and default settings.
α and β diversity and differential abundance analysis
Raw taxonomic abundance tables from Bracken (v.2.7)134 were loaded in R. Features with total counts <20, relative abundance <0.01% or prevalence in fewer than 10% of samples were excluded. Samples with high host DNA alignment rates (MHK271) were also removed. CLR transformation of filtered relative abundances was performed. Shannon entropy was calculated using the mia (v.1.12.0; https://github.com/microbiome/mia) R package. Relative abundance data were analyzed using mixOmics128 (v.6.28.0) and Bray–Curtis and Jaccard distance matrices were calculated using vegan (v.2.6-8). Differential abundance of species was assessed using ALDEx2 (v.1.36.0)129. Features with adjusted P values < 0.05 were deemed significant.
Functional pathway analysis
Functional profiling was conducted using HUMAnN3 (v.3.0)76. Reads were processed in paired-end mode. Raw counts were loaded in R and low-abundance pathways with total abundance <0.001 across all samples were removed. Differential abundance of microbial pathways between groups was analyzed using ANCOMBC2 (v.2.0.0)135. Significant pathways with q-values <0.1 were identified and log2 fold change values were used to assess the direction and magnitude of differential abundance.
For β-glucuronidase activity analysis, relative abundances corresponding to EC:3.2.1.31 were extracted from HUMAnN3 output. Values for each sample were normalized by dividing by the median value of the FMT-YF group within each cohort. Statistical significance between groups was assessed using the Wilcoxon rank-sum test.
Fecal microbiota transplantation
For FMT, animals underwent a 10-day course of antibiotics/antimycotic (Ab/Am) treatment before transplantation. The regimen included ampicillin in drinking water at 1 g l−1 (Sandoz, 00781) and oral gavage of amphotericin-B (Sigma, A9528) at 1 mg kg−1 day−1, metronidazole (MFR VIONA PHARMACEUTICALS, 096988) at 100 mg kg−1 day−1, vancomycin (Alvogen, Vet-Rx-MW 090160) at 50 mg kg−1 day−1 and neomycin (MFR VETONE, 510570) at 100 mg kg−1 day−1. Starting on the third day after the end of the Ab/Am treatment, FMT was performed biweekly for 3 weeks via oral gavage. Donor stool samples were collected from separate cohorts of more than 30 young or estropausal female mice that were not included in the microbial profiling analyses. Samples were pooled and prepared by suspending in saline (20 mg ml−1), followed by vortexing and centrifugation to obtain the supernatant. Each recipient animal received 100 µl of the fecal suspension. Two independent cohorts were enrolled in the study, each using independently prepared pools of FMT materials. Sample collection for subsequent processing and analysis occurred 3 days after the final FMT session.
Ovarian RNA-seq sample preparation
Ovaries were collected and immediately snap-frozen, and stored at −80 °C. For RNA extraction, 600 µl of TRIzol reagent (Ambion, 15596018) was added to the ovaries in Lysing Matrix D tubes (MP, 6913500) and tissues were homogenized with a BeadBug 6 microtube homogenizer (Benchmark Scientific, D1036) at 3,500 rpm for 30 s for three cycles in the cold room. Total RNA was purified using the Direct-Zol RNA Miniprep (Zymo Research, R2052), following the manufacturer’s instructions. Integrity and quality of the purified total RNA were assessed on the 4200 TapeStation system (Agilent, G2991A) using High Sensitivity RNA ScreenTapes (Agilent, 5067–5579). Subsequently, mRNA-seq libraries were prepared and sequenced by Novogene to obtain PE150 sequencing reads on the NovaSeq X Plus platform.
FMT cohort ovarian RNA-seq data analysis
Data quality control and pre-processing
Paired-end 150-bp reads were hard-trimmed to remove the first six bases and keep up to 100 bp using Fastx Trimmer (v.0.0.14; http://hannonlab.cshl.edu/fastx_toolkit/) and processed using Trim Galore (v.0.6.6; https://github.com/FelixKrueger/TrimGalore) using default parameters. Trimmed reads were mapped to the mm39 genome reference using STAR136 (v.2.7.9a). Read counts were assigned to genes from the mm39 genome reference using subread137 (v.2.0.3). After assigning read counts to genes, downstream processing was performed in R (v.4.1.2). Low-count genes (genes with <1 count in at least 6 out of 14 RNA-seq libraries) were removed as recommended by DESeq2 (ref. 138) developers. R package sva126 (v.3.42.0) was used to estimate surrogate variables and the removeBatchEffect function from limma125 (v.3.50.1) was used to regress out the effects of surrogate variables and RNA integrity differences from raw read counts. To remove extreme values within the read counts, Winsorize function from DescTools (v.0.99.50; https://andrisignorell.github.io/DescTools/) was used to adjust the values above the 99.9th percentile to the values at the 99.9th percentile. DESeq2 (ref. 138) package (v.1.34.0) was used for downstream analyses of the RNA-seq data. Genes with FDR < 5% were considered statistically significant. Normalized counts from DESeq2 were used as input for PCA. PCA was performed using the prcomp function in R. For the MDS analysis, we calculated the distance between samples using the inverse of Spearman’s rank correlation coefficient (1 − Rho) as the distance metric. These distances were then input into the cmdscale function in R to compute the MDS coordinates.
Gene Ontology analysis of RNA-seq dataset
We used the GSEA55 approach through the phenoTest package. Gene Ontology (GO) term annotations (ENS.GO.BP, ENS.GO.MF, ENS.GO.CC and GTRD gene sets) were sourced from ENSEMBL (Ensembl 108) and Molecular Signature Database139. The t-statistic from DESeq2 (ref. 138) was used to rank genes for functional enrichment analysis. For readability, only the top ten most-significant pathways with negative normalized enrichment score (NES) and top ten most-significant pathways with positive NES are shown in figures if more than that passed the FDR < 5% significance threshold for GO terms and FDR < 10% for GTRD TF targets.
Deconvolution analysis of immune cell type proportion
To estimate any differences in the immune cell proportions between FMT-YF and FMT-EF groups of the FMT cohort, single-cell RNA-seq reference-based deconvolution analyses were performed, using CSCDRNA140 and Granulator (https://github.com/Novartis/granulator). We downloaded the cell type annotated Seurat object of a publicly available murine ovarian single-cell RNA-seq dataset (Open Science Framework dataset ID 924fz)56. In the CSCDRNA analysis, the min.p parameter was set to 0.3 following the authors’ recommendation140, to efficiently exclude the 30% least significant portion of marker genes for each cell type. This threshold was chosen to increase efficiency and reduce noise. For Granulator, the qprogwc method141 was used with default parameters to estimate the immune cell proportions within the ovarian bulk RNA-seq dataset. Cell proportions were tested for significance using a Wilcoxon rank-sum test.
Pre-processing and filtering candidate transcription factor-bound genes
For potential TFs identified from GSEA55 analysis against GTRD63 TF targets, peak information was obtained for Cistrome.org. Peak annotation was performed using HOMER142 (v.4.11.1). Genes with peaks annotated in the transcription start site promoter of protein-coding genes were considered as direct target genes. Obtained gene lists were used to perform GSEA through the phenoTest (v.1.42.0) package.
Pre-processing of public ovarian aging datasets
To evaluate the correlation between ovarian aging and FMT gene expression profiles of the ovaries, we pre-processed and analyzed two publicly available ovarian aging datasets (2–3-month-old versus 12-month-old female mice; BioProject dataset PRJNA100222 and Genome Sequence Archive dataset CRA003645)70,71. For each dataset, reads were pre-processed and aligned to the mm39 genome reference using the same methods as for the FMT cohort ovarian RNA-seq dataset. Low-count genes (genes with <1 count in at least four of seven RNA-seq libraries) were removed. sva126 (v.3.42.0) and removeBatchEffect function from limma125 (v.3.50.1) were used to estimate surrogate variables and regress out the effects of surrogate variables, respectively. DESeq2 (ref. 138) package (v.1.34.0) was used for downstream analyses. P values for common genes between the two datasets were combined and corrected using Fisher’s method and Benjamini–Hochberg correction method, respectively. FDR < 1 × 10−5 was used to catalog gene sets that are robustly up- or downregulated with aging. Obtained gene lists were used to perform GSEA through the phenoTest package.
Ovarian tissue IL-6 immunohistochemistry and analysis
The same ovarian tissues used for H&E staining were processed for IL-6 immunohistochemistry. Tissues were fixed in Bouin’s solution (Sigma, HT10132) for 24 h at room temperature and transferred to 70% ethanol for storage. Paraffin embedding and sectioning were performed by the USC Norris Comprehensive Cancer Center Translational Pathology Core Facility. IL-6 immunostaining (Abcam, ab6672) with hematoxylin counterstaining and diaminobenzidine (DAB) detection was conducted by the USC Immunohistochemistry Laboratory.
Slides were imaged on a Keyence BZ-X All-in-One Fluorescence Microscope. For each ovary, three sections were analyzed. Image analysis was performed in ImageJ2 (v.2.14.0/1.54f) using the following workflow: images were color-deconvoluted with the H-DAB vector, hematoxylin (colour_1) and DAB (colour_2) signals were thresholded at 230 and 210, respectively, and the positive area was measured. The percentage of DAB signal was calculated as the DAB-positive area divided by the hematoxylin-positive area. All measurements were performed in a blinded manner. Mean values across the three sections per ovary were used for downstream analysis. Statistical significance between groups was assessed using the nonparametric Wilcoxon rank-sum test.
Fertility assay
Fertility assay was performed as previously described73. FMT female mice were randomly paired with 4-month-old males at a 1:1 ratio 3 days after the last day of FMT administration until the birth of the first litter. The number of pups in the first litter and the latency, defined as the duration between the pairing date and the birth date of the first litter, were recorded and analyzed. Statistical difference in latency in the first litter after pairing was evaluated using a log-rank test. For pairs that did not produce pups within the 60-day timeframe, the latency was recorded as 60 days. The effect size was reported as the percent difference in median time between groups, calculated as the difference in median time divided by the average of the two medians.
Stool menaquinone concentration quantification
Frozen stool samples were shipped to the Vitamin K Laboratory at the Tufts University Human Nutrition Research Center on Aging, where stool menaquinone concentrations were measured using LC–MS, as detailed elsewhere143. Measurements were performed in a blinded manner.
Serum untargeted metabolomics
Sample processing
Serum samples were stored at −80 °C until they were thawed on ice for analysis. Metabolites were extracted by adding 80 µl of ice-cold methanol to 20 µl of serum, followed by incubation on ice with vortexing every 5 min. The samples were then centrifuged at 21,300g for 10 min at 4 °C.
Liquid chromatography–tandem mass spectrometry
Untargeted metabolomics was performed at the USC Alfred E. Mann Multi-Omics Mass Spectrometry Core. A volume of 10 µl from each sample was injected onto a SeQuant ZIC-pHILIC Column (5 µm, 2.1 × 150 mm; Millapore Sigma) using a Vanquish Flex UHPLC System (Thermo Fisher Scientific). Chromatography was conducted at a flow rate of 100 µl min−1 with an optimized gradient of solvent A (water/20 mM ammonium carbonate, pH 9.8) and solvent B (acetonitrile) as follows: 0 min/97% B, 1 min/97% B, 28 min/75% B, 39 min/50% B, 44 min/20% B, 45 min/20% B, 45 min/97% B and 60 min/97% B. The chromatography system was coupled to a heated electrospray ionization source and a hybrid quadrupole-orbitrap mass spectrometer (Q Exactive, Thermo Fisher Scientific). Mass spectra were acquired in full MS/data-dependent MS2 mode, alternating between positive and negative ion polarities.
Source parameters were set as follows: a source voltage of 3.5 kV, source temperature of 275 °C, sheath gas of 45, auxiliary gas of 10, sweep gas of 2 and capillary temperature of 320 °C. Full scans were performed across an m/z range of 70–1,050 with the following settings: an automated gain control (AGC) target of 5 × 106, a maximum injection time of 100 ms, and a resolution of 70,000 at m/z 200. Up to five consecutive MS/MS scans were acquired per full scan with the following parameters: quadrupole isolation of charge states 1–7; isolation window of 0.6 m/z; normalized collision energies of 20, 30 and 40; dynamic exclusion of 30 s; AGC target of 1 × 105; maximum injection time of 100 ms; and resolution of 17,500 at m/z 200.
Data analysis
Raw data were processed using Compound Discoverer (v.3.3 SP3, Thermo Fisher Scientific) for metabolite identification, retention time alignment and unknown compound detection across all samples. Metabolites were identified by matching exact masses (Δ ppm of >10 or <−10) and fragmentation spectra to the mzCloud database (ddMS2; ‘mzCloud Best Match’ >50). For quality control, only ions with a peak rating >4 in at least six samples within a group were retained. A variance stabilization normalization (VSN) normalized filtered metabolic matrix via normalizeVSN function of limma125 (v.3.50.1) was used for PCA. The filtered metabolic matrix was then scaled and LDA was performed via the MASS (v.7.3-60.2) R package. Metabolites with a consistent direction of log2 fold change between two batches were subsequently retained for abundance analysis.
Causal mediation analysis
Causal mediation analysis to identify specific features and ASVs from shotgun metagenomics and 16S rRNA amplicon sequencing datasets from the FMT cohort acting as mediators between the treatment (FMT-YF versus FMT-EF) and outcome (changes in the ovarian transcriptome profiles) was performed using mediation package144 (v.4.5.0) in R. Feature table, taxonomic reference and differential abundance analysis results generated from QIIME2 (ref. 123) were imported to R using qiime2R (https://github.com/jbisanz/qiime2R). ASVs identified as significant in differential abundance analyses were tested. CLR-transformed ASV counts were used for the mediation analyses. For the outcome variable, the first dimension from the MDS analysis of the FMT ovarian mRNA-seq dataset was used. Linear models were constructed for both the mediator and the outcome, incorporating the treatment as an independent variable in each. The mediate function from the mediation package was then employed to estimate the average causal mediation effects, average direct effects and total effects.
Statistics and reproducibility
No statistical method was used to predetermine sample size. Sample sizes were chosen based on previous studies and practical considerations. Animals were randomly assigned to experimental groups when feasible. Blinding was used for outcome assessment where feasible. Follicle counting was performed by three blinded observers. IL-6 immunohistochemistry and serum hormone quantifications were performed in a blinded manner. All statistical analyses were performed in R (v.4.2.4). Unless otherwise indicated, comparisons between two groups were performed using the nonparametric Wilcoxon rank-sum test. For fertility latency analyses, statistical differences were evaluated using a log-rank test, with nonbreeding pairs censored at 60 days. Data are presented as boxplots showing the median, 25th and 75th percentiles, with whiskers representing 1.5× the interquartile range. Individual datapoints are overlaid when feasible and are shown in the corresponding figures. Potential outliers were assessed using Grubbs’ test. Values with a test P value < 0.05 were excluded from subsequent analyses; any exclusions were noted in the corresponding figure legends. Experiments were performed with independent biological samples as described for each cohort, and key findings were reproduced across independent cohorts when applicable. Specific statistical tests, exact sample numbers and the unit of replication are reported in the relevant figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw FASTQ files have been deposited to the Sequence Read Archive under accession PRJNA1076509. Raw microscopy images of ovarian H&E staining, and hematoxylin and diaminobenzidine staining in immunohistochemistry (IL-6) are available on Figshare (https://doi.org/10.6084/m9.figshare.25749588 (ref. 145), https://doi.org/10.6084/m9.figshare.25749594 (ref. 146), https://doi.org/10.6084/m9.figshare.25749591 (ref. 147), https://doi.org/10.6084/m9.figshare.28462274 (ref. 148) and https://doi.org/10.6084/m9.figshare.29882735 (ref. 149)). Processed serum metabolomics data can be found in Supplementary Table 3. Raw data can be found on Metabolomics Workbench, project ST003750. All R code was run using R v.4.1.2. Any additional information required to reanalyze the data reported in this work paper is available from the corresponding author upon request. Source Data are provided with this paper.
Code availability
All scripts used to analyze the datasets are available on the Benayoun laboratory GitHub at https://github.com/BenayounLaboratory/Ovarian_Aging_Microbiome.
References
Davis, S. R. et al. Menopause. Nat. Rev. Dis. Primers 1, 15004 (2015).
Broekmans, F. J., Soules, M. R. & Fauser, B. C. Ovarian aging: mechanisms and clinical consequences. Endocr. Rev. 30, 465–493 (2009).
Faddy, M. J. & Gosden, R. G. A model conforming the decline in follicle numbers to the age of menopause in women. Hum. Reprod. 11, 1484–1486 (1996).
Fu, C. et al. Association of reproductive factors with dementia: a systematic review and dose-response meta-analyses of observational studies. eClinicalMedicine 43, 101236 (2022).
Shuster, L. T., Rhodes, D. J., Gostout, B. S., Grossardt, B. R. & Rocca, W. A. Premature menopause or early menopause: long-term health consequences. Maturitas 65, 161–166 (2010).
Muka, T. et al. Association of age at onset of menopause and time since onset of menopause with cardiovascular outcomes, intermediate vascular traits, and all-cause mortality: a systematic review and meta-analysis. JAMA Cardiol. 1, 767–776 (2016).
Ossewaarde, M. E. et al. Age at menopause, cause-specific mortality and total life expectancy. Epidemiology 16, 556–562 (2005).
Hong, J. S. et al. Age at menopause and cause-specific mortality in South Korean women: Kangwha Cohort Study. Maturitas 56, 411–419 (2007).
Knobil, E. The neuroendocrine control of the menstrual cycle. Recent Prog. Horm. Res. 36, 53–88 (1980).
Santoro, N., Roeca, C., Peters, B. A. & Neal-Perry, G. The menopause transition: signs, symptoms, and management options. J. Clin. Endocrinol. Metab. 106, 1–15 (2021).
Randolph, J. F. Jr. et al. Change in follicle-stimulating hormone and estradiol across the menopausal transition: effect of age at the final menstrual period. J. Clin. Endocrinol. Metab. 96, 746–754 (2011).
Kelsey, T. W., Wright, P., Nelson, S. M., Anderson, R. A. & Wallace, W. H. A validated model of serum anti-Mullerian hormone from conception to menopause. PLoS ONE 6, e22024 (2011).
Weiss, G., Skurnick, J. H., Goldsmith, L. T., Santoro, N. F. & Park, S. J. Menopause and hypothalamic-pituitary sensitivity to estrogen. JAMA 292, 2991–2996 (2004).
Xiong, J. et al. FSH blockade improves cognition in mice with Alzheimer’s disease. Nature 603, 470–476 (2022).
Mao, L., Wang, L., Bennett, S., Xu, J. & Zou, J. Effects of follicle-stimulating hormone on fat metabolism and cognitive impairment in women during menopause. Front. Physiol. 13, 1043237 (2022).
Xiong, J. et al. FSH and ApoE4 contribute to Alzheimer’s disease-like pathogenesis via C/EBPβ/δ-secretase in female mice. Nat. Commun. 14, 6577 (2023).
Luckey, T. D. Introduction to intestinal microecology. Am. J. Clin. Nutr. 25, 1292–1294 (1972).
Savage, D. C. Microbial ecology of the gastrointestinal tract. Annu. Rev. Microbiol. 31, 107–133 (1977).
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Biagi, E. et al. Ageing and gut microbes: perspectives for health maintenance and longevity. Pharmacol. Res. 69, 11–20 (2013).
Zapata, H. J. & Quagliarello, V. J. The microbiota and microbiome in aging: potential implications in health and age-related diseases. J. Am. Geriatr. Soc. 63, 776–781 (2015).
Parker, A. et al. Fecal microbiota transfer between young and aged mice reverses hallmarks of the aging gut, eye, and brain. Microbiome 10, 68 (2022).
Kundu, P. et al. Neurogenesis and prolongevity signaling in young germ-free mice transplanted with the gut microbiota of old mice. Sci. Transl. Med. 11, eaau4760 (2019).
Wu, J. et al. Association between premature ovarian insufficiency and gut microbiota. BMC Pregnancy Childbirth 21, 418 (2021).
Qi, X. et al. Gut microbiota-bile acid-interleukin-22 axis orchestrates polycystic ovary syndrome. Nat. Med. 25, 1225–1233 (2019).
Weger, B. D. et al. The mouse microbiome is required for sex-specific diurnal rhythms of gene expression and metabolism. Cell Metab. 29, 362–382.e368 (2019).
Lombardi, P., Goldin, B., Boutin, E. & Gorbach, S. L. Metabolism of androgens and estrogens by human fecal microorganisms. J. Steroid Biochem. 9, 795–801 (1978).
Adlercreutz, H., Martin, F., Jarvenpaa, P. & Fotsis, T. Steroid absorption and enterohepatic recycling. Contraception 20, 201–223 (1979).
Eriksson, H. & Gustafsson, J. A. Steroids in germfree and conventional rats. Distribution and excretion of labelled pregnenolone and corticosterone in male and female rats. Eur. J. Biochem. 15, 132–139 (1970).
Finch, C. E. The menopause and aging, a comparative perspective. J. Steroid Biochem. Mol. Biol. 142, 132–141 (2014).
Whitehead, J. C. et al. A clinical frailty index in aging mice: comparisons with frailty index data in humans. J. Gerontol. A Biol. Sci. Med. Sci. 69, 621–632 (2014).
Wallace, W. H. & Kelsey, T. W. Human ovarian reserve from conception to the menopause. PLoS ONE 5, e8772 (2010).
Hirshfield, A. N. Development of follicles in the mammalian ovary. Int. Rev. Cytol. 124, 43–101 (1991).
Tehrani, F. R., Mansournia, M. A., Solaymani-Dodaran, M. & Azizi, F. Age-specific serum anti-Mullerian hormone levels: estimates from a large population-based sample. Climacteric 17, 591–597 (2014).
Broer, S. L., Broekmans, F. J., Laven, J. S. & Fauser, B. C. Anti-Mullerian hormone: ovarian reserve testing and its potential clinical implications. Hum. Reprod. Update 20, 688–701 (2014).
Felicio, L. S., Nelson, J. F. & Finch, C. E. Prolongation and cessation of estrous cycles in aging C57BL/6J mice are differentially regulated events. Biol. Reprod. 34, 849–858 (1986).
Johnson, J. S. et al. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 10, 5029 (2019).
Gupta, S. et al. Amplicon sequencing provides more accurate microbiome information in healthy children compared to culturing. Commun. Biol. 2, 291 (2019).
Knight, R. et al. Best practices for analysing microbiomes. Nat. Rev. Microbiol. 16, 410–422 (2018).
Willis, A. D. Rarefaction, α diversity, and statistics. Front. Microbiol. 10, 2407 (2019).
Wang, Y. & LeCao, K. A. Managing batch effects in microbiome data. Brief. Bioinform. 21, 1954–1970 (2020).
Gamez-Macias, P. E. et al. Intestinal permeability, gut inflammation, and gut immune system response are linked to aging-related changes in gut microbiota composition: a study in female mice. J. Gerontol. A Biol. Sci. Med. Sci. 79, glae045 (2024).
de la Cuesta-Zuluaga, J. et al. Age- and sex-dependent patterns of gut microbial diversity in human adults. mSystems 4, e00261-19 (2019).
Ghosh, T. S., Shanahan, F. & O’Toole, P. W. Toward an improved definition of a healthy microbiome for healthy aging. Nat. Aging 2, 1054–1069 (2022).
Luo, M. et al. E. coli Nissle 1917 ameliorates mitochondrial injury of granulosa cells in polycystic ovary syndrome through promoting gut immune factor IL-22 via gut microbiota and microbial metabolism. Front. Immunol. 14, 1137089 (2023).
Biagi, E. et al. Gut Microbiota and extreme longevity. Curr. Biol. 26, 1480–1485 (2016).
Wang, N. et al. Enriched taxa were found among the gut microbiota of centenarians in East China. PLoS ONE 14, e0222763 (2019).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Murali, R. et al. Deregulated metabolic pathways in ovarian cancer: cause and consequence. Metabolites 13, 560 (2023).
Ji, Z. et al. Deregulation of lipid metabolism: the critical factors in ovarian cancer. Front. Oncol. 10, 593017 (2020).
Borman, S. M., Christian, P. J., Sipes, I. G. & Hoyer, P. B. Ovotoxicity in female Fischer rats and B6 mice induced by low-dose exposure to three polycyclic aromatic hydrocarbons: comparison through calculation of an ovotoxic index. Toxicol. Appl. Pharmacol. 167, 191–198 (2000).
Van Kempen, T. A., Milner, T. A. & Waters, E. M. Accelerated ovarian failure: a novel, chemically induced animal model of menopause. Brain Res. 1379, 176–187 (2011).
Mayer, L. P., Devine, P. J., Dyer, C. A. & Hoyer, P. B. The follicle-deplete mouse ovary produces androgen. Biol. Reprod. 71, 130–138 (2004).
Boehme, M. et al. Microbiota from young mice counteracts selective age-associated behavioral deficits. Nat. Aging 1, 666–676 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Morris, M. E. et al. A single-cell atlas of the cycling murine ovary. eLife 11, e77239 (2022).
Jew, B. et al. Accurate estimation of cell composition in bulk expression through robust integration of single-cell information. Nat. Commun. 11, 1971 (2020).
Rea, I. M. et al. Age and age-related diseases: role of inflammation triggers and cytokines. Front. Immunol. 9, 586 (2018).
Michaud, M. et al. Proinflammatory cytokines, aging, and age-related diseases. J. Am. Med. Dir. Assoc. 14, 877–882 (2013).
Chen, Y. et al. Circulating interleukin-6 mediates PM(2.5)-induced ovarian injury by suppressing the PPARγ pathway. Research 7, 0538 (2024).
Lovasco, L. A. et al. Accelerated ovarian aging in the absence of the transcription regulator TAF4B in mice. Biol. Reprod. 82, 23–34 (2010).
Baumann, C. et al. Acute irradiation induces a senescence-like chromatin structure in mammalian oocytes. Commun. Biol. 6, 1258 (2023).
Yevshin, I., Sharipov, R., Valeev, T., Kel, A. & Kolpakov, F. GTRD: a database of transcription factor binding sites identified by ChIP-seq experiments. Nucleic Acids Res. 45, D61–D67 (2017).
Zheng, R. et al. Cistrome Data Browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res. 47, D729–D735 (2019).
Mei, S. et al. Cistrome Data Browser: a data portal for ChIP-seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 45, D658–D662 (2017).
Huang, H. Y. et al. Association of polymorphisms for nuclear receptor coactivator 1 gene with egg production traits in the maternal line of Shaobo hens. Br. Poult. Sci. 52, 328–332 (2011).
Wang, M. et al. The USP7 inhibitor P5091 induces cell death in ovarian cancers with different P53 status. Cell. Physiol. Biochem. 43, 1755–1766 (2017).
Briley, S. M. et al. Reproductive age-associated fibrosis in the stroma of the mammalian ovary. Reproduction 152, 245–260 (2016).
Zhang, Z., Schlamp, F., Huang, L., Clark, H. & Brayboy, L. Inflammaging is associated with shifted macrophage ontogeny and polarization in the aging mouse ovary. Reproduction 159, 325–337 (2020).
Zhou, Z. et al. Temporal transcriptomic landscape of postnatal mouse ovaries reveals dynamic gene signatures associated with ovarian aging. Hum. Mol. Genet. 30, 1941–1954 (2021).
Yang, Q. et al. NADase CD38 is a key determinant of ovarian aging. Nat. Aging 4, 110–128 (2024).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Mehalko, K. et al. Lack of accelerated ovarian aging in a follicle-stimulating hormone receptor haploinsufficiency model. Transl. Med. Aging 7, 1–8 (2023).
Wrzosek, L. et al. Transplantation of human microbiota into conventional mice durably reshapes the gut microbiota. Sci. Rep. 8, 6854 (2018).
Gong, X., Gutala, R. & Jaiswal, A. K. Quinone oxidoreductases and vitamin K metabolism. Vitam. Horm. 78, 85–101 (2008).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. eLife 10, e65088 (2021).
Dekkers, K. F. et al. An online atlas of human plasma metabolite signatures of gut microbiome composition. Nat. Commun. 13, 5370 (2022).
Pilley, S. E. et al. A metabolic atlas of mouse aging. Cell Metab. 38, 1–20 (2025).
Marquez Campos, E., Stehle, P. & Simon, M. C. Microbial metabolites of flavan-3-Ols and their biological activity. Nutrients 11, 2260 (2019).
He, Y. et al. Lactic acid bacteria alleviate polycystic ovarian syndrome by regulating sex hormone related gut microbiota. Food Funct. 11, 5192–5204 (2020).
Dikalov, S., Panov, A. & Dikalova, A. Critical role of mitochondrial fatty acid metabolism in normal cell function and pathological conditions. Int. J. Mol. Sci. 25, 6498 (2024).
Wang, Z. H. et al. Targeting mitochondria for ovarian aging: new insights into mechanisms and therapeutic potential. Front. Endocrinol. 15, 1417007 (2024).
Zhang, I. W. et al. Palmitoylcarnitine impairs immunity in decompensated cirrhosis. JHEP Rep. 6, 101187 (2024).
Xu, X. L. et al. Estrogen biosynthesis and signal transduction in ovarian disease. Front. Endocrinol. 13, 827032 (2022).
Stachenfeld, N. S. Hormonal changes during menopause and the impact on fluid regulation. Reprod. Sci. 21, 555–561 (2014).
Liu, T., Bowen, R. L., Wilson, A. C. & Atwood, C. S. Estropause, sex hormones and metal homeostasis in the mouse brain. Front. Neurol. 13, 841822 (2022).
Schele, E. et al. The gut microbiota reduces leptin sensitivity and the expression of the obesity-suppressing neuropeptides proglucagon (Gcg) and brain-derived neurotrophic factor (Bdnf) in the central nervous system. Endocrinology 154, 3643–3651 (2013).
Fan, K. C. et al. Altered gut microbiota in older adults with mild cognitive impairment: a case-control study. Front. Aging Neurosci. 15, 1162057 (2023).
Ellis, J. L. et al. Dietary vitamin K is remodeled by gut microbiota and influences community composition. Gut Microbes 13, 1–16 (2021).
Tarkesh, F., Namavar Jahromi, B., Hejazi, N. & Tabatabaee, H. Beneficial health effects of Menaquinone-7 on body composition, glycemic indices, lipid profile, and endocrine markers in polycystic ovary syndrome patients. Food Sci. Nutr. 8, 5612–5621 (2020).
Sibayama-Imazu, T. et al. Induction of apoptosis in PA-1 ovarian cancer cells by vitamin K2 is associated with an increase in the level of TR3/Nur77 and its accumulation in mitochondria and nuclei. J. Cancer Res. Clin. Oncol. 134, 803–812 (2008).
Harshman, S. G. & Shea, M. K. The role of vitamin K in chronic aging diseases: inflammation, cardiovascular disease, and osteoarthritis. Curr. Nutr. Rep. 5, 90–98 (2016).
Sampathkumar, N. K. et al. Widespread sex dimorphism in aging and age-related diseases. Hum. Genet. 139, 333–356 (2020).
Podcasy, J. L. & Epperson, C. N. Considering sex and gender in Alzheimer disease and other dementias. Dialogues Clin. Neurosci. 18, 437–446 (2016).
Vina, J. & Lloret, A. Why women have more Alzheimer’s disease than men: gender and mitochondrial toxicity of amyloid-beta peptide. J. Alzheimers Dis. 20, S527–S533 (2010).
Varghese, M. et al. Female adipose tissue has improved adaptability and metabolic health compared to males in aged obesity. Aging 12, 1725–1746 (2020).
Austad, S. N. & Fischer, K. E. Sex differences in lifespan. Cell Metab. 23, 1022–1033 (2016).
Lemaitre, J. F. et al. Sex differences in adult lifespan and aging rates of mortality across wild mammals. Proc. Natl Acad. Sci. USA 117, 8546–8553 (2020).
Markle, J. G. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088 (2013).
Steegenga, W. T. et al. Sexually dimorphic characteristics of the small intestine and colon of prepubescent C57BL/6 mice. Biol. Sex Differ. 5, 11 (2014).
Org, E. et al. Sex differences and hormonal effects on gut microbiota composition in mice. Gut Microbes 7, 313–322 (2016).
Vemuri, R. et al. The microgenderome revealed: sex differences in bidirectional interactions between the microbiota, hormones, immunity and disease susceptibility. Semin. Immunopathol. 41, 265–275 (2019).
Kaliannan, K. et al. Estrogen-mediated gut microbiome alterations influence sexual dimorphism in metabolic syndrome in mice. Microbiome 6, 205 (2018).
Hogan, K. A., Chini, C. C. S. & Chini, E. N. The multi-faceted ecto-enzyme CD38: roles in immunomodulation, cancer, aging, and metabolic diseases. Front. Immunol. 10, 1187 (2019).
Camacho-Pereira, J. et al. CD38 dictates age-related NAD decline and mitochondrial dysfunction through an SIRT3-dependent mechanism. Cell Metab. 23, 1127–1139 (2016).
Gomes, A. P. et al. Declining NAD(+) induces a pseudohypoxic state disrupting nuclear-mitochondrial communication during aging. Cell 155, 1624–1638 (2013).
Chini, C. C. S. et al. CD38 ecto-enzyme in immune cells is induced during aging and regulates NAD(+) and NMN levels. Nat. Metab. 2, 1284–1304 (2020).
Covarrubias, A. J. et al. Senescent cells promote tissue NAD(+) decline during ageing via the activation of CD38(+) macrophages. Nat. Metab. 2, 1265–1283 (2020).
Cuny, H., Kristianto, E., Hodson, M. P. & Dunwoodie, S. L. Simultaneous quantification of 26 NAD-related metabolites in plasma, blood, and liver tissue using UHPLC-MS/MS. Anal. Biochem. 633, 114409 (2021).
Sebastiani, P. et al. Metabolite signatures of chronological age, aging, survival, and longevity. Cell Rep. 43, 114913 (2024).
Jang, C. et al. Metabolite exchange between mammalian organs quantified in pigs. Cell Metab. 30, 594–606 e593 (2019).
Wright, L. E. et al. Comparison of skeletal effects of ovariectomy versus chemically induced ovarian failure in mice. J. Bone Miner. Res. 23, 1296–1303 (2008).
National Toxicology, P. Toxicology and carcinogenesis studies of 4-vinyl-1-cyclohexene diepoxide (CAS no. 106-87-6) in F344/N rats and B6C3F1 mice (dermal studies). Natl Toxicol. Program Tech. Rep. Ser. 362, 1–249 (1989).
Shimizu, K. et al. Normalization of reproductive function in germfree mice following bacterial contamination. Exp. Anim. 47, 151–158 (1998).
Munyoki, S. K. et al. The microbiota extends the reproductive lifespan by safeguarding the ovarian reserve. Cell Host Microbe 33, 1731–1747.e1738 (2025).
Mekada, K. et al. Genetic differences among C57BL/6 substrains. Exp. Anim. 58, 141–149 (2009).
Pedersen, T. & Peters, H. Proposal for a classification of oocytes and follicles in the mouse ovary. J. Reprod. Fertil. 17, 555–557 (1968).
Myers, M., Britt, K. L., Wreford, N. G., Ebling, F. J. & Kerr, J. B. Methods for quantifying follicular numbers within the mouse ovary. Reproduction 127, 569–580 (2004).
Winship, A. L., Sarma, U. C., Alesi, L. R. & Hutt, K. J. Accurate follicle enumeration in adult mouse ovaries. J. Vis. Exp. 164, e61782 (2020).
Mohammadghasemi, F., Jahromi, S. K., Hajizadeh, H., Homafar, M. A. & Saadat, N. The protective effects of exogenous melatonin on nicotine-induced changes in mouse ovarian follicles. J. Reprod. Infertil. 13, 143–150 (2012).
Roig, I., Brieno-Enriquez, M. A. & Caldes, M. G. Meiosis in a bottle: new approaches to overcome mammalian meiocyte study limitations. Genes 2, 152–168 (2011).
Ongaro, L. et al. Development of a highly sensitive ELISA for measurement of FSH in serum, plasma, and whole blood in mice. Endocrinology 162, bqab014 (2021).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Leek, J. T. & Storey, J. D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007).
Hornung, R., Boulesteix, A. L. & Causeur, D. Combining location-and-scale batch effect adjustment with data cleaning by latent factor adjustment. BMC Bioinform. 17, 27 (2016).
Rohart, F., Gautier, B., Singh, A. & Le Cao, K. A. mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752 (2017).
Fernandes, A. D. et al. Unifying the analysis of high-throughput sequencing datasets: characterizing RNA-seq, 16S rRNA gene sequencing and selective growth experiments by compositional data analysis. Microbiome 2, 15 (2014).
Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 42, D643–D648 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Lu, J. et al. Metagenome analysis using the Kraken software suite. Nat. Protoc. 17, 2815–2839 (2022).
Lin, H. & Peddada, S. D. Multigroup analysis of compositions of microbiomes with covariate adjustments and repeated measures. Nat. Methods 21, 83–91 (2024).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Karimnezhad, A. More accurate estimation of cell composition in bulk expression through robust integration of single-cell information. Bioinform Adv. 2, vbac049 (2022).
Gong, T. & Szustakowski, J. D. DeconRNASeq: a statistical framework for deconvolution of heterogeneous tissue samples based on mRNA-Seq data. Bioinformatics 29, 1083–1085 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Karl, J. P., Fu, X., Dolnikowski, G. G., Saltzman, E. & Booth, S. L. Quantification of phylloquinone and menaquinones in feces, serum, and food by high-performance liquid chromatography-mass spectrometry. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 963, 128–133 (2014).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38 (2014).
Benayoun, B. A. & Kim, M. Normal ovarian aging histology. figshare https://doi.org/10.6084/m9.figshare.25749588 (2024).
Benayoun, B. A. et al. Ovarian histology from VCD-treated and control mice. figshare https://doi.org/10.6084/m9.figshare.25749594 (2024).
Benayoun, B. A. & Kim, M. Ovarian histology from isochronic and heterochronic FMT recipient adult mice. figshare https://doi.org/10.6084/m9.figshare.25749591 (2024).
Kim, M. Hematoxylin and eosin staining images of ovaries from FMT cohort mice. figshare https://doi.org/10.6084/m9.figshare.28462274 (2025).
Kim, M. Raw immunohistochemistry images of IL6 in mouse ovary. figshare https://doi.org/10.6084/m9.figshare.29882735 (2025).
Acknowledgements
We thank V. Ansere and J. Bravo for feedback on our paper. We also thank D. Valenzano for his insightful comments and guidance on microbiome data analysis. We thank R. Vaca for processing the ovarian immunohistochemistry samples.
This work was supported by the GCRLE-2020 post-doctoral fellowship from the Global Consortium for Reproductive Longevity and Equality at the Buck Institute, made possible by the Bia-Echo Foundation to M.K., USC Provost’s Undergraduate Research Fellowship to J.W., NIA T32 AG052374 predoctoral fellowship and Diana Jacobs Kalman/AFAR Scholarships for Research in the Biology of Aging to R.J.L., and Pew Biomedical Scholar award no. 00034120 from the Pew Charitable Trust to B.A.B. Ovarian histological analysis was performed by the Translational Pathology Core at the USC Norris Comprehensive Cancer Center (supported by National Cancer Institute P30 CA014089) and Immunohistochemistry Laboratory at USC. Serum hormone quantification was performed by the University of Virginia Center for Research in Reproduction Ligand Assay and Analysis Core (supported by the Eunice Kennedy Shriver National Institute of Child Health and Human Development/National Institutes of Health grant R24HD102061). Computational analyses were performed using the Center for Advanced Research Computing resources at the USC. Stool vitamin K concentrations were supported by the USDA Agricultural Research Service under Cooperative Agreement no. 58-1950-7-707. Any opinions, findings, conclusions or recommendations expressed in this publication are those of the authors and do not necessarily reflect the views of the USDA. We thank W. Cohn, the Assistant Director of the USC Mann Multi-Omics Mass Spectrometry Core, for running the metabolomics samples on their instruments.
Author information
Authors and Affiliations
Contributions
M.K. and B.A.B. designed the study. M.K. performed live animal experiments. M.K. and R.J.L. dissected mouse tissues. M.K., J.W. and Y.K. analyzed ovarian histology data. A.X. generated the R shiny application. M.L., X.F. and S.L.B. quantified stool MK concentrations. S.E.P., M.K. and P.J.M. performed and analyzed metabolomics data. M.K. and J.W. performed computational analyses. M.K. and B.A.B. wrote the paper with input from all authors. All authors edited and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Eldin Jašarević, Katherine Kennedy, Lindsay Wu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of ovarian health state of young and estropausal female mice.
a, Schematic diagram of ovarian health index calculation. b, Representative hematoxylin and eosin (H&E)-stained ovarian section annotated to illustrate the morphological criteria used for follicle classification. Marked regions indicate follicles at different developmental stages: primordial (Pr), primary (P), secondary (S), antral (A), and corpus luteum (CL). c, Representative images of ovarian hematoxylin-eosin stained sections for young vs. estropausal female mice. Scale bar: 50 µm. d, Boxplots of primordial, primary, secondary and antral follicle and corpus luteum counts per section (p values ~0.0421, ~0.0442, ~0.0129, ~0.0146 and ~0.0133, respectively). e, Boxplots of serum concentrations of FSH (p value ~0.0952) and Inhibin A (p value ~0.0952). Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (d) and (e). YF: Young female; EF: Estropausal female.
Extended Data Fig. 2 Characterization of fecal microbial profiles of young and estropausal female mice.
a, Principal coordinate analysis results of Bray–Curtis dissimilarity (top panel) and Jaccard (bottom panel) indices. YF: Young female; EF: Estropausal female.
Extended Data Fig. 3 Characterization of ovarian health state of VCD cohort mice.
a, Schematic diagram of the experimental setup of VCD cohort. b, Representative images of ovarian hematoxylin-eosin stained sections. Scale bar: 50 µm. c–f, Boxplots of (c) primordial, primary, secondary and antral follicle, and corpus luteum counts per section (p values ~0.0833, ~0.0227, ~0.0073 and ~0.0248, respectively), (d) combined follicle counts (p value ~0.0117), (e) serum concentrations of AMH, FSH and Inhibin A (p values ~0.0079, ~0.0079 and ~0.0079, respectively), and (f) ovarian health index (p value ~0.0104) of control and VCD-injected mice. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (c–f).
Extended Data Fig. 4 Characterization of fecal microbial profiles of VCD cohort mice.
a, Principal component analysis result of CLR-transformed and batch-corrected ASV counts from control and VCD-injected mice. b, Principal coordinate analysis results of Bray–Curtis dissimilarity (top panel) and Jaccard (bottom panel) indices. c, d, Boxplots of normalized observed features and Shannon entropy indices of (c) females (p values ~0.6905 and 1, respectively), and (d) males (p values ~0.4286 and ~0.4286, respectively). Median from control-injected mice was used to normalize indices. e, Boxplot of weight changes from pre-injection to 100 days post-injection with either vehicle control or VCD in female and male VCD cohorts (p values 1 and 1, respectively). f, Differential abundance analysis results of microbial genera of control vs. VCD-injected female mice using ALDEx2. g, Functional abundance prediction analysis of control vs. VCD-injected female mice using PICRUSt2. h, Boxplot of beta-glucuronidase activity from PICRUSt2 (p value ~0.8413). Median from CTL females was used to normalize indices. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (c–e) and (h) and Benjamini–Hochberg corrected p value of Wilcoxon test are reported for (f) and (g).
Extended Data Fig. 5 Characterization of fecal microbial profiles of young and old male mice.
a, Schematic diagram of the experimental setup of male aging cohorts. b, Principal component analysis result of CLR-transformed and batch-corrected ASV counts from young and old male mice. Animals from three independent cohorts, n=14 per group (variation due to animal death or abnormal health issues prior to experiment). c, Principal coordinate analysis results of Bray–Curtis dissimilarity (top panel) and Jaccard (bottom panel) indices. d, Boxplots of observed features and Shannon entropy indices of young and old male mice (p values ~0.9633 and ~0.2456, respectively). Medians from young males for each cohort were used to normalize indices per cohort. e, Differential abundance analysis results of microbial genera of young and old male mice using ALDEx2. f, Functional abundance prediction analysis of young vs. old male mice using PICRUSt2. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (d) and Benjamini–Hochberg corrected p value of Wilcoxon test are reported for (e) and (f). YM: Young male; OM: Old male.
Extended Data Fig. 6 Bulk RNA-seq analysis of ovaries from young and estropausal female mice fecal microbiota transplantation recipient mice.
a, Multidimensional scaling (MDS) analysis result of RNA expression between young and estropausal female FMT recipient mice (FMT-YF and FMT-EF, respectively). b, Heat map of significant (FDR < 5%) genes differentially expressed in FMT-YF vs. FMT-EF mice. c, Top GSEA enriched terms from Gene Ontology Cellular Component (GO CC). Only up to top ten most-upregulated and top ten most-downregulated gene sets are plotted for readability. d, Dot plots of immune cell proportions in percentages between young and estropausal fecal microbiota transplantation recipient mice (FMT-YF and FMT-EF, respectively) predicted by deconvolution analyses using CSCDRNA (left panel) and Granulator (right panel). e, Top GSEA enriched terms from Gene Transcription Regulation Database (GTRD – TF targets). Only up to top ten most-upregulated and top ten most-downregulated gene sets are plotted for readability. f, GSEA enrichment score plots Ncoa1-peak (left panel) and Usp7-peak (right panel) genes (p values ~0.0138 and ~1e-04, respectively). Benjamini–Hochberg–adjusted p values derived from permutation-based GSEA statistics are reported. g, Bar plots of normalized log2 counts of Ncoa1 and Usp7 expression.
Extended Data Fig. 7 Characterization of ovarian health state of FMT cohort mice.
a, Representative images of ovarian hematoxylin-eosin stained sections. Scale bar: 50 µm. (b–e) Boxplots of (b) combined ovarian follicle counts (p value ~0.0041), (c) primordial, primary, secondary and antral follicle and corpus luteum counts per section (p values ~0.8689, ~0.5760, ~0.0889, ~0.0061 and ~0.5981, respectively), (d) serum concentrations of AMH, FSH (Multiplex assay), FSH (Ultra-sensitive assay), FSH combined and INHBA (p values ~0.0597, ~0.3176, 1, ~0.6701 and ~0.4428, respectively), and (e) weight percentage difference between day 0 and 16 of FMT regimen (p value ~0.5392). Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (c–e).
Extended Data Fig. 8 Characterization of fecal microbial profiles of FMT cohort mice.
a, Principal component analysis result of CLR-transformed and batch-corrected counts from young and estropausal female mice fecal microbiota transplantation recipient mice (FMT-YF and FMT-EF, respectively). b, Principal coordinate analysis results of Bray–Curtis dissimilarity (left panel) and Jaccard (right panel) indices. c, Boxplots of observed features and Shannon entropy indices of FMT-YF and FMT-EF mice (p values ~0.8981 and 1, respectively). d, Differential abundance analysis results of microbial species of FMT-YF vs. FMT-EF mice using ALDEx2. e, Venn diagram of overlapping microbial genera between genera that increase in abundance with female aging (“UP in EF”) and increase in abundance in FMT-EF recipient mice (“UP in FMT-EF”). f, Functional abundance prediction analysis of FMT-YF vs. FMT-EF mice using PICRUSt2. g, Boxplot of beta-glucuronidase activity from PICRUSt2 (p value ~0.0513). Median from FMT-YF group was used to normalize indices. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown. Significance in nonparametric two-sided Wilcoxon rank-sum tests are reported for (c) and (g), and Benjamini–Hochberg corrected p value of Wilcoxon test are reported for (d) and (f). YF: Young female; EF: Estropausal female.
Extended Data Fig. 9 Stool vitamin K quantification of FMT cohort mice.
a, Stool concentrations of combined vitamin K (MK6-MK13) of FMT-YF and FMT-EF. b, Fold change in stool concentrations of vitamin K (MK6-MK13) of FMT-YF and FMT-EF. Boxplots show the median (center line), the 25th and 75th percentiles (bounds of the box), with whiskers extending to 1.5× the interquartile range (IQR). Individual datapoints are shown.
Extended Data Fig. 10 Causal mediation analysis of microbial species and ovarian transcriptome changes in fecal microbiota transplantation recipient mice.
a, Graphical representation of causal mediation analysis. b, Causal mediation analysis results of 16S amplicon ASVs that showed significant mediation effects on changes in the ovarian transcriptome in fecal microbiota transplantation recipient mice. Bars represent model-derived mediation estimates (Average Causal Mediation Effect, Average Direct Effect, and Total Effect). Vertical lines denote 95% nonparametric bootstrap confidence intervals. Statistical significance is denoted as n.s., not significant; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. c, Heat map of Spearman correlation analysis between CLR-transformed shotgun metagenomics count data and serum hormone quantification data. #: p value < 0.1; *: p value < 0.05. ACME: Average Causal Mediated Effect; ADE: Average Direct Effect.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7 and Tables 1 and 2.
Supplementary Table 3 (download XLSX )
MS2-filtered metabolite abundance matrix.
Source Data Supplementary Fig. 1 (download XLSX )
Statistical source data for Supplementary Fig. 1.
Source Data Supplementary Fig. 2 (download XLSX )
Statistical source data for Supplementary Fig. 2.
Source Data Supplementary Fig. 3 (download XLSX )
Statistical source data for Supplementary Fig. 3.
Source Data Supplementary Fig. 4 (download XLSX )
Statistical source data for Supplementary Fig. 4.
Source Data Supplementary Fig. 5 (download XLSX )
Statistical source data for Supplementary Fig. 5.
Source Data Supplementary Fig. 6 (download XLSX )
Statistical source data for Supplementary Fig. 6.
Source Data Supplementary Fig. 7 (download XLSX )
Statistical source data for Supplementary Fig. 7.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, M., Wang, J., Pilley, S.E. et al. Estropausal gut microbiota transplant improves measures of ovarian function in adult mice. Nat Aging 6, 682–702 (2026). https://doi.org/10.1038/s43587-026-01069-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-026-01069-3