Abstract
After birth, the heart undergoes a shift in energy metabolism and cytoarchitecture to enhance efficient energy production and cardiac contraction, which is essential for postnatal development and growth. However, the precise mechanisms regulating this process remain elusive. Here we show that the RNA modification enzyme Mettl1 is a critical regulator of postnatal metabolic reprogramming and cardiomyocyte maturation in mice, primarily through its influence on the translation of the rate-limiting ketogenesis enzyme Hmgcs2. Our findings reveal that ketogenesis is vital for the postnatal transition of fuel from glucose to fatty acids in cardiomyocytes, achieved by modulating tricarboxylic acid cycle–related enzymatic activity via lysine β-hydroxybutyrylation protein modification. Loss of Mettl1 results in aberrant metabolic reprogramming and cardiomyocyte immaturity, leading to heart failure, although some clinical features can be rescued by β-hydroxybutyrate supplementation. Our study provides mechanistic insights into how Mettl1 regulates metabolic reprogramming in neonatal cardiomyocytes and highlights the importance of ketogenesis in cardiomyocyte maturation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The raw sequencing data generated in this study are available at the National Center for Biotechnology Informationʼs Gene Expression Omnibus (GSE263137). Source data are provided with this paper. Other data that support the findings of this study are provided in Extended Data and Supplementary Information files. For RNA-seq and Ribo-Seq analysis, the reference genomes (GRCm38 (Ensembl release 102) and Rnor_6.0 (Ensembl release 104)) used for read alignment are from the Ensembl database.
Code availability
All the software and algorithms employed in this study are presented in the Methods section and are available without any access restrictions.
References
Maroli, G. & Braun, T. The long and winding road of cardiomyocyte maturation. Cardiovasc. Res. 117, 712–726 (2021).
Puente, B. N. et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157, 565–579 (2014).
Guo, Y. & Pu, W. T. Cardiomyocyte maturation: new phase in development. Circ. Res. 126, 1086–1106 (2020).
Singh, B. N. et al. Proliferation and maturation: Janus and the art of cardiac tissue engineering. Circ. Res. 132, 519–540 (2023).
Li, Z. et al. Postnatal state transition of cardiomyocyte as a primary step in heart maturation. Protein Cell 13, 842–862 (2022).
Karbassi, E. et al. Cardiomyocyte maturation: advances in knowledge and implications for regenerative medicine. Nat. Rev. Cardiol. 17, 341–359 (2020).
Laflamme, M. A. & Murry, C. E. Heart regeneration. Nature 473, 326–335 (2011).
Martin, O. J. et al. A role for peroxisome proliferator-activated receptor γ coactivator-1 in the control of mitochondrial dynamics during postnatal cardiac growth. Circ. Res. 114, 626–636 (2014).
Gaspar, J. A. et al. Unique metabolic features of stem cells, cardiomyocytes, and their progenitors. Circ. Res. 114, 1346–1360 (2014).
Bartelds, B. et al. Perinatal changes in myocardial metabolism in lambs. Circulation 102, 926–931 (2000).
Cardoso, A. C. et al. Mitochondrial substrate utilization regulates cardiomyocyte cell cycle progression. Nat. Metab. 2, 167–178 (2020).
Aubert, G. et al. The failing heart relies on ketone bodies as a fuel. Circulation 133, 698–705 (2016).
Bedi, K. C. Jr. et al. Evidence for intramyocardial disruption of lipid metabolism and increased myocardial ketone utilization in advanced human heart failure. Circulation 133, 706–716 (2016).
Horton, J. L. et al. The failing heart utilizes 3-hydroxybutyrate as a metabolic stress defense. JCI Insight 4, e124079 (2019).
Uchihashi, M. et al. Cardiac-specific Bdh1 overexpression ameliorates oxidative stress and cardiac remodeling in pressure overload-induced heart failure. Circ. Heart Fail. 10, e004417 (2017).
Talman, V. et al. Molecular atlas of postnatal mouse heart development. J. Am. Heart Assoc. 7, e010378 (2018).
Yamada, S. et al. TEAD1 trapping by the Q353R–Lamin A/C causes dilated cardiomyopathy. Sci. Adv. 9, eade7047 (2023).
Delaughter, D. M. et al. Single-cell resolution of temporal gene expression during heart development. Dev. Cell 39, 480–490 (2016).
Quaife-Ryan, G. A. et al. Multicellular transcriptional analysis of mammalian heart regeneration. Circulation 136, 1123–1139 (2017).
Zhang, H. et al. AP-1 activation mediates post-natal cardiomyocyte maturation. Cardiovasc. Res. 119, 536–550 (2023).
Sakamoto, T. et al. A critical role for estrogen-related receptor signaling in cardiac maturation. Circ. Res. 126, 1685–1702 (2020).
Wickramasinghe, N. M. et al. PPARdelta activation induces metabolic and contractile maturation of human pluripotent stem cell-derived cardiomyocytes. Cell Stem Cell 29, 559–576 (2022).
Guo, Y. et al. Sarcomeres regulate murine cardiomyocyte maturation through MRTF-SRF signaling. Proc. Natl Acad. Sci. USA 118, e2008861118 (2021).
Zhou, P. et al. Dynamic changes in P300 enhancers and enhancer–promoter contacts control mouse cardiomyocyte maturation. Dev. Cell 58, 898–914 (2023).
Alfar, E. A., El-Armouche, A. & Guan, K. MicroRNAs in cardiomyocyte differentiation and maturation. Cardiovasc. Res. 114, 779–781 (2018).
Poon, E. N. et al. Integrated transcriptomic and regulatory network analyses identify microRNA-200c as a novel repressor of human pluripotent stem cell-derived cardiomyocyte differentiation and maturation. Cardiovasc. Res. 114, 894–906 (2018).
Dorn, L. E. et al. The N6-methyladenosine mRNA methylase METTL3 controls cardiac homeostasis and hypertrophy. Circulation 139, 533–545 (2019).
Mathiyalagan, P. et al. FTO-dependent N6-methyladenosine regulates cardiac function during remodeling and repair. Circulation 139, 518–532 (2019).
Han, Y. et al. Loss of m6A methyltransferase METTL5 promotes cardiac hypertrophy through epitranscriptomic control of SUZ12 expression. Front. Cardiovasc. Med. 9, 852775 (2022).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Paredes, A. et al. γ-Linolenic acid in maternal milk drives cardiac metabolic maturation. Nature 618, 365–373 (2023).
Fujimoto, Y. et al. Long-chain fatty acids induce lipid droplet formation in a cultured human hepatocyte in a manner dependent of Acyl-CoA synthetase. Biol. Pharm. Bull. 29, 2174–2180 (2006).
Wang, Y. et al. Polyunsaturated fatty acids promote the rapid fusion of lipid droplets in Caenorhabditis elegans. J. Biol. Chem. 298, 102179 (2022).
Wu, D. et al. Prohibitin 2 deficiency impairs cardiac fatty acid oxidation and causes heart failure. Cell Death Dis. 11, 181 (2020).
Kienesberger, P. C. et al. Early structural and metabolic cardiac remodelling in response to inducible adipose triglyceride lipase ablation. Cardiovasc. Res. 99, 442–451 (2013).
Cheng, L. et al. Cardiomyocyte-restricted peroxisome proliferator-activated receptor-ẟ deletion perturbs myocardial fatty acid oxidation and leads to cardiomyopathy. Nat. Med. 10, 1245–1250 (2004).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 49, D1541–d7 (2021).
Xie, Z. et al. Metabolic regulation of gene expression by histone lysine β-hydroxybutyrylation. Mol. Cell 62, 194–206 (2016).
Koronowski, K. B. et al. Ketogenesis impact on liver metabolism revealed by proteomics of lysine β-hydroxybutyrylation. Cell Rep. 36, 109487 (2021).
Ho, K. L. et al. Increased ketone body oxidation provides additional energy for the failing heart without improving cardiac efficiency. Cardiovasc. Res. 115, 1606–1616 (2019).
Zhuang, T. et al. ALKBH5-mediated m6A modification of IL-11 drives macrophage-to-myofibroblast transition and pathological cardiac fibrosis in mice. Nat. Commun. 15, 1995 (2024).
Song, H. et al. METTL3 and ALKBH5 oppositely regulate m6A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy 15, 1419–1437 (2019).
Wu, L. et al. Inhibition of METTL3 ameliorates doxorubicin-induced cardiotoxicity through suppression of TFRC-mediated ferroptosis. Redox Biol. 72, 103157 (2024).
Shi, J. et al. NAT10 is involved in cardiac remodeling through ac4C-mediated transcriptomic regulation. Circ. Res. 133, 989–1002 (2023).
Wang, K. et al. PIWI-interacting RNA HAAPIR regulates cardiomyocyte death after myocardial infarction by promoting NAT10-mediated ac4C acetylation of Tfec mRNA. Adv. Sci. (Weinh.) 9, e2106058 (2022).
Ma, W. et al. N-Acetyltransferase 10 represses Uqcr11 and Uqcrb independently of ac4C modification to promote heart regeneration. Nat. Commun. 15, 2137 (2024).
Xu, T. et al. Loss of NAT10 reduces the translation of Kmt5a mRNA through ac4C modification in cardiomyocytes and induces heart failure. J. Am. Heart Assoc. 13, e035714 (2024).
Chen, X. Z. et al. TMEM11 regulates cardiomyocyte proliferation and cardiac repair via METTL1-mediated m7G methylation of ATF5 mRNA. Cell Death Differ. 30, 1786–1798 (2023).
Yu, S. et al. The m7G methyltransferase Mettl1 drives cardiac hypertrophy by regulating SRSF9-mediated splicing of NFATc4. Adv. Sci. (Weinh.) 11, e2308769 (2024).
Lopaschuk, G. D. et al. Cardiac energy metabolism in heart failure. Circ. Res. 128, 1487–1513 (2021).
Murashige, D. et al. Comprehensive quantification of fuel use by the failing and nonfailing human heart. Science 370, 364–368 (2020).
Schugar, R. C. et al. Cardiomyocyte-specific deficiency of ketone body metabolism promotes accelerated pathological remodeling. Mol. Metab. 3, 754–769 (2014).
Miyamoto, J. et al. Ketone body receptor GPR43 regulates lipid metabolism under ketogenic conditions. Proc. Natl Acad. Sci. USA 116, 23813–23821 (2019).
Gibson, K. M. et al. Fatal cardiomyopathy associated with 3-hydroxy-3-methylglutaryl-CoA lyase deficiency. J. Inherit. Metab. Dis. 17, 291–294 (1994).
Leung, A. A. et al. A case of dilated cardiomyopathy associated with 3-hydroxy-3-methylglutaryl-coenzyme A (HMG CoA) lyase deficiency. Case Rep. Med. 2009, 183125 (2009).
Han, H. et al. N7-methylguanosine tRNA modification promotes esophageal squamous cell carcinoma tumorigenesis via the RPTOR/ULK1/autophagy axis. Nat. Commun. 13, 1478 (2022).
Yan, Y. et al. The cardiac translational landscape reveals that micropeptides are new players involved in cardiomyocyte hypertrophy. Mol. Ther. 29, 2253–2267 (2021).
Lin, S. et al. Nucleotide resolution profiling of m7G tRNA modification by TRAC-Seq. Nat. Protoc. 14, 3220–3242 (2019).
Schwartz, S. et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159, 148–162 (2014).
Acknowledgements
We thank members of the Huang laboratory for advice and support. This work is supported by the National Key R&D Program of China (2021YFA1302500); the National Natural Science Foundation of China (U23A20396 and 82270252 to Z.-P.H. and 92268202 to J.O.); the Guangzhou Municipal Science and Technology Bureau (202206010139 to Z.-P.H.); and the Non-profit Central Research Institute Fund of the Chinese Academy of Medical Sciences (2023-PT320-03 to Y.L.).
Author information
Authors and Affiliations
Contributions
T.D. and Y.H. performed most of the experiments. T.D. and H.H. performed bioinformatic analyses. T.X., Y.Y. and J.W. built viral vectors and designed siRNAs/gRNAs used in this study. S.L. and D.C. provided Mettl1-flox and Mettl1-KI-flox mice used in this study. Y.L. isolated and maintained cardiomyocytes used in this study. C.L., X.L., Y.D. and J.O. performed quality control of all data and revised the manuscript. Z.-P.H. supervised the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Gary Lopaschuk, Mirko Völkers and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 M1cKO mice with immature cardiomyocyte develop heart failure.
a. Schematic of construction mice with cardiomyocyte-specific Mettl1 deletion. b-c. Western blotting of Mettl1 and Cre protein (b), and quantification of Mettl1 protein (c) in mouse hearts at indicated time points (n = 3 hearts for each group). d. Time course of change in heart-to-body weight ratio (Hw/Bw) of control and M1cKO mouse hearts (nctrl-4wk = 10 mice, nM1cKO-4wk = 10 mice, nctrl-6wk = 12 mice, nM1cKO-6wk = 12 mice, nctrl-9wk = 11 mice, nM1cKO-9wk = 7 mice). e-g. Time course of change in mRNA expression of Anp, Bnp, and Acta1 measured by qRT-PCR of control and M1cKO mouse hearts (n = 4 hearts for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant. Uncropped blots for b are provided in Source data.
Extended Data Fig. 2 RNA-seq and metabolomics analysis of M1cKO mouse hearts.
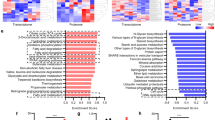
a. KEGG pathway enrichment analysis using the differentially expressed genes (DEGs) in 4-week-old M1cKO mouse hearts. b-c. Principal component analysis (PCA) of metabolites in negative (b) and positive (c) ion modes in 4-week-old control and M1cKO mouse hearts. d-e. Volcano plot of differential metabolites detected in negative (d) and positive (e) ion modes (including unidentified ones) in 4-week-old control and M1cKO mouse hearts. f. Grouping of metabolic hits identified by LC-MS/MS analysis in 4-week-old control and M1cKO mouse hearts (n = 5 hearts for each group). g. Quantification of citrate by LC-MS/MS analysis in control and M1cKO mouse hearts (n = 4 hearts for each group). h. Quantification of medium-chain fatty acids (MCFAs) and long-chain fatty acids (LCFAs) by LC-MS/MS analysis in control and M1cKO mouse hearts (n = 4 hearts for each group). i-j. Quantification of cholesterol (i) and TG (52:3) (j) by LC-MS/MS analysis in control and M1cKO mouse hearts (n = 4 hearts for each group). k. Quantification of C20:3 (M + 16) in control and M1KD NRVMs after [U-13C16] palmitate labeling for 24 h, as revealed by metabolic flux assay (n = 3 hearts for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant.
Extended Data Fig. 3 Mettl1 alters the accumulation of lipid substances but not through enzymes involved in TG turnover.
a. The mean fluorescence intensity (MFI) of BODIPY measured by flow cytometry analysis in control and M1KD NRVMs under BSA or OAPA treatment (n = 3 biological replicates for each group). b. Heat map of normalized triglycerides (TG) and cholesteryl esters (CE) concentration identified by combining liquid chromatography with tandem mass spectrometry (LC-MS/MS) in control and M1KD NRVMs under OAPA treatment. c. mRNA expression of TG synthesis genes and TG lipolysis genes measured by qRT-PCR in control and M1KD NRVMs under OAPA treatment (n = 3 biological replicates for each group). d-e. Western blotting and quantification of TG synthesis proteins and TG lipolysis proteins in control and M1KD NRVMs under OAPA treatment (d) (n = 3 biological replicates for each group) and in control and M1cKO mouse hearts (e) (n = 3 hearts for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant. Uncropped blots for d and e are provided in Source data.
Extended Data Fig. 4 Mettl1 regulates fatty acid metabolism in H9C2 cardiomyocytes.
a. Sequence confirmation of CRISPR/Cas9-mediated gene editing in Mettl1 in H9C2 CMs. b. Western blot analysis of the Mettl1 in WT and Mettl1-knockout (M1-null) H9C2 CMs (n = 1 biological replicate for each group, observed in 3 independent experiments). c. Quantification of indicated lipids by LC-MS/MS in WT and M1-null H9C2 CMs (n = 4 biological replicates for each group). d. Quantification of indicated metabolites by LC-MS/MS in WT and M1-null H9C2 CMs (n = 4 biological replicates for each group). e-f. Representative images of BODIPY staining indicating the accumulation of neutral lipid droplets (green dots) in WT and M1-null H9C2 CMs under BSA or OAPA treatment (e) and quantification of BODIPY intensity (f). The nucleus was stained with DAPI. Scale bar, 20 µm. g. The mean fluorescence intensity (MFI) of BODIPY measured by flow cytometry analysis in WT and M1-null H9C2 CMs under BSA or OAPA treatment. h-i. Triglyceride (h) and cholesterol (i) levels in WT and M1-null H9C2 CMs under OAPA treatment. j. mRNA expression of slc27a1 measured by qRT-PCR in WT and M1-null H9C2 CMs under OAPA treatment. k. Measurement of fluorescence-labeled long-chain fatty acids uptaken by WT and M1-null H9C2 CMs under OAPA treatment. l. Measurement of intracellular fluorescence-labeled glucose uptake by WT and M1-null H9C2 CMs under OAPA treatment. m. Extracellular lactate levels in WT and M1-null H9C2 CMs under OAPA treatment. n. ATP contents in WT and M1-null H9C2 CMs under OAPA treatment. n = 3 biological replicates for each group in (f-n). o. KEGG pathway enrichment analysis using the differentially expressed genes (DEGs) in M1-null H9C2 CMs under OAPA treatment. p. Volcano plot of DEGs in M1-null H9C2 CMs under OAPA treatment. Statistical analysis was evaluated using unpaired two-tailed Student’s t-test in c, d, f, h-n; and one-way ANOVA with Tukey’s multiple comparisons test in g. The values are mean ± s.d. Uncropped blots for b are provided in Source data.
Extended Data Fig. 5 Mettl1 regulates tRNA m7G modification but has minimal impact on other methylation reactions.
a. Quantification of SAM metabolism-related metabolites by LC-MS/MS analysis in control and M1cKO mouse hearts (n = 4 hearts for each group). b. Dot blotting and quantification of N6-methyladenosine (m6A) and m7G levels in total RNA from control and M1cKO mouse hearts (n = 3 hearts for each group). c. Western blotting of protein with mono-methylation (left panels) (n = 2 hearts for each group, observed in 3 independent experiments), di-methylation (middle panels) (n = 3 hearts for each group, observed in 3 independent experiments) and tri-methylation (right panels) (n = 3 hearts for each group, observed in 3 independent experiments) of lysine residues in control and M1cKO mouse hearts. d. Western blotting of protein with symmetrical (left panels) and asymmetric (right panels) di-methylation of arginine residues in control and M1cKO mouse hearts (n = 3 hearts for each group, observed in 3 independent experiments). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant. Uncropped blots for c-d are provided in Source data.
Extended Data Fig. 6 Mettl1 does not mediate translational regulation of FAO genes.
a. Western blotting and quantification of fatty acid oxidation proteins in 4-week-old control and M1cKO mouse hearts. b. mRNA expression of fatty acid oxidation genes measured by qRT-PCR in 4-week-old control and M1cKO mouse hearts. c. TE analysis of fatty acid oxidation genes based on qRT-PCR and western blotting in 4-week-old control and M1cKO mouse hearts. d. Western blotting and quantification of fatty acid oxidation proteins in control and M1cKO mouse hearts at P7. e. mRNA expression of fatty acid oxidation genes measured by qRT-PCR in control and M1cKO mouse hearts at P7. f. TE analysis of fatty acid oxidation genes based on qRT-PCR and western blotting in control and M1cKO mouse hearts at P7. n = 3 hearts for each group in (a-f). g. GSEA plot of oxidative phosphorylation pathway in RNA-Seq data upon control and M1cKO mouse hearts at P7. h. mRNA expression of oxidative phosphorylation genes measured by qRT-PCR in control and M1cKO mouse hearts at P7 (n = 4 hearts for each group). i. Western blotting and quantification of Uqcrc2 in control and M1cKO mouse hearts at P7 (n = 3 hearts for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant. Uncropped blots for a, d and i are provided in Source data.
Extended Data Fig. 7 Mettl1 regulates the translation of Hmgcs2 but not Hmgcs2 enzyme activity and ketone oxidation.
a. Relative MS intensity of indicated proteins in P1, P4, P9, P23 mouse hearts (nP1 = 14 hearts, n4 = 14 hearts, nP9 = 10 hearts, nP23 = 10 hearts, missing values are eliminated) (GSE119530). b. Relative total enzyme activity of Hmgcs2 in control and M1cKO hearts (n = 3 hearts for each group). c. Western blotting and quantification of Hmgcs2 in control and M1cKO mouse hearts (n = 3 hearts for each group). d. Relative enzyme activity of Hmgcs2 normalized to the corresponding protein levels in control and M1cKO hearts (n = 3 hearts for each group). e-f. Western blotting and quantification of enzymes involved in ketone oxidation in control and M1cKO mouse hearts (e) (n = 3 hearts for each group) and in control and M1KD NRVMs (f) (n = 3 biological replicates for each group). g. Fraction of labeled acetylcarnitine from [U-13C]β-HB in control and M1cKO mouse hearts, as revealed by metabolic flux assay (n = 3 hearts for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. NS, not significant. Uncropped blots for c, e and f are provided in Source data.
Extended Data Fig. 8 Hmgcs2 overexpression rescues the impaired fatty acid metabolism and immaturity in CM induced by loss of Mettl1.
a. The mean fluorescence intensity (MFI) of BODIPY measured by flow cytometry analysis in OAPA-treated control and M1KD NRVMs supplied with PBS or β-HB (n = 4 biological replicates for each group). b. Western blot analysis of Hmgcs2 in NRVMs infected with Ad-GFP or Ad-Hmgcs2 (Ad-H2) (n = 3 biological replicates for each group, observed in 3 independent experiments). c-d. Representative images of nile red staining indicating the accumulation of neutral lipid droplets (red dots) in OAPA-treated control and M1KD NRVMs infected with Ad-GFP or Ad-H2 (c) and quantification of nile red intensity (d). The nucleus was stained with DAPI. Scale bar, 20 µm. n = 3 biological replicates for each group. e. The mean fluorescence intensity (MFI) of nile red measured by flow cytometry analysis in OAPA-treated control and M1KD NRVMs infected with Ad-GFP or Ad-H2 (n = 4 biological replicates for each group). f-g. Measurement of ATP contents (f) (n = 3 biological replicates for each group) and intracellular fluorescence-labeled glucose (g) (n = 4 biological replicates for each group) in OAPA-treated control and M1KD NRVMs infected with Ad-GFP or Ad-H2. h. Representative confocal images of OAPA-treated control and M1KD NRVMs infected with Ad-GFP or Ad-H2 stained for α-actinin and DAPI. Scale bars, 20 μm. i-j. Quantification of sarcomere length (i) and sarcomere network (j) in confocal images (n = 50 CMs from 3 biological replicates). k. The mean fluorescence intensity (MFI) of Ca2+ measured by flow cytometry analysis in OAPA-treated control and M1KD NRVMs infected with Ad-GFP or Ad-H2 (n = 4 biological replicates for each group). l. Western blot analysis of Hmgcs2 in H9C2 CMs infected with lenti-vector or lenti-Hmgcs2 (lenti-H2) (n = 1 biological replicate for each group, observed in 3 independent experiments)). m-n. Representative images of BODIPY staining indicating the accumulation of neutral lipid droplets (green dots) in OAPA-treated WT and M1-null H9C2 CMs infected with lenti-vector or lenti-H2 (m) and quantification of BODIPY intensity (n). The nucleus was stained with DAPI. Scale bar, 20 µm. o. The mean fluorescence intensity (MFI) of BODIPY measured by flow cytometry analysis in OAPA-treated WT and M1-null H9C2 CMs infected with lenti-vector or lenti-H2. p-q Measurement of ATP contents (p) and intracellular fluorescence-labeled glucose (q) in OAPA-treated WT and M1-null H9C2 CMs infected with lenti-vector or lenti-H2. n = 3 biological replicates for each group in (n-q). r. Western blotting of protein K-bhb in PBS or β-HB-treated NRVMs (n = 2 biological replicates for each group, observed in 3 independent experiments). s. Oxygen consumption rate (OCR) detected by O2K analysis in PBS or β-HB-treated NRVMs (supplemented with OAPA) (n = 3 biological replicates for each group). t. Quantification of basal and maximal OCRs in the O2K analysis data (n = 3 biological replicates for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test in t; and one-way ANOVA with Tukey’s multiple comparisons test in a, d-g, i-k, and n-q. The values are mean ± s.d. Uncropped blots for b, l and r are provided in Source data.
Extended Data Fig. 9 β-HB treatment promotes K-bhb modifications and rescues decreased activities of TCA cycle-related enzymes in M1-null H9C2 CMs.
a-c. Relative enzyme activities of mitochondrial IDH (IDHm) (a), succinyl-CoA synthetase (SCS) (b) and cytoplasmic MDH (c) in OAPA-treated WT and M1-null H9C2 cardiomyocytes (n = 3 biological replicates for each group). d-g. Immunoprecipitation assays (d) and quantification (e-g) of PBS or β-HB-treated Mettl1-null H9C2 cardiomyocytes. IP: anti-K-hbb; IB: Idh2, Sucla2 and Mdh1. n = 3 biological replicates for each group. h-j. Relative enzyme activities of mitochondrial IDH (IDHm) (h), succinyl-CoA synthetase (SCS) (i) and cytoplasmic MDH (j) in PBS or β-HB-treated Mettl1-null H9C2 cardiomyocytes (n = 3 biological replicates for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test. The values are mean ± s.d. Uncropped blots for d are provided in Source data.
Extended Data Fig. 10 β-HB supplementation rescues the impaired fatty acid metabolism and immaturity in Hmgcs2-KD NRVMs.
a. mRNA expression of Hmgcs2 measured by qRT-PCR in NRVMs transfected with negative control siRNA (siNC) or Hmgcs2 siRNA (siH2) (control and Hmgcs2-KD NRVMs hereafter). n = 3 biological replicates for each group. b. Western blotting and quantification of Hmgcs2 in control and Hmgcs2-KD NRVMs (n = 3 biological replicates for each group). c-d. Representative images of BODIPY staining indicating the accumulation of neutral lipid droplets (green dots) in OAPA-treated control and Hmgcs2-KD NRVMs supplemented with PBS or β-HB (c) and quantification of BODIPY intensity (d). The nucleus was stained with DAPI. Scale bar, 20 µm; n = 3 biological replicates for each group. e. Measurement of ATP contents in OAPA-treated control and Hmgcs2-KD NRVMs supplemented with PBS or β-HB (n = 3 biological replicates for each group). f. Representative confocal images of OAPA-treated control and Hmgcs2-KD NRVMs supplemented with PBS or β-HB stained for α-actinin and DAPI. Scale bars, 20 μm. g-h. Quantification of sarcomere length (g) and sarcomere network (h) in confocal image data (n = 50 CMs from 3 biological replicates). i-l. Immunoprecipitation assays (i) and quantification (j-l) of OAPA-treated control and Hmgcs2-KD NRVMs supplemented with PBS or β-HB. IP: anti-K-hbb; IB: Idh2, Sucla2 and Mdh1. n = 3 biological replicates for each group. m-o. Relative enzyme activities of mitochondrial IDH (IDHm) (m), succinyl-CoA synthetase (SCS) (n) and cytoplasmic MDH (o) in control and Hmgcs2-KD NRVMs supplemented with PBS or β-HB (n = 3 biological replicates for each group). Statistical analysis was evaluated using unpaired two-tailed Student’s t-test in a and b; and one-way ANOVA with Tukey’s multiple comparisons test in d, e, g, h, and j-o. The values are mean ± s.d. Uncropped blots for b and i are provided in Source data.
Supplementary information
Supplementary information (download PDF )
Supplementary Figs. 1–7.
Supplementary Tables 1–18 (download XLSX )
Supplementary tables for this paper.
Supplementary Data 1 (download PDF )
Source data for supplementary figures.
Supplementary Data (download PDF )
Unprocessed blot and gel images.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 8 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 9 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig./Table 10 (download XLSX )
Statistical source data.
Source Data Extended Data Fig./Table 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Du, T., Han, Y., Han, H. et al. The tRNA methyltransferase Mettl1 governs ketogenesis through translational regulation and drives metabolic reprogramming in cardiomyocyte maturation. Nat Cardiovasc Res 3, 1438–1453 (2024). https://doi.org/10.1038/s44161-024-00565-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-024-00565-2
This article is cited by
-
tRNA methylation drives early postnatal cardiomyocyte maturation
Nature Cardiovascular Research (2024)