Abstract
Metabolic dysfunction–associated steatotic liver disease (MASLD), potentially ameliorated by bariatric-metabolic surgery, remains a global health concern in the absence of approved drugs. Protein post-translational modifications (PTMs) are crucial for MASLD. However, the functional significance of lysine crotonylation (Kcr) remains unclear. We aimed to investigate the mechanisms by Kcr-regulated IDH1 in the tricarboxylic acid (TCA) cycle and MASLD development. Herein, we reported a quantitative proteomics analysis of global crotonylome upon MASLD and Post-bariatric. Specifically, decreases in K58cr, K151cr, K212cr and K345cr of IDH1 upon MASLD were observed. PCAF and SIRT7 dynamically regulated the IDH1 Kcr. Abolishment of IDH1 Kcr impaired TCA cycle by decreasing IDH1 enzymatic activity. Male mice with liver-specific expression of crotonylation-mimic mutants of IDH1 were resistant to HFD-induced obesity, insulin resistance, glucose intolerance and MASLD. Our findings unravel the mechanisms of IDH1 Kcr and indicate that targeting PCAF/SIRT7-IDH1 Kcr and metabolites may be a promising strategy for MASLD therapy.
Similar content being viewed by others
Introduction
Metabolic dysfunction–associated steatotic liver disease (MASLD) is the most common cause of chronic liver disease worldwide1,2,3. MASLD is associated with obesity and encompasses a range of pathologies, including steatosis, metabolic dysfunction-associated steatohepatitis (MASH), fibrosis, and cirrhosis. MASLD also increases the risk of hepatocellular carcinoma (HCC)2,3,4. Accumulating evidences reveal that the protein post-translational modifications (PTMs) play a key role in MASLD pathogenesis5. The aberrant PTMs alterations (e.g., acetylation, ubiquitination and lactylation) control hepatic lipid metabolism, oxidative stress, and DNA repair to boost MASLD progression6,7. Metabolic dysfunction also induces lasting epigenetic changes by regulating small metabolites, the donor molecules for PTMs8,9,10. Weight loss with diet or bariatric-metabolic surgery is effective for treating MASLD, but drugs specifically approved MASLD are not available11,12,13. The lack of understanding of the molecular mechanisms underlying the complex interplay between epigenetic and metabolic factors impedes the development of improved therapeutic interventions for MASLD.
Lysine crotonylation (Kcr), a metabolically regulated PTMs, is involved in key cellular processes relevant to physiology and diseases14. Kcr affects protein functions through diverse mechanisms, including by regulating enzymatic activity, subcellular localization, and crosstalk with other PTMs and by controlling protein-protein and protein-DNA interactions14,15,16. Kcr is regulated by enzymes that deposit or remove modifications. Histone acetyltransferase p300/CBP17, MOF18, PCAF19, GCN520, HBO121 and TIP6022 have also been identified to possess crotonyltransferase activity. The deacetylases, including HDAC1/2/3/6/7/8, SIRT1/2/3/6/7 and CDYL, are able to remove Kcr16,23,24,25,26. Emerging evidences have shown that Kcr is crucial for regulating cellular metabolism in HCC27,28 However, few relevant literatures on the crotonylome during the MASLD development have been reported.
Isocitrate dehydrogenases (IDH) 1 and 2 are abundant in liver and can convert isocitrate to α-ketoglutarate (α-KG) by reducing NADP+ and serve to regulate redox status, lipogenesis, glucose, and amino acid metabolism29. Mutation of IDH1 plays a pathogenic role in cancers, converting α-KG to the oncometabolite 2-hydroxyglutarate (2-HG)30,31. Recent studies highlight the function of IDH1 that can be regulated by acetylation and phosphorylation modification32,33. Acetylation of IDH1 significantly down-regulates its catalytic activity and the rate of α-KG production in colorectal cancer cells32. Proteomics studies showed that enzymes in the tricarboxylic acid (TCA) cycle, including IDH1, are extensively crotonylated15,19,34. However, crotonylation as a new type of post-translational modification, IDH1 Kcr and its function in MASLD have not yet been reported30,34,35,36.
In this study, we used a quantitative proteomics approach to gain a global view of the crotonylome alterations in response to MASLD and bariatric-metabolic surgery. We identified crotonylated IDH1 as a core target in MASLD. We also confirmed that PCAF and SIRT7 act as regulatory enzymes of IDH1 Kcr, and the Kcr of IDH1 is involved in regulating lipid accumulation in hepatocytes via boosting the TCA cycle. Moreover, the disruption of IDH1 Kcr in hepatocytes aggravates MASLD in HFD-fed mice. MASLD-hepatic tissues from both patients and diet-fed mice exhibited significantly reduced PCAF/SIRT7-mediated IDH1 Kcr. Our results identify the role of IDH1 Kcr in MASLD and suggest a potential pharmaceutical target for MASLD.
Results
Global analysis of Kcr in HFD-induced MASLD or post-bariatric surgery mice
Kcr has been reported to function in several physiological and pathological processes15,23,37. It is important in regulating cellular metabolism and may contribute to the progression of HCC27,28. However, the function of Kcr in MASLD development and prognosis remains obscure. MASLD was induced in 6-week-old male C57BL/6 J mice by high-fat diet (HFD) feeding for 12 weeks and then assigned to either HFD sham group or vertical sleeve gastrectomy (VSG)-improved MASLD group. To determine the weight-loss-independent effects, we included a PAIR-FED sham group that was pair-fed to the VSG group to match their caloric intake during post-surgery period. In addition, a sham group of mice were fed a normal chow diet (NCD) for the duration of this study. High-fat feeding accelerated MASLD progression, increased liver lipid accumulation, and elevated liver function markers compared to the normal chow diet (NCD)-treated mice (Fig. 1a–c and Supplementary Fig. 1–2). VSG surgery effectively alleviated MASLD progression, improved liver function markers, insulin resistance, glucose intolerance, and significantly reduced body weight, liver fat accumulation, cholesterol, and hyperlipidemia than HFD-induced MASLD mice (Fig. 1a–c and Supplementary Fig. 1–2). While both VSG and pair-fed mice maintained comparable weight loss post-intervention, only the VSG group exhibited a marked reduction in intrahepatic lipid deposition (Fig. 1b). This distinct outcome underscores that the therapeutic benefits of VSG on MASLD pathogenesis, including improvements in metabolic and hepatic parameters, are mediated through mechanisms independent of caloric restriction induced by surgery. In general, VSG attenuates HFD-induced obesity and MASLD.
a Schematic representation of experimental workflow showing 4D label-free quantification of Kcr in NCD, HFD, PAIR-FED and VSG mouse livers. Created in BioRender. Liu, S. (2025) https://BioRender.com/vr8ptir. b Representative H&E staining of liver sections 8 weeks post-operation in NCD, HFD, PAIR-FED, and VSG group mice (n = 5/group). NAS scores were derived from H&E staining. Scale bars: 100 μm. c Body weight was recorded weekly post-operation in NCD, HFD, PAIR-FED, and VSG group mice (n = 5/group). d Pie chart illustrating the distribution of the number of identified Kcr sites per protein. e Map illustrating the subcellular distribution of Kcr proteins. f Icelogo representation illustrating flanking sequence preferences for all Kcr sites. g Motif analysis of all identified Kcr proteins. h Bar graphs illustrating representative ontology annotations enriched in the Kcr proteome. Data are shown as the mean ± SD. P values were obtained using one-way ANOVA followed by Tukey’s multiple comparisons test. Source data are provided as a Source Data file.
We analyzed the liver samples from normal and MASLD mice models (Fig. 1a and Supplementary Fig. 3). The inventory of crotonylated peptides detected in liver tissue was evaluated to gain further insights into their role in MASLD. The modified peptides and sites enriched in these samples were analyzed by nanoflow liquid chromatography-tandem mass spectrometry (LC-MS/MS). In this study, a total of 17,408 unique Kcr sites across 3913 proteins were identified, with 14,684 Kcr sites from 3188 quantified. Among these Kcr proteins, 1247 (31.9%) had a single Kcr site and 975 (24.9%) had more than six Kcr sites (Fig. 1d and Supplementary Data 1–2). We further examined the subcellular distribution of Kcr. 43% Kcr sites were identified in the cytoplasm, 13% were found in the nucleus and 18% were in the mitochondria (Fig. 1e). We next evaluated the amino acid sequences flanking the identified Kcr sites against all human background sequences using iceLogo (Fig. 1f and Supplementary 3). Consistently, analysis using Motif-X algorithms identified KExxxK, KExxxR and AKxxxxxxK as significantly overrepresented hotspots for Kcr sites (Fig. 1g and Supplementary Data 4). We further analyzed the potential Kcr-regulated intracellular pathways. GO enrichment analysis revealed that Kcr proteins are involved in diverse biological processes and molecular functions, such as carboxylic acid metabolic process, peptide metabolic process and monocarboxylic acid metabolic process (Fig. 1h and Supplementary Data 5).
Quantitative analysis of Kcr proteome in mouse liver tissue
We next quantified the changes of protein Kcr in response to HFD-fed MASLD mice relative to total protein abundance in normal liver tissues or VSG relative to total protein abundance in PAIR-FED liver tissues. The cutoff ratio for significant Kcr changes between each two groups was set to above 1.5 or below 0.67. Upon MASLD, 170 sites in 150 proteins were up-regulated and 396 Kcr sites in 321 proteins were down-regulated (Fig. 2a and Supplementary Data 1). GO enrichment analysis revealed that up-regulated Kcr proteins are enriched in hormone catabolic process and monocarboxylic acid metabolic process, whereas down-regulated Kcr proteins are mainly involved in carboxylic acid metabolic and biosynthetic process (Fig. 2b, c and Supplementary Data 6). After VSG surgery, 369 sites in 283 proteins were up-regulated and 117 Kcr sites in 94 proteins were down-regulated (Fig. 2d and Supplementary Data 1). GO enrichment analysis revealed that up-regulated Kcr proteins are enriched in carboxylic acid metabolic and catabolic process, whereas down-regulated Kcr proteins are mainly involved in cell junction assembly and lipid translocation (Fig. 2e, f and Supplementary Data 7). Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis also revealed that MASLD-related Kcr proteins and VSG-related Kcr are involved in diverse biological processes, including fatty acid degradation and TCA cycle (Supplementary Fig. 4a–f and Supplementary Data 8–10).
a Histogram showing the ratio distribution of quantifiable Kcr proteins and sites between HFD and NCD mice livers. b Bar graphs showing GO pathway associated with all identified Kcr proteins between HFD and NCD mouse livers. c Bar graphs showing GO pathway associated with up-regulated (left) or down-regulated (right) Kcr proteins between HFD and NCD mice liver. d Histogram showing the ratio distribution of quantifiable Kcr proteins and sites between VSG and PAIR-FED mice liver. e Bar graphs showing GO pathway associated with all identified Kcr proteins between VSG and PAIR-FED mice liver. f Bar graphs showing GO pathway associated with up-regulated (left) or down-regulated (right) Kcr proteins between VSG and PAIR-FED mice livers. g Venn diagram showing the overlap between HFD-downregulated Kcr substrates and VSG-upregulated substrate Kcr substrates. h Overview of the TCA cycle based on crotonylome.
Next, we compared MASLD-related and VSG-related Kcr proteins. The results show that 99 overlapped proteins down-regulated in the MASLD group and up-regulated in the VSG group, and only 4 overlapped proteins changed reversely (Fig. 2g and Supplementary Fig. 4g). We then analyzed the interaction network of MASLD-regulated or bariatric surgery-related Kcr proteome on the basis of STRING database. A number of highly connected subnetworks of Kcr proteins were identified using MCODE. We found that the changes in TCA cycle pathway, especially its key enzymes, such IDH1, IDH2 and SUCLG2, were obviously downregulated upon MASLD and up-regulated after VSG (Fig. 2h and Supplementary Fig. 5–6). We then selected IDH1 as the representative target to further investigate the biological consequence of MASLD-regulated Kcr.
IDH1 Kcr decreases in response to HFD-induced MASLD
To validate Kcr of IDH1, total cellular extracts from HepG2 or AML12 cells were immunoprecipitated with anti-PanKcr followed by immunoblotting with anti-IDH1. Meanwhile, the reciprocal immunoprecipitation with anti-IDH1 followed by immunoblotting with the antibody against PanKcr confirmed that IDH1 was crotonylated in HepG2 or AML12 cells (Fig. 3a, b). Quantitative analysis of Kcr modification revealed a significant decrease in IDH1 Kcr levels in the livers of MASLD mice compared to those of normal controls (Fig. 2h). Palmitic acid (PA) and oleic acid (OA)-treated hepatocytes were used to construct MASLD model in vitro, in which total cholesterol (TC) and triglycerides (TG) were increased (Supplementary Fig. 7a, b). Immunoprecipitation assays were performed, followed by immunoblotting with the indicated antibodies. Indeed, the level of IDH1 Kcr decreased upon PA + OA-induced MASLD model (Fig. 3c, d). In contrast, PA + OA did not affect Kac of IDH1 (Fig. 3d), reinforcing the notion that IDH1 Kcr is preferably responded to PA + OA-induced MASLD models. Examination of the site specificity of MASLD-related Kcr of IDH1 revealed a total of 19 Kcr sites on IDH1 from our LC-MS/MS results (Fig. 3e). Upon MASLD progression, the levels of K58cr, K151cr, K212cr and K345cr decreased, respectively, whereas the levels on other IDH1 Kcr sites did not change significantly (Fig. 3e and Supplementary Fig. 6). We compared the differences in IDH1 sites among different species and found that K58, K151, K212, and K345 are highly conserved among different species (Fig. 3f). We substituted each of four lysine (K) residues with arginine (R) and performed immunoprecipitation assays in AML12 and HEK293T cells transfected MYC-tagged WT or point mutant of IDH1. We found the Kcr of IDH1 within K58R, K151R, K212R and K345R were decreased (Fig. 3g). We then generated quadruple-sites mutant by mutating the indicated IDH1 K to R. Immunoprecipitation assays of HEK293T cells transfected MYC-tagged WT or IDH1-4KR showed that IDH1 Kcr decreased with quadruple-sites mutants (Fig. 3h).
a Immunoprecipitation in HepG2 or AML12 cells with anti-PanKcr or anti-IgG followed by immunoblotting with anti-IDH1. b Immunoprecipitations in HepG2 or AML12 cells with anti-IDH1 or anti-IgG followed by immunoblotting with anti-PanKcr. c Immunoprecipitations in HepG2 or AML12 cells treated with or without PA + OA for 24 h using anti-PanKcr or anti-IgG followed by immunoblotting with anti-IDH1. d Immunoprecipitations in HepG2 or AML12 cells treated with or without PA + OA using anti-IDH1 followed by immunoblotting with anti-PanKcr or anti-IgG. e Schematic diagram of IDH1. f Sequence alignment homology around the four crotonylation sites in different species IDH1. g K58, K151, K212 and K345 are key Kcr sites of IDH1 in vivo. Immunoprecipitation assays in HEK293T cells upon overexpression of the indicated Myc-tagged constructs with anti-Myc followed by immunoblotting with anti-Myc and anti-PanKcr. h Immunoprecipitation assays in HEK293T cells upon overexpressing Myc-tagged IDH1 WT or 4KR with anti-Myc followed by immunoblotting with anti-Myc and anti-PanKcr. i The specificity of antibodies against IDH1-K58cr, IDH1-K151cr, IDH1-K212cr and IDH1-K345cr was verified by dot blot assays. The nitrocellulose membrane was spotted with the indicated amounts of uncrotonylated or crotonylated IDH1 peptides and immunoblotted with the indicated antibodies. j Verification of the specificity of anti-IDH1-K58cr, anti-IDH1-K151cr, anti-IDH1-K212cr and anti-IDH1-K345cr by western blotting. Immunoprecipitation assays were performed in AML12 cells overexpressing the indicated Myc-tagged IDH1 constructs with anti-Myc followed by immunoblotting with the indicated antibodies, respectively. Source data are provided as a Source Data file.
To further confirm the LC-MS/MS results, we generated polyclonal antibodies specifically recognizing IDH1 K58cr, IDH1 K151cr, IDH1 K212cr, or IDH1 K345cr, respectively. The specificity of these antibodies was verified by dot blotting assays using corresponding peptides with or without Kcr modification (Fig. 3i). Western blotting of total cell lysates showed that both crotonylated IDH1 and unmodified IDH1 run at approximately 47 kDa (Supplementary Fig. 7c). Moreover, immunoprecipitation assays in AML12 and HEK293T cells transfected with MYC-tagged WT IDH1 or point-mutant IDH1-K58R, IDH1-K151R, IDH1-K212R or IDH1-K345R confirmed that K to R mutagenesis abolished the recognition by respective IDH1 Kcr antibodies (Fig. 3j, Supplementary Fig. 7d). Moreover, we used the polyclonal antibodies to test the Kcr of IDH1K58, K151, K212 and K345 in cell lines respectively (Supplementary Fig. 7e). Intriguingly, IDH1 Kcr decreased significantly in PA + OA-induced hepatocytes compared with bovine serum albumin (BSA)-induced hepatocytes (Supplementary Fig. 7e). Meanwhile, the quantity of IDH1 protein showed no change in the PA + OA-induced MASLD hepatocytes models. The above results confirm that it is the Kcr of IDH1 rather than the protein of IDH1 that plays a significant role in the progression of MASLD in cellular level.
IDH1 Kcr is mediated by PCAF and SIRT7
Crotonyltransferases and their counteracting decrotonylases are important for controlling various cellular processes23. Given the fact that many acetyltransferases possess crotonyl-transferase activity38, we transiently transfected HEK293T cells with p300, CBP, PCAF or GCN5. Then the cell lysates were subjected to immunoblotting with a PanKcr antibody. As shown in Fig. 4a, we found that the ectopic expression of PCAF, not p300/CBP or GCN5, promoted the Kcr of IDH1, suggesting that IDH1 Kcr is a function of PCAF. The endogenous Kcr of IDH1 was reduced by the reduction of PCAF (Fig. 4b). In addition, there was an interaction between IDH1 and PCAF (Supplementary Fig. 8a). These results demonstrate that PCAF might represent a physiological crotonyltransferase for IDH1.
a Immunoprecipitation assays with anti-Myc in HEK293T cells co-transfected with Myc-tagged IDH1 and HA-tagged HATs followed by immunoblotting with antibodies against the indicated proteins. b Immunoprecipitation assays with anti-IDH1 in HEK293T cells transfected with control siRNA or siRNA targeting PCAF followed by immunoblotting with anti-PanKcr or anti-IgG. c HEK293T cells were expressed Myc-tagged IDH1 and then treated with 3 μM TSA or 5 mM NAM for 12 h, immunoprecipitation assays with anti-Myc followed by immunoblotting with antibodies against the indicated proteins. d Immunoprecipitation assays with anti-Myc in HEK293T cells transfected with Myc-tagged IDH1 and FLAG-tagged SIRTs followed by immunoblotting with antibodies against the indicated proteins. e Immunoprecipitation assays with anti-IDH1 in HEK293T cells transfected with control siRNA or siRNA targeting SIRT7 followed by immunoblotting with anti-PanKcr or anti-IgG. f Immunoprecipitation assays with anti-Myc in HEK293T cells co-transfected with Myc-tagged IDH1, HA-tagged PCAF and FLAG-tagged SIRT7, followed by immunoblotting with the indicated antibodies. g Schematic diagram of dynamic regulation of IDH1 Kcr by PCAF and SIRT7. Source data are provided as a Source Data file.
To identify the potential physiological decrotonylase(s) for IDH1, we compared the treatment using nicotinamide (NAM), a sirtuin deacetylase (SIRT) inhibitor with that using a histone deacetylase inhibitor, trichostatin A (TSA) and found that the former effectively increased IDH1 Kcr (Fig. 4c). Moreover, NAM also rescued the reduction in IDH1 Kcr induced by PA + OA (Supplementary Fig. 8b). Furthermore, the ectopic expression of SIRT7, but not other SIRTs or HDACs, was found to decrease PCAF-mediated Kcr of IDH1 (Fig. 4d, f and Supplementary Fig. 8c). In contrast, the depletion of endogenous SIRT7 using short interfering RNA (siRNA) increased Kcr of IDH1 (Fig. 4e). Moreover, there was also an interaction between IDH1 and SIRT7 (Supplementary Fig. 8a), supporting the notion that SIRT7 is a bona fide decrotonylase for IDH1.
To further characterize the role of PCAF and/or SIRT7 upon IDH1 Kcr regulation during the development of MASLD, we examined their levels in the PA + OA-treated hepatocytes. The results showed that either mRNA levels or protein levels of PCAF were lower in PA + OA-treated than in BSA-treated AML12 cells and HepG2 cells, whereas the levels of SIRT7 showed an inverse trend (Supplementary Fig. 9a, b). We further validated PCAF/SIRT7-mediated IDH1 Kcr downregulation in mouse primary hepatic cells (MPHs) under steatotic conditions (Supplementary Fig. 10a, b). Intriguingly, this regulatory pattern was absent in both RAW264.7 macrophages and LX-2 hepatic stellate cells (Supplementary Fig. 10c, d), suggesting cell type-specific regulation of IDH1 Kcr. Taken together, Kcr of IDH1 was modulated by decrotonylase SIRT7 and crotonyltransferase PCAF (Fig. 4g).
IDH1 Kcr is crucial for TCA cycle
IDH1 is a key enzyme involved in TCA cycle that catalyzes the conversion of isocitrate to α-ketoglutarate (α-KG), with NADP+ as electron receptor to generate NADPH30. We first examined whether Kcr affected the enzymatic activity of IDH1. We conducted K58, K151, K212 and K345 simultaneously mutated to glutamine (4KQ) to mimick crotonylated IDH1. Wild-type IDH1, IDH1-4KR and IDH1-4KQ were reconstituted separately in AML12 cells in which endogenous IDH1 was knocked down by siRNA. The knockdown efficiency of IDH1 and equal expression levels of transfected constructs were validated by western blotting (Supplementary Fig. 11a). Importantly, the knockdown of IDH1 was associated with a significant decrease in the enzymatic activity of IDH1, an effect that could be completely rescued by overexpression of siRNA-resistant wild-type IDH1 or IDH1-4KQ, but not IDH1-4KR (Fig. 5a). To investigate the metabolic consequences of IDH1 Kcr, we measured IDH1 upstream and downstream metabolites in AML12 cells subjected to five experimental conditions: siControl + Vector, siIDH1 + Vector, siIDH1 +WT, siIDH1 + 4KR and siIDH1 + 4KQ. IDH1 depletion triggered significant accumulation of upstream metabolites citrate and isocitrate, consistent with impaired oxidative decarboxylation activity (Fig. 5a, b and Supplementary Fig. 11b). While both wild-type IDH1 and the 4KQ mutant restored metabolic homeostasis, the 4KR mutant failed to rescue this phenotype. Conversely, α-KG levels exhibited compensatory declines (Fig. 5c), whereas fumarate remained unaltered across the groups (Supplementary Fig. 11c). Notably, the 4KR mutant uniquely disrupted cellular redox balance, as evidenced by an increase in NADP+/NADPH ratio and a reduction in ATP production compared to wild-type IDH1 (Fig. 5d, e). Changes in metabolites indicate that IDH1 Kcr affects IDH1 activity and, consequently, the TCA cycle.
a AML12 cells were treated with specific siRNA targeting IDH1 3’ UTR or control siRNA for 24 h before transfection with the indicated Myc-tagged constructs. IDH1 activity assay with purified indicated Myc-tagged immunoprecipitates from AML 12 cells (n = 3/group). b, c Histogram displaying Isocitrate (b) and α-KG (c) levels in AML12 cells (n = 3/group) described in (a). d, e Histogram displaying NADP+/NADPH ratio (d) and ATP levels (e) in AML12 cells (n = 3/group) described in (a). f Schematic diagram of 13C-labeled D-13C6 glucose in glycolysis and the TCA cycle. g Histogram showing changes in TCA cycle (upper) or glycolysis (lower)-related metabolites among IDH1-WT and IDH1 mutants (n = 3/group). h The Oxygen consumption rate (OCR) in AML12 cells (n = 6/group) with endogenous IDH1 knockdown and reconstituted expression of IDH1 WT, IDH1 4KR or 4KQ. i Quantification of Basal respiration, maximal respiration, and ATP production (n = 6/group). j Representative fluorescence images reflecting ROS level in AML12 cells described in (a), scale bar: 50 μm. k Fatty acid beta oxidation (FAO) efficiency in AML12 cells (n = 3/group) described in (a). Data are shown as the mean ± SD. P values were obtained using one-way ANOVA followed by Tukey’s multiple comparisons test (a–e, h, i, k) or Sidak test (g). Source data are provided as a Source Data file.
Consistent with the above metabolic changes, we quantified 13C-labled metabolites via mass spectrometry from AML12 cells that were fed 13C6-glucose for 12 h (Fig. 5f). These metabolites were designated in the study as the metabolite’s name followed by ‘M + n’ (‘M’ stands for molecular mass; ‘n’ indicates the number of heavy carbons 13C in the metabolite). In IDH1-knockdown AML12 cells, IDH1-4KR overexpression significantly reduced α-KG M + 2 levels compared to the IDH1-WT group, while isocitrate M + 2 showed a non-significant increasing trend (Fig. 5g and Supplementary Data 11). These findings demonstrate that IDH1 Kcr inhibition impairs enzymatic activity, leading to decreased isocitrate-to-α-KG conversion and consequent TCA cycle dysfunction.
Next, we took Seahorse analysis to detect cellular energy metabolic capacity in AML12 cells (Fig. 5h, i and Supplementary Fig. 11d, e). We evaluated mitochondrial oxidative phosphorylation (OXPHOS) activity by measuring the oxygen consumption rate (OCR). Compared with the control group, the knockdown of IDH1 exhibited lower basal respiration, maximal respiration, and ATP production level, suggesting that IDH1 deficiency impairs oxygen consumption rate and triggers mitochondrial exhaustion. However, both the overexpression of IDH-WT and IDH1-4KQ in AML12 cells after knockdown of IDH1 brought a sharp upturn of oxygen consumption, while the overexpression of IDH1-4KR did not, indicating that the capacity of IDH1 mediating TCA cycle may have been limited while the crotonylation decreased. Meanwhile, quantitative analysis of glycolytic activity through extracellular acidification rate (ECAR) demonstrated that in rescue experiments following endogenous IDH1 depletion, both wild-type IDH1 and the 4KQ mutant moderately impaired glycolysis, glycolytic capacity, whereas the 4KR mutant showed no such effects. Moreover, analysis of the basal OCR/ECAR ratio demonstrated that IDH1 Kcr significantly modulates the balance between OXPHOS and glycolysis (Supplementary Fig. 11f). Together, these findings indicate that the loss of IDH1 Kcr impairs mitochondrial function more profoundly than glycolytic activity.
The NADP+-dependent IDH1 is an important metabolic enzyme that generates NADPH. NADPH is required for reduced glutathione, which is the major antioxidant against reactive oxygen species (ROS). We additionally measured ROS levels and found that crotonylation null mutant (KR mutation) increased ROS production (Fig. 5j). Based on these findings, we observed that reduced crotonylation of IDH1 leads to decreased protease activity, causing mitochondrial dysfunction. This reduction in oxygen consumption results in inefficient mitochondrial electron transport, leading to increased ROS production and promoting MASLD development.
We also measured the fatty acid beta-oxidation (FAO) efficiency in AML12 cells and found that IDH1 4KR mutation can decrease the efficiency of FAO (Fig. 5k). Based on these results, we concluded that decreased IDH1 Kcr diminishes its enzymatic activity, leading to mitochondrial dysfunction. This reduction in oxygen consumption results in inefficient mitochondrial electron transport, elevated ROS production, and ultimately contributes to MASLD progression.
We next examined whether Kcr of IDH1 affected its subcellular distribution. To this end, AML12 cells were transfected with constructs encoding wild-type IDH1, IDH1-4KR and IDH1-4KQ respectively, and subsequently subjected to immunofluorescence staining after been fixed. The results showed that wild-type IDH1, IDH1-4KR and IDH1-4KQ were similarly found in nucleus, cytoplasm, and mitochondria, suggesting that Kcr on these sites has no obvious effect on the subcellular localization of IDH1 (Supplementary Fig. 11g).
Previous studies show that active IDH1 enzyme form homodimers and enhance its activity30,39. To further determine whether IDH1 Kcr affects its dimer formation, we conducted co-transfection of mutations, including Myc-IDH1-WT/4KR/4KQ and FLAG-IDH1 in HEK293T cells. The results showed that the abolishment of IDH1 Kcr promoted IDH1 dimer disruption to form monomers (Supplementary Fig. 11h). Taken together, abrogation of IDH1 Kcr blocked its dimer formation and impaired enzymatic activity, supporting that Kcr of IDH1 is critical for TCA cycle during MASLD progression.
Downregulation of IDH1 Kcr promotes hepatic lipid accumulation
To determine whether IDH1 Kcr may affect biological processes in the liver, we first examined the effect of IDH1 Kcr on cellular lipotoxicity of PA + OA-simulated AML12 cells. Wild-type IDH1, IDH1-K58R, IDH1-K151R, IDH1-K212R, IDH1-K345R, IDH1-4KR and IDH1-4KQ were reconstituted respectively in AML12 cells in which endogenous IDH1 was knocked down by siRNA (Fig. 6a–c). The knockdown efficiency of IDH1 and equal expression levels of transfected constructs were validated by western blotting (Fig. 6c). Without PA + OA treatment, the knockdown of IDH1 in AML12 cells led to an increase in the lipid deposition, an effect that could be largely rescued by overexpression of wild-type IDH1, IDH1-K58R, IDH1-K151R, IDH1-K212R, IDH1-K345R, IDH1-4KR and IDH1-4KQ to equal extent. On the contrary, upon treating with PA + OA, all these IDH1-KR mutants failed to ameliorate the elevated lipid deposition in IDH1 deficient cells when compared with wild-type IDH1 and IDH1-4KQ (Fig. 6a–c), supporting that IDH1 Kcr protects liver cells from cell lipid accumulation.
a AML12 cells were treated with control or IDH1 siRNA for 24 h, followed by expression of Myc-tagged Vector, IDH1-WT, IDH1-K58R, IDH1-K151R, IDH1-K212R, IDH1-K345R, IDH1-4KR and IDH1-4KQ. After another 24 h, cells were treated with BSA (top) or 0.5 mM PA + OA (bottom) for 24 h, then fixed and stained with Oil Red O. Scale bars: 100 μm. b Quantification of the Oil Red O staining area as described in (a) (n = 6/group). c The expression of the indicated proteins was measured in AML12 cells as described in (a). d AML12 cells were treated with control or PCAF siRNA for 48 h before cells were treated with BSA (top) or 0.5 mM PA + OA (bottom) for another 24 h, then fixed and stained with Oil Red O. Scale bars: 100 μm; e Quantification of the Oil Red O staining area as described in (d) (n = 3/group). f The expression of the indicated proteins was measured in AML12 cells as described in (d). g FLAG-tagged SIRT7 were expressed in AML12 cells for 48 h, followed by treatment with BSA (top) or 0.5 mM PA + OA (bottom) for 24 h, fixed and stained with Oil Red O. Scale bars: 100 μm. h Quantification of the Oil Red O staining area as in (g) (n = 3/group). i The expression of the indicated proteins was measured in AML12 cells as described in (g). Data are shown as the mean ± SD. P values were obtained using one-way ANOVA followed by Tukey’s multiple comparisons test (b) or two-tailed unpaired Student’s t-test (e, h). All experimental data were verified at least three independent experiments. Source data are provided as a Source Data file.
In addition, we sought to verify the role of PCAF and SIRT7 in IDH1 Kcr mediated cellular lipid accumulation. We detected that PCAF knockdown contributes to an increase in the lipid deposition in AML12 cells with or without PA + OA treatment (Fig. 6d–f). Consistently, SIRT7 overexpression showed the same tendency in AML12 cells with or without PA + OA treatment (Fig. 6g–i). These results suggested that block of IDH1 Kcr enhances cellular lipotoxicity as well as PCAF knockdown or SIRT7 overexpression.
IDH1-4KR exacerbates lipid accumulation and disrupts TCA cycle function in HFD-fed mice
To confirm the role of IDH1 Kcr in MASLD development, we generated adeno-associated virus (AAV)8-IDH1-WT, AAV8-IDH1-4KR, and AAV8-IDH1-4KQ expression vectors and injected them through the tail veins of C57BL/6 J mice, which were then fed a HFD respectively (Fig. 7a, b and Supplementary Fig. 12a). The knockdown efficiency of IDH1 Kcr of transfected constructs were validated by western blotting (Fig. 7c, Supplementary Fig. 12a). Strikingly, IDH1-4KQ mice had a lower body weight than the corresponding control mice (Fig. 7b, d). These mice also displayed significantly improved glucose tolerance and decreased insulin resistance (Fig. 7e, f). Moreover, HFD-induced hepatic steatosis was improved in IDH1-4KQ mice, and these mice also had a lower liver weight than the control mice (Fig. 7g–k).
a Schematic of the HFD-induced MASLD model. b Photos of AAV8-TBG-ZsGreen, AAV8-TBG-IDH1-WT, AAV8-TBG-IDH1-4KR, and AAV8-TBG-IDH1-4KQ mice. c Representative western blotting analysis of liver FLAG-IDH1 and α-Tubulin expression levels in the mice described in (a). d Body weight was measured weekly in AAV8-ZsGreen, AAV8-IDH1-WT, AAV8-IDH1-4KR, or AAV8-IDH1-4KQ mice (n = 8/group). e Blood glucose concentration was determined using the insulin tolerance test (ITT) in 22-week-old mice (n = 8/group). IP: IDH1-WT vs. IDH1-4KR; IIP: IDH1-WT vs. IDH1-4KQ. f Blood glucose concentration was determined using the intraperitoneal glucose tolerance test (IPGTT) in 22-week-old mice (n = 8/group). IP: IDH1-WT vs. IDH1-4KR; IIP: IDH1-WT vs. IDH1-4KQ. g Representative images of liver morphology (top) and liver sections stained with H&E (middle), or Oil Red O (bottom) are shown. Scale bars: 100 μm. h, i NAS score (h) and quantification of the Oil Red O staining of liver sections (i) from each group (n = 8/group). The stained area of each liver was normalized to the control group average. j, k Liver weight (j) and liver index (k) of the mice (n = 8/group) described in (a) were measured. l IDH1 activity assay in the livers of the mice (n = 8/group) described in (a). m, n Histogram showing Isocitrate (m) and α-KG (n) levels in the livers of mice (n = 8/group) indicated in (a). o Histogram displaying NADP+/NADPH ratio in the livers of mice (n = 8/group) indicated in (a). p FAO efficiency in the livers of the mice (n = 8/group) described in (a). q–s Histogram showing hepatic total cholesterol (TC) (q), triglyceride (TG) (r) and Free fatty acids (FFA) (s) concentrations in the livers of mice (n = 8/group) indicated in (a). Data are shown as the mean ± SD. P values were obtained using one-way ANOVA followed by Tukey’s multiple comparisons test. Source data are provided as a Source Data file.
We tested the effects of Kcr on IDH1 enzymatic activity in vivo. IDH1 enzymatic activity was significantly higher in the IDH1-WT group than in the Vector group, while IDH1-4KR overexpression reduced activity versus IDH1-WT, and IDH1-4KQ group markedly enhanced it (Fig. 7l). We then measured IDH1 upstream and downstream metabolites and FAO efficiency in mouse liver tissues. Isocitrate levels were elevated in the Vector group compared to the IDH1-WT group, further increased in the IDH1-4KR group, and significantly decreased in the IDH1-4KQ group (Fig. 7m). Conversely, α-KG levels were reduced in the Vector and IDH1-4KR groups compared to the IDH1-WT group, but significantly increased in the IDH1-4KQ group (Fig. 7n). Above all, isocitrate and α-KG levels indicated IDH1 enzymatic activity. Elevated IDH1 activity in mouse liver tissue corresponded to increased α-KG and decreased isocitrate levels, whereas reduced IDH1 activity showed the opposite trend. NADP+/NADPH ratio showed a progressive increase in the Vector group relative to the IDH1-WT group, with further elevation in the IDH1-4KR group, while the IDH1-4KQ group exhibited a significant reduction (Fig. 7o). Compared to IDH1-WT mice, IDH1-4KR mice exhibited lower FAO efficiency, whereas IDH1-4KQ mice showed the reverse (Fig. 7p). TC, TG, and FFA levels in mouse liver tissue showed a consistent pattern, elevated in the Vector group compared to the IDH1-WT group, significantly increased with IDH1-4KR overexpression, and significantly decreased with IDH1-4KQ overexpression (Fig. 7q–s). The blood analysis results in mice were consistent with the liver tissue findings (Supplementary Fig. 12b). These data support a physiological protective role for IDH1 Kcr in HFD-induced obesity and MASLD.
PCAF/SIRT7-mediated IDH1 Kcr is impaired in the livers both MASLD mice and patients
To further validate the LC-MS/MS results and confirm the expression patterns of PCAF/SIRT7-IDH1 Kcr axis in MASLD, we established a second mouse cohort independently replicated with the first cohort shown in Supplementary Fig. 1 (Supplementary Fig. 13, n = 5 per group). Comprehensive analysis of liver tissues from both cohorts demonstrated that IDH1 expression remained stable at both mRNA and protein levels in HFD-fed MASLD and post-bariatric surgery conditions (Fig. 8a–d). Specifically, MASLD induced a loss of IDH1 Kcr at K58, K151, K212, and K345, which was rescued by VSG treatment (Fig. 8a-d). We also detected decreased PCAF levels and increased SIRT7 levels in the livers of HFD-fed MASLD mice than in the livers from the corresponding controls, as well as increased PCAF levels and decreased SIRT7 levels in the livers of VSG mice than in the livers from PAIR-FED group (Fig. 8a–d). These in vivo findings not only corroborated our previous observations in PA + OA-treated cells, but also aligned with reported metabolic characteristics of Sirt7 KO mice, which exhibit resistance to HFD-induced obesity and hepatic steatosis, contrasting sharply with other sirtuin deficiencies40. Collectively, our results demonstrate that impaired PCAF/SIRT7-regulated IDH1 Kcr is mechanistically linked to MASLD progression at the animal level and can be therapeutically targeted by VSG intervention.
a Protein lysates from the livers of NCD, HFD, VSG, and PAIR-FED mice (n = 5/group) from first cohort were extracted for western blotting using IDH1-K58cr, IDH1-K151cr, IDH1-K212cr, IDH-K345cr, PCAF, SIRT7, IDH1 and α-Tubulin antibodies. b Relative mRNA levels of Pcaf (left), Sirt7 (middle), and Idh1(right) in NCD, HFD, VSG and PAIR-FED mice from first cohort. c Protein lysates from the livers of NCD, HFD, VSG, and PAIR-FED mice (n = 5/group) from second cohort were extracted for western blotting using IDH1-K58cr, IDH1-K151cr, IDH1-K212cr, IDH-K345cr, PCAF, SIRT7, IDH1 and α-Tubulin antibodies. d Relative mRNA levels of Pcaf (left), Sirt7 (middle), Idh1 (right) in NCD, HFD, VSG and PAIR-FED mice from second cohort. Data are shown as the mean ± SD. P values were obtained using one-way ANOVA followed by Tukey’s multiple comparisons test. All experimental data were verified at least three independent experiments. Source data are provided as a Source Data file.
To comprehensively evaluate the role of IDH1 Kcr in MASH progression, we employed two distinct liver fibrosis models: Gubra Amylin NASH (GAN) diet-induced obese MASH mice (Fig. 9a, b, d and f) and methionine choline-deficient (MCD) diet-induced lean MASH mice with liver fibrosis (Fig. 9g, h, j and l). Strikingly, both models recapitulated our key findings from MASLD mice. Compared to NCD mice, neither GAN nor MCD diet-induced MASH mice exhibited significant changes in IDH1 protein or mRNA levels, whereas IDH1 Kcr was significantly diminished (Fig. 9c, e, i and k). Consistent with our previous observations, PCAF was downregulated whereas SIRT7 was upregulated in both obese (GAN) and lean (MCD) MASH mice (Fig. 9c, e, i and k). These parallel results across three different diseases models (HFD-MASLD, GAN-MASH, MCD-MASH) strongly suggest that dysregulation of the PCAF/SIRT7-IDH1 Kcr axis represents a conserved molecular mechanism contributing to MASLD progression into MASH.
a Schematic overview of the GAN-MASH models. Created in BioRender. Liu, S. (2025) https://BioRender.com/vr8ptir. b Photos of NCD and GAN group mice at 34 weeks. c Protein lysates from the livers of NCD and GAN mice were extracted for western blotting using IDH1-K58cr, IDH1-K151cr, IDH1-K212cr, IDH-K345cr, IDH1, PCAF, SIRT7, α-SMA and α-Tubulin antibodies. d Body weight of NCD and GAN group mice (n = 10/group) at 34 weeks. e Relative mRNA levels of Pcaf, Sirt7, and Idh1 in NCD and GAN mice. f Representative images of liver tissues from NCD and GAN group mice (n = 10/group) were presented, including H&E-stained sections, Oil Red O staining, Masson’s trichrome staining, Sirius red staining, NAS scores, and relative Oil Red O area. Scale bars: 100 μm. g Schematic overview of the MCD-MASH models. Created in BioRender. Liu, S. (2025) https://BioRender.com/vr8ptir. h Photos of NCD and MCD group mice at 14 weeks. i Protein lysates from the livers of NCD and MCD mice were extracted for western blotting using IDH1-K58cr, IDH1-K151cr, IDH1-K212cr, IDH-K345cr, IDH1, PCAF, SIRT7 α-SMA and α-Tubulin antibodies. j Body weight of NCD and MCD group mice (n = 10/group) at 14 weeks. k Relative mRNA levels of Pcaf, Sirt7, and Idh1 in NCD and MCD mice. l Representative images of liver tissues from NCD and MCD group mice were presented, including H&E-stained sections, Oil Red O staining, Masson’s trichrome staining, Sirius red staining, NAS scores, and relative Oil Red O area. Scale bars: 100 μm. Data are shown as the mean ± SD. P values were obtained using two-tailed unpaired Student’s t-test. Data were verified in three (c, e, i, k) independent experiments. Source data are provided as a Source Data file.
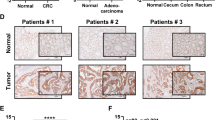
To confirm the clinical relevance of our findings, we next analyzed liver tissues from healthy donors, MASLD patients, and MASH patients (Fig. 10a). Western blotting analysis of key molecular markers revealed that IDH1 Kcr levels showed a progressive decline from healthy controls to MASLD and further to MASH patients, suggesting a potential role in disease progression (Fig. 10b). Additionally, we observed a consistent pattern of decreased PCAF and increased SIRT7 expression in patient samples (Fig. 10b, c and Supplementary Fig. 14a), which mirrored the corresponding changes in mRNA and protein levels observed in our cellular and murine models. Moreover, we studied the association between the elevated hepatic SIRT7 levels and the MASLD progression in publicly available transcriptome profiles of two independent cohorts comprising patients with the full histological spectrum of MASLD (Fig. 10d and Supplementary Fig. 14b) (GSE130970 and GSE135251). However, PCAF showed no significant change, suggesting SIRT7-dependent IDH1 Kcr downregulation during the MASLD progression.
a Representative images of liver tissues from healthy controls, MASLD and MASH patients were presented, including H&E-stained sections, Oil Red O staining, Masson’s trichrome staining and Sirius red staining. Scale bars: 100 μm. b Protein lysates from the liver tissues from healthy, MASLD and MASH patients were extracted for western blotting using IDH1-K58cr, IDH1-K151cr, IDH1-K212cr, IDH-K345cr, IDH1, PCAF, SIRT7, α-SMA and α-Tubulin antibodies. c Relative mRNA expression levels of PCAF and SIRT7 in liver tissues from healthy and MASLD patients (n = 16/group). d Hepatic PCAF and SIRT7 levels were associated to MASLD progression in two independent cohorts (GSE135251, GSE130970). e Schematic outline illustrates the mechanisms that the PCAF/SIRT7-mediatded IDH1 Kcr alleviates MASLD progression by enhancing the TCA cycle. Created in BioRender. Liu, S. (2025) https://BioRender.com/rxqskmj. Data are shown as the mean ± SD. P values were obtained using two-tailed unpaired Student’s t-test. All data were verified in three independent experiments. Source data are provided as a Source Data file.
Our multi-layered evidence from cellular, animal, and clinical models demonstrates that PCAF/SIRT7-regulated IDH1 Kcr underlies MASLD pathogenesis, while VSG intervention rescues this defect. These results position the PCAF/SIRT7-IDH1 Kcr axis as an evolutionarily conserved mechanism-based therapeutic strategy and diagnostic biomarker for MASLD/MASH progression.
Discussion
The widespread prevalence of MASLD, the attendant risks for metabolic and hepatic complications8,13, and the lack of approved therapies are significant hurdles, but there is no solid and mechanistic foundation for tackling this challenge. In this study, we revealed a significantly altered protein Kcr in the livers of HFD-induced MASLD mice and in the livers of VSG-improved hepatic steatosis. We identified 17,408 Kcr sites in 3913 proteins, representing by far the first crotonylome dataset associated with MASLD (Fig. 1). Our study expands the depth of Kcr proteome and provides a comprehensive picture of Kcr of proteins in diverse cellular pathways. Importantly, Kcr of enzymes in cellular metabolism, especially TCA cycle, glycolysis, and fatty acid metabolism, exhibits a response to MASLD progression. Therefore, our findings indicate a direct link between the metabolic enzyme Kcr and MASLD pathogenesis, demonstrating that Kcr serves as a crucial modulator in both cellular metabolism and epigenetic regulation during MASLD progression. MASLD is the fastest-growing cause of HCC2 and the key role of Kcr in HCC has been identified27,28, but its function in non-malignant conditions like MASLD remains unanswered.
It is reported that the alterations in dysregulated TCA cycle activity may play a central role in the pathogenesis of MASLD41,42. Among these proteins in TCA cycle, the core target of Kcr modification, IDH1, was also identified, which is a highly conserved protein among species and can convert isocitrate to α-KG in the TCA cycle. We found that Kcr affected IDH1 activity as well as the formation of IDH1 dimer, though it did not affect its localization (Fig. 5a, Supplementary Fig. 11g, h). More importantly, with the block of IDH1 Kcr in MASLD, the IDH1 activity was partially lost, which led to the reduction of the downstream metabolite α-KG. However, this phenomenon was rescued after the increase of IDH1 Kcr, suggesting that the change of α-KG content might be a potential target for the treatment of MASLD. Loss of IDH1 Kcr resulted in reduced NADPH production, which subsequently elevated ROS levels. Concurrently, OCR was diminished, while ECAR was increased, indicating a metabolic shift toward glycolysis. Furthermore, the inhibition of FAO led to the accumulation of free fatty acid (FFA), TC and TG. These combined effects exacerbated the progression of MASLD.
IDH1 can be regulated by various PTMs including phosphorylation, acetylation and ubiquitination32,43,44. For example, IDH1 has been reported to undergo phosphorylation at Y135 and Y139, which is co-clustered with R132H and R132C mutants and associated with several cancers33. In addition, recent studies indicate that SIRT2-dependent IDH1 K224 deacetylation regulates cellular metabolism and inhibits liver metastasis of colorectal cancer32. In our study, we found that K58cr, K151cr, K212cr and K345cr are involved in TCA cycle during HFD-induced or bariatric surgery-improved MASLD. We are also aware that K58, K151, K212 and K345 have been described previously as ubiquitylated (K58 and K345 are also acetylated and K151 is also SUMOylated) by proteomic studies, though their functions have not been characterized43,45. We have performed Immunoprecipitation assays in AML12 cells and found that PA + OA did not affect the level of IDH1 ubiquitination, SUMOylation and acetylation (Fig. 3d and Supplementary Fig. 7f), reinforcing the notion that IDH1 Kcr was largely performed under steatotic conditions. Owing to the fact that our data regarding the function of Kcr on these sites are mainly derived from K to R (or Q) mutagenesis analysis, we could not fully exclude the possibility that other modifications of these sites may also play a role during MASLD. Further characterization of the regulator that targets IDH1 in response to MASLD and deciphering their mechanistic details would help to better clarify the function of Kcr of IDH1 on these sites. Clearly, how multiple PTMs coordinately regulate the function of IDH1 in different cell metabolism events remains to be further investigated.
Next, we demonstrated PCAF and SIRT7 are the main regulators of Kcr of IDH1 on these sites, we are aware that the level of IDH1 Kcr could be regulated by other factors, such as crotonyltransferases and/or decrotonylases, which could also contribute to the regulation of dynamics of IDH1 Kcr during MASLD progression. In addition, beased on patients’ results (Fig. 10a–d and Supplementary Fig. 14c) and by querying published clinical data sets showed a clear upregulation of SIRT7 and no significant change of PCAF, suggesting that targeting SIRT7-dependent IDH1 decrotonylation might provide a potential therapeutic approach against MASLD development (Fig. 10d). Similarly, using The Cancer Genome Atlas (TCGA) database, we found that the hepatic SIRT7 mRNA levels were upregulated in HCC (Supplementary Fig. 14c). The role of SIRT7-dependent IDH1decrotonylation in the progression from early MASLD to MASH, as well as from non-malignant hepatic diseases to HCC, requires further investigation.
Bariatric surgery ameliorates obesity and MASLD, but the molecular mechanism of VSG-improved MASLD remains unclear46. The comparison of liver in mice before and after bariatric surgery showed that MASLD-associated Kcr changes could be partially reversible. Especially, the protein levels of PCAF and SIRT7 and Kcr of IDH1 were changed by VSG intervention (Fig. 8), suggesting that there exists a potential target for explaining the reduction of hepatic steatosis after VSG. We are aware that other Kcr proteins also play an important role after VSG. More similar proteomic analyses combined with focused functional analyses in the future might help decode the crotonylome and the role for Kcr in the regulation of multiple metabolic parameters following VSG. In addition, whether other bariatric-metabolic surgeries, like Roux-en-Y gastric bypass (RYGB), exhibit similar effects on MASLD is unclear. Likewise, the epigenetic remodeling signature after bariatric-metabolic surgery provides a data set of epigenetic organ remodeling in mice and may reinforce our understanding of liver regeneration.
Collectively, we demonstrated that IDH1 is the core target of HFD-induced and bariatric surgery-improved MASLD-associated Kcr signature. Mechanistically, the crotonylation-mimic mutant of IDH1 enhances its enzymatic activity and protects cells against hepatic steatosis, whereas the crotonylation-null mutant of IDH1 reduces the Kcr of IDH1 and impairs TCA cycle function. PCAF and SIRT7 are key regulators of IDH1 Kcr. Given the great need for therapeutic targets and pharmacotherapies for MASLD, these findings highlight anti-IDH1 Kcr as a potential pharmaceutical target for the treatment of MASLD.
Methods
Cell culture and transfection
AML12, HepG2 and HEK293T cells were obtained from the American Type Culture Collection (ATCC, Manassas, Virginia, USA). AML12 cells were maintained in Dulbecco’s Modified Eagle Medium/Nutrient Mixture F-12 (DMEM: F-12) (Gibco) supplemented with 10% fetal bovine serum (FBS) (Hyclone), ITS supplement (5 μg/mL insulin, 5 μg/mL transferrin, 5 ng/mL selenium; Collaborative Biomedical Products, Bedford, MA), 40 ng/mL dexamethasone, and 1% (100 U/mL-100 μg/mL) penicillin-streptomycin (HyClone). HepG2 cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (ThermoFisher Scientific) supplemented with 10% FBS and 1% penicillin-streptomycin (HyClone). HEK293T cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (Gibco) supplemented with 10% FBS and 1% penicillin-streptomycin (HyClone). RAW264.7 cells were maintained in DMEM (Gibco) containing 10% FBS and 1% penicillin-streptomycin (HyClone). LX-2 cells were cultured in DMEM (Gibco) containing 10% FBS, 1% penicillin-streptomycin (HyClone), and 2 mM L-glutamine. Mouse primary hepatic cells (MPHs) were cultured in William’s E medium supplemented with 10% FBS, 1% penicillin-streptomycin (HyClone) and primary hepatocyte maintenance supplement (Gibco). The cells were incubated at 37 °C in a humidified chamber containing 5% CO2. To induce lipid deposition, cells were treated with 0.5-mM palmitic acid (PA) and oleic acid (OA) for 24 h at a ratio of 1:2(V/V) PA: OA. All transfections were carried out using LipofectamineTM 3000 transfection reagent (ThermoFisher Scientific) according to the manufacturer’s recommendations. Cells were transfected with siRNA oligonucleotides using LipofectamineTM RNAiMAX (ThermoFisher Scientific) with the final concentration at 25 nM. Sequences of siRNA oligonucleotides are listed in Supplementary Data 13.
Hepatocyte isolation and culture
Hepatocytes were isolated by a two-step collagenase perfusion technique with modifications. Briefly, the inferior vena cava (IVC) was cannulated with a 24-gauge 3/4-inch angiocatheter (BD) and the portal vein was cut. The liver was perfused via the inferior vena cava with 100 mL of Liver Perfusion Medium (LP, Invitrogen) at 37 °C, followed by perfusion with 100 mL of collagenase type IV (Wellington) in HBSS containing Ca2+ and Mg2+ (GIBCO). After the liver was digested, it was dissected out and cut into small pieces and passed through a 100 μm strainer (Falcon). Hepatocytes were separated from non-parenchymal cells (NPCs) by low-speed centrifugation (50 g × 5 mins × 3, brake = 2), and further purified by Percoll gradient centrifugation (50 % v/v, Sigma)47,48.
Patients and clinical specimens
Human liver samples were obtained from 41 individuals (15 men, 26 women, aged more than 18 years) who underwent laparoscopic cholecystectomy at the Beijing Friendship Hospital (Beijing, China). Written informed consent was obtained from all patients before their enrollment, and the study protocol was approved by the Human Institutional Review Board of Beijing Friendship Hospital (approval numbers 2025-P2-052-02). Liver samples from individuals who engaged in excessive alcohol consumption (> 140 g for men or > 70 g for women, per week) or drug abuse, or who had a viral infection (e.g., hepatitis B virus or hepatitis C virus), were excluded. A total of 22 patients were defined as exhibiting characteristic fatty liver changes. Samples without fibrosis were included in the simple steatosis group, while samples showing fibrosis were classified as the MASH group. Studies using human specimens were approved by the Beijing Friendship Hospital.
Mice models
All mice used were the C57BL/6 J background. Six-week-old male mice (weight, 18–20 g) were purchased from HFK Bio Inc (Beijing, China). Mice were randomly divided into different groups and housed in a specific pathogen-free environment with the temperature ranging from 20 °C to 25 °C under a 12 h:12 h light/dark cyIcle with ad libitum access to water. To establish MASLD models, mice were fed a high-fat diet (HFD) (protein, 18.1%; fat, 61.6%; and carbohydrates, 20.3%; Research Diet, New Jersey, USA) for 12 weeks or 20 weeks49. Control group mice were fed a normal chow diet (NCD) (protein, 18.3%; fat, 10.2%; and carbohydrates, 71.5%; HFK Bio Inc, Beijing, China). The high-fat mice were randomized into the vertical sleeve gastrectomy (VSG), PAIR-FED, and High fat diet (HFD) groups (n = 5/cohort, two complete experimental repeats at 1-month intervals). In the PAIR-FED group, the mice underwent a sham operation and received the same type and quantity of food as that given to the VSG group mice on the previous day50. In the HFD group, the mice underwent a sham operation and received a high-fat diet freely. The NCD group mice underwent a sham operation and were continuously fed a normal chow diet (n = 5/cohort, two complete experimental repeats at 1-month intervals). To establish MASH models, mice were fed Gubra Amylin NASH (GAN) diet (high transfat, 44%; high cholesterol, 2%; high fructose, 22%, Dyets, USA) for 28 weeks (n = 10)51. We also fed mice with methionine-choline-deficient (MCD) diet (21% fat, 63% carbohydrate, 16% protein, Research Diet, New Jersey, USA, A02082002BR) for 6 weeks (n = 10) to establish lean MASH combined liver fibrosis mice52. For hepatocyte-specific IDH1 overexpression in C57BL/6 mice, mice were injected via the tail vein with adeno-associated virus 8 (AAV8)-thyroxine binding globulin promoter (TBG)-ZsGreen, AAV8-IDH1-WT, AAV8-IDH1-4KR and AAV8-IDH1-4KQ. Mice were administered with 3 × 1011 genome copies/mouse of AAV in 100 μL phosphate buffered saline (PBS) at 6 weeks, followed by feeding high-fat diet for 22 weeks; for each AAV8-IDH1 group, n = 8 mice. Body weight was measured weekly, and intraperitoneal glucose tolerance tests (IPGTT) and insulin tolerance tests (ITT) were conducted following a 12-hour fast. At the end of the study, mice were euthanized by sodium pentobarbital injection (100 mg/kg, i.p.), with subsequent collection of liver, blood, and adipose tissues. The animal studies were performed in compliance with ethical guidelines for animal studies and approved by the Institutional Animal Care and Ethics Committee of Beijing Friendship Hospital (approval numbers 20-2033).
Mice surgery
Perioperative adaptation and intervention were performed according to a previously published protocol50. All mice received sustained-release meloxicam (1 mg/kg, Mobic®, Boehringer Ingelheim, Germany) before surgery for pain control and Avertin (20 mg/mL, Sigma-Aldrich, USA) for anesthesia. In mice undergoing VSG, the lateral 80% of the stomach was isolated by an electric scalpel, followed by cutting parallel to the smaller curvature and using double-layer sutures (continuous full-thickness and Lembert sutures) to construct the sleeve. Sham surgery had analogous manipulation and isolation of the stomach but without excision. Mice were maintained on a liquid diet (Two Cal® HN, Abbott Nutrition, USA) until 3 days postoperatively. Solid HFD or NCD were reintroduced to mice on postoperative day 4. Their body weight was measured at a fixed time every week for 8 weeks postoperatively.
Protein extraction and trypsin digestion
The sample was grinded with liquid nitrogen into cell powder and then transferred to a 5 mL centrifuge tube. After that, four volumes of lysis buffer (1% Triton X-100, 1% protease inhibitor cocktail, 3 μM TSA and 50 mM NAM) was added to the cell powder, followed by sonication three minutes on ice using a high intensity ultrasonic processor (Scientz). The remaining debris was removed by centrifugation at 12,000 g at 4 °C for 10 min. The supernatant was collected, and the protein concentration was determined with BCA kit according to the manufacturer’s instructions.
For digestion, the sample was slowly added to the final concentration of 20% (m/v) TCA to precipitate protein, then vortexed to mix and incubated for 2 h at 4 °C. The precipitate was collected by centrifugation at 4500 g for 5 min at 4 °C. The precipitated protein was washed with 200 mM triethylammonium bicarbonate (TEAB) and ultrasonically dispersed. Trypsin was added at 1:50 trypsin-to-protein mass ratio for the first digestion overnight. The sample was reduced with 5 mM dithiothreitol for 60 min at 37 °C and alkylated with 11 mM iodoacetamide for 45 min at room temperature in darkness. Finally, the peptides were desalted by Strata X SPE column.
HPLC fractionation
The tryptic peptides were fractionated into fractions by high-pH reverse-phase HPLC using a Thermo Betasil C18 column (particle size, 5 μm; inside diameter, 4.6 mm; length, 250 mm). Briefly, peptides were first separated with a 2% to 60% acetonitrile gradient in 10 mM ammonium bicarbonate (pH 10) over 60 min into 60 fractions. Then, the peptides were combined into nine fractions and dried by vacuum centrifugation.
Kcr peptide enrichment
Tryptic peptides were dissolved in the NETN buffer (100-mM NaCl, 1-mM EDTA, 50-mM Tris-HCl, and 0.5% NP-40; pH 8.0) and then incubated with pre-washed crotonyllysine (Kcr) antibody conjugated agarose beads (Lot number PTM503, PTM Bio) at a ratio of 15 μL of beads per milligram of protein at 4 °C overnight. The antibody beads were washed four times with the NETN buffer and twice with deionized water. The beads-bound Kcr peptides were then eluted three times using 0.1% trifluoroacetic acid (TFA). Before being subjected to LC-MS/MS analysis, the eluted peptides were cleaned with C18 ZipTips (Millipore), according to the manufacturer’s instructions.
LC-MS/MS analysis
The tryptic peptides were dissolved in solvent A, directly loaded onto a home-made reversed-phase analytical column (25 cm length, 100 μm i.d.). The mobile phase consisted of solvent A (0.1% formic acid, 2% acetonitrile/in water) and solvent B (0.1% formic acid in acetonitrile). Peptides were separated with following gradient: 0–40 min, 7–24%B, 40–52 min, 24–32%B, 52–56 min, 32–80%B, 56–60 min, 80%B, and all at a constant flow rate of 450 nl/minon a NanoElute UHPLC system (Bruker Daltonics).
The peptides were subjected to capillary source followed by the timsTOF Pro mass spectrometry. The electrospray voltage applied was 1.65 kV. Precursors and fragments were analyzed at the TOF detector, with a MS/MS scan range from 100–1700. The timsTOF Pro was operated in parallel accumulation serial fragmentation (PASEF) mode. Precursors with charge states 0–5 were selected for fragmentation, and 10 PASEF-MS/MS scans were acquired per cycle. The dynamic exclusion was set to 24 s.
Database searching and protein quantification
The resulting MS/MS data were processed using MaxQuant search engine (v.1.6.15.0). Tandem mass spectra were searched against the Mus_musculus_10090_SP_20210721.fasta (17089 entries) concatenated with reverse decoy and contaminants database. Trypsin/P was specified as cleavage enzyme allowing up to 2 missing cleavages. Min. peptide length was set as 7 and max. number of modification per peptide was set as 5. The mass tolerance for precursor ions was set as 20 ppm in first search and 20 ppm in main search, and the mass tolerance for fragment ions was set as 20 ppm. Carbamidomethyl on Cys was specified as fixed modification. Acetylation on protein N-terminal, oxidation on Met and Kcr were specified as variable modifications. False discovery rate (FDR) of protein, peptide and PSM was adjusted to <1%.
Bioinformatics analysis
The motif characteristics of modification sites were analyzed by software MoMo based on motif-x algorithm. The peptide sequences composed of 10 amino acids upstream and downstream of all identified modification sites were selected; The analysis background are the peptide sequences composed of 10 amino acids upstream and downstream of all potential modification sites. When the number of peptide segments of a characteristic sequence is greater than 20 and the statistical test p value is less than 0.000001, it is considered that the sequence is one of the motifs of the modified peptide segments.
The amino acid sequence heatmap is a 2D data matrix where every row is an amino acid and every column is a position, wherein the gradient shows if log10 P values correlate with the color. Fisher’s exact test method calculated the position and amino acid-specific P values. The IceLogo software (https://iomics.ugent.be/icelogoserver/) was used to examine the properties of amino acids surrounding modification sites; the statistical test was set to “t-test,” P values were set to “<0.05,” and the “Choosing scoring system” was set to “percentage difference.” GO functional and KEGG pathway enrichment were performed using DAVID 6.853,54. The interacting genes/proteins database (STRING; http://string-db.org/) was retrieved for PPI analysis. The Cytoscape (version 3.0) software was used to display the network55.
Immunoprecipitation and western blotting
Briefly, cellular lysates were prepared by incubating the cells in lysis buffer [50-mM Tris-HCl, 150-mM NaCl, 0.5% NP-40, and 2-mM EDTA (pH 7.5)] or BC100 buffer [20-mM Tris-HCl (pH 8.0), 100-mM NaCl, 1-mM EDTA, and 0.5% Triton] containing protease inhibitor cocktail for 20 min at 4 °C, followed by centrifugation at 14,000 g for 15 min at 4 °C. Protein concentration of the lysates was determined using a BCA protein assay kit (Pierce) according to the manufacturer’s protocol. Overall, 5% (1:20) cellular extracts were used for input. For immunoprecipitation, 500-μg of protein lysates was incubated with 2-μg of specific antibodies for 12 h at 4 °C with constant rotation; 60-μL of 50% protein A or G agarose beads were then added, and the incubation was continued for an additional 2 h. Beads were then washed five times using the lysis buffer. Between washes, the beads were collected by centrifugation at 500 g for 5 min at 4 °C. The precipitated proteins were eluted from the beads by resuspending the beads in 2× SDS–polyacrylamide gel electrophoresis (PAGE) loading buffer and boiling for 10 min. The resultant materials from immunoprecipitation or cell lysates were resolved using 10% SDS-PAGE gels and transferred onto nitrocellulose membranes. For western blotting, membranes were incubated with appropriate antibodies for 1 h at room temperature or overnight at 4 °C followed by incubation with a secondary antibody. Immunoreactive bands were visualized using a western blotting luminol reagent (Santa Cruz Biotechnology) according to the manufacturer’s recommendation15. The antibodies used were listed in Supplementary Data 13.
Real-time reverse transcription-PCR (RT-PCR) assay
Total cellular RNA was isolated with Trizol reagent (Invitrogen) and used for first-strand cDNA synthesis with the Reverse Transcription System (Roche). Quantitation of all gene transcripts was performed by qPCR using a Power SYBR Green PCR Master Mix (Roche) and an ABIPRISM 7500 sequence detection system (Applied Biosystems) with the expression of β-actin as the internal control. Primers used for real-time PCR are listed in Supplementary Data 12.
D-Glucose-13C6 labeling metabolite flux assay
For D-Glucose-13C6 labeling metabolite flux assay, the culture medium was composed of glucose free DMEM (Gibco) supplemented with 2 g/L D-Glucose-13C6, 10% FBS, and 1% penicillin‒streptomycin. The medium was adjusted to pH 7.0 − 7.4 and filtered and sterilized for usage. Cells were seeded at a density of approximately 2 × 106 cells per 10 cm dish. Labeling medium was used to replace unlabeled medium when cells grew to ~60% confluence. An unlabeled culture was prepared in parallel by adding equal concentrations of unlabeled glucose to the media to identify unlabeled metabolites. After incubation for 12 h, the medium was quickly and completely aspirated, the plates were immediately placed on dry ice, and 400 μL of 80% methanol (cooled to –80 °C) was added and incubated for 20 min on dry ice. Next, we sonicated the cell samples, transferred the cell lysate/methanol mixture to a 2 mL tube on dry ice, centrifuged at 18,000 g for 15 min at 4 °C, and again transferred the metabolite-containing supernatant to a new 2 mL tube. Then, 100 μL 80% methanol was added to resolve the samples. The metabolite residues were re-dissolved in 80% methanol and used for LC-MS/MS analysis56.
Metabolomics LC-MS/MS Analysis
Q Exactive orbitrap mass spectrometer coupled with Ultimate 3000 (Thermo Fisher Scientific, USA) was used for untargeted analysis. BEH amide column (2.1 mm × 100 mm, Waters, USA) was applied to LC separation. In the positive ion mode, mobile phases A and B contained 10 mM ammonium acetate in 95% and 50% acetonitrile respectively. Formic acid was used to adjust pH to 3.0. In the negative ion mode, pH of the mobile phases was adjusted to 9.0 using ammonium hydroxide solution. Mass resolutions of 70,000 and 17,500 were applied for precursors and fragments. Data dependent MS/MS acquisition with fragmenting the top ten most intense precursors was performed in the analysis. MS/MS spectra were acquired with stepped NCE of 15, 30 and 45. Raw data from the UPLC-TQMS system were collected, stored, identified, and quantified by MassLynx software (Waters). The following statistics were performed by open R software (v4.1.1).
Measurement of oxygen consumption rate (OCR) and extracellular acidification rate (ECAR)
OCR and ECAR were determined by the Agilent Seahorse XF Analyzer following the manufacturer’s protocol. For OCR measurement, 1 × 105 cells were plated in Seahorse XF24 assay plates and incubated in XF base medium (Agilent, 102353-100) overnight. The mitochondrial function was measured using the Agilent Seahorse XF cell mito stress test kit (Agilent, 103015-100). OCR was determined following the sequential addition of oligomycin (2.5 μM), FCCP (2 μM), and rotenone (0.5 μM) plus antimycin A (0.5 μM) to the system. For ECAR measurement, 3 × 104 cells were plated in SeahorseXF96 assay plates and incubated overnight. Agilent Seahorse XFp glycolytic rate assay kit (Agilent, 103344-100) was used to measure the glycolytic rate. ECAR was determined following the sequential addition of glucose (10 mM), oligomycin (1.5 μM), and 2-DG (50 mM) to the system. The cell number was used for data normalization57.
IDH1 activity assay
IDH1 activity was determined by the commercially available IDH1 activity detection kit (Sigma, MAK062). AML12 cells in 6-well plate were transfected with indicated plasmids for 48 h, then 1 × 106 cells were analyzed following instructions. Mouse liver tissues were analyzed following instructions39.
Measurement of isocitrate, α-KG, NADP+/NADPH, ATP and FAO
The Isocitrate of AML12 cells and mouse liver tissue were determined by isocitrate assay kit (Sigma). α-KG of AML12 cells and mouse liver tissue were determined by α-KG assay kit (Sigma). NADP+/NADPH of AML12 cells were determined by NADP+/NADPH assay kit (WST-8 method) (Beyotime). ATP of AML12 cells were determined by ATP assay kit (Beyotime). Fatty acid beta-oxidation (FAO) of AML12 were determined by FAO colorimetric Assay Kit (ELAB Science). All these detections were performed following instructions39.
Measurement of triglyceride, cholesterol and free fatty acid (FFA)
Triglycerides and cholesterol were measured in cell lysates, plasma and mouse liver tissue using a Triglyceride ELISA kit (LabAssay) and Cholesterol ELISA kit (LabAssay), respectively. Briefly, cells, plasma and mice livers were collected, sonicated, and centrifuged at 10,000 g at 4 °C for 10 min. The supernatant was incubated with Enzyme Buffer solution for 15 min at room temperature, and the absorbance was read at 600–700 nm using a plate reader. FFA of mouse liver tissue was determined by FFA assay kit (Solibao). All these detections were performed following instructions.
Immunofluorescence staining and confocal microscopy
AML12 cells were transfected with indicated plasmids for 48 h. Cells grown on coverslips were washed three times with PBS,and then fixed in 4% (w/v) paraformaldehyde for 10 min. Cells were permeabilized with 0.3% Triton X-100 in PBS for 30 min, blocked with 0.8% bovine serum albumin (BSA), incubated with appropriate primary antibodies coupled to anti-Myc tag (Abcam, 1:100) at 4 °C overnight, and then incubated with a secondary FITC/TRITC-conjugated antibody (Sigma-Aldrich, 1:1000) at room temperature for 30 min in the dark. Cells were then washed for four times, and a final concentration of 4’, 6-diamidino-2-phenylindole dihydrochloride (DAPI) (Sigma-Aldrich) was included in the final wash to stain nuclei. Confocal laser scanning microscopy images were obtained using a Zeiss LSM 700 microscope equipped with argon laser. The images were captured and analyzed using Adobe Photoshop CS4.
ROS detection
2’,7’-Dichlorodihydroflfluorescein diacetate (DCFH-DA, 10 μM, 4091-99-0, MCE, USA) or 2’-(4-ethoxyphenyl)-6-(4-methylpiperazin-1-yl)-1H,3’H-2,5’-bibenzo[d]imidazole trihydrochloride (Hoechst 33342, 10 μg/mL, 23491-52-3, MCE, USA) was loaded into 1 × 106 cells/100 μL, incubated at 37 °C for 20 min and washed twice with 1 × PBS. ROS production was detected with Immunofluorescence staining.
Oil red O staining
AML12 cells cultured in the presence or absence of PA and OA for 48 h were fixed with 2% paraformaldehyde in phosphate-buffered saline (PBS) for 15 min and treated with 0.18% Oil red O reagent (Shanghai Gefan Biotechnology)/60% isopropanol in ultra-pure water for 20 min at room temperature. Liver tissues were embedded with optimal cutting temperature compound, frozen and sliced at 5 μm. Sections were rinsed with 60% isopropanol (Shenbo Chemical Industry) followed by 15 min of Oil red O staining (Shanghai Gefan Biotechnology), along with isopropanol differentiation. Following H&E staining, cells were washed with distilled water and phosphate-buffered saline (PBS) before 10 min of fixation in 4% paraformaldehyde. Cells were subjected to PBS wash, 1 min of rinsing with 60% isopropanol, 10 min of Oil red O staining, and distilled water wash. After that, the colour reaction was terminated, and the cells were observed under an optical microscope. Oil red O staining images were analyzed using ImageJ software, with the %Area metric used to quantify intracellular lipid droplet content. %Area represents the proportion of the stained area relative to the total cell area, with higher values indicating greater lipid droplet content58.
Intraperitoneal glucose tolerance test (IPGTT)/Insulin tolerance test (ITT)
IPGTT: At 7 weeks postoperatively, mice from each group were subjected to IPGTT. Mice were fasted for 8 h before the meal (during the night) and then given 1.5 g/kg of glucose intraperitoneally. Then, to measure the glucose level, approximately 150 µL of blood sample was collected from the tail vein of each mouse at different time intervals (0, 30, 60, 90, and 120 min). The glucose–time curve was calculated with GraphPad Prism software using the area under the curve (AUC) values.
ITT: ITT was performed 7 weeks postoperatively. For this test, mice were fasted for 4 h before the meal (during the night) and intraperitoneally injected with 0.75 U/kg of Humulin R. The blood samples were collected from the tail vein after 0, 30, 60, 90, and 120 min of insulin injection to determine blood glucose concentration, and GraphPad Prism software was used to calculate the AUC.
Small-animal MRI scanning
Mice were anesthetized using isoflurane (induction: 4.5% isoflurane, 2 L/min O2; maintenance: 1.0–3.0% isoflurane, 1 L/min O2) for the duration of the scans (approximately 30 min). Adipose tissue was measured using MRI (PharmaScan 7 T, Bruker®, Billerica, USA). Image acquisition was synchronized with the respiratory cycle to minimize physiological artifacts59. Fat content was calculated by ImageJ.
Histological analysis
Liver samples of mice were fixed in formalin overnight and transferred into 70% ethanol. Then, the samples were paraffin-embedded for hematoxylin and eosin (H&E) staining60. PicroSirius Red staining was performed using a 0.1% Direct Red 80 solution (VWR #B21693.14) in saturated aqueous solution of picric acid (Merck #621). Masson’s trichrome staining was subsequently conducted using Light Green SF Yellowish (VWR #B23330.09) according to established protocols61,62. For histological assessment, two blinded experienced pathologists independently evaluated H&E-stained sections to determine NAFLD activity scores (NAS). The NAS was recorded as the unweighted sum of the scores for steatosis (0–3), lobular inflammation (0–3), and hepatocellular ballooning (0–2), thus ranging from 0 to 8. NAS of ≥5 correlated with a diagnosis of MASH, and biopsies with scores of less than 3 were diagnosed as “not MASH”63.
Statistical analysis
Data are presented as mean ± SD. GraphPad Prism 9.0 was chosen to analyze the results from at least three independent experiments. P < 0.05 was statistically significant. Statistical significance was assessed using a two-tailed t-test or Wilcoxon’s signed-rank test for comparisons between two groups, or one-way analysis of variance (ANOVA) followed by Tukey’s multiple comparisons test for comparisons involving multiple groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The publicly available data used in this study is available in the GEO database under accession code GSE135251, GSE130970 and TCGA database (https://portal.gdc.cancer.gov/projects/TCGA-LIHC). The mass spectrometry proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD065361 (https://www.iprox.cn/page/project.html?id=IPX0012364000)64,65. The D-Glucose-13C6 labeling metabolite flux data generated in this study have been deposited in the OMIX, China National Center for Bioinformation/ Beijing Institute of Genomics, Chinese Academy of Sciences under accession code OMIX010635. All data supporting the findings described in this manuscript are available in the article and in the Supplementary Information and from the corresponding author upon request. Source data are provided with this paper.
References
Powell, E. E., Wong, V. W.-S. & Rinella, M. Non-alcoholic fatty liver disease. Lancet 397, 2212–2224 (2021).
Loomba, R., Friedman, S. L. & Shulman, G. I. Mechanisms and disease consequences of nonalcoholic fatty liver disease. Cell 184, 2537–2564 (2021).
Rinella, M. E. et al. A multisociety Delphi consensus statement on new fatty liver disease nomenclature. J. Hepatol. 79, 1542–1556 (2023).
Quek, J. et al. Global prevalence of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis in the overweight and obese population: a systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 8, 20–30 (2023).
Gallego-Duran, R. Romero-Gomez M. Epigenetic mechanisms in non-alcoholic fatty liver disease: An emerging field. World J. Hepatol. 7, 2497–2502 (2015).
Eslam, M. et al. A new definition for metabolic dysfunction-associated fatty liver disease: An international expert consensus statement. J. Hepatol. 73, 202–209 (2020).
Wu, X. et al. Targeting protein modifications in metabolic diseases: molecular mechanisms and targeted therapies. Signal Transduct. Target Ther. 8, 220 (2023).
Samuel, V. T. & Shulman, G. I. Nonalcoholic Fatty Liver Disease as a Nexus of Metabolic and Hepatic Diseases. Cell Metab. 27, 22–41 (2018).
Bhat, N. & Mani, A. Dysregulation of Lipid and Glucose Metabolism in Nonalcoholic Fatty Liver Disease. Nutrients 15, 2323 (2023).
Sharma, U. & Rando, O. J. Metabolic Inputs into the Epigenome. Cell Metab. 25, 544–558 (2017).
Perdomo, C. M., Cohen, R. V., Sumithran, P., Clément, K. & Frühbeck, G. Contemporary medical, device, and surgical therapies for obesity in adults. Lancet 401, 1116–1130 (2023).
Geerts, A. & Lefere, S. Bariatric surgery for non-alcoholic fatty liver disease: Indications and post-operative management. Clin. Mol. Hepatol. 29, S276–s285 (2023).
Bayoumi, A., Gronbaek, H., George, J. & Eslam, M. The Epigenetic Drug Discovery Landscape for Metabolic-associated Fatty Liver Disease. Trends Genet 36, 429–441 (2020).
Tan, M. J. et al. Identification of 67 Histone Marks and Histone Lysine Crotonylation as a New Type of Histone Modification. Cell 146, 1015–1027 (2011).
Yu, H. et al. Global crotonylome reveals CDYL-regulated RPA1 crotonylation in homologous recombination-mediated DNA repair. Sci. Adv. 6, eaay4697 (2020).
Liu, S. et al. Chromodomain Protein CDYL Acts as a Crotonyl-CoA Hydratase to Regulate Histone Crotonylation and Spermatogenesis. Mol. Cell 67, 853–866.e855 (2017).
Sabari, B. R. et al. Intracellular Crotonyl-CoA Stimulates Transcription through p300-Catalyzed Histone Crotonylation. Mol. Cell 58, 203–215 (2015).
Liu, X. G. et al. MOF as an evolutionarily conserved histone crotonyltransferase and transcriptional activation by histone acetyltransferase-deficient and crotonyltransferase-competent CBP/p300. Cell Discov. 3, 17016 (2017).
Xu, W. et al. Global profiling of crotonylation on non-histone proteins. Cell Res. 27, 946–949 (2017).
Liao, W. et al. Persistent high glucose induced EPB41L4A-AS1 inhibits glucose uptake via GCN5 mediating crotonylation and acetylation of histones and non-histones. Clin. Transl. Med. 12, e699 (2022).
Xiao, Y. H. et al. HBO1 is a versatile histone acyltransferase critical for promoter histone acylations. Nucleic Acids Res. 49, 8037–8059 (2021).
Song, X. et al. Dynamic crotonylation of EB1 by TIP60 ensures accurate spindle positioning in mitosis. Nat. Chem. Biol. 17, 1314–1323 (2021).
Yang, P. et al. Crotonylation and disease: Current progress and future perspectives. Biomed. Pharmacother. 165, 115108 (2023).
Wan, J., Liu, H., Chu, J. & Zhang, H. Functions and mechanisms of lysine crotonylation. J. Cell Mol. Med. 23, 7163–7169 (2019).
Fellows, R. et al. Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat. Commun. 9, 105 (2018).
Yu, A. Q. et al. SIRT7-Induced PHF5A Decrotonylation Regulates Aging Progress Through Alternative Splicing-Mediated Downregulation of CDK2. Front Cell Dev. Biol. 9, 710479 (2021).
Lao, Y. et al. Glutaryl-CoA dehydrogenase suppresses tumor progression and shapes an anti-tumor microenvironment in hepatocellular carcinoma. J Hepatol. 81, 847–861 (2024).
Zhang, D. et al. Global crotonylome reveals hypoxia-mediated lamin A crotonylation regulated by HDAC6 in liver cancer. Cell Death Dis. 13, 717 (2022).
Li, S. et al. Mutation of IDH1 aggravates the fatty acid‑induced oxidative stress in HCT116 cells by affecting the mitochondrial respiratory chain. Mol. Med Rep. 19, 2509–2518 (2019).
Chen, D. et al. Mutant and Wild-Type Isocitrate Dehydrogenase 1 Share Enhancing Mechanisms Involving Distinct Tyrosine Kinase Cascades in Cancer. Cancer Discov. 9, 756–777 (2019).
Pirozzi, C. J. & Yan, H. The implications of IDH mutations for cancer development and therapy. Nat. Rev. Clin. Oncol. 18, 645–661 (2021).
Wang, B. et al. SIRT2-dependent IDH1 deacetylation inhibits colorectal cancer and liver metastases. EMBO Rep. 21, e48183 (2020).
Huang, K.-l. et al. Spatially interacting phosphorylation sites and mutations in cancer. Nat.Commun. 12, 2313 (2021).
Zheng, Y. et al. Modulation of cellular metabolism by protein crotonylation regulates pancreatic cancer progression. Cell Rep. 42, 112666 (2023).
Chen, D. et al. Lysine acetylation restricts mutant IDH2 activity to optimize transformation in AML cells. Mol. Cell 81, 3833–3847.e3811 (2021).
Jiang, G., Li, C., Lu, M., Lu, K. & Li, H. Protein lysine crotonylation: past, present, perspective. Cell Death Dis. 12, 703 (2021).
Guo, Z. et al. Hypoxia-induced downregulation of PGK1 crotonylation promotes tumorigenesis by coordinating glycolysis and the TCA cycle. Nat. Commun. 15, 6915 (2024).
Zheng, X. et al. Histone acetyltransferase PCAF up-regulated cell apoptosis in hepatocellular carcinoma via acetylating histone H4 and inactivating AKT signaling. Mol. Cancer 12, 96 (2013).
Tateishi, K. et al. Extreme Vulnerability of IDH1 Mutant Cancers to NAD+ Depletion. Cancer Cell 28, 773–784 (2015).
Yoshizawa, T. et al. SIRT7 controls hepatic lipid metabolism by regulating the ubiquitin-proteasome pathway. Cell Metab. 19, 712–721 (2014).
Sunny, N. E., Parks, E. J., Browning, J. D. & Burgess, S. C. Excessive hepatic mitochondrial TCA cycle and gluconeogenesis in humans with nonalcoholic fatty liver disease. Cell Metab. 14, 804–810 (2011).
Kang, W., Suzuki, M., Saito, T. & Miyado, K. Emerging Role of TCA Cycle-Related Enzymes in Human Diseases. Int. J. Mol. Sci. 22, 13057 (2021).
Weeks, J. et al. Evaluating Mechanisms of IDH1 Regulation through Site-Specific Acetylation Mimics. Biomolecules 11, 740 (2021).
Zhuo, F. F. et al. Lycorine promotes IDH1 acetylation to induce mitochondrial dynamics imbalance in colorectal cancer cells. Cancer Lett. 573, 216364 (2023).
Narayan, S., Bader, G. D. & Reimand, J. Frequent mutations in acetylation and ubiquitination sites suggest novel driver mechanisms of cancer. Genome Med. 8, 55 (2016).
Tian, P. et al. Metabolic and bariatric surgery in China: A summary of the Greater China Metabolic and Bariatric Surgery Database and comparison with other international registry databases. Diabetes Obes. Metab. 25, 27–33 (2023).
Peng, W. C. et al. Inflammatory Cytokine TNFα Promotes the Long-Term Expansion of Primary Hepatocytes in 3D Culture. Cell 175, 1607–1619.e1615 (2018).
Wang, H. et al. Inhibition of phospholipase D1 ameliorates hepatocyte steatosis and non-alcoholic fatty liver disease. JHEP Rep. 5, 100726 (2023).
Hou, T. et al. Cytoplasmic SIRT6-mediated ACSL5 deacetylation impedes nonalcoholic fatty liver disease by facilitating hepatic fatty acid oxidation. Mol. Cell 82, 4099–4115.e4099 (2022).
Bruinsma, B. G., Uygun, K., Yarmush, M. L. & Saeidi, N. Surgical models of Roux-en-Y gastric bypass surgery and sleeve gastrectomy in rats and mice. Nat. Protoc. 10, 495–507 (2015).
Song, Y. et al. Tyrosine kinase receptor B attenuates liver fibrosis by inhibiting TGF-β/SMAD signaling. Hepatology 78, 1433–1447 (2023).
Li, X. et al. The methyltransferase METTL3 negatively regulates nonalcoholic steatohepatitis (NASH) progression. Nat. Commun. 12, 7213 (2021).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Cline, M. S. et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2, 2366–2382 (2007).
Trefely, S. et al. Quantitative subcellular acyl-CoA analysis reveals distinct nuclear metabolism and isoleucine-dependent histone propionylation. Mol. Cell 82, 447–462.e446 (2022).
Ma, B. et al. Albumosomes formed by cytoplasmic pre-folding albumin maintain mitochondrial homeostasis and inhibit nonalcoholic fatty liver disease. Signal Transduct. Target Ther. 8, 229 (2023).
Xiao, M. C. et al. TRIB3-TRIM8 complex drives NAFLD progression by regulating HNF4α stability. J. Hepatol. 80, 778–791 (2024).
Pierard, M. et al. Interactions of exercise training and high-fat diet on adiponectin forms and muscle receptors in mice. Nutr. Metab. (Lond.) 13, 75 (2016).
Guan, C. J. et al. ZNF774 is a potent suppressor of hepatocarcinogenesis through dampening the NOTCH2 signaling (vol 325, pg 127, 2019). Oncogene 39, 2844–2844 (2020).
Ye, Q. et al. Deficiency of gluconeogenic enzyme PCK1 promotes metabolic-associated fatty liver disease through PI3K/AKT/PDGF axis activation in male mice. Nat. Commun. 14, 1402 (2023).
Lin, Z. et al. RING finger protein 13 protects against nonalcoholic steatohepatitis by targeting STING-relayed signaling pathways. Nat. Commun. 14, 6635 (2023).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Acknowledgements
We thank Fengxiang Lu from Capital Medical University for scientific editing of this manuscript, Dong Zhang and Guangyong Sun from Beijing Chao-Yang Hospital for providing critical comments on the manuscript, Xinyan Zhao from Beijing Friendship Hospital for performing histological analysis, Yang Li from Peking University for providing experimental and technical support, Peng Jin from Ruijin Hospital for conducting data analysis. We thank Jingjie PTM biolab (Hangzhou) Co. Ltd. for MS analysis. We thank the Metabolomics Facility Center of Metabolomics and Lipidomics at the National Protein Science Technology Center of Tsinghua University for LC-MS/MS experiments. This research was funded by the National Natural Science Foundation of China (82200964 to H.Y., 82103484 to C.G., 81872296 to S.L., and 82070580 to Z.Z.) and the National key clinical specialist construction Programs, Technologies Research and Development Program (2015BAI13B09 to Z.Z.), National Key R&D Program of China (2022YFA1304800 to S.L.), Capital’s Funds for Health Improvement and Research (2020-1-2021 and 2024-1-1192 to Z.Z.), Beijing Natural Science Foundation (7214218 to C.G.), Beijing Postdoctoral Research Foundation (2022-ZZ-004 to H.Y.), China Postdoctoral Science Foundation (2023M732411 to H.Y.), Science and Technology Project of Beijing Education Committee (KM202110025017 to C.G.), Beijing Hospitals Authority Youth Program (QML20230116 to C.G.) and Beijing Friendship Hospital Youth Program (yyqcjh2023-6 to C.G.).
Author information
Authors and Affiliations
Contributions
Author contributions: Z.Z., H.Y., and Shanshan L. planned the project and analyzed data. Y. J., Shanshan L., and H.Y. wrote the manuscript. Shanshan L., Y. Z., L. Z., H.Y., Y. J., L.W. carried out molecular experiments. C.G., and L.W. carried out and analyzed the proteomic experiments. K.L., D.Y., Y.M., provided experimental assistance. K.L., L.Z., C.G., Shumeng L., Z.Z. discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, S., Ji, Y., Wei, L. et al. Crotonylation of IDH1 alleviates MASLD progression by enhancing the TCA cycle. Nat Commun 16, 7961 (2025). https://doi.org/10.1038/s41467-025-62731-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62731-9